Abstract
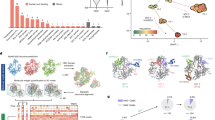
It was recently shown that bacteria use, apart from CRISPR–Cas and restriction systems, a considerable diversity of phage resistance systems1,2,3,4, but it is largely unknown how phages cope with this multilayered bacterial immunity. Here we analysed groups of closely related Bacillus phages that showed differential sensitivity to bacterial defence systems, and discovered four distinct families of anti-defence proteins that inhibit the Gabija, Thoeris and Hachiman systems. We show that these proteins Gad1, Gad2, Tad2 and Had1 efficiently cancel the defensive activity when co-expressed with the respective defence system or introduced into phage genomes. Homologues of these anti-defence proteins are found in hundreds of phages that infect taxonomically diverse bacterial species. We show that the anti-Gabija protein Gad1 blocks the ability of the Gabija defence complex to cleave phage-derived DNA. Our data further reveal that the anti-Thoeris protein Tad2 is a ‘sponge’ that sequesters the immune signalling molecules produced by Thoeris TIR-domain proteins in response to phage infection. Our results demonstrate that phages encode an arsenal of anti-defence proteins that can disable a variety of bacterial defence mechanisms.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Data that support the findings of this study are available in the article and Supplementary Tables 1–15. IMG and MGV accession numbers, protein sequences and nucleotide sequences are available in Supplementary Tables 8–14. Coordinates and structure factors for cbTad1–1″–3′-gcADPR, SPO1 Tad2 apo, SPO1 Tad2–1″–3′-gcADPR, SPO1 Tad2–1″–2′-gcADPR and Had1 have been deposited in the Protein Data Bank under the accession codes 8SMD, 8SME, 8SMF, 8SMG and 8TTO, respectively. The genome sequences of the phages SPO1L1–SPO1L5 and SPβL1–SPβL8 have been deposited with GenBank under accession codes OQ921336–OQ921348, respectively. Source data are provided with this paper.
References
Dy, R. L., Richter, C., Salmond, G. P. C. & Fineran, P. C. Remarkable mechanisms in microbes to resist phage infections. Annu. Rev. Virol. 1, 307–331 (2014).
Bernheim, A. & Sorek, R. The pan-immune system of bacteria: antiviral defence as a community resource. Nat. Rev. Microbiol. 18, 113–119 (2020).
Hampton, H. G., Watson, B. N. J. & Fineran, P. C. The arms race between bacteria and their phage foes. Nature 577, 327–336 (2020).
Tal, N. & Sorek, R. SnapShot: bacterial immunity. Cell 185, 578 (2022).
Samson, J. E., Magadán, A. H., Sabri, M. & Moineau, S. Revenge of the phages: defeating bacterial defences. Nat. Rev. Microbiol. 11, 675–687 (2013).
Atanasiu, C., Su, T.‐J., Sturrock, S. S. & Dryden, D. T. F. Interaction of the ocr gene 0.3 protein of bacteriophage T7 with EcoKI restriction/modification enzyme. Nucleic Acids Res. 30, 3936–3944 (2002).
Walkinshaw, M. et al. Structure of Ocr from bacteriophage T7, a protein that mimics B-form DNA. Mol. Cell 9, 187–194 (2002).
Drozdz, M., Piekarowicz, A., Bujnicki, J. M. & Radlinska, M. Novel non-specific DNA adenine methyltransferases. Nucleic Acids Res. 40, 2119–2130 (2012).
Thavalingam, A. et al. Inhibition of CRISPR-Cas9 ribonucleoprotein complex assembly by anti-CRISPR AcrIIC2. Nat. Commun. 10, 2806 (2019).
Lu, W.-T., Trost, C. N., Müller-Esparza, H., Randau, L. & Davidson, A. R. Anti-CRISPR AcrIF9 functions by inducing the CRISPR–Cas complex to bind DNA non-specifically. Nucleic Acids Res. 49, 3381–3393 (2021).
Bondy-Denomy, J. et al. Multiple mechanisms for CRISPR–Cas inhibition by anti-CRISPR proteins. Nature 526, 136–139 (2015).
