Abstract
Type A γ-aminobutyric acid receptors (GABAARs) are pentameric ligand-gated chloride channels that mediate fast inhibitory signalling in neural circuits1,2 and can be modulated by essential medicines including general anaesthetics and benzodiazepines3. Human GABAAR subunits are encoded by 19 paralogous genes that can, in theory, give rise to 495,235 receptor types. However, the principles that govern the formation of pentamers, the permutational landscape of receptors that may emerge from a subunit set and the effect that this has on GABAergic signalling remain largely unknown. Here we use cryogenic electron microscopy to determine the structures of extrasynaptic GABAARs assembled from α4, β3 and δ subunits, and their counterparts incorporating γ2 instead of δ subunits. In each case, we identified two receptor subtypes with distinct stoichiometries and arrangements, all four differing from those previously observed for synaptic, α1-containing receptors4,5,6,7. This, in turn, affects receptor responses to physiological and synthetic modulators by creating or eliminating ligand-binding sites at subunit interfaces. We provide structural and functional evidence that selected GABAAR arrangements can act as coincidence detectors, simultaneously responding to two neurotransmitters: GABA and histamine. Using assembly simulations and single-cell RNA sequencing data8,9, we calculated the upper bounds for receptor diversity in recombinant systems and in vivo. We propose that differential assembly is a pervasive mechanism for regulating the physiology and pharmacology of GABAARs.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Atomic coordinates for the GABAAR models reported here have been deposited to the PDB with accession codes 7QN5-9 and 7QNA-E, and the cryo-EM density maps have been deposited in the Electron Microscopy Data Bank with accession codes EMD-14067–EMD-14076. Raw movies of all datasets have been deposited in the Electron Microscopy Public Image Archive with accession codes EMPIAR-10909–EMPIAR-10914. Coordinate files used during model building can be found in the PDB under the accession codes 7A5V and 6HUG. scRNA-seq data used in the study are available from Allen Brain Map: https://portal.brain-map.org/atlases-and-data/rnaseq/human-multiple-cortical-areas-smart-seq.Source data are provided with this paper.
Code availability
The scripts for scRNA-seq analysis and for running and analysing the receptor assembly simulations are available at www.github.com/andrijasente/GABA-differential-assembly.
References
Sigel, E. & Steinmann, M. E. Structure, function, and modulation of GABAA receptors. J. Biol. Chem. 287, 40224–40231 (2012).
Sieghart, W. & Sperk, G. Subunit composition, distribution and function of GABAA receptor subtypes. Curr. Top. Med. Chem. 2, 795–816 (2002).
Sieghart, W. & Savic, M. M. International Union of Basic and Clinical Pharmacology. CVI: GABAA receptor subtype- and function-selective ligands: key issues in translation to humans. Pharmacol. Rev. 70, 836–878 (2018).
Zhu, S. et al. Structure of a human synaptic GABAA receptor. Nature 559, 67–72 (2018).
Phulera, S. et al. Cryo-EM structure of the benzodiazepine-sensitive α1β1γ2S tri-heteromeric GABAA receptor in complex with GABA. eLife 7, e39383 (2018).
Laverty, D. et al. Cryo-EM structure of the human α1β3γ2 GABAA receptor in a lipid bilayer. Nature 565, 516–520 (2019).
Masiulis, S. et al. GABAA receptor signalling mechanisms revealed by structural pharmacology. Nature 565, 454–459 (2019).
Tasic, B. et al. Shared and distinct transcriptomic cell types across neocortical areas. Nature 563, 72–78 (2018).
Hodge, R. D. et al. Conserved cell types with divergent features in human versus mouse cortex. Nature 573, 61–68 (2019).
Wisden, W., Laurie, D. J., Monyer, H. & Seeburg, P. H. The distribution of 13 GABAA receptor subunit mRNAs in the rat brain. I. Telencephalon, diencephalon, mesencephalon. J. Neurosci. 12, 1040–1062 (1992).
