Abstract
Group 2 innate lymphoid cells (ILC2s) regulate inflammation and immunity in mammalian tissues1,2. Although ILC2s are found in cancers of these tissues3, their roles in cancer immunity and immunotherapy are unclear. Here we show that ILC2s infiltrate pancreatic ductal adenocarcinomas (PDACs) to activate tissue-specific tumour immunity. Interleukin-33 (IL33) activates tumour ILC2s (TILC2s) and CD8+ T cells in orthotopic pancreatic tumours but not heterotopic skin tumours in mice to restrict pancreas-specific tumour growth. Resting and activated TILC2s express the inhibitory checkpoint receptor PD-1. Antibody-mediated PD-1 blockade relieves ILC2 cell-intrinsic PD-1 inhibition to expand TILC2s, augment anti-tumour immunity, and enhance tumour control, identifying activated TILC2s as targets of anti-PD-1 immunotherapy. Finally, both PD-1+ TILC2s and PD-1+ T cells are present in most human PDACs. Our results identify ILC2s as anti-cancer immune cells for PDAC immunotherapy. More broadly, ILC2s emerge as tissue-specific enhancers of cancer immunity that amplify the efficacy of anti-PD-1 immunotherapy. As ILC2s and T cells co-exist in human cancers and share stimulatory and inhibitory pathways, immunotherapeutic strategies to collectively target anti-cancer ILC2s and T cells may be broadly applicable.
Similar content being viewed by others
Main
ILC2s are innate antigen-independent lymphocytes that regulate immunity to pathogens and commensals in tissues. Although ILC2s have been detected in cancers, their role in tumour immunity is unclear.
TILC2s infiltrate pancreatic cancers
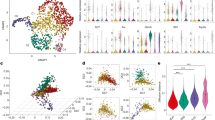
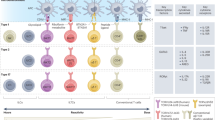
To investigate the role of ILC2s in cancer, we analysed tumour-infiltrating lymphocytes in unselected primary human PDACs. We found intratumoral cells that lacked immune cell lineage markers (lineage−) but expressed markers of ILCs (CD25 and CD127)2 and ILC2s (IL33 receptor (ST2, also known as IL1RL1 or IL33R) and GATA3) (Fig. 1a, Extended Data Fig. 1a, Supplementary Table 1). These putative TILC2s were enriched in ‘hot’ tumours (enriched in CD8+ T cells) from rare long-term PDAC survivors4 when compared with ‘cold’ tumours from short-term survivors. In addition, higher TILC2 frequencies correlated with longer survival (Fig. 1b, Extended Data Fig. 1b). Higher bulk RNA expression of the ILC2-activating cytokine IL33 in tumours, but not of any other ILC-activating cytokine, was associated with longer survival (Fig. 1c, Extended Data Fig. 1c, Supplementary Table 2). Furthermore, expression of IL33, but not of other ILC-activating cytokines, correlated with higher intratumoral immune cytolytic activity (Fig. 1c, Extended Data Fig. 1c). Although these data assess RNA and not protein expression, they suggest that IL33 and TILC2s activate anti-tumour immunity in human PDAC.
a, Gating, frequency, and phenotype of ILCs from unselected patients with PDAC. Grey curves, isotype controls; numbers, mean fluorescence intensity. b, Frequency (top) and survival association (bottom) of ILC2s in microarrays of tumour samples from short- and long-term PDAC survivors. c, Bulk tumour IL33 mRNA association with survival and tumour cytolytic index (CYT) in short- and long-term PDAC survivors. d, Gating and frequency of ILCs in mice with PDAC. e, Intratumoral ILC frequency and number in Rag2−/− PDAC mice treated with anti-CD90.2 or isotype (Iso) antibodies. f, Gating, frequency, and number of ILCs in Il33+/+ and Il33−/− PDAC mice. High and low in b, c defined as higher or lower, respectively, than the median for the cohort. HR, hazard ratio. d–f, Data were collected 14 (d, f) and 10 (e) days after tumour implantation, pooled from two or more independent experiments with n ≥ 3 per group; each point indicates one mouse analysed separately. n, number of tumours from individual patients or mice. Horizontal bars show medians. P values determined by one-way ANOVA with Tukey’s (a) and Kruskal–Wallis multiple comparison (d) post tests, two-tailed Mann–Whitney test (b, e, f), two-sided log-rank test (b, c, survival curves), and linear regression (c).
We next looked for ILCs in tumours from mice in which PDAC development is driven by mutated Kras and p53 (autochthonous KPC mice)5 and orthotopic mouse models of PDAC (PDAC mice6,7). In both models, we detected TILC2s that were phenotypically similar to those in human PDACs and to mouse ILC2s2,8 (Fig. 1d, Extended Data Fig. 1d–f). The frequency of mouse ILC2s was increased in tumours, but not in adjacent organs (Fig. 1d, Extended Data Fig. 1g), consistent with their tissue residency9, and were depleted in Rag2−/− mice10,11 by targeting the lymphocyte antigen CD90.2 (Fig. 1e, Extended Data Fig. 1h). Therefore, ILC2s are conserved cells that expand locally in mouse and human PDACs.
To identify the signals that expand TILC2s, we found that IL33 was the most highly expressed ILC-activating cytokine in tumours in both PDAC and KPC mice12 (Extended Data Fig. 2a). IL33 was heterogeneously expressed in both human and mouse PDACs (Extended Data Fig. 2b, c) and maximally expressed in intratumoral myeloid cells13,14 (Extended Data Fig. 2d, e). To understand the role of IL33 and TILC2s in PDAC immunity, we studied TILC2 dependency on IL33 in IL33High PDAC mice, to model IL33High, ILC2-enriched hot tumours in long-term human PDAC survivors. The expansion and function of TILC2s were IL33-dependent, as Il33−/− PDAC mice had reduced TILC2 number, frequency (Fig. 1f, Extended Data Fig. 2f) and cytokine production (Extended Data Fig. 2g) when compared to Il33+/+ PDAC mice. Recombinant IL33 (rIL33) expanded ILCs in ILC-proficient Rag2−/− PDAC mice, but not in ILC-deficient mice that lack both Rag2 and the gene encoding the gamma subunit of the IL-2 receptor (Il2rg, also known as γc) (Rag2−/−γc−/− PDAC mice) (Extended Data Fig. 2h, i). Collectively, these data show that IL33 expands TILC2s in PDACs.
TILC2s boost tumour immunity in tissues
As ILC2s have tissue-specific phenotypes15, we hypothesized that TILC2s have tissue-specific effects on PDAC immunity. To test this, we compared the effects of IL33 deficiency on pancreatic and skin tumour growth (pancreatic TILC2s express ST2 whereas skin TILC2s do not15,16; Extended Data Fig. 2j). Compared with Il33+/+ mice, Il33−/− mice with orthotopic PDAC had larger tumours, accelerated tumour growth, and worse survival (Fig. 2a). By contrast, mice with subcutaneous PDACs showed no IL33-dependent phenotype (Fig. 2b, Extended Data Fig. 2k). Although these mice were fully backcrossed onto identical genetic backgrounds, we confirmed that the differences were not due to potential minor genetic mismatches by observing larger tumours in Il33−/− mice compared to Il33+/+ littermates (Extended Data Fig. 2l). These anti-tumour effects depended on IL33 produced by host haematopoietic cells, as chimaeric mice transplanted with Il33−/− bone marrow had larger tumours than did control mice (Extended Data Fig. 2m–o). RNA sequencing (RNA-seq) of purified CD45+ intratumoral immune cells from Il33+/+ and Il33−/− mice with orthotopic PDACs showed that PDAC immune cells from Il33−/− mice had diminished transcriptional signatures of T cell activation and MHC class I antigen processing (Extended Data Fig. 3a), suggesting that Il33−/− PDAC mice might have a defect in T cell priming. Consistently, Il33−/− mice with orthotopic but not subcutaneous PDACs had lower frequencies of global and activated tumour-infiltrating CD8+ T cells and reduced central memory CD8+ T cells (TCM) in draining lymph nodes (DLN) but not distant lymph nodes, with no consistent changes in other immune cell frequencies (Fig. 2c, Extended Data Fig. 3b–e). Depletion of all T cells prevented the increase in tumour size in Il33−/− compared to Il33+/+ mice (Fig. 2d), and rIL33-treated Rag2−/− PDAC mice showed no differences in tumour weight compared with untreated mice (Extended Data Fig. 4a), confirming that the anti-tumour effects of IL33 were mediated by T cells. Orthotopic tumours from Il33−/− and Il33+/+ PDAC mice also had similar histology and collagen and fibroblast content (Extended Data Fig. 4b–d), and rIL33 had no effects on tumour cells in vitro (Extended Data Fig. 4e–g), showing that IL33 had no direct effects on tumour or stromal cells. Together, these data show that IL33 activates tissue-specific cancer immunity by potentially activating TILC2s to prime CD8+ T cells.
a, b, Tumour weight, tumour volume, and survival of Il33+/+ and Il33−/− mice with orthotopic (a) or subcutaneous (b) PDACs. c, Frequency of all (left) and IFN-γ producing (right) CD8+ T cells in orthotopic Il33+/+ and Il33−/− PDAC tumours. d, Tumour weight in T cell-depleted Il33+/+ and Il33−/− mice with orthotopic PDACs. e, Frequency of tumour rejection (orthotopic and subcutaneous) and tumour weight (orthotopic) in Il33+/+ and Il33−/− mice with KPC-OVA PDACs. f, Experimental design (left), frequency of tumour rejection (middle), and tumour weight (right) of KPC-OVA PDAC tumours in iCOS-T mice with intact or depleted ILC2s. g, Gating (top) and frequency (bottom) of OVA-specific CD8+ T cells in DLNs of iCOS-T mice with intact or depleted ILC2s in orthotopic KPC-OVA PDACs. Data were collected 14 days (a, c, d), 28 days (b), 42 days (e), and 8 days (f, g) after implantation. Mean ± s.e.m; horizontal bars show median. Data pooled from two or more independent experiments with n ≥ 4 per group; n and data points denote individual mice analysed separately. P values determined by two-tailed Mann–Whitney test (a–g), two-sided log-rank test (a, b, survival curves), two-way ANOVA with Sidak’s multiple comparison test (a, b, tumour volumes), and χ2 test (e, f, per cent rejection).