Stanley, S. Y. & Maxwell, K. L. Phage-encoded anti-CRISPR defenses. Annu. Rev. Genet. 52, 445–464 (2018).
Li, Y. & Bondy-Denomy, J. Anti-CRISPRs go viral: the infection biology of CRISPR-Cas inhibitors. Cell Host Microbe 29, 704–714 (2021).
Jia, N. & Patel, D. J. Structure-based functional mechanisms and biotechnology applications of anti-CRISPR proteins. Nat. Rev. Mol. Cell Biol. 22, 563–579 (2021).
Davidson, A. R. et al. Anti-CRISPRs: protein inhibitors of CRISPR-Cas systems. Annu. Rev. Biochem. 89, 309–332 (2020).
Otsuka, Y. & Yonesaki, T. Dmd of bacteriophage T4 functions as an antitoxin against Escherichia coli LsoA and RnlA toxins. Mol. Microbiol. 83, 669–681 (2012).
Blower, T. R., Evans, T. J., Przybilski, R., Fineran, P. C. & Salmond, G. P. C. Viral evasion of a bacterial suicide system by RNA–based molecular mimicry enables infectious altruism. PLoS Genet. 8, e1003023 (2012).
Doron, S. et al. Systematic discovery of antiphage defense systems in the microbial pangenome. Science 359, eaar4120 (2018).
Gao, L. et al. Diverse enzymatic activities mediate antiviral immunity in prokaryotes. Science 369, 1077–1084 (2020).
Cohen, D. et al. Cyclic GMP–AMP signalling protects bacteria against viral infection. Nature 574, 691–695 (2019).
Millman, A. et al. Bacterial retrons function in anti-phage defense. Cell 183, 1551–1561 (2020).
Bernheim, A. et al. Prokaryotic viperins produce diverse antiviral molecules. Nature 589, 120–124 (2021).
Millman, A. et al. An expanded arsenal of immune systems that protect bacteria from phages. Cell Host Microbe 30, 1556–1569 (2022).
Whiteley, A. T. et al. Bacterial cGAS-like enzymes synthesize diverse nucleotide signals. Nature 567, 194–199 (2019).
Lau, R. K. et al. Structure and mechanism of a cyclic trinucleotide-activated bacterial endonuclease mediating bacteriophage immunity. Mol. Cell 77, 723–733 (2020).
Millman, A., Melamed, S., Amitai, G. & Sorek, R. Diversity and classification of cyclic-oligonucleotide-based anti-phage signalling systems. Nat. Microbiol. 5, 1608–1615 (2020).
Ofir, G. et al. Antiviral activity of bacterial TIR domains via immune signalling molecules. Nature 600, 116–120 (2021).
Tal, N. et al. Cyclic CMP and cyclic UMP mediate bacterial immunity against phages. Cell 184, 5728–5739 (2021).
Kronheim, S. et al. A chemical defence against phage infection. Nature 564, 283–286 (2018).
Bobonis, J. et al. Bacterial retrons encode phage-defending tripartite toxin–antitoxin systems. Nature 609, 144–150 (2022).
Huiting, E. et al. Bacteriophages inhibit and evade cGAS-like immune function in bacteria. Cell 186, 864–876 (2023).
Hobbs, S. J. et al. Phage anti-CBASS and anti-Pycsar nucleases subvert bacterial immunity. Nature 605, 522–526 (2022).
Leavitt, A. et al. Viruses inhibit TIR gcADPR signalling to overcome bacterial defence. Nature 611, 326–331 (2022).
Rosenthal, R., Toye, P. A., Korman, R. Z. & Zahler, S. A. The prophage of SP beta c2dcitK1, a defective specialized transducing phage of Bacillus subtilis. Genetics 92, 721–739 (1979).
Tucker, R. G. Acquisition of thymidylate synthetase activity by a thymine-requiring mutant of Bacillus subtilis following Infection by the temperate phage φ3. J. Gen. Virol. 4, 489–504 (1969).
