Abstract
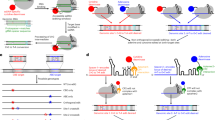
The CRISPR prime editor PE2 consists of a Streptococcus pyogenes Cas9 nickase (nSpCas9) fused at its C-terminus to a Moloney murine leukemia virus reverse transcriptase (MMLV-RT). Here we show that separated nSpCas9 and MMLV-RT proteins function as efficiently as intact PE2 in human cells. We use this Split-PE system to rapidly identify and engineer more compact prime editor architectures that also broaden the types of RTs used for prime editing.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
Plasmids encoding constructs used in this study have been deposited at Addgene and will be available at https://www.addgene.org/Keith_Joung (Addgene plasmid nos. 190104–190112). All targeted amplicon sequencing data have been deposited at the National Center of Biotechnology Information’s Sequence Read Archive and can be accessed via http://www.ncbi.nlm.nih.gov/bioproject/861237 (ref. 40).
Code availability
Custom Python scripts that were generated for CRISPResso analyses are provided in the Supplementary Information (Supplementary Note 2).
References
Anzalone, A. V. et al. Search-and-replace genome editing without double-strand breaks or donor DNA. Nature 576, 149–157 (2019).
Chen, P. J. et al. Enhanced prime editing systems by manipulating cellular determinants of editing outcomes. Cell 184, 5635–5652 (2021).
Li, X. et al. Highly efficient prime editing by introducing same-sense mutations in pegRNA or stabilizing its structure. Nat. Commun. 13, 1669 (2022).
Xu, W. et al. A design optimized prime editor with expanded scope and capability in plants. Nat Plants 8, 45–52 (2022).
Wang, Y. et al. BE-PIGS: a base-editing tool with deaminases inlaid into Cas9 PI domain significantly expanded the editing scope. Signal Transduct. Target Ther. 4, 36 (2019).
Petri, K. et al. CRISPR prime editing with ribonucleoprotein complexes in zebrafish and primary human cells. Nat. Biotechnol. 40, 189–193 (2022).
Kleinstiver, B. P. et al. Broadening the targeting range of Staphylococcus aureus CRISPR–Cas9 by modifying PAM recognition. Nat. Biotechnol. 33, 1293–1298 (2015).
Tsai, S. Q. et al. GUIDE-seq enables genome-wide profiling of off-target cleavage by CRISPR–Cas nucleases. Nat. Biotechnol. 33, 187–197 (2015).
Kleinstiver, B. P. et al. High-fidelity CRISPR–Cas9 nucleases with no detectable genome-wide off-target effects. Nature 529, 490–495 (2016).
Gu, J. et al. Substitution of Asp114 or Arg116 in the fingers domain of Moloney murine leukemia virus reverse transcriptase affects interactions with the template-primer resulting in decreased processivity. J. Mol. Biol. 305, 341–359 (2001).
Das, D. et al. A directed approach to improving the solubility of Moloney murine leukemia virus reverse transcriptase. Protein Sci. 10, 1936–1941 (2001).
Katano, Y. et al. Generation of thermostable Moloney murine leukemia virus reverse transcriptase variants using site saturation mutagenesis library and cell-free protein expression system. Biosci. Biotechnol. Biochem. 81, 2339–2345 (2017).
Cote, M. L. et al. Murine leukemia virus reverse transcriptase: structural comparison with HIV-1 reverse transcriptase. Virus Res. 134, 186–202 (2008).
Das, D. et al. The crystal structure of the monomeric reverse transcriptase from Moloney murine leukemia virus. Structure 12, 819–829 (2004).
Bock, D. et al. In vivo prime editing of a metabolic liver disease in mice. Sci. Transl. Med. 14, eabl9238 (2022).
Ann Ran, F. et al. In vivo genome editing using Staphylococcus aureus Cas9. Nature 520, 186–191 (2015).