We next tested whether the tissue-specific effects of IL33 depended on CD8+ T cells by contrasting the rejection phenotypes of KPC cells expressing the CD8+ T cell rejection antigen ovalbumin (KPC-OVA cells) at different tissue sites. Notably, 70% of Il33+/+ mice rejected orthotopic KPC-OVA tumours, whereas 0% of Il33−/− mice did. By contrast, 100% of Il33+/+ and Il33−/− mice rejected subcutaneous KPC-OVA tumours (Fig. 2e). To assess whether this phenotype resulted from ILC2 deficiency and ineffective CD8+ T cell priming, we acutely depleted ILC2s and examined antigen-specific CD8+ T cells in DLNs in the iCOS-T mouse, in which diphtheria toxin depletes ILC2s while sparing ICOS+CD4+ T cells17 (Fig. 2f, Extended Data Fig. 5a). ILC2 depletion recapitulated the Il33−/− phenotype; mice with orthotopic KPC-OVA tumours showed a lower rate of tumour rejection and larger tumour size, whereas those with subcutaneous tumours showed no differences (Fig. 2f), with an anticipated varied phenotype compared to Il33−/− mice due to differences in time of rejection assessment and depletion efficacy. Tetramer analysis in mice with orthotopic KPC-OVA tumours depleted of ILC2s revealed a reduced frequency of OVA-specific CD8+ T cells in DLNs and spleens, and a reduced frequency of CD8+ TCM cells in DLNs (as seen in Il33−/− mice) (Fig. 2g, Extended Data Fig. 5b, c). Therefore, ILC2 deficiency partially phenocopied IL33 deficiency. Although direct effects of IL33 on CD8+ T cells cannot be ruled out, we found no ST2 expression on intratumoral CD8+ T cells (Extended Data Fig. 5d). To summarize, these loss-of-function experiments suggest that the IL33–TILC2 axis primes tissue-specific CD8+ T cell PDAC immunity.
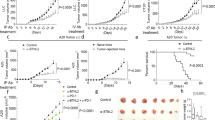
Next, we investigated whether rIL33 treatment had similar tissue-specific anti-tumour effects. Treatment with rIL33 prevented tumour establishment in mice with orthotopic PDACs and prolonged survival, but had no effects on mice with subcutaneous PDACs, resulting in progressive tumour growth and ulceration requiring euthanasia (Fig. 3a). rIL33 had similar tissue-specific anti-tumour effects in KPC-OVA PDAC mice (Extended Data Fig. 6a). Similarly, rIL18, a cytokine that preferentially activates IL18R+ skin ILC2s15, restricted the growth of subcutaneous PDACs infiltrated by IL18R+ ILCs, but not of orthotopic PDACs, which lack IL18R+ ILCs (Fig. 3b, Extended Data Fig. 6b). rIL33 selectively expanded ILC2s in DLNs and tumours (Fig. 3c) but not in spleens of mice with orthotopic PDACs or in any organs in mice with subcutaneous PDACs (Extended Data Fig. 6c, d). ILC2 expansion was accompanied by enhanced intratumoral CD8+ T cell cytokine capacity and PD-1 upregulation (Extended Data Fig. 6e), with no consistent changes in other intratumoral immune cells (Extended Data Fig. 6f), although potential modulation of their function cannot be ruled out. Consistent with indirect priming of anti-tumour CD8+ T cells by ILC2s, rIL33 treatment doubled the number of intratumoral CD103+ dendritic cells (DCs) (Fig. 3d, Extended Data Fig. 6g), which prime and recruit CD8+ T cells into PDACs7. To determine whether the effects of rIL33 depended on ILC2s, we administered rIL33 to PDAC-bearing Rorafl/flIl7rCre/+ mice, which are constitutively deficient in ILC2s17. ILC2 deficiency (Extended Data Fig. 6h) abrogated the efficacy of rIL33 (Fig. 3e) and attenuated increases in CD103+ DCs in tumours (Fig. 3f). rIL33 also had no anti-tumour effects (Fig. 3g) and failed to induce PD-1 expression in intratumoral CD8+ T cells (Extended Data Fig. 6i) in CD103+ DC-deficient Batf3−/− mice, showing that CD103+ DCs are essential for rIL33-mediated tumour control. To test whether TILC2s produced chemokines to recruit DCs into tumours, we used single-cell RNA-seq (scRNA-seq; Extended Data Fig. 7a–c, Supplementary Table 3) and found that rIL33-activated TILC2s retained markers of ILC2 identity but exhibited distinct transcriptional profiles (Extended Data Fig. 8a–e), and selectively expressed Ccl5 (Extended Data Fig. 8f). CCL5 recruits CD103+ DCs into tumours18 and induced efficient DC migration in vitro (Fig. 3h). Together, these data suggest that rIL33 expands TILC2s to produce CCL5, potentially recruit CD103+ DCs into tumours, and activate CD8+ T cells to induce therapeutic tumour immunity.
a, Tumour weight, tumour volume, and survival in orthotopic and subcutaneous PDAC mice treated with vehicle or rIL33. b, Tumour weight and volume in mice with orthotopic or subcutaneous PDACs treated with vehicle or rIL18. c, Gating, frequency, and number of ILC2s in rIL33-treated mice with orthotopic PDACs (DLN vehicle, n = 13, top; tumour vehicle, n = 12, bottom; DLN rIL33, n = 14, top; tumour rIL33, n = 15, bottom). d, Gating and frequency of CD103+ DCs in tumours from rIL33-treated mice with orthotopic PDACs. e, f, Tumour weight and volume (e) and frequency of CD103+ DCs (f) in tumours from rIL33-treated wild-type (WT) and ILC2-deficient mice with orthotopic PDACs. g, Tumour volume in rIL33-treated WT and CD103+ DC-deficient Batf3−/− mice with orthotopic PDACs. h, Migration of purified DCs towards CCL5. Data were collected 5 (c, d) or 7 (e, f) weeks after tumour implantation. Median ± s.e.m; horizontal bars show median. Data were pooled from at least two independent experiments, with n ≥ 3 per group; n and data points denote individual mice analysed separately or individual replicates (h). P values determined by two-sided log-rank test (a, survival curves), two-way ANOVA (a, b, e, g, tumour volume), and two-tailed Mann–Whitney test (a–f, h).
PD-1 blockade activates TILC2s
As stimulating ILC2s with rIL33 had anti-tumour effects, we searched for strategies to further activate ILC2s. Recent data have shown that, like T cells, ILC2s regulate their activity through coinhibitory immune checkpoint pathways. Specifically, the immune checkpoint PD-1 regulates mouse ILC2 development and marks effector ILCs19, and when PD-1 is genetically deficient or inhibited with a blocking antibody (anti-PD-1), IL33-activated ILC2s show increased expansion and effector function in mice and humans20. PD-1+ ILC2s are also found in human tumours3. However, inhibiting immune checkpoints on ILC2s for cancer therapy has been unexplored.
Using scRNA-seq (Extended Data Fig. 7a–c), we found that PD-1 was the only detectable coinhibitory molecule expressed at baseline by TILC2s (Extended Data Fig. 9a). Treatment with rIL33 upregulated PD-1 on a fraction of TILC2s but not on DLN ILC2s (Extended Data Fig. 9b), suggesting that PD-1 may functionally restrain activated TILC2s. We therefore tested whether combining rIL33 with anti-PD-1 treatment could cooperatively activate TILC2s to enhance anti-tumour efficacy. Consistent with the expression of PD-1 only on rIL33-activated TILC2s, anti-PD-1 alone induced a partial anti-tumour response (Fig. 4a, as previously reported in PDACs7) but did not appreciably alter TILC2 frequencies (Fig. 4b, Extended Data Fig. 9c). A combination of rIL33 and anti-PD-1 maximally expanded ILC2s in tumours and DLNs (Fig. 4b) and enhanced tumour control compared to anti-PD-1 alone (Fig. 4a). To investigate whether anti-PD-1 activated ILC2s by cell-intrinsic PD-1 blockade, we compared the single-cell transcriptional profiles of TILC2s and DLN ILC2s following in vivo treatment. Whereas TILC2s retained the transcriptional and cellular identities of ILC2s irrespective of treatment (Extended Data Fig. 9d), rIL33 and anti-PD-1 induced a unique transcriptional phenotype in TILC2s in PDAC mice compared to all other treatment conditions and tissue sites (Fig. 4c), and increased expression of ILC2 markers, canonical (amphiregulin (Areg))15 and non-canonical (Cxcl2)21 ILC2 effector molecules, cellular activation machinery (Junb, Fosl2, and Ybx1), and coinhibitory immune checkpoints (Extended Data Fig. 9e–i). Finally, the anti-tumour effects of dual therapy were abrogated in ILC2-deficient mice (Fig. 4d), which shows that ILC2s are necessary for the efficacy of dual anti-PD-1 and rIL33 therapy. These results suggest that anti-PD-1 amplifies activated TILC2s by possibly inhibiting the PD-1 pathway on ILC2s, and not on T cells.
a, b, Tumour volume (vehicle, n = 8; rIL33, n = 13; anti-PD-1 + rIL33, n = 14) and survival (vehicle, n = 9; rIL33, n = 10; anti-PD-1 + rIL33, n = 15) (a) and gating, frequency, and number of ILC2s (b) in treated PDAC mice. c, scRNA-seq (n = 7,022 single ILC2s) in treated PDAC mice in a nonlinear representation of the top 15 principal components; cells are coloured by cluster (left) or treatment and tissue (right). d, Tumour volume in wild-type (WT) and ILC2-deficient PDAC mice treated with anti-PD-1 and rIL33. e, TILC2s were sort-purified from rIL33-treated WT or Pdcd1−/− PDAC mice and transferred into ILC2-deficient PDAC mice, and tumour volumes were measured. f–h, TILC2s were sort-purified from rIL33-treated PDAC CD45.1 donor mice and transferred into ILC2-deficient CD45.2 PDAC recipient mice, which were then treated with anti-PD-1. f, Experimental design, tumour volume and tumour weight; g, frequency of CD45.1 and CD45.2 cells; h, frequency of T cells in recipient mice 9 weeks after cell transfer (TILC2−: all groups, n = 8; TILC2+: spleen, n = 9; DLN, n = 7; tumour, n = 7). Frequencies in g represent percentage of live donor- or recipient-derived immune cells. i, Tumour volume (vehicle, n = 13; other groups, n = 10) and survival (vehicle and anti-PD-1, n = 15; rIL33, n = 24; rIL33 + anti-PD-1, n = 26) of treated PDAC mice (KPC 52 cells). ILC2 deficient, Rorafl/flIl7rCre. Data were collected 5 weeks (b), 10 days (c), or 6 weeks (d) after orthotopic tumour cell implantation. Median ± s.e.m; horizontal bars show median. Data pooled from at least two independent experiments with n ≥ 3 per group; n and data points denote individual mice analysed separately. Data for scRNA-seq represent pooled purified single cells from biological replicates (vehicle n = 10, rIL33 n = 5, anti-PD-1 + rIL33 n = 5). P values determined by two-way ANOVA with Tukey’s multiple comparison post test (a, d–f, i, tumour volume), two-tailed Mann–Whitney test (b, d, f, g, h), and two-sided log-rank test (a, i, survival curves).