Noyer-Weidner, M., Jentsch, S., Pawlek, B., Günthert, U. & Trautner, T. A. Restriction and modification in Bacillus subtilis: DNA methylation potential of the related bacteriophages Z, SPR, SP beta, phi 3 T, and rho 11. J. Virol. 46, 446–453 (1983).
Stewart, C. R. et al. The genome of Bacillus subtilis bacteriophage SPO1. J. Mol. Biol. 388, 48–70 (2009).
Tesson, F. et al. Systematic and quantitative view of the antiviral arsenal of prokaryotes. Nat. Commun. 13, 2561 (2022).
Cheng, R. et al. A nucleotide-sensing endonuclease from the Gabija bacterial defense system. Nucleic Acids Res. 49, 5216–5229 (2021).
Stokar-Avihail, A. et al. Discovery of phage determinants that confer sensitivity to bacterial immune systems. Cell 186, 1863–1876 (2023).
Antine, S. P. et al. Structural basis of Gabija anti-phage defense and viral immune evasion. Nature https://doi.org/10.1038/s41586-023-06855-2 (2023).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Peters, J. M. et al. A comprehensive, CRISPR-based functional analysis of essential genes in bacteria. Cell 165, 1493–1506 (2016).
Chen, I.-M. A. et al. IMG/M v.5.0: an integrated data management and comparative analysis system for microbial genomes and microbiomes. Nucleic Acids Res. 47, D666–D677 (2019).
Nayfach, S. et al. Metagenomic compendium of 189,680 DNA viruses from the human gut microbiome. Nat. Microbiol. 6, 960–970 (2021).
Athukoralage, J. S. et al. An anti-CRISPR viral ring nuclease subverts type III CRISPR immunity. Nature 577, 572–575 (2020).
Bondy-Denomy, J., Pawluk, A., Maxwell, K. L. & Davidson, A. R. Bacteriophage genes that inactivate the CRISPR/Cas bacterial immune system. Nature 493, 429–432 (2013).
Uribe, R. V. et al. Discovery and characterization of Cas9 inhibitors disseminated across seven bacterial phyla. Cell Host Microbe 25, 233–241 (2019).
Manik, M. K. et al. Cyclic ADP ribose isomers: production, chemical structures, and immune signaling. Science 377, eadc8969 (2022).
MacLean, R. C. & San Millan, A. The evolution of antibiotic resistance. Science 365, 1082–1083 (2019).
Kortright, K. E., Chan, B. K., Koff, J. L. & Turner, P. E. Phage therapy: a renewed approach to combat antibiotic-resistant bacteria. Cell Host Microbe 25, 219–232 (2019).
Nobrega, F. L., Costa, A. R., Kluskens, L. D. & Azeredo, J. Revisiting phage therapy: new applications for old resources. Trends Microbiol. 23, 185–191 (2015).
Federici, S. et al. Targeted suppression of human IBD-associated gut microbiota commensals by phage consortia for treatment of intestinal inflammation. Cell 185, 2879–2898 (2022).
Hussain, F. A. et al. Rapid evolutionary turnover of mobile genetic elements drives bacterial resistance to phages. Science 374, 488–492 (2021).
LeGault, K. N. et al. Temporal shifts in antibiotic resistance elements govern phage-pathogen conflicts. Science 373, eabg2166 (2021).
Gilchrist, C. L. M. & Chooi, Y.-H. clinker & clustermap.js: automatic generation of gene cluster comparison figures. Bioinformatics 37, 2473–2475 (2021).
Kropinski, A. M., Mazzocco, A., Waddell, T. E., Lingohr, E. & Johnson, R. P. in Bacteriophages: Methods and Protocols Vol. 1 (eds Kropinski, A. M. & Cloike, M. R. J.) 69–76 (Humana, 2009).
Baym, M. et al. Inexpensive multiplexed library preparation for megabase-sized genomes. PLoS ONE 10, e0128036 (2011).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.journal 17, 10 (2011).
Nurk, S. et al. Assembling genomes and mini-metagenomes from highly chimeric reads. Res. Comput. Mol. Biol. 7821, 158–170 (2013).