Yu, S. F. et al. Human foamy virus replication: a pathway distinct from that of retroviruses and hepadnaviruses. Science 271, 1579–1582 (1996).
Wohrl, B. M. Structural and functional aspects of foamy virus protease-reverse transcriptase. Viruses 11, 598 (2019).
Berkhout, B. et al. Identification of an active reverse transcriptase enzyme encoded by a human endogenous HERV-K retrovirus. J. Virol. 73, 2365–2375 (1999).
Lee, Y. N. et al. Reconstitution of an infectious human endogenous retrovirus. PLoS Pathog. 3, e10 (2007).
Mills, D. A. et al. Splicing of a group II intron involved in the conjugative transfer of pRS01 in lactococci. J. Bacteriol. 178, 3531–3538 (1996).
Mohr, S. et al. Thermostable group II intron reverse transcriptase fusion proteins and their use in cDNA synthesis and next-generation RNA sequencing. RNA 19, 958–970 (2013).
Dai, L. et al. ORF-less and reverse-transcriptase-encoding group II introns in archaebacteria, with a pattern of homing into related group II intron ORFs. RNA 9, 14–19 (2003).
Blocker, F. J. et al. Domain structure and three-dimensional model of a group II intron-encoded reverse transcriptase. RNA 11, 14–28 (2005).
Stamos, J. L. et al. Structure of a thermostable group II intron reverse transcriptase with template-primer and its functional and evolutionary implications. Mol. Cell 68, 926–939 (2017).
Zhao, C. et al. Crystal structures of a group II intron maturase reveal a missing link in spliceosome evolution. Nat. Struct. Mol. Biol. 23, 558–565 (2016).
Zhao, C. et al. An ultraprocessive, accurate reverse transcriptase encoded by a metazoan group II intron. RNA 24, 183–195 (2018).
Kelley, L. A. et al. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protoc. 10, 845–858 (2015).
Liu, P. et al. Improved prime editors enable pathogenic allele correction and cancer modelling in adult mice. Nat. Commun. 12, 2121 (2021).
Liu, B. et al. A split prime editor with untethered reverse transcriptase and circular RNA template. Nat. Biotechnol. 40, 779–786 (2022).
Truong, D. J. et al. Development of an intein-mediated split–Cas9 system for gene therapy. Nucleic Acids Res. 43, 6450–6458 (2015).
Levy, J. M. et al. Cytosine and adenine base editing of the brain, liver, retina, heart and skeletal muscle of mice via adeno-associated viruses. Nat. Biomed. Eng. 4, 97–110 (2020).
Toro, N. et al. Multiple origins of reverse transcriptases linked to CRISPR–Cas systems. RNA Biol. 16, 1486–1493 (2019).
Hopp, T. P. et al. A short polypeptide marker sequence useful for recombinant protein identification and purification. Bio/Technology 6, 1204–1210 (1988).
Rohland, N. et al. Cost-effective, high-throughput DNA sequencing libraries for multiplexed target capture. Genome Res. 22, 939–946 (2012).
Gibson, D. G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Kleinstiver, B. P. et al. Engineered CRISPR–Cas9 nucleases with altered PAM specificities. Nature 523, 481–485 (2015).
Gramlich, M. et al. Antisense‐mediated exon skipping: a therapeutic strategy for titin‐based dilated cardiomyopathy. EMBO Mol. Med. 7, 562–576 (2015).
Clement, K. et al. CRISPResso2 provides accurate and rapid genome editing sequence analysis. Nat. Biotechnol. 37, 224–226 (2019).
Grünewald, J. et al. Datasets. Sequence Read Archive (SRA). http://www.ncbi.nlm.nih.gov/bioproject/861237 (2022).