Nusser, Z., Sieghart, W. & Somogyi, P. Segregation of different GABAA receptors to synaptic and extrasynaptic membranes of cerebellar granule cells. J. Neurosci. 18, 1693–1703 (1998).
Pearce, R. A. Physiological evidence for two distinct GABAA responses in rat hippocampus. Neuron 10, 189–200 (1993).
Baude, A., Sequier, J. M., McKernan, R. M., Olivier, K. R. & Somogyi, P. Differential subcellular distribution of the alpha 6 subunit versus the alpha 1 and beta 2/3 subunits of the GABAA/benzodiazepine receptor complex in granule cells of the cerebellar cortex. Neuroscience 51, 739–748 (1992).
Farrant, M. & Nusser, Z. Variations on an inhibitory theme: phasic and tonic activation of GABAA receptors. Nat. Rev. Neurosci. 6, 215–229 (2005).
Brickley, S. G. & Mody, I. Extrasynaptic GABAA receptors: their function in the CNS and implications for disease. Neuron 73, 23–34 (2012).
Scholze, P. et al. Two distinct populations of α1α6-containing GABAA-receptors in rat cerebellum. Front. Synaptic Neurosci. 12, 591129 (2020).
Barrera, N. P. et al. Atomic force microscopy reveals the stoichiometry and subunit arrangement of the α4β3δ GABAA receptor. Mol. Pharmacol. 73, 960–967 (2008).
Patel, B., Mortensen, M. & Smart, T. G. Stoichiometry of delta subunit containing GABAA receptors. Br. J. Pharmacol. 171, 985–994 (2014).
Wagoner, K. R. & Czajkowski, C. Stoichiometry of expressed α4β2δ γ-aminobutyric acid type A receptors depends on the ratio of subunit cDNA transfected. J. Biol. Chem. 285, 14187–14194 (2010).
Botzolakis, E. J. et al. Comparison of gamma-aminobutyric acid, type A (GABAA), receptor αβγ and αβδ expression using flow cytometry and electrophysiology: evidence for alternative subunit stoichiometries and arrangements. J. Biol. Chem. 291, 20440–20461 (2016).
Zhou, X. et al. High-level production and purification in a functional state of an extrasynaptic gamma-aminobutyric acid type A receptor containing α4β3δ subunits. PLoS ONE 13, e0191583 (2018).
Eaton, M. M. et al. γ-Aminobutyric acid type A α4, β2, and δ subunits assemble to produce more than one functionally distinct receptor type. Mol. Pharmacol. 86, 647–656 (2014).
Nakane, T. et al. Single-particle cryo-EM at atomic resolution. Nature 587, 152–156 (2020).
Saras, A. et al. Histamine action on vertebrate GABAA receptors: direct channel gating and potentiation of GABA responses. J. Biol. Chem. 283, 10470–10475 (2008).
Wongsamitkul, N. et al. α Subunits in GABAA receptors are dispensable for GABA and diazepam action. Sci. Rep. 7, 15498 (2017).
Taylor, P. M. et al. Identification of amino acid residues within GABAA receptor beta subunits that mediate both homomeric and heteromeric receptor expression. J. Neurosci. 19, 6360–6371 (1999).
Bianchi, M. T., Clark, A. G. & Fisher, J. L. The wake-promoting transmitter histamine preferentially enhances alpha-4 subunit-containing GABAA receptors. Neuropharmacology 61, 747–752 (2011).
Lee, H. J. et al. A pharmacological characterization of GABA, THIP and DS2 at binary α4β3 and β3δ receptors: GABA activates β3δ receptors via the β3+δ− interface. Brain Res. 1644, 222–230 (2016).
Miller, P. S. & Aricescu, A. R. Crystal structure of a human GABAA receptor. Nature 512, 270–275 (2014).
Hoerbelt, P. et al. Mutagenesis and computational docking studies support the existence of a histamine binding site at the extracellular β3+ β3− interface of homooligomeric β3 GABAA receptors. Neuropharmacology 108, 252–263 (2016).
Falch, E. & Krogsgaard-Larsen, P. The binding of the GABA agonist [3H]THIP to rat brain synaptic membranes. J. Neurochem. 38, 1123–1129 (1982).