Procedure & Checklist – Preparing Multiplexed Microbial Libraries Using SMRTbell Express Template Prep Kit 2.0 (PacBio), version 8 https://www.pacb.com/wp-content/uploads/Procedure-Checklist-–-Preparing-Multiplexed-Microbial-Libraries-Using-SMRTbell-Express-Template-Prep-Kit-2.0.pdf (2021).
Hyatt, D. et al. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinform. 11, 119 (2010).
Mazzocco, A., Waddell, T. E., Lingohr, E. & Johnson, R. P. in Bacteriophages: Methods and Protocols Vol. 1 (eds Kropinski, A. M. & Cloike, M. R. J.) 81–85 (Humana, 2009).
Steinegger, M. & Söding, J. MMseqs2 enables sensitive protein sequence searching for the analysis of massive data sets. Nat. Biotechnol. 35, 1026–1028 (2017).
Deatherage, D. E. & Barrick, J. E. in Engineering and Analyzing Multicellular Systems (eds Sun, L. & Shou, W.) 165–188 (Humana, 2014).
Adler, B. A. et al. Broad-spectrum CRISPR-Cas13a enables efficient phage genome editing. Nat. Microbiol. 7, 1967–1979 (2022).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Nguyen, L.-T., Schmidt, H. A., von Haeseler, A. & Minh, B. Q. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 32, 268–274 (2015).
Letunic, I. & Bork, P. Interactive Tree Of Life (iTOL) v5: an online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 49, W293–W296 (2021).
Zhou, W. et al. Structure of the human cGAS-DNA complex reveals enhanced control of immune surveillance. Cell 174, 300–311 (2018).
Kabsch, W. XDS. Acta Crystallogr. D 66, 125–132 (2010).
Evans, P. R. & Murshudov, G. N. How good are my data and what is the resolution? Acta Crystallogr. D 69, 1204–1214 (2013).
Mirdita, M. et al. ColabFold: making protein folding accessible to all. Nat. Methods 19, 679–682 (2022).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr. D 75, 861–877 (2019).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
Karplus, P. A. & Diederichs, K. Linking crystallographic model and data quality. Science 336, 1030–1033 (2012).
Weiss, M. S. Global indicators of X-ray data quality. J. Appl. Crystallogr. 34, 130–135 (2001).
Acknowledgements
We thank the members of the laboratory of R.S. for comments on the manuscript and discussion; Y. Peleg and S. Albeck for assistance with protein expression; Y. Fridmann-Sirkis for help with surface plasmon resonance analysis; and H. Keren-Shaul and D. Pilzer for help with PacBio sequencing. R.S. was supported, in part, by the European Research Council (grant no. ERC-AdG GA 101018520), the Israel Science Foundation (MAPATS grant 2720/22), the Deutsche Forschungsgemeinschaft (SPP 2330, Grant 464312965), the Ernest and Bonnie Beutler Research Program of Excellence in Genomic Medicine, and the Knell Family Center for Microbiology. E.Y. is supported by the Clore Scholars Program, and, in part, by the Israeli Council for Higher Education via the Weizmann Data Science Research Center. P.J.K. was supported, in part, by the Pew Biomedical Scholars programme and The Mathers Foundation. S.J.H. is supported through a Cancer Research Institute Irvington Postdoctoral Fellowship (number CRI3996). X-ray data were collected at the Northeastern Collaborative Access Team beamlines 24-ID-C and 24-ID-E (P30 GM124165), and used a Pilatus detector (S10RR029205), an Eiger detector (S10OD021527) and the Argonne National Laboratory Advanced Photon Source (DE-AC02-06CH11357), and at beamline 8.2.1 of the Advanced Light Source, a US Department of Energy Office of Science User Facility under contract number DE-AC02-05CH11231 and supported in part by the Howard Hughes Medical Institute, the ALS-ENABLE programme and the NIGMS grant P30 GM124169-01.