Acknowledgements
Support for this work was provided by the National Institutes of Health (RM1 HG009490 and R35 GM118158 to J.K.J.). J.K.J. is additionally supported by the Desmond and Ann Heathwood MGH Research Scholar Award and the Robert B. Colvin, M.D., Endowed Chair in Pathology. J.G. was funded by the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation), Projektnummer 416375182. The project described was supported by a Career Development Award from the American Society of Gene & Cell Therapy (awarded to J.G.). The content is solely the responsibility of the authors and does not necessarily represent the official views of the American Society of Gene & Cell Therapy. K.P. was funded by the DFG, Projektnummer 417577129. We thank M. K. Clement for technical advice; R. Zhou, J. Y. Hsu, A. Schmidts, J. F. Angstman and P. Exconde for discussions and technical advice; and L. P. Pottenplackel for assistance with editing the manuscript.
Author information
Authors and Affiliations
Contributions
B.R.M., R.N.S., C.J.W., P.K.C., E.J.B.H. and J.G. performed the laboratory experiments. J.G., R.S. and K.P. performed the computational analyses. J.G., B.R.M, R.N.S. and J.K.J. conceived of and designed the study. J.K.J. and J.G. supervised the work. J.G., R.N.S. and J.K.J. wrote the initial manuscript draft, with the help of L. P. Pottenplackel, and all authors contributed to the writing of the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
J.K.J. and two other investigators who work on the National Institutes of Health award supporting this research, but are not authors on this publication, are co-founders of and have a financial interest in SeQure Dx, Inc., a company developing technologies for gene editing target profiling. J.K.J. also has, or had during the course of this research, financial interests in several companies developing gene editing technology: Beam Therapeutics, Blink Therapeutics, Chroma Medicine, Editas Medicine, EpiLogic Therapeutics, Excelsior Genomics, Hera Biolabs, Monitor Biotechnologies, Nvelop Therapeutics (f/k/a ETx, Inc.), Pairwise Plants, Poseida Therapeutics and Verve Therapeutics. J.K.J.ʼs interests were reviewed and are managed by Massachusetts General Hospital and Mass General Brigham in accordance with their conflict of interest policies. J.K.J. is a co-inventor on various patents and patent applications that describe gene editing and epigenetic editing technologies. K.P. has a financial interest in SeQure Dx, Inc. K.P.ʼs interests and relationships have been disclosed to Massachusetts General Hospital and Mass General Brigham in accordance with their conflict of interest policies. P.K.C. is a paid consultant to and has financial interests in Nvelop Therapeutics (f/k/a ETx, Inc.). J.G., B.R.M. and J.K.J. are co-inventors on a patent application that has been filed by Mass General Brigham/Massachusetts General Hospital on engineered bipartite PE architectures, reduced size PEs and non-MMLV-RT PE architectures.
Peer review
Peer review information
Nature Biotechnology thanks Jia Chen and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–10 and Supplementary Notes 1 and 2
Supplementary Table 1
Editing frequencies in comparison experiments between intact and split PE architectures.
Supplementary Table 2
Editing frequencies and PBS/RTT lengths for a subset of comparison experiments between intact and split PE architectures.
Supplementary Table 3
Constructs used in this study.
Supplementary Table 4
gRNA and amplicon sequences, next-generation sequencing barcodes and read depth of individual experiments.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Grünewald, J., Miller, B.R., Szalay, R.N. et al. Engineered CRISPR prime editors with compact, untethered reverse transcriptases. Nat Biotechnol 41, 337–343 (2023). https://doi.org/10.1038/s41587-022-01473-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41587-022-01473-1
This article is cited by
-
Enhancing prime editor flexibility with coiled-coil heterodimers
Genome Biology (2024)
-
Prime editing using CRISPR-Cas12a and circular RNAs in human cells
Nature Biotechnology (2024)
-
Structural basis for pegRNA-guided reverse transcription by a prime editor
Nature (2024)
-
Enhancing prime editor activity by directed protein evolution in yeast
Nature Communications (2024)
-
Genome-wide profiling of prime editor off-target sites in vitro and in vivo using PE-tag
Nature Methods (2023)