Wafford, K. A. & Ebert, B. Gaboxadol—a new awakening in sleep. Curr. Opin. Pharmacol. 6, 30–36 (2006).
Hanchar, H. J. et al. Ethanol potently and competitively inhibits binding of the alcohol antagonist Ro15-4513 to α4/6β3δ GABAA receptors. Proc. Natl Acad. Sci. USA 103, 8546–8551 (2006).
Wallner, M., Hanchar, H. J. & Olsen, R. W. Low-dose alcohol actions on α4β3δ GABAA receptors are reversed by the behavioral alcohol antagonist Ro15-4513. Proc. Natl Acad. Sci. USA 103, 8540–8545 (2006).
Olsen, R. W., Hanchar, H. J., Meera, P. & Wallner, M. GABAA receptor subtypes: the “one glass of wine” receptors. Alcohol 41, 201–209 (2007).
Linden, A. M. et al. Ro 15-4513 antagonizes alcohol-induced sedation in mice through αβγ2-type GABAA receptors. Front. Neurosci. 5, 3 (2011).
Storustovu, S. I. & Ebert, B. Pharmacological characterization of agonists at delta-containing GABAA receptors: functional selectivity for extrasynaptic receptors is dependent on the absence of γ2. J. Pharmacol. Exp. Ther. 316, 1351–1359 (2006).
Brown, N., Kerby, J., Bonnert, T. P., Whiting, P. J. & Wafford, K. A. Pharmacological characterization of a novel cell line expressing human α4β3δ GABAA receptors. Br. J. Pharmacol. 136, 965–974 (2002).
Marti-Solano, M. et al. Combinatorial expression of GPCR isoforms affects signalling and drug responses. Nature 587, 650–656 (2020).
Rudolph, U. & Knoflach, F. Beyond classical benzodiazepines: novel therapeutic potential of GABAA receptor subtypes. Nat. Rev. Drug Discov. 10, 685–697 (2011).
Bencsits, E., Ebert, V., Tretter, V. & Sieghart, W. A significant part of native γ-aminobutyric acidA receptors containing α4 subunits do not contain γ or δ subunits. J. Biol. Chem. 274, 19613–19616 (1999).
Levitan, E. S. et al. Structural and functional basis for GABAA receptor heterogeneity. Nature 335, 76–79 (1988).
Brickley, S. G., Cull-Candy, S. G. & Farrant, M. Single-channel properties of synaptic and extrasynaptic GABAA receptors suggest differential targeting of receptor subtypes. J. Neurosci. 19, 2960–2973 (1999).
Mertens, S., Benke, D. & Mohler, H. GABAA receptor populations with novel subunit combinations and drug binding profiles identified in brain by α5- and δ-subunit-specific immunopurification. J. Biol. Chem. 268, 5965–5973 (1993).
Fritschy, J. M. et al. Five subtypes of type A gamma-aminobutyric acid receptors identified in neurons by double and triple immunofluorescence staining with subunit-specific antibodies. Proc. Natl Acad. Sci. USA 89, 6726–6730 (1992).
Duggan, M. J., Pollard, S. & Stephenson, F. A. Immunoaffinity purification of GABAA receptor α-subunit iso-oligomers. Demonstration of receptor populations containing α1 α2, α1 α3, and α2 α3 subunit pairs. J. Biol. Chem. 266, 24778–24784 (1991).
Owens, D. F. & Kriegstein, A. R. Is there more to GABA than synaptic inhibition? Nat. Rev. Neurosci. 3, 715–727 (2002).
Datta, D., Arion, D. & Lewis, D. A. Developmental expression patterns of GABAA receptor subunits in layer 3 and 5 pyramidal cells of monkey prefrontal cortex. Cereb. Cortex 25, 2295–2305 (2015).
Fatemi, S. H., Folsom, T. D., Rooney, R. J. & Thuras, P. D. Expression of GABAA α2-, β1- and ε-receptors are altered significantly in the lateral cerebellum of subjects with schizophrenia, major depression and bipolar disorder. Transl. Psychiatry 3, e303 (2013).