Author information
Authors and Affiliations
Contributions
The study was conceptualized and designed by E.Y., A. Leavitt and R.S. E.Y. built and executed the computational pipeline and analysed the data. A. Leavitt isolated the phages and conducted all of the in vivo experiments unless stated otherwise. A. Lu and P.J.K. carried out the structural analysis of Tad1 and Tad2. A.E.R. and P.J.K. determined and analysed the structure of Had1. C.A. and G.A. carried out the biochemical experiments with cell lysates and led the mechanistic characterization of the Tad2 activity. I.O. designed and conducted all of the phage knock-in experiments and the knockout of gad2 from the phage SPβL7. J.G. designed and generated the knockdown clones. DNA cleavage experiments were carried out by S.P.A., S.E.M. and P.J.K. S.J.H. helped with the structural analysis, characterization of the Tad2 activity, and analysis of Had1 oligomerization. The study was supervised by G.A. and R.S. The manuscript was written by E.Y. and R.S. All authors contributed to editing the manuscript and support the conclusions.
Corresponding authors
Ethics declarations
Competing interests
R.S. is a scientific co-founder of and adviser for BiomX and Ecophage. The other authors declare no competing interests.
Peer review
Peer review information
Nature thanks David Taylor and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Genome comparisons of phages from the SPbeta and SBSphiJ groups.
Genome comparison of (a) eleven phages from the SPbeta group and (b) eight phages from the SBSphiJ group. Amino acid sequence similarity is marked by grey shading. Genome similarity was visualized using clinker56.
Extended Data Fig. 2 Phages from the same family are differentially sensitive to bacterial defense systems.
Results of phage infection experiments with (a) eleven phages of the SPbeta group, (b) six phages of the SPO1 group, and (c) eight phages of the SBSphiJ group. Data represent plaque-forming units per ml (PFU/ml) of phages infecting control cells (“no system”), and cells expressing the respective defense systems. Shown is the average of three technical replicates, with individual data points overlaid. The Thoeris and Hachiman data presented here are the same as those presented in Figs. 3b and 5b, respectively.
Extended Data Fig. 3 Gad1 proteins inhibit Gabija mediated defense.
(a) Multiple sequence alignment of the original Gad1 from phage phi3T and five Gad1 homologs that were chosen for experimental verification. Conserved residues are in purple. (b) Results of phage infection experiments with eleven phages of the SPbeta group. Data represent plaque-forming units per ml (PFU/ml) of phages infecting control cells (“no system”), cells expressing the Gabija system (“Gabija”), and cells co-expressing the Gabija system and a Gad1 homolog. Shown is the average of three technical replicates, with individual data points overlaid. The SPbeta data presented here are the same as those presented in Fig. 2d.
Extended Data Fig. 4 Gad2 inhibits Gabija mediated defense.
(a) Phylogeny and distribution of Gad2 homologs. Homologs that were tested experimentally are indicated on the tree by cyan diamonds. (b) An Alphafold2 model for the structure of Gad2 from phage SPbetaL7. (c) Mutations in the predicted nucleotidyltransferase active site in Gad2 result in loss of anti-defense activity. Data represent plaque-forming units per ml (PFU/ml) of phage SPbeta infecting cells co-expressing the Gabija system and WT or mutated Gad2 from Brevibacillus laterosporus, as well control cells expressing neither Gabija nor Gad2 (“Control”) and cells expressing the Gabija system without Gad2 (“No Gad2”). Shown is the average of three technical replicates, with individual data points overlaid. (d) SDS-PAGE analysis of Ni-NTA co-purified GajAB with Shewanella phage 1/4 Gad1 and Brevibacillus laterosporus Gad2 demonstrates that Gad2 does not stably interact with the GajAB complex. Asterisk indicates minor contamination with the E. coli protein ArnA. Data are representative of three independent experiments. For gel source data, see Supplementary Fig. 1. (e) SDS-PAGE analysis of purified Brevibacillus laterosporus Gad2. Asterisk indicates contamination with the E. coli protein ArnA. Data are representative of three independent experiments. For gel source data, see Supplementary Fig. 1. (f) Biochemical reconstitution of GajAB DNA degradation demonstrates that Gad2 does not directly inhibit GajAB cleavage of a 56-bp target DNA. Data are representative of three independent experiments. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 5 Tad2 proteins inhibit Thoeris mediated defense.