PD-1 inhibits cell-intrinsic TILC2 function
To investigate whether interrupting the PD-1 pathway on activated TILC2s contributed to the anti-tumour effects of dual therapy, we transferred sort-purified rIL33-activated PD-1 proficient (wild-type) or PD-1 deficient (Pdcd1−/−) TILC2s into tumour-bearing ILC2-deficient mice (Fig. 4e, Extended Data Fig. 10a). Transfer of wild-type TILC2s had no anti-tumour efficacy in established tumours, but transfer of Pdcd1−/− TILC2s restricted tumour growth, indicating that interrupting PD-1 signalling on TILC2s can enhance tumour control (Fig. 4e). We next tested whether rIL33-activated PD-1+ TILC2s could directly amplify the efficacy of anti-PD-1 therapy in established tumours. We transferred sort-purified rIL33-activated congenic CD45.1+ TILC2s into CD45.2+ ILC2-deficient mice with established tumours, and treated the mice with anti-PD-1 after the transfer (Fig. 4f). Transferred TILC2s were more than 97% PD-1+ (Extended Data Fig. 10b), accumulated in the tumours and DLNs but not spleens of anti-PD-1 treated recipient mice, and persisted for up to 9 weeks post-transfer (Fig. 4g). Transfer of PD-1+ TILC2s augmented the efficacy of anti-PD-1 treatment, restricted tumour growth (Fig. 4f), and increased T cell frequencies in the tumours and DLNs, but not spleens, of recipient mice (Fig. 4h). These data show that blocking PD-1 signalling on rIL33-activated TILC2s directly amplified the anti-tumour efficacy of anti-PD-1 treatment.
To examine the efficacy of rIL33 and anti-PD-1 treatment in IL33Low, anti-PD-1-resistant tumours, we selected an aggressive cold PDAC tumour model (KPC 52 mice) that generates IL33Low tumours (Extended Data Fig. 2b), has 50% fewer CD8+ T cells than IL33High tumours, and has a median survival of only 2 weeks (Extended Data Fig. 10c), to mimic the immunological and survival features of patients with IL33Low PDAC tumours who show short-term survival. Although KPC 52 PDAC mice do not exhibit the sequential steps of PDAC tumorigenesis from pre-invasive neoplasias to invasive PDAC that are seen in spontaneous KPC mice, they recapitulated the anti-PD-1 resistance seen in spontaneous KPC mice and human PDACs (Fig. 4i). Combination treatment with rIL33 and anti-PD-1 reduced tumour volume by over 50% in these mice, with a nearly 50% improvement in survival (Fig. 4i). Finally, to assess the potential to use dual rIL33 and anti-PD-1 therapy to treat patients with PDAC, we investigated the co-occurrence of PD-1+ TILC2s and PD-1+ T cells in human PDACs. Nearly 60% of human PDACs had low frequencies of PD-1+ TILC2s and PD-1+ T cells, with a significant correlation between the two cell types (Extended Data Fig. 10d), which suggests that they frequently co-occur in human PDAC. In addition, IL33 mRNA correlated substantially with PDCD1 mRNA, which encodes PD-1 (Extended Data Fig. 10e). PD-1 expression has been associated with longer survival22, which suggests that the IL33–PD-1 axis may positively impact survival in individuals with PDAC. In summary, rIL33-activated ILC2s can amplify responses to anti-PD-1 in both tumours that are partially sensitive to PD-1 and those that are PD-1-resistant.
Discussion
Our results suggest that ILC2s can be activated as a broader strategy to prime CD8+ T cells in cancers (Extended Data Fig. 10f). However, given the tissue-specific phenotypes of ILC2s, more work is required to determine whether activating them will have similar effects in cancers arising in different tissues. Given the divergent effects of ILC2s on tumour immunity in different tissues, our findings also underscore the need for pre-clinical cancer studies to be performed in native organs to accurately reflect the local immune environments.
Immune checkpoints modulate ILC2 function, but the ability to harness ILC2s with immune checkpoint blockade for cancer therapy has remained unclear. We have shown that blocking PD-1 on activated ILC2s promotes anti-tumour effects, suggesting that ILC2s may partially contribute to the efficacy of PD-1 pathway blockade in human cancers. More broadly, this highlights that differential responses to immune checkpoint blockade may depend on tissue-specific factors. Refining strategies to identify ILC2s in human cancers will clarify their prognostic and predictive potential.
As activated ILC2s (Extended Data Fig. 10g) and T cells share several immune modulatory molecules and co-exist in human cancers, a broader array of checkpoints could be co-targeted on ILC2s and T cells in tumours. Further investigations to collectively target ILC2s and T cells for cancer immunotherapy are therefore warranted.
Methods
Mice
C57BL/6 (wild-type, WT, CD45.2), C57BL/6 CD45.1, Rag2−/−, Rag2−/−γc−/−, Batf3−/−, and Pdcd1−/− mice were purchased from Jackson Labs. Il33−/− and Il33Cit/+ mice were a gift from M. J. Rosen. Cd4Cre/+Icosfl-Dtr/+ and Rorafl/flIl7rCre/+ mice were a gift from A. N. J. McKenzie and have been previously described17,23. For all experiments, 6−12-week-old mice were matched by age and sex and randomly assigned to specific treatment groups, with at least two independent experiments performed throughout. Pdx1CreLSL-KrasG12D/+LSL-Trp53R172H/+ (KPC) mice have been previously described5. Sample sizes for experiments were determined without formal power calculations. Animals were bred and maintained in a specific pathogen-free animal facility, and all experiments were conducted in accordance with an Institutional Animal Care and Use Committee (IACUC) approved protocol at Memorial Sloan Kettering Cancer Center (MSKCC) and in compliance with all relevant ethical regulations.
Cell lines and animal procedures
All tumour cell lines were derived from KPC mice. KPC 4662 cells from Pdx1CreLSL-KrasG12D/+LSL-Trp53R172H/+ mice (a gift from R. H. Vonderheide) were transfected with GFP and used for all experiments unless indicated otherwise. KPC 8-1, 18-3, and 52 cells derived from Ptf1aCreLSL-KrasG12D/+LSL-Trp53R172H/+ mice were a gift from C. Iacobuzio-Donahue. KPC 4662 cells engineered to express OVA were previously described24 (a gift from R. H. Vonderheide). All cell lines were authenticated as bona fide PDAC cell lines based on histopathologic verification by a dedicated pancreatic cancer pathologist. Orthotopic tumours established with KPC 4662 cells were IL33High and transiently decreased in size in response to anti-PD-1 therapy initiated at the time of implantation (anti-PD-1 partial sensitivity). Orthotopic tumours established with KPC 52 cells were IL33Low and did not decrease in size in response to anti-PD-1 therapy initiated at the time of implantation (anti-PD-1 resistant). All cell lines were regularly tested using the MycoAlert Mycoplasma Detection Kit (Lonza). Orthotopic PDAC tumours were established as previously described6. In brief, mice were anaesthetized using a ketamine–xylazine cocktail, and a small (7-mm) incision was made into the left abdominal side. Tumour cells (106 KPC cells per mouse; 1.25 × 105 KPC-OVA cells per mouse) were suspended in Matrigel (Becton Dickinson), diluted 1:1 with cold phosphate-buffered saline (PBS) (total volume of 50 μl), and injected into the tail of the pancreas using a 26-gauge needle. Successful injection was verified by the appearance of a fluid bubble without intraperitoneal leakage. The abdominal wall was closed with absorbable Vicryl RAPIDE sutures (Ethicon), and the skin was closed with wound clips (Roboz). For subcutaneous PDAC tumours, tumour cells (106 KPC cells per mouse; 1.25 × 105 KPC-OVA cells per mouse) were resuspended in sterile PBS (Fisher Scientific) and implanted subcutaneously. Mice were euthanized at the indicated time points and processed for histology or flow cytometry. Autochthonous KPC mice were euthanized when tumours were detectable by ultrasound. Tumour volumes were measured using serial ultrasound (Vevo 2100 Linear Array Imaging and Vivo LABORATORY Version 3.1.1, Fuji Film Visual Sonics) for orthotopic tumours as previously described25. For subcutaneous tumours, tumour length and width were measured every 2–3 days using calipers, and tumour volumes were calculated as volume = length/2 × width2. For survival analyses, survival was determined by a tumour volume of ≥500 mm3 or mouse health requiring euthanasia as defined by institutional IACUC guidelines. No mouse tumours exceeded IACUC-defined maximal tumour volumes of ≥2 cm3. No blinding was performed in experimental mouse interventions, as knowledge of the treatment groups was required.
T cell depletion
CD4+ and CD8+ cells were depleted by intraperitoneal (i.p.) injection of 250 μg of anti-mouse CD4 antibody (clone GK1.5, Bio X Cell, InVivoPlus) and 250 μg of anti-mouse CD8a antibody (clone 2.43, Bio X Cell, InVivoPlus). Control mice were treated with rat IgG2b isotype control (clone LTF-2, Bio X Cell, InVivoPlus). Mice were treated daily for 3 days before tumour implantation, and then every 3 days for the duration of the experiment. CD4+ and CD8+ T cell depletion was confirmed by flow cytometric analysis of tumours and secondary lymphoid organs (>85% depletion).
ILC depletion
ILCs were depleted in Rag2−/− mice by i.p. injection of 300 μg of anti-mouse CD90.2 (clone 30-H12, Bio X Cell) on days 0, 1, 3, 6, 9, and 13 following tumour implantation as previously described26. ILC2s were depleted in Cd4Cre/+Icosfl-DTR/+ experimental mice and Cd4Cre/+Icos+/+ control mice treated by i.p. injection of diphtheria toxin (Sigma-Aldrich) at a dosage of 25 ng per gram of mouse body weight. Mice were treated the day before tumour implantation and then every other day thereafter for a total of five doses, as previously described17. ILC2 depletion was confirmed by flow cytometric analysis of tumours (Extended Data Fig. 5a).
Bone marrow chimaeras
Bone marrow was removed from CD45.2 congenically labelled donor mice, filtered through a 70-mm filter, centrifuged, and resuspended in sterile PBS to a concentration of 108 live cells per 200 μl. CD45.1 congenically labelled C57BL/6J recipient mice were irradiated (5.5 Gy × 2, 6 h apart) 24 h before bone marrow transplant and were maintained on endofloxacin water for 4 weeks after irradiation. A single-cell suspension of CD45.2 bone marrow chimaera in sterile PBS (108 live cells per recipient mouse) was transplanted to each recipient mouse by retroorbital injection. Reconstitution was confirmed by flow cytometry of the peripheral blood at 4 and 8 weeks post transplantation. Tumour implantation experiments were performed at 12 weeks post transplantation.