Mueller, T. M., Remedies, C. E., Haroutunian, V. & Meador-Woodruff, J. H. Abnormal subcellular localization of GABAA receptor subunits in schizophrenia brain. Transl. Psychiatry 5, e612 (2015).
Murshudov, G. N. et al. REFMAC5 for the refinement of macromolecular crystal structures. Acta Crystallogr. D Biol. Crystallogr. 67, 355–367 (2011).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
Afonine, P. V. et al. Real-space refinement in PHENIX for cryo-EM and crystallography. Acta Crystallogr. D Struct. Biol. 74, 531–544 (2018).
Reeves, P. J., Kim, J. M. & Khorana, H. G. Structure and function in rhodopsin: a tetracycline-inducible system in stable mammalian cell lines for high-level expression of opsin mutants. Proc. Natl Acad. Sci. USA 99, 13413–13418 (2002).
Elegheert, J. et al. Lentiviral transduction of mammalian cells for fast, scalable and high-level production of soluble and membrane proteins. Nat. Protoc. 13, 2991–3017 (2018).
Dostalova, Z. et al. Human α1β3γ2L γ-aminobutyric acid type A receptors: high-level production and purification in a functional state. Protein Sci. 23, 157–166 (2014).
Miller, P. S. et al. Structural basis for GABAA receptor potentiation by neurosteroids. Nat. Struct. Mol. Biol. 24, 986–992 (2017).
Uchanski, T. et al. Megabodies expand the nanobody toolkit for protein structure determination by single-particle cryo-EM. Nat. Methods 18, 60–68 (2021).
Russo, C. J. & Passmore, L. A. Electron microscopy: ultrastable gold substrates for electron cryomicroscopy. Science 346, 1377–1380 (2014).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Tegunov, D. & Cramer, P. Real-time cryo-electron microscopy data preprocessing with Warp. Nat. Methods 16, 1146–1152 (2019).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Asarnow, D., Palovcak, E. & Cheng, Y. asarnow/pyem: UCSF pyem v0.5 (Zenodo, 2019).
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Zivanov, J., Nakane, T. & Scheres, S. H. W. Estimation of high-order aberrations and anisotropic magnification from cryo-EM data sets in RELION-3.1. IUCrJ 7, 253–267 (2020).
Zivanov, J., Nakane, T. & Scheres, S. H. W. A Bayesian approach to beam-induced motion correction in cryo-EM single-particle analysis. IUCrJ 6, 5–17 (2019).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Ramlaul, K., Palmer, C. M., Nakane, T. & Aylett, C. H. S. Mitigating local over-fitting during single particle reconstruction with SIDESPLITTER. J. Struct. Biol. 211, 107545 (2020).
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014).
Naydenova, K. & Russo, C. J. Measuring the effects of particle orientation to improve the efficiency of electron cryomicroscopy. Nat. Commun. 8, 629 (2017).
Goddard, T. D. et al. UCSF ChimeraX: meeting modern challenges in visualization and analysis. Protein Sci. 27, 14–25 (2018).
Waterhouse, A. et al. SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic Acids Res. 46, W296–W303 (2018).
Weininger, D. SMILES, a chemical language and information system. 1. Introduction to methodology and encoding rules. J. Chem. Inf. Comput. Sci. 28, 31–36 (1988).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Nicholls, R. A., Fischer, M., McNicholas, S. & Murshudov, G. N. Conformation-independent structural comparison of macromolecules with ProSMART. Acta Crystallogr. D Biol. Crystallogr. 70, 2487–2499 (2014).
Forman, S. A. A hydrophobic photolabel inhibits nicotinic acetylcholine receptors via open-channel block following a slow step. Biochemistry 38, 14559–14564 (1999).
Sente, A. & Naydenova, K. Scripts for differential assembly diversifies GABAA receptor structures and signaling (v1.0.1). Zenodo https://doi.org/10.5281/zenodo.5884871 (2022).