(a) Multiple sequence alignment of the original Tad2 from phage SPO1, and 5 Tad2 homologs that were chosen for experimental verification. Conserved residues are in purple. Black arrows indicate residues that are involved in 1″–3′ gcADPR binding. (b) Results of phage infection experiments with six phages of the SPO1 group. Data represent plaque-forming units per ml (PFU/ml) of phages infecting control cells (“no system”), cells expressing the Thoeris system (“Thoeris”), and cells co-expressing the Thoeris system and a Tad2 homolog. Shown is the average of three technical replicates, with individual data points overlaid.
Extended Data Fig. 6 Tad2 binds 1″–3′ gcADPR.
(a) Incubation of Tad2 with 1″–3′ gcADPR in vitro does not yield observable degradation products. Representative HPLC traces of 1″–3′ gcADPR incubated with buffer, Tad2, or with the enzyme Cap-Clip known to cleave diphosphate linkages as a positive control. (b) Size-exclusion chromatography of 1″–3′ gcADPR-bound or apo state Tad2. 1″–3′ gcADPR-bound Tad2 shows a substantial shift compared to Tad2 in the apo state. (c) Surface plasmon resonance binding sensorgrams for Tad2 at five concentrations of 1″–3′ gcADPR. The black lines are the global fits using the instrument’s evaluation software. ka = 3.42E + 05 ± 5.2E + 02 (1/Ms), kd = 0.00798 ± 1E-05 (1/s). (d) Surface plasmon resonance binding sensorgrams for Tad2 at multiple concentrations of cADPR.
Extended Data Fig. 7 Size-exclusion chromatography of Tad2 and various standards.
Observed peak demonstrates that Tad2 forms a homomultimer.
Extended Data Fig. 8 Comparison of Tad2 and Tad1 in the apo and ligand-bound states.
(a) Overview of the crystal structure of SPO1 Tad2 in the apo state in front and top view. (b,c) Overview and detailed binding pocket views of adenine interactions (left) and ribose/phosphate interactions (right) of the crystal structures of SPO1 Tad2 in complex with 1″–3′ gcADPR (b) or 1″–2′ gcADPR (c). (d) Overview of the crystal structure (PDB: 7UAV) of cbTad1 in the apo state in front view and top view. (e,f) Overview and detailed binding pocket views of adenine interactions (left) and ribose/phosphate interactions (right) of the crystal structures of cbTad1 in complex with 1″–3′ gcADPR (e) or 1″–2′ gcADPR (f, PDB: 7UAW).
Extended Data Fig. 9 Had1 proteins inhibit Hachiman-mediated defense.
(a) Results of phage infection experiments with eight phages of the SBSphiJ group. Data represent plaque-forming units per ml (PFU/ml) of phages infecting control cells (“no system”), cells expressing the Hachiman system (“Hachiman”), and cells co-expressing the Hachiman system and a Had1 homolog. Shown is the average of three technical replicates, with individual data points overlaid. (b) Structure-guided sequence alignment of Had1 homologs colored by BLOSUM62 score. (c) SDS-PAGE and (d) SEC-MALS analysis of purified Had1. Full-length Had1 elutes as a single species that is consistent with a homodimeric complex (predicted homodimer 12.5 kDa, observed 12.6 kDa). Data are representative of three independent experiments. For gel source data, see Supplementary Fig. 1.
Supplementary information
Supplementary Fig. 1
Uncropped gel images.
Supplementary Tables 1
Supplementary Tables 1–6.
Supplementary Tables 2
Supplementary Tables 7–14.
Supplementary Table 15
Supplementary Table 15.
Source data
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yirmiya, E., Leavitt, A., Lu, A. et al. Phages overcome bacterial immunity via diverse anti-defence proteins. Nature 625, 352–359 (2024). https://doi.org/10.1038/s41586-023-06869-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-023-06869-w
This article is cited by
-
A never-ending defence fight
Nature Reviews Microbiology (2024)
-
Structural basis of Gabija anti-phage defence and viral immune evasion
Nature (2024)
-
Inhibitors of bacterial immune systems: discovery, mechanisms and applications
Nature Reviews Genetics (2024)
-
An OLD protein teaches us new tricks: prokaryotic antiviral defense
Nature Communications (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.