Recombinant IL33, IL18, and PD-1 blockade
For rIL33, mice were treated with i.p. injections of 500 ng of carrier-free recombinant mouse IL33 (R&D Systems) in sterile PBS daily for 7 days, and then every 2 days thereafter as previously described16. For rIL18, mice were treated with i.p. injection of 2 μg of carrier-free recombinant mouse IL-18 (R&D Systems) in sterile PBS at days 3, 7, 11, and 15 after tumour inoculation as previously described27. The chimaeric anti-mouse PD-1 antibody (4H2) used in this study was engineered as a mouse IgG1 isotype monoclonal antibody (mAb) and was shown to bind to CHO transfectants expressing PD-1 and to block binding of PD-L1 and PD-L2 to these cells. The affinity of 4H2 for mouse PD-1, determined by surface plasmon resonance using PD-1-Fc, was 4.68 × 10−9 M. The antibody was produced and purified at Bristol Myers Squibb (BMS). Each batch was certified to have <0.5 EU/mg endotoxin and be of >95% purity. All dosing solutions were prepared in PBS. Mice were treated with i.p. injection of 250 μg anti-PD-1 every 2 days. Transient reduction in tumour size but subsequent regrowth while on continuous anti-PD-1 treatment was defined as a partial response. No reduction in tumour size while on continuous anti-PD-1 was defined as resistance.
Human samples
All tissues were collected at MSKCC following study protocol approval by the MSKCC Institutional Review Board. Informed consent was obtained from all patients. The study was performed in strict compliance with all institutional ethical regulations. All tumour samples were surgically resected primary PDACs.
Tissue microarray
Tissue microarrays (TMAs) were constructed from tumour and adjacent non-tumour cores from formalin-fixed, paraffin-embedded tissue blocks from short-term survivors (n = 45 tumours, 5 normal tissues) and long-term survivors (n = 51 tumours, 5 normal tissues) of PDAC as previously described4. Patient subsets were randomly selected to undergo tissue microarray construction. Patients treated with neoadjuvant therapy were excluded. All tumours were subjected to pathological re-review and histological confirmation by two expert PDAC pathologists before analysis. Long-term survivors were defined as patients with overall survival of >3 years from surgery and short-term survivors as patients with survival >3 months and <1 year from surgery, to exclude perioperative mortalities. ILC2High and ILC2Low were defined as greater or lesser, respectively, than the median ILC2 frequency for the entire TMA cohort.
Tumour transcriptomic profiling
Patient subsets were randomly selected to undergo transcriptomic profiling as previously described4. Patients in the TMA cohort with tumour tissue available for transcriptomic assessment were included in analyses in Fig. 1b to allow protein confirmation of RNA expression. Extracted RNA was qualified on an Agilent BioAnalyzer and quantified by fluorometry (Ribogreen). Preparation of RNA for whole-transcriptome expression analysis was done using the WT Pico Reagent Kit (Affymetrix). Reverse transcription was initiated at the poly-A tail as well as throughout the entire length of RNA to capture both coding and multiple forms of non-coding RNA. RNA amplification was achieved using low-cycle PCR followed by linear amplification using T7 in vitro transcription technology. The cRNA was then converted to biotinylated sense-strand DNA hybridization targets. The prepared target was hybridized to GeneChip Human Transcriptome Array 2.0 (Affymetrix). Washes were performed using the GeneChip Hybridization, Wash and Stain Kit using a Fluidics Station 450/250. Arrays were scanned using the GeneChip Scanner 3000. Data analysis for the array was done using Affymetrix Expression Console Software (SST-RMA algorithm to summarize the signal from array probesets). Immune cytolytic activity was determined as previously described28.
Cell isolation
Mouse and human PDAC tumours and adjacent pancreata were mechanically dissociated and incubated in collagenase (collagenase II for mouse tumours, collagenase IV for human tumours, both 5 mg/ml; Worthington Biochemical Corp., Fisher Scientific), DNase I (0.5 mg/ml; Roche Diagnostics), and Hank’s balanced salt solution (Gibco, Fisher Scientific) for 30 min at 37 °C. Digestion was then quenched with fetal bovine serum (FBS, Life Technologies), and cells were filtered sequentially through 100- and 40-μm nylon cell strainers (Falcon, Fisher Scientific). Lymph nodes were mechanically disassociated and filtered through 100- and 40-μm nylon cell strainers (Falcon, Fisher Scientific) using PBS with 1% FBS (Life Technologies). Spleens were mechanically dissociated and filtered through 70- and 40-μm nylon cell strainers (Falcon, Fisher Scientific) using PBS with 1% FBS, followed by RBC lysis (RBC lysis buffer, Thermo Fisher Scientific). Mouse Fc receptors were blocked with FcεRIII/II-specific antibody (1 μg per 1 × 106 cells; clone 2.4G2, Bio X Cell).
ILC2 adoptive transfer
CD45.1 C57BL/6 or Pdcd1−/− orthotopic PDAC mice were treated with 500 ng of carrier-free recombinant mouse IL33 (R&D Systems) in sterile PBS daily for 10 days. Live, CD45+, lineage−, CD90+, CD25+, ST2+ TILC2s were sort-purified to 98% purity at day 10 post-implantation using an Aria Cell sorter (BD Biosciences). TILC2s (5 × 105 cells) were immediately transferred to orthotopic PDAC tumour-bearing Rorafl/flIl7rCre/+ CD45.2 mice on days 7 and 14 post-tumour implantation via i.p. injection. Control mice received equivalent volumes of PBS via i.p. injection. Anti-PD-1 treatment in recipient mice was initiated on the day of ILC2 cell transfer. Tissues were collected at indicated time points.
Flow cytometry
Single-cell suspensions were stained using antibody cocktails in the dark at 4 °C, washed, and analysed on a FACS LSR Fortessa (BD Biosciences). Mouse ILCs were defined as live, CD45+, lineage− (CD3, CD5, NK1.1, CD11b, CD11c, CD19, FcεR1), CD25+, CD127+ cells, as previously described2,8. Mouse immune cells were defined as follows: ILC2s: live, CD45+, lineage−, CD25+, ST2+ cells; central memory T cells (TCM): live, CD45+, CD3+, NK1.1−, CD8+, CD62l+, CD44+; dendritic cells (DC): live, CD45+, CD3−, NK1.1−, Gr1−, F4/80−, CD11c+, MHC-II+; B cells: live, CD45+, CD3−, CD19+; T cells: live, CD45+, CD3+; CD4+ T cells: live, CD45+, CD3+, CD4+; CD8+ T cells: live, CD45+, CD3+, CD8+; regulatory T cells: live, CD45+, CD3+, CD4+, FoxP3+; tumour-associated macrophages: live, CD45+, CD11b+, F4\80+, GR1−; myeloid-derived suppressor cells (MDSCs): live, CD45+, CD3−, CD11b+, F4\80−, GR1+. Mouse cells were stained with the following antibodies: from Biolegend, CD45 (clone 30-F11, Pacific Blue), CD45.1 (clone A20, BV711), NK1.1 (clone PK136, APC), Gr-1 (clone RB6-8C5, BV605), CD103 (clone 2E7, BV711); from BD Biosciences, CD5 (clone 53-7.3, APC), CD11c (clone HL3, APC), NK1.1 (clone PK136, BV605), CD4 (clone RM4-5, BV786), CD62L (clone MEL-14, APC), CD19 (clone 1D3, BV510), Ly6C (clone AL-21, PerCP-Cy5.5), Ly6G (clone 1A8, AF700), PD-1 (clone J43 BV605), TNF-α (clone MP6-XT22, BV510), IFN-γ (clone XMG1.2, APC-Cy7), CD90.2 (clone 53-2.1, BV786), T-bet (clone Q4-46, BV711), RORγ-t (clone Q31-378, BV786), GATA3 (clone L50-823, PE-Cy7), and IL4 (clone 11B11, BV650); from Thermo Fisher Scientific CD3 (clone 17A2, Alexa Fluor 700), CD11b (clone M1/70, APC), CD11b (clone M1/70, PerCP-Cy5.5), CD8 (clone 53-6.7, Alexa Fluor 700), CD19 (clone 1D3, Alexa Fluor 700), FcεR1 (clone MAR-1, APC), F4/80 (clone BM8, PE-Cy5), CD3 (clone 145-2C11, PE-Cy7), MHC-II (clone M5/114.15.2, Alexa Fluor 700), CD44 (clone IM7, PerCP-Cy5.5), CD127 (clone A7R34, FITC), CD25 (clone PC61.5, PerCP-Cy5.5), IL5 (clone TRFK5, PE), CD11c (clone N418, FITC), ST2 (clone RMST2-2, PE-Cy7), and FOXP3 (clone FJK-16S, APC); and from MBL international, SINFEKL tetramer (catalogue # TB-5001-1, PE).
Human ILCs were defined as live, CD45+, lineage− (CD3, CD5, CD56, CD11b, CD11c, CD16, CD19, TCRα/β, FcεR1), CD25+, CD127+ cells as previously described8. Human cells were stained with the following antibodies: from BD Biosciences, GATA3 (clone L50-823, BV711), T-bet (clone O4-46, BV650), RORγ-T (clone Q21-559, PE); from Biolegend, CRTH2 (clone BM16, PE-Cy7), CD11b (clone ICRF44, APC), CD56 (clone NCAM16.2, BV650), CD25 (clone BC96, PerCP-Cy5.5), CD45 (clone HI30, Pacific Blue), TCRα/β (clone IP26, APC); from Thermo Fisher Scientific, CD16 (clone CB16, APC), CD11c (clone 3.9, APC), CD127 (clone RDR5, FITC), CD3 (clone OKT3, Alexa Fluor 700), ST2 (clone hIL33Rcap, PE), CD5 (clone L17F12, APC), CD19 (clone HIB19, AF700), FcεR1 (clone AER-37, APC). All samples for flow cytometry were prospectively collected from unselected patients with PDAC.
To examine intracellular cytokine production, single-cell suspensions of tumours were stimulated for 6 h ex vivo with phorbol 12-myristate (PMA, 100 ng/ml) and ionomycin (1 ng/ml) in the presence of brefeldin A (10 μg/ml) (all from Sigma-Aldrich) at 37 °C. Cells were then surface-stained, fixed, permeabilized, and stained for cytokine production using the Fixation and Permeabilization Buffer Kit per the manufacturer’s recommendations (Invitrogen, Thermo Fisher Scientific). Appropriate isotype controls were used as indicated. Analysis was performed using FlowJo (versions 9 and 10, Tree Star).