Acknowledgements
We thank M. Madan Babu, D. Barford, T. Blundell, R. Calinescu, T. Flock, A. B. Kleist, A. Murzin, D. Malinverni, C. J. Russo, N. Unwin and the members of Aricescu laboratory for helpful discussions and comments on this work; V. B. Kasaragod for help with model building; T. Nakane and K. Yamashita for help with data processing; J. Grimmett and T. Darling for support with scientific computing; S. Chen, G. Cannone, G. Sharov, A. Yeates and B. Ahsan for electron microscopy support; T. Uchanski, E. Pardon and J. Steyaert for the Mb192 megabody. Cryo-EM datasets were collected at the MRC-LMB and Cambridge University Department of Biochemistry EM (BiocEM) facilities. We acknowledge funding from the UK Medical Research Council (MR/L009609/1 and MC_UP_1201/15 to A.R.A.), National Institute for General Medical Sciences (1R01-GM135550 to K.W.M. and A.R.A.), the Department of Anesthesia, Critical Care & Pain Medicine at Massachusetts General Hospital (to K.W.M.), MRC-LMB and Cambridge Trust (LMB Cambridge Scholarship to A.S. and K.N.), School of Clinical Medicine, University of Cambridge (MB/PhD fellowship to A.S.), and Boehringer Ingelheim Fonds (PhD Fellowship to J.M.). The cryo-EM facility at the Department of Biochemistry is funded by the Wellcome Trust (206171/Z/17/Z and 202905/Z/16/Z) and the University of Cambridge.
Author information
Authors and Affiliations
Contributions
A.S., K.W.M. and A.R.A. conceived the project. A.S., Y.J. and R.D. generated and characterized cell lines. A.S. and S.M. purified proteins and prepared the cryo-EM samples. A.S., S.W.H. and D.Y.C. collected the cryo-EM data. A.S., K.N. and S.M. processed the cryo-EM data. A.S., T.M., J.M. and A.R.A. built and refined the atomic models. R.D. and K.W.M. designed, performed and analysed the electrophysiology experiments. X.Z. and K.W.M. designed, performed and analysed the radioligand binding assays. A.S. designed and performed the bioinformatics analyses. A.S. and K.N. designed, performed and analysed the simulations for receptor assembly. A.S., K.N., R.D., K.W.M. and A.R.A. wrote the manuscript, with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Margot Ernst and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Quality of cryo-EM maps and models.
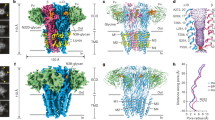
Each panel contains masked, unmasked, phase-randomised and map vs model Fourier shell correlation (FSC) curves (top), particle orientation distribution (middle) and local resolution-coloured maps for each structure. a, α4β3δ "apo" (+HEPES). b, β3δ "apo" (+HEPES). c, α4β3δ + GABA + histamine. d, β3δ + histamine. e, α4β3δ + histamine. f, α4β3γ2 + GABA. g, β3γ2 + GABA. h, α4β3δ + THIP + histamine. i, β3δ + THIP + histamine. j, α1β3γ2 + Ro15-4513. The probability density function for a certain orientation ranges between 0 (blue, low) and 0.0001 (red, high).
Extended Data Fig. 2 Survey of ligands bound in agonist pockets in different datasets.
a, α4β3δ and β3δ GABAARs solved in the absence of any known modulators. In both subtypes, HEPES is bound in the β3+/β3− agonist pocket under loop C. b, α4β3δ and β3δ GABAARs solved in the presence of 0.2 mM GABA and 1 mM histamine. c, α4β3γ2 and β3γ2 GABAARs solved in the presence of 0.2 mM GABA. d, α4β3δ and β3δ GABAARs solved in the presence of 1 mM THIP and 1 mM histamine. e, Summary table illustrating ligand vs interface compatibility. Empty spaces are left in cases where insufficient data is available to make a conclusion. For the GABA binding to β3+/γ2− pocket, a question mark indicates variable occupancy. f, Comparison of pore profiles of α4β3δ + GABA + histamine, α4β3δ + histamine, α1β3γ2 + alprazolam (6HUO) and α1β3γ2 + bicuculline (6HUK).
Extended Data Fig. 3 Ligand coordination in the “agonist pockets” under the loops C at various inter-subunit interfaces.
All pockets in which a ligand was observed in our structures are shown with a single representative example. Potential hydrogen bonds are displayed as dashed yellow lines. Histamine and GABA in the β3+/α4− pocket are taken from the α4β3δ + GABA + histamine structure. THIP in the β3+/α4− and δ+/β3− pocket is taken from the α4β3δ + THIP + histamine structure. Histamine in the β3+/β3− pocket is taken from the α4β3δ + histamine structure. HEPES in the β3+/β3− pocket is taken from the "apo" α4β3δ structure. GABA in the β3+/γ2− pocket is taken from the α4β3γ2 + GABA structure. Chain IDs represented in the panels are given as capital letters next to the subunit type.
Extended Data Fig. 4 Characterizing GABA and histamine effects on GABAA receptors composed of α4, β3 and δ subunits.
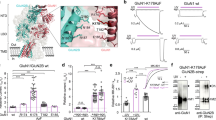
a–c, Unlike α4β3δ receptors (a), β3δ (b) and β3 (c) receptors are not gated by 200 μM GABA. Representative current traces from three successive pulses separated by 1 min: (1) 4 s pulse of 10 mM GABA plus 30 μM Etomidate (total current, purple); (2) 8 s pulse of 200 μM GABA (green), and (3) repeat of pulse (1) (pink). These pulses are applied to: a, cells co-expressing α4, β3 and δ subunits; b, cells co-expressing β3 and δ subunits; c, cells expressing the β3 subunit alone. d, e, Representative currents normalized to 10 mM GABA currents in the same cell were obtained with varying concentration of (a) GABA (0.001–10 μM, n = 38 cells with 3-8 cells for each concentration) or (b) histamine (0.03–10 mM, n = 31 cells with 3-8 cells for each concentration). In panel e, orange dashed line denotes the baseline. f, Hill plot of peak amplitude obtained with the normalized currents from (a) and (b) plotted against agonist concentration. The EC50 for GABA was 69.3 nM (95% CI 40.5–118.3 nM) and for histamine was 821.2 μM (95% CI 340.5–1981.0 μM) and Hill coefficient of 0.9 (95% CI 0.5–1.3) and 1.1 (95% CI 0.3–2.0) respectively. g, Scatter plot of peak amplitudes of currents shown in Fig. 2d by GABA (10 mM, n = 7 cells) and histamine (10 mM, n = 6 cells) alone or co-applied (n = 8 cells) in the first pulse as a percent of currents elicited by 10 mM GABA + 30 μM Etomidate. One-way ANOVA P < 0.0001. h–k, Scatter plots of the time constants (τ) and amplitudes for the slow and fast phases of deactivation that were obtained by fitting a double exponential to the falling phase of currents such as those in Fig. 2e (n = 4 cells for each experiments). In every paired experiment, the direction of change was consistent. In the presence of histamine fast and slow time constants decreased significantly (P = 0.03, two-tailed paired t test) and the fraction of receptors with slow deactivation increased (P = 0.04, two-tailed paired t test). Significance levels compared to controls were determined by ordinary one–way ANOVA or by the Paired Student’s t test and are denoted as **** P < 0.0001 and * P < 0.05. Data are presented as mean ± SD.
Extended Data Fig. 5 GABA and histamine crosstalk happens at the level of individual α4β3δ receptors.