Immunohistochemistry
Tissues were fixed in paraformaldehyde (Fisher Scientific) for 24 h and embedded in paraffin. The tissue sections were deparaffinized with EZPrep buffer (Ventana Medical Systems), then antigen retrieval was performed with CC1 buffer (Ventana Medical Systems). Sections were blocked for 30 min with Background Buster solution (Innovex), followed by avidin-biotin blocking for 8 min (Ventana Medical Systems). Mouse IL33 (AF3626, R&D Systems), mouse smooth muscle actin (Abcam), and human IL33 (AF3625, R&D Systems) antibodies were applied, and sections were incubated for 4 h, followed by a 60-min incubation with biotinylated rabbit anti-goat IgG (Vector labs), or biotinylated goat anti-rabbit IgG (Vector labs) at 1:200 dilution. Detection was performed with a DAB detection kit (Ventana Medical Systems) according to the manufacturer’s instructions. Any section containing cells demonstrating cytoplasmic or nuclear positivity for IL33 was designated to have positive staining. Slides were counterstained with Masson’s trichrome, or haematoxylin and eosin, and coverslipped with Permount (Fisher Scientific). All histologic sections were evaluated by an independent PDAC pathologist.
Mouse immunofluorescence
IL33/CD11b/CK19/Iba1 immunofluorescence
Multiplex immunofluorescent staining was performed using a Discovery XT processor (Ventana Medical Systems) as described29.
IL33
First, sections were incubated with anti-mIL33 (R&D Systems, catalogue # AF3626, 1 μg/ml) for 4 h, followed by 60 min incubation with biotinylated horse anti-goat IgG (Vector Laboratories) at 1:200 dilution. Detection was performed with Streptavidin-HRP D (part of DABMap kit, Ventana Medical Systems), followed by incubation with Tyramide Alexa Fluor 488 (Invitrogen) prepared according to the manufacturer’s instructions with predetermined dilutions.
CD11b
Next, sections were incubated with anti-CD11b (Abcam, clone EPR1544) for 5 h, followed by 60 min incubation with biotinylated goat anti-rabbit IgG (Vector Laboratories) at 1:200 dilution. Detection was performed with Streptavidin-HRP D (part of DABMap kit, Ventana Medical Systems), followed by incubation with Tyramide Alexa 594 (Invitrogen) prepared according to the manufacturer’s instructions with predetermined dilutions.
CK19
Next, slides were incubated with anti-CK19 (Abcam, clone EP1580Y) for 5 h, followed by 60 min incubation with biotinylated goat anti-rabbit (Vector Laboratories) at 1:200 dilution. Detection was performed with Streptavidin-HRP D (part of DABMap kit, Ventana Medical Systems), followed by incubation with Tyramide Alexa Fluor 546 (Invitrogen) prepared according to the manufacturer’s instructions with predetermined dilutions.
Iba1
Finally, sections were incubated with anti-Iba1 (Wako, catalogue #019-19741) for 5 h, followed by 60 min incubation with biotinylated goat anti-rabbit IgG (Vector Laboratories) at 1:200 dilution. Detection was performed with Streptavidin-HRP D (part of DABMap kit, Ventana Medical Systems), followed by incubation with Tyramide Alexa 647 (Invitrogen) prepared according to the manufacturer’s instructions with predetermined dilutions. After staining, slides were counterstained with DAPI (Sigma-Aldrich) for 10 min and coverslipped with Mowiol.
Human immunofluorescence
Tissue sections were deparaffinized with proprietary Leica Bond buffer (Leica Biosystems), and antigen retrieval was performed with Leica Bond ER2 buffer (Leica Biosystems). First, sections were incubated with anti-PD-1 (Cell Marque, clone NAT105) for 1 h, followed by detection with Bond Polymer Refine Detection kit (Leica Biosystems) and Tyramide Alexa Fluor 488 (Invitrogen). Next, sections were incubated with anti-CD3 (DAKO, catalogue#A0452) for 1 h, followed by detection with Bond Polymer Refine Detection kit (Leica Biosystems) and Tyramide CF594 (Biotum). Next, sections were incubated with anti-GATA3 (Cell Marque, clone L50-823) for 1 h, followed by detection with Bond Polymer Refine Detection kit (Leica Biosystems) and CF 543 (Biotum). Finally, sections were incubated with anti-CD45 (DAKO, clone 2B11 + PD7/26) for 1 h, followed by detection with Bond Polymer Refine Detection kit (Leica Biosystems) and Tyramide Alexa Fluor 647 (Invitrogen). All detections were prepared according to the manufacturer’s instructions with predetermined dilutions. After staining, slides were counterstained with DAPI (Sigma-Aldrich) for 10 min and coverslipped with Mowiol.
Digital image processing and analysis
Slides were digitized using Panoramic Flash 250 (3Dhistech, Budapest, Hungary) using a Zeiss 20×/0.8NA objective and custom filters for A488, A546, A594, and A647. Each core was exported into a multi-channel tiff file and analysed using a custom macro written in FIJI/ImageJ. For quantification, each nucleus was segmented using the DAPI channel after appropriate processing and background subtraction. Then for each nucleated cell, the presence or absence of the other markers was assessed after setting appropriate thresholds for each marker. The number of cells with specific combinations of markers was tallied. ILC2s were defined as CD45+ CD3− GATA3+ nucleated cells, PD-1-expressing ILC2s were defined as CD45+ CD3− GATA3+ PD-1+ nucleated cells, and PD-1-expressing T cells were defined as CD45+ CD3+ PD-1+ nucleated cells. For each patient, the frequency of each cell type as a fraction of all nucleated cells was calculated in triplicate cores, followed by determination of the mean frequency of triplicate cores to calculate the final cellular frequency per patient.
RNA sequencing
Mouse
Tissues from mice with orthotopic PDACs (n = 6) were collected and dissociated into single-cell suspensions as described above. Tumour-infiltrating leukocytes were positively selected by magnetically activated cell sorting using mouse CD45 MicroBeads (Miltenyi Biotec). Purification of magnetically activated sorted cells was confirmed by flow cytometry and was >95%. RNA was isolated from the sorted cells using an RNeasy Plus Mini Kit (Qiagen). Poly(A) capture and paired-end RNA-seq were performed by the MSKCC Integrated Genomics Core Facility. Specifically, after RiboGreen quantification and quality control by Agilent BioAnalyzer, 500 ng of total RNA underwent polyA selection and TruSeq library preparation according to instructions provided by Illumina (TruSeq Stranded mRNA LT Kit, catalogue # RS-122-2102), with eight cycles of PCR. Samples were barcoded and run on a HiSeq 4000 in a 100 bp/100 bp paired-end run, using the HiSeq 3000/4000 SBS Kit (Illumina). An average of 83 million paired reads was generated per sample. Ribosomal reads represented at most 0.03% of the total reads generated, and the percentage of mRNA bases averaged 76.6%. The expression data set was loaded into Gene Set Enrichment Analysis (GSEA) 3.0. Gene set databases for antigen presentation and T cell mediated immunity were selected from MSIGDB v6.1, with a false discovery rate of ≤0.25 to facilitate exploratory discovery. GSEA was run with 1,000 permutations. Three gene set databases met this threshold: GO: 0002474 (antigen processing and presentation of peptide antigen via MHC class I); GO: 0002711 (positive regulation of t cell mediated immunity); and GSE19825 (naive vs day 3 effector CD8 T cell up).
Single-cell RNA sequencing
Library preparation for single-cell immune profiling, sequencing, and post-processing of the raw data were performed at the Epigenomics Core at Weill Cornell Medicine.
Single-cell RNA library preparation and sequencing
Single-cell suspensions of fluorescence-activated cell (FAC)-sorted ILC2 cells from pancreatic KPC tumours and mesenteric DLNs from mice treated with vehicle, rIL33 alone, or rIL33 + anti-PD-1 were prepared as described above. scRNA-seq libraries were prepared according to 10X Genomics specifications (Chromium Single Cell V(D)J User Guide PN-1000006, 10X Genomics). Four independent cellular suspensions (85–90% viable) at a concentration between 90 and 200 cells/μl were loaded onto to the 10X Genomics Chromium platform to generate Gel Beads-in-Emulsion (GEM), targeting about 2,000 single cells per sample. After GEM generation, the samples were incubated at 53 °C for 45 min in a C1000 Touch Thermal cycler with 96-Deep Well Reaction Module (Bio-Rad) to generate polyA cDNA barcoded at the 5′ end by the addition of a template switch oligo (TSO) linked to a cell barcode and Unique Molecular Identifiers (UMIs). GEMs were broken, and the single-strand cDNA was cleaned up with DynaBeads MyOne Silane Beads (Thermo Fisher Scientific). The cDNA was amplified for 16 cycles (98 °C for 45 s; 98 °C for 20 s, 67 °C for 30 s, 72 °C for 1 h). The quality of the cDNA was assessed using an Agilent Bioanalyzer 2100, obtaining a product of about 1,200 bp. Fifty nanograms of cDNA was enzymatically fragmented, end repaired, A-tailed, subjected to a double-sided size selection with SPRIselect beads (Beckman Coulter), and ligated to adaptors provided in the kit. A unique sample index for each library was introduced through 14 cycles of PCR amplification using the indexes provided in the kit (98 °C for 45 s; 98 °C for 20 s, 54 °C for 30 s, 72 °C for 20 s × 14 cycles; 72 °C for 1 min; held at 4 °C). Indexed libraries were subjected to a second double-sided size selection, and libraries were then quantified using Qubit fluorometric quantification (Thermo Fisher Scientific). The quality was assessed on an Agilent Bioanalyzer 2100, obtaining an average library size of 450 bp. No treatment samples had concentrations below detectable limits, and cDNA amplification was done with 18 cycles and sample index with 16 cycles. Libraries were diluted to 10 nM and clustered using a NovaSeq600 on a paired-end read flow cell and sequenced for 28 cycles on R1 (10X barcode and the UMIs), followed by 8 cycles of I7 Index (sample Index), and 89 bases on R2 (transcript), obtaining about 100 million clusters per sample, except for tumours from vehicle-treated mice (clustered at about 10 million). Primary processing of sequencing images was done using Illumina’s Real Time Analysis software (RTA). 10X Genomics Cell Ranger Single Cell Software suite v3.0.2 (https://support.10xgenomics.com/single-cell-gene-expression/software/pipelines/latest/what-is-cell-ranger) was used to perform sample demultiplexing, alignment to mouse genomic reference mm10, filtering, UMI counting, single-cell 5′ end gene counting, and quality control using the manufacturer’s parameters. Data from approximately 11,000 single cells that passed quality control were obtained with approximately 41,000 mean reads per cell (48% sequencing saturation).
scRNA-seq data processing
The Seurat R package version 3.1 pipeline was used to identify clusters on combined data sets30. First, individual data sets were read into R as count matrices and converted into Seurat objects, selecting on genes expressed in ≥3 cells and on cells with at least 200 detected genes. A standard pre-processing workflow was then used to filter cells based on excluding cells with either more than 2,500 or fewer than 200 unique genes expressed, and cells with greater than 5% mitochondrial gene content.