To address the complexity arising from the presence of multiple receptor subtypes, we investigated the modulation of 100 nM GABA currents by 300 μM histamine in cells expressing different combinations of α4, β3 and δ subunits. We first established that 100 nM GABA robustly activates currents in α4β3δ cells (n = 6 cells) (a), barely in α4β3 (n = 5 cells) (b) and not in β3δ (n = 6 cells) (c) or β3 cells (n = 4 cells) (d). Histamine at 300 μM robustly enhances the GABA current in α4β3δ cells but only modestly in the α4β3, β3δ and β3 cells. The modest enhancement is attributable to histamine currents alone (e.g., from the β3δ or β3-homomeric subtypes). Representative current traces from three successive pulses separated by 1 min: (1) 8 s pulse of 100 nM GABA (dark green); (2) 8 s pulse of 100 nM GABA with a 4 s notch of 300 μM histamine added after two seconds (blue), and (3) 8 s pulse of 100 nM GABA (light green). Finally, to open all receptor isoforms, a normalization pulse of (10 mM GABA + 30 μM etomidate) was applied. e, Histamine enhances GABA current strongly only in α4β3δ receptors (P < 0.0001, one-way ANOVA). Scatter plot shows peak current amplitudes of the blue traces in panels a–d, normalized to the total cell current (10 mM GABA + 30 μM etomidate). Data are presented as mean ± SD. Sample size is the same as in panels a–d.
Extended Data Fig. 6 Ro15-4513 cannot bind to the α4+/δ− interface.
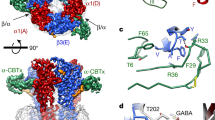
a, b, Two views of Ro15-4513 bound in the α1+/γ2− benzodiazepine pocket of an α1β3γ2 receptor. c, d, Equivalent views of an α4β3δ receptor show that clashes with the side chains of R135 (α4+), E71 and H92 (δ−) prevent Ro15-4513 (and, likely, any benzodiazepine ligand) from binding to the α4+/δ− interface in the same mode as observed in α1β3γ2.
Extended Data Fig. 7 Single-cell expression patterns of GABAAR subunits in the human cortex.
a, Grouping cortical cell types by their GABAAR subunit expression. The majority of cell types (31) expresses a combination of 11 different GABAAR subunits, including α1-5, β1-3, δ and γ2-3. Cell type "Exc L5-6 THEMIS THTPA" expresses the highest number of subunits (14): α1-5, β1-3, δ, ε, γ1-3 and ρ1. Expression in this panel is binarized such that any subunit with a trimmed mean of summed intron and exon counts larger than 0 count per million (CPM) is considered as expressed (Methods). b, Heat map showing trimmed means of CPMs for every GABAAR subunit across individual cell types in the human cortex. Cell types are clustered by their gene expression profiles.
Extended Data Fig. 8 Cryo-EM data processing strategy.
a, A typical cryo-EM data processing workflow. The numbers in this case are derived from the α4β3δ "apo" (+HEPES) dataset. b, Soft mask used to separate receptors with 2 or 3 Nb25 bound, which largely (but not completely) separates α4β3δ and β3δ receptors. The separation is incomplete in all cases because Nb25 may also bind weakly to the β3+/α4− interface. c, To further separate α4β3δ and β3δ receptors, a soft mask around the putative position of the vestibule glycan on the α4 subunit and the putative position of the N149 glycan on the β3 subunit is used. d, Soft mask around the α4 vestibule glycan used to separate α4β3γ2 from β3γ2 receptor subtypes.
Supplementary information
Supplementary Information
This file contains Supplementary Figures; Supplementary Tables; Supplementary Methods; Supplementary Discussion and Supplementary References
Rights and permissions
About this article
Cite this article
Sente, A., Desai, R., Naydenova, K. et al. Differential assembly diversifies GABAA receptor structures and signalling. Nature 604, 190–194 (2022). https://doi.org/10.1038/s41586-022-04517-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-022-04517-3
This article is cited by
-
Pathology of pain and its implications for therapeutic interventions
Signal Transduction and Targeted Therapy (2024)
-
De novo variants in GABRA4 are associated with a neurological phenotype including developmental delay, behavioral abnormalities and epilepsy
European Journal of Human Genetics (2024)
-
Inhibition of acyl-CoA binding protein (ACBP) by means of a GABAARγ2-derived peptide
Cell Death & Disease (2024)
-
Atypical antipsychotics antagonize GABAA receptors in the ventral tegmental area GABA neurons to relieve psychotic behaviors
Molecular Psychiatry (2023)
-
A pentameric TRPV3 channel with a dilated pore
Nature (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.