Following filtering, the samples were merged, and the gene expression measurements for retained cells were log-transformed, normalized by total expression per cell, and scaled to 10,000 molecules per cell. The top 2,000 highly variable genes across the single cells were then identified, and principal component (PC) analysis was conducted. After examining jackstraw and elbow plots, we selected the top 15 PCs for clustering using K-nearest neighbour (KNN) clustering with cluster resolution set at 0.4, identifying 6–8 clusters in all samples-combined and tumour-combined merged data sets. Nonlinear dimensional reduction with UMAP was used to visualize the data sets, also using the top 15 PCs. Differential gene expression for gene marker discovery across the clusters was performed using the Wilcoxon rank sum test as used in the Seurat package. Pairwise comparison using Wilcoxon rank sum test was performed with the Holm P value adjustment method to compare gene expression between samples.
In vitro assays
KPC 4662 cells were cultured for 1 week in a 96-well flat-bottomed plate (Falcon) in complete medium: RPMI with l-glutamine (Gibco, Thermo Fisher Scientific) with 10% fetal bovine serum (Life Technologies), 100 units/ml of penicillin, 100 μg/ml of streptomycin, and rIL33 at concentrations of 0, 10, 100, and 500 ng/ml. Culture medium and cytokines were replenished every 48 h. Viability was measured using a colorimetric tetrazolium salt assay (Cell Counting Kit, Dojindo Molecular Technologies) per the manufacturer’s instructions and read on a Synergy HT Multi-Detection Microplate Reader (Biotek). Cells were collected and stained for Annexin V (Thermo Fisher Scientific), Ki-67 (clone SolA15, Thermo Fisher Scientific), and ST2 (clone RMST2, Thermo Fisher Scientific). For all in vitro experiments, 2–3 technical replicates were performed per independent experiment.
In vitro dendritic cell migration assays
Mouse splenic DCs were isolated and enriched using a mouse pan DC isolation kit according to the manufacturer’s protocol (Miltenyi Biotech). Flow cytometry was used to assess DC purity (>70% CD11c+ of live cells). Cells were plated in complete RPMI medium at 5 × 105 cells/ml with 50 ng/ml of recombinant mouse GM-CSF (Biolegend) overnight. Next, chemotaxis of splenic DCs was analysed by transwell migration assays. RPMI (600 μl) with or without 100 ng/ml of recombinant mouse CCL5 (Biolegend) was added to the lower chambers of a 6.5-mm Transwell plate with 5.0-μm pore polycarbonate membrane inserts (Sigma-Aldrich). RPMI (200 μl) was also added to the upper chambers, and plates were allowed to equilibrate at 37 °C in 5% CO2 for 15 min. Splenic DCs (1 × 105 cells in 100 μl RPMI) were then loaded into the upper chambers and incubated at 37 °C in 5% CO2 for 2 h. After incubation, membrane inserts were carefully removed, and cells were collected from the lower chambers. Migrated DCs were incubated with DAPI and CD11c antibodies for 20 min at 4 °C, and Precision Count Beads (Biolegend) were added to quantify the number of live migrated CD11c+ cells using flow cytometry according to the manufacturer’s protocol.
Statistics
Data are expressed as median. As we observed many statistically significant effects in the data without a priori sample size calculations, no statistical methods were used to determine sample size. Comparisons between two groups were performed using unpaired Mann–Whitney test with the Benjamini–Krieger–Yekutieli false discovery approach for multiple time point comparisons (two-tailed). Comparisons among multiple groups were performed using one-way ANOVA test followed by Kruskal–Wallis multiple comparison post-test. Comparisons among multiple groups across multiple time points were performed using two-way ANOVA test. Correlations between two variables were calculated using linear regression. Survival curves were compared by two-sided log-rank test. Tumour incidences were compared by χ2 test. All alpha levels were 0.05, with P < 0.05 considered a significant difference. Statistical analyses were performed using Prism 7.0 (GraphPad Software).
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this paper.
Data availability
Source code for immune quantification is available in Supplementary Data 1. Bulk RNA-seq data are available under Gene Expression Omnibus (GEO) accession number GSE129388. scRNA-seq data are available under GEO accession number GSE136720. Source data are provided for all experiments. All other data are available from the corresponding author upon reasonable request.
References
Vivier, E., van de Pavert, S. A., Cooper, M. D. & Belz, G. T. The evolution of innate lymphoid cells. Nat. Immunol. 21, 790–794 (2016).
Vivier, E. et al. Innate lymphoid cells: 10 years on. Cell 174, 1054–1066 (2018).
Salimi, M. et al. Activated innate lymphoid cell populations accumulate in human tumour tissues. BMC Cancer 18, 341 (2018).
Balachandran, V. P. et al. Identification of unique neoantigen qualities in long-term survivors of pancreatic cancer. Nature 551, 512–516 (2017).
Hingorani, S. R. et al. Trp53R172H and KrasG12D cooperate to promote chromosomal instability and widely metastatic pancreatic ductal adenocarcinoma in mice. Cancer Cell 7, 469–483 (2005).
Pylayeva-Gupta, Y., Lee, K. E., Hajdu, C. H., Miller, G. & Bar-Sagi, D. Oncogenic Kras-induced GM-CSF production promotes the development of pancreatic neoplasia. Cancer Cell 21, 836–847 (2012).
Li, J. et al. Tumor cell-intrinsic factors underlie heterogeneity of immune cell infiltration and response to immunotherapy. Immunity 49, 178–193.e7 (2018).
Brestoff, J. R. et al. Group 2 innate lymphoid cells promote beiging of white adipose tissue and limit obesity. Nature 519, 242–246 (2015).
Gasteiger, G., Fan, X., Dikiy, S., Lee, S. Y. & Rudensky, A. Y. Tissue residency of innate lymphoid cells in lymphoid and nonlymphoid organs. Science 350, 981–985 (2015).
Kirchberger, S. et al. Innate lymphoid cells sustain colon cancer through production of interleukin-22 in a mouse model. J. Exp. Med. 210, 917–931 (2013).
Monticelli, L. A. et al. Arginase 1 is an innate lymphoid-cell-intrinsic metabolic checkpoint controlling type 2 inflammation. Nat. Immunol. 17, 656–665 (2016).
Neesse, A. et al. CTGF antagonism with mAb FG-3019 enhances chemotherapy response without increasing drug delivery in murine ductal pancreas cancer. Proc. Natl Acad. Sci. USA 110, 12325–12330 (2013).
Hardman, C. S., Panova, V. & McKenzie, A. N. J. IL-33 citrine reporter mice reveal the temporal and spatial expression of IL-33 during allergic lung inflammation. Eur. J. Immunol. 43, 488–498 (2013).
Talabot-Ayer, D. et al. The mouse interleukin (Il)33 gene is expressed in a cell type- and stimulus-dependent manner from two alternative promoters. J. Leukoc. Biol. 91, 119–125 (2012).
Ricardo-Gonzalez, R. R. et al. Tissue signals imprint ILC2 identity with anticipatory function. Nat. Immunol. 19, 1093–1099 (2018).
Dalmas, E. et al. Interleukin-33-activated islet-resident innate lymphoid cells promote insulin secretion through myeloid cell retinoic acid production. Immunity 47, 928–942.e7 (2017).
Oliphant, C. J. et al. MHCII-mediated dialog between group 2 innate lymphoid cells and CD4+ T cells potentiates type 2 immunity and promotes parasitic helminth expulsion. Immunity 41, 283–295 (2014).
Böttcher, J. P. et al. NK cells stimulate recruitment of cDC1 into the tumor microenvironment promoting cancer immune control. Cell 172, 1022–1037.e14 (2018).
Yu, Y. et al. Single-cell RNA-seq identifies a PD-1hi ILC progenitor and defines its development pathway. Nature 539, 102–106 (2016).
Taylor, S. et al. PD-1 regulates KLRG1+ group 2 innate lymphoid cells. J. Exp. Med. 214, 1663–1678 (2017).
Kim, J. et al. Intratumorally establishing type 2 innate lymphoid cells blocks tumor growth. J. Immunol. 196, 2410–2423 (2016).
Diana, A. et al. Prognostic value, localization and correlation of PD-1/PD-L1, CD8 and FOXP3 with the desmoplastic stroma in pancreatic ductal adenocarcinoma. Oncotarget 7, 40992–41004 (2016).
Donovan, C. et al. Roles for T/B lymphocytes and ILC2s in experimental chronic obstructive pulmonary disease. J. Leukoc. Biol. 105, 143–150 (2019).
Evans, R. A. et al. Lack of immunoediting in murine pancreatic cancer reversed with neoantigen. JCI Insight 1, 88328 (2016).
Sastra, S. A. & Olive, K. P. Quantification of murine pancreatic tumors by high-resolution ultrasound. Methods Mol. Biol. 980, 249–266 (2013).
Monticelli, L. A. et al. Innate lymphoid cells promote lung-tissue homeostasis after infection with influenza virus. Nat. Immunol. 12, 1045–1054 (2011).
Ma, Z. et al. Augmentation of immune checkpoint cancer immunotherapy with IL18. Clin. Cancer Res. 22, 2969–2980 (2016).
Rooney, M. S., Shukla, S. A., Wu, C. J., Getz, G. & Hacohen, N. Molecular and genetic properties of tumors associated with local immune cytolytic activity. Cell 160, 48–61 (2015).
Yarilin, D. et al. Machine-based method for multiplex in situ molecular characterization of tissues by immunofluorescence detection. Sci. Rep. 5, 9534 (2015).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Acknowledgements
We thank J. Novak, J. Moore and E. Patterson for editorial assistance; B. Medina, G. Vitiello, J. Zhang, S. Zeng, F. Rossi, J. Loo, N. Param, J. Maltbaek, O. Grbovic-Huezo, Y. Senbabaoglu, M. Gigoux, R. Giese and S. Budhu for helpful discussions and technical assistance; and the Epigenomics Core of Weill Cornell Medical College for technical assistance with scRNA-seq. This work was supported by the V Foundation Convergence Scholar Grant (J.A.M., J.D.W., V.P.B.), the Stand Up to Cancer Convergence Award (J.D.W., V.P.B.), the National Cancer Institute K12CA184746-01A1 (V.P.B.), the Damon Runyon Clinical Investigator Award (V.P.B.), the Ben and Rose Cole Pria Foundation Scholar Award (V.P.B.), the Sarah Min and Matthew Pincus Pancreatic Cancer Immunotherapy Award (V.P.B.), an administrative supplement to NIH P30-CA008748 (S.D.L., V.P.B.), NIH R01 CA204228, NIH P30CA023108 (S.D.L.), Swim Across America and the Ludwig Institute for Cancer Research (J.D.W., T.M.), and the Parker Institute for Cancer Immunotherapy (J.D.W., T.M.). Services by the MSKCC Small-Animal Core Facility and Integrated Genomics Core were funded by the National Cancer Institute Cancer Center Support Grant (P30 CA008748-48), Cycle for Survival, and the Marie-Josée and Henry R. Kravis Center for Molecular Oncology.
Author information
Authors and Affiliations
Contributions
V.P.B. conceived the study. J.A.M, J.L., L.A.R., S.D.L., J.D.W., R.P.D., T.M. and V.P.B. designed all the experiments. J.A.M, J.L., L.A.R., J.R., J.Z., A.R. and V.P.B. performed all the experiments. B.G. assisted with generation of bone marrow chimaeras. J.L., Z.S. and D.R. analysed the scRNA-seq results. G.A. and U.B. performed the pathologic analyses. E.E., Y.P. and D.A.T. generated and assisted in experiments on autochthonous KPC mice. M. Gururajan provided technical assistance with the PD-1 blocking antibody. M. Gönen provided statistical oversight. J.A.M., J.L., L.A.R., J.R., S.D.L., R.P.D., T.M. and V.P.B. analysed all the data. All authors interpreted the data. J.A.M. and V.P.B. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
V.P.B. is a recipient of an immuno-oncology translational research grant from Bristol-Myers Squibb and is an inventor on a patent application related to work on neoantigen modelling. S.D.L. is a member of the scientific advisory board of Nybo Pharmaceuticals, and co-founder of Episteme Prognostics. J.D.W. is a consultant for Adaptive Biotech, Advaxis, Amgen, Apricity, Array BioPharma, Ascentage Pharma, Astellas, Bayer, Beigene, Bristol-Myers Squibb, Celgene, Chugai, Elucida, Eli Lilly, F Star, Genentech, Imvaq, Janssen, Kleo Pharma, Linneaus, MedImmune, Merck, Neon Therapuetics, Ono, Polaris Pharma, Polynoma, Psioxus, Puretech, Recepta, Trieza, Sellas Life Sciences, Serametrix, Surface Oncology and Syndax; is a recipient of research support from Bristol-Myers Squibb, Medimmune, Merck Pharmaceuticals and Genentech; has equity in Potenza Therapeutics, Tizona Pharmaceuticals, Adaptive Biotechnologies, Elucida, Imvaq, Beigene, Trieza and Linneaus; and has received an honorarium from Esanex. T.M. is a consultant for Immunos Therapeutics and Pfizer; is a co-founder with equity in Imvaq Therapeutics; receives research funding from Bristol-Myers Squibb, Surface Oncology, Kyn Therapeutics, Infinity Pharmaceuticals Inc., Peregrine Pharmaceuticals Inc., Adaptive Biotechnologies, Leap Therapeutics Inc. and Aprea; and is an inventor on patent applications related to work on Oncolytic Viral Therapy, Alpha Virus Based Vaccine, Neo Antigen Modeling, CD40, GITR, OX40, PD-1 and CTLA-4. M. Gururajan is an employee of Bristol-Myers Squibb and has financial interest in the company.
Additional information
Peer review information Nature thanks Richard Locksley and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Identification of IL33-dependent ILCs in pancreatic cancer.
a, Gating strategy to identify human ILCs. The first plot was pre-gated on live (DRAQ7−) cells and singlets. Lineage 1 cocktail: CD5, CD11b, CD11c, CD16, FcεR1. Lineage 2 cocktail: CD3, CD19, TCRα/β. ILCs were identified as lineage− CD56− CD25+ CD127+ cells. FMO, fluorescence minus one. b, Representative image of immunofluorescence of ILC2s in tumour tissue microarrays from short- and long-term PDAC survivors (n = 96). Arrows, putative ILC2s. c, Top, overall survival of patients with more (high) or less (low) than the median intratumoral mRNA level of ILC-stimulating cytokines. Bottom, correlation between expression of ILC-activating cytokines and immune cytolytic index (CYT) in short- and long-term survivors of human PDAC. Curves were fit by linear regression. n = 25. d, Gating strategy to identify mouse ILCs. The first plot was pre-gated on live (DRAQ7−) cells and singlets. Lineage 1 cocktail: CD5, CD11b, CD11c, FcεR1. Lineage 2 cocktail: CD3, CD19. ILCs were identified as lineage− NK1.1− CD25+ CD127+, and ILC2s were identified as lineage− NK1.1− CD25+ ST2+ cells. Gating on orthotopic PDAC mice shown. e, Intratumoral ILC frequency in orthotopic PDAC mice established with KPC cell lines 8-1, 18-3, and in autochthonous KPC mice with spontaneous PDAC (KPCSpont). Composite ILC frequencies from Fig. 1d and others are included for comparison (KPC 4662). f, Phenotype of ILCs in PDAC mice. Grey curves, isotype controls; numbers, mean fluorescence intensity. g, Expansion kinetics of ILCs in tissues from PDAC mice. h, Changes in non-ILC cell frequency in Rag2−/− PDAC mice treated with anti-CD90.2 or isotype antibodies. Data were collected 14 days (d–f), 10 days (h), or at the indicated time points after tumour implantation. n indicates individual mice analysed separately in at least two independent experiments with n ≥ 2 per group. Median ± s.e.m; horizontal bars show median. P values determined by two-sided log-rank test (c, top), linear regression (c, bottom), or two-tailed Mann–Whitney test (g). P values in g indicate tumour comparisons to all other tissues.
Extended Data Fig. 2 Host-derived IL33 activates pancreatic ILC2s.
a, mRNA expression of ILC1- (IL12, IL15, IL18), ILC2- (IL25, IL33, TSLP), and ILC3-inducer cytokines (IL23) and the IL33 receptor (ST2) in orthotopic PDAC tumours (left) and autochthonous PDAC tumours in KPC mice from a previously published mRNA microarray (right)12. b, Representative IL33 immunohistochemistry (IHC) of IL33Low and IL33High human (tissue microarray, n = 96) and mouse PDAC (n = 3 per group). c, Frequency of patients with PDAC demonstrating IL33 positivity by IHC in a human PDAC tumour microarray. d, Multiplexed immunofluorescence for IL33, ductal marker CK19, and myeloid markers CD11b and Iba in mouse PDAC (top). Arrows, IL33-expressing cells. IL33 mean fluorescence intensity (MFI) in non-immune (CD45−), immune (CD45+), macrophage (TAM), and monocytic and granulocytic myeloid-derived suppressor cell (M-MDSC and G-MDSC) populations in tumours from IL33Cit reporter PDAC mice (bottom). e, Representative IL33 protein expression shown by IHC in orthotopic PDAC tumours in Il33+/+ (WT) mice, and non-tumour-bearing pancreata in Il33–/– mice (n = 3 per group). f, ILC frequency (top) and cell number (bottom) in organs and DLNs of Il33+/+ and Il33−/− orthotopic PDAC mice. g, Gating and frequency of IL4 and IL5 expression in intratumoral ILCs in Il33+/+ and Il33−/− orthotopic PDAC mice. h, i, ILC2 (h) and immune cell frequencies (i) in orthotopic Rag2–/– and Rag2–/–γc–/– PDAC mice with or without treatment with rIL33. j, Frequency of ST2+ tumour ILCs in mice with subcutaneous (SQ) and orthotopic PDACs. k, Tumours in orthotopic and subcutaneous PDAC mice. l, Tumour weight in Il33+/+ and Il33−/− littermate PDAC mice. m, Experimental schema of bone-marrow chimaeras to evaluate contribution of haematopoietic cell-derived IL33 to tumour control. n, o, Haematopoietic cell reconstitution (n) and tumour weight (o) in irradiated CD45.1 congenic mice reconstituted with either CD45.2 Il33+/+ or CD45.2 Il33−/− bone marrow. Data were collected 14 (a, b, d-g, j, k (orthotopic), l, o), 28 (k (subcutaneous)), or 10 (h, i) days after tumour implantation. Horizontal bars show median. n indicates individual mice analysed separately in at least two independent experiments with n ≥ 2 per group. P values determined by one-way ANOVA (a) or two-tailed Mann–Whitney test (d, f–h, j, l, o).
Extended Data Fig. 3 Host-derived IL33 activates pancreatic T cell immunity.
a, Gene set enrichment analysis of bulk RNA-seq from purified CD45+ immune cells from Il33+/+ and Il33−/− PDAC mice. Enrichment plots and enrichment scores are shown for three gene sets comparing expression in Il33−/− to expression in Il33+/+ mice (n = 3 mice per group). FDR, false discovery rate. b, c, Gating of CD8+ T cells (b) and frequencies of various immune cell types (c, left) and CD4+ T cell lineages (c, right) in Il33+/+ and Il33−/− orthotopic PDAC mice. d, Frequency of T central memory (TCM) cells (CD45+ CD3+ CD8+ CD44+ CD62L+) in tumour DLNs and non-tumour draining distant lymphoid organs (inguinal lymph node and spleen) in Il33+/+ and Il33−/− orthotopic PDAC mice. e, Frequency of CD8+ T cells in subcutaneous PDAC tumours. NK, natural killer cells; NKT, natural killer T cells; Treg, regulatory T cells; MDSC, myeloid-derived suppressor cells; DC, dendritic cells. Data were collected 14 days after tumour implantation or at the time points indicated. Median ± s.e.m; horizontal bars show median. n indicates individual mice analysed separately in at least two independent experiments with n ≥ 2 per group. P values determined by one-way ANOVA (d).
Extended Data Fig. 4 IL33 and ILCs do not directly induce tumour cell death.
a, Tumour weight in Rag2−/− and Rag2−/−γc−/− PDAC mice treated with vehicle or rIL33. b, Representative haematoxylin and eosin-stained sections (left) with histologic tumour cell differentiation status in Il33+/+ and Il33−/− PDAC mice (right). c, Trichrome staining in tumours from Il33+/+ and Il33−/− PDAC mice (n = 3 per group). d, Immunohistochemistry for smooth muscle actin in tumours from Il33+/+ and Il33−/− PDAC mice (n = 3 per group). e, Intratumoral ST2 expression on KPC cells in Il33+/+ and Il33−/− orthotopic PDAC mice. f, ST2 expression on live KPC cells following rIL33 treatment in vitro (DRAQ7 stains dead cells) (n = 3 per group). g, KPC cell number, viability, proliferation (Ki-67), and apoptosis (annexin) following rIL33 treatment in vitro (n = 3–6 per group). Horizontal bars show median. n in a–e indicates individual mice analysed separately in at least two independent experiments with n ≥ 3 per group. n in f, g indicates technical replicates and is representative of at least two independent experiments. P value determined by two-tailed Mann–Whitney test (a).
Extended Data Fig. 5 ILC2s prime antigen-specific CD8+ T cells.
a, Gating and frequency of TILC2s in ILC2-intact (diphtheria toxin (DT)-treated Cd4Cre/+Icos+/+) and ILC2-depleted (DT-treated Cd4Cre/+Icosfl.DTR/+) mice. b, Gating and frequency of OVA-specific CD8+ T cells in spleens from ILC2-intact and ILC2-depleted mice. OVA-specific T cells were detected as SIINFEKL-tetramer+ cells. c, Gating and frequency of TCM cells (CD45+ CD3+ CD8+ CD44+ CD62L+) in tumour, DLNs, and spleens from ILC2-intact and ILC2-depleted mice. d, ST2 expression on CD45+ CD3+ CD8+ T cells after tumour implantation in PDAC mice. Data were collected 14 days after tumour implantation or at the time points indicated. Median ± s.e.m; horizontal bars show median. n indicates individual mice analysed separately in at least two independent experiments with n ≥ 2 per group. P values determined by two-tailed Mann–Whitney test (a–c) and two-way ANOVA with Tukey’s multiple comparison post test (d, indicating comparison of tumour ILCs to all other groups).
Extended Data Fig. 6 Immunophenotyping in rIL33-treated PDAC mice.
a, Tumour establishment of orthotopic and subcutaneous KPC-OVA PDAC tumours in vehicle (veh) and rIL33-treated mice. b, Gating (left) and frequency (right) of IL18R1 expression on tumour ILCs in subcutaneous (SQ) and orthotopic PDAC mice. c, Gating (left) and frequency (right) of splenic ILC2s following rIL33 treatment in orthotopic PDAC mice. d, Gating (left), frequency (middle), and number (right) of TILC2s following rIL33 treatment in subcutaneous PDAC mice. e, Gating (left) and frequency (right) of cytokine and PD-1 expression on tumour CD8+ T cells following rIL33 treatment in orthotopic PDAC mice. f, Frequency of immune cells in vehicle- and rIL33-treated orthotopic PDAC mice. g, Gating strategy for identification of CD103+ DCs. h, Gating (left; tumours) and frequency (right) of ILC2s in tumours and DLNs from wild-type (WT) or ILC2-deficient (Rorafl/flIl7rCre) PDAC mice following rIL33 treatment. i, Gating (left) and frequency (right) of PD-1+ CD8+ T cells in tumours from rIL33-treated wild-type (WT) and Batf3−/− mice. Data were collected 6 (a), 4 (b), 2 (c, e, f), 5 (d), 7 (h), and 3 (i) weeks after tumour implantation. Horizontal bars show median. n indicates individual mice analysed separately in at least two independent experiments with n ≥ 2 per group. P values determined by χ2 test (a), two-tailed Mann–Whitney test (all else).
Extended Data Fig. 7 scRNA-seq of tumour and DLN ILC2s in PDAC mice.
a, Experimental design for in vivo treatment, purification, and single-cell analysis of ILC2s. b, c, Quality metrics. b, Scatter plots showing, for each cell, the relationship between the number of UMIs and the number of genes. c, Violin plots showing the distribution of the number of genes (left), number of UMIs (middle), and percentage of normalized reads from mitochondrial genes (right) in each treatment group (columns), and each tissue (rows). Each dot represents a single cell. For each treatment group and tissue, data represent pooled purified single cells from biological replicates of n = 10 (vehicle), n = 5 (rIL33), and n = 5 (anti-PD-1 + rIL33) PDAC mice.
Extended Data Fig. 8 Activated ILC2s from tumours and DLNs have distinct transcriptional features.
a–d, Single-cell analysis of 1,634 rIL33-activated tumour and DLN ILC2s (experimental design as in Extended Data Fig. 7a). UMAP plots show single cells (dots) in a nonlinear representation of the top 15 principal components. Expression of ILC2 (Gata3, Id2, Rora) and ILC3 (gene, Rorc; protein, RORγT) transcription factors (TFs) (a), ILC2 surface markers (b), and ILC clusters and tissues (tumour and DLN) (c). Expression of the ILC-1 transcription factor Tbx21 (T-bet) was undetectable. d, e, Differentially expressed genes by cluster (d) and tissue (e). f, Distribution of Ccl5 expression from ILC2s in tumours and DLNs; violin plots show distribution with minima, maxima, and circle indicating median. Each dot in a–c represents a single cell. For each treatment group and tissue, data represent pooled purified single cells from biological replicates of n = 5 rIL33-treated PDAC mice. P value in f by two-sided pairwise Wilcoxon rank sum test.
Extended Data Fig. 9 Combined anti-PD-1 and rIL33 treatment induces a unique transcriptional profile in TILC2s.
a, Expression of coinhibitory immune checkpoints in TILC2s in vehicle-treated PDAC mice by scRNA-seq. b, Gating and frequency of PD-1+ ILC2s in vehicle- and rIL33-treated PDAC mice. c, ILC2 frequency in treated PDAC mice. Corresponding tumour volumes, cell number, and scRNA-seq are shown in Fig. 4a–c. d, scRNA-seq of ILC2s from treated PDAC mice. Expression of ILC1 (gene, Tbx21; protein, T-bet), ILC2 (Gata3, Id2, Rora), and ILC3 (gene, Rorc; protein, Rorγt) transcription factors (TFs) in purified tumour and DLN ILC2s. Corresponding UMAP plots by cluster and treatment are depicted in Fig. 4c. e–g, Top differentially expressed genes by treatment and tissue (e), cluster (f), and distribution of expression for select differentially expressed genes by treatment and tissue (g). h, UMAP plots of 3,415 single TILC2s in a nonlinear representation of the top 15 principal components. i, Differentially expressed genes in TILC2s by treatment. Each dot in d, h represents a single cell; in d–i, for each treatment group and tissue, data represent pooled purified single cells from biological replicates of n = 10 (vehicle), n = 5 (rIL33), and n = 5 (anti-PD-1 + rIL33) PDAC mice (number of single cells for tumour: vehicle n = 28, rIL33 n = 752, rIL33 + anti-PD-1 n = 2,635; for DLN: rIL33 n = 882, rIL33 + anti-PD-1 n = 2,725). Violin plots show distribution with minima, maxima, and circle indicating median. Horizontal bars show median. P values by two-tailed Mann–Whitney test (b, c) and two-sided pairwise Wilcoxon rank sum test (g).
Extended Data Fig. 10 Activated TILC2s express PD-1 and co-exist with PD-1+ T cells.
a, b, Orthotopic PDAC mice (C57BL/6 WT, Pdcd1−/−, CD45.1) were treated with 500 ng of carrier-free rIL33 daily for 10 days (experimental designs shown in Fig. 4e, f). Live, CD45+, lineage−, CD90+, CD25+, ST2+ TILC2s were sort-purified to 98% purity at day 10 post-implantation. TILC2s (5 × 105 cells) were immediately transferred to orthotopic PDAC tumour-bearing ILC2-deficient (Rorafl/flIl7rCre/+) CD45.2 mice on days 7 and 14 post-tumour implantation via i.p. injection. Control mice received equivalent volumes of PBS via i.p. injection. a, Representative plots for TILC2 sort-purification (top) and post-sort purity (bottom). b, Representative plots showing PD-1 expression on sort-purified TILC2s from wild-type and CD45.1 mice in the experimental designs outlined in Fig. 4e, f. c, Survival and intratumoral CD8+ T cell frequency of orthotopic KPC 4662 and KPC 52 PDAC tumours; horizontal bars in c show median. d, Frequency of PD-1+ ILC2s (left) and correlation with PD-1+ T cells (right) in human PDACs. e, Linear regression analysis of IL33 and PDCD1 mRNA in bulk tumour transcriptomes from short- and long-term human PDAC survivors (left) and survival association of PD-1+ cells in tumour tissue microarrays of short- and long-term PDAC survivors (right); high and low defined as higher or lower than the median for the cohort. f, Model linking the IL33–TILC2 axis to T cell immunity in PDAC. ©2019, Memorial Sloan Kettering Cancer Center. g, Distribution of expression of costimulatory molecules in untreated TILC2s by scRNA-seq. Experimental design as shown in Extended Data Fig. 7a; data represent pooled purified single cells from biological replicates of n = 10 (vehicle). Data are representative of purity and PD-1 expression on sorted TILC2s in two independent experiments with n ≥ 4 per group (a, b). n and data points denote individual mice and patients analysed separately. P values determined by two-tailed Mann–Whitney test (c), two-sided log rank test (c, e, survival curves) and linear regression (d, e).
Supplementary information
Supplementary Tables
Supplementary Tables: Contains three tables with data on – (1) the clinicopathological characteristics of PDAC patients in the cohort analysed by flow cytometry (Fig. 1a); (2) the clinicopathological characteristics of PDAC patients in transcriptome cohort (Fig. 1c, Extended Data Fig. 1c); and (3) ILC2 single-cell RNA sequencing cellular and transcriptomic metrics (Fig. 4c, Extended Data Figs. 7-9, 10g).
Supplementary Data
Supplementary Data: Source code for immune quantification.
Source data
Rights and permissions
About this article
Cite this article
Moral, J.A., Leung, J., Rojas, L.A. et al. ILC2s amplify PD-1 blockade by activating tissue-specific cancer immunity. Nature 579, 130–135 (2020). https://doi.org/10.1038/s41586-020-2015-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2015-4
This article is cited by
-
Cytolytic too: Granzyme B-expressing human ILC2s mediate tumor rejection
Cell Research (2024)
-
A diversity of novel type-2 innate lymphoid cell subpopulations revealed during tumour expansion
Communications Biology (2024)
-
Group 3 innate lymphoid cells in intestinal health and disease
Nature Reviews Gastroenterology & Hepatology (2024)
-
Combination of IL-33 with PD-1 blockade augment mILC2s-mediated anti-tumor immunity
Cancer Immunology, Immunotherapy (2024)
-
Group 2 innate lymphoid cells and their surrounding environment
Inflammation and Regeneration (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.