Abstract
A decline in stem cell function impairs tissue regeneration during ageing, but the role of the stem-cell-supporting niche in ageing is not well understood. The small intestine is maintained by actively cycling intestinal stem cells that are regulated by the Paneth cell niche1,2. Here we show that the regenerative potential of human and mouse intestinal epithelium diminishes with age owing to defects in both stem cells and their niche. The functional decline was caused by a decrease in stemness-maintaining Wnt signalling due to production of Notum, an extracellular Wnt inhibitor, in aged Paneth cells. Mechanistically, high activity of mammalian target of rapamycin complex 1 (mTORC1) in aged Paneth cells inhibits activity of peroxisome proliferator activated receptor α (PPAR-α)3, and lowered PPAR-α activity increased Notum expression. Genetic targeting of Notum or Wnt supplementation restored function of aged intestinal organoids. Moreover, pharmacological inhibition of Notum in mice enhanced the regenerative capacity of aged stem cells and promoted recovery from chemotherapy-induced damage. Our results reveal a role of the stem cell niche in ageing and demonstrate that targeting of Notum can promote regeneration of aged tissues.
Similar content being viewed by others
Main
Tissue turnover and regenerative capacity decrease with ageing in many tissue types4,5,6. The intestinal epithelium is one of the fastest renewing tissues in the human body and has previously been reported to regenerate without loss of self-renewal in long-term in vitro organoid culture7. However, complications in the gastrointestinal system increase with age8,9,10, and intestines of old mice regenerate more slowly after radiation-induced damage11, which suggests reduced stem cell activity.
To assess possible ageing-induced changes in the human intestinal epithelium, we used the capacity of intestinal-stem-cell (ISC)-containing epithelial crypts to form clonogenic organoids7 as an in vitro assay of intestinal regenerative potential. We observed a significant age-induced reduction in the organoid-forming capacity of colonic crypts derived from biopsies from healthy human donors (Fig. 1a). As the heterogeneous human colon material does not allow downstream analysis of stem-cell-intrinsic and -extrinsic effects, we next analysed the effects of age on mouse small-intestinal epithelium. Crypts from old (more than 24-month-old) mice formed significantly fewer organoids than those isolated from young (3-to-9-month-old) mice (Extended Data Fig. 1a). Notably, regenerative growth of de novo crypts was also diminished in the organoids formed by the old crypts (Fig. 1b, Extended Data Fig. 1b, c), which indicates a reduction in stem cell function. Furthermore, the reduced crypt formation observed during serial passage of secondary crypt domains demonstrated that the decline in epithelial regeneration was due to alterations intrinsic to the epithelium (Extended Data Fig. 1d, e).
a, Organoid-forming capacity of human colonic crypts (n = 24). Solid and dotted lines show linear regression and 95% confidence interval. Inset, crypts from donors of more than 60 years of age show significantly lower organoid-forming capacity. b, Regenerative growth of crypts from young (Y) and old (O) mice. Organoids derived from young mice (n = 6) generate more de novo crypt domains (arrowheads) in primary cultures (5–9 days after isolation). Representative images from day seven of culture. Scale bar, 50 μm. Student’s paired t-test. c, Cellular frequencies analysed by flow cytometry (n = 30 young, n = 26 old). For fluorescence-activated cell sorting (FACS) gating strategy, see Supplementary Fig. 1. d, Clonogenicity of young and old Lgr5hi stem cells co-cultured with young and old Paneth cells. n, number of mice analysed. P values shown for comparison of average for young Lgr5hi cells co-cultured with young Paneth cells. Young mice aged between 3 and 9 months; old mice over 24 months of age in all experiments. Unless otherwise mentioned, in box plots, the line represents median, the box shows interquartile range and whiskers show the range. All other data are represented as mean ± s.d. and conditions compared by two-tailed unpaired Student’s t-test. P values shown in corresponding panels. P <0.05 is considered significant.
Intestinal tissue renewal is largely maintained by the LGR5-expressing ISCs, which are located between Paneth cells at the crypt base. ISCs divide regularly and produce transit-amplifying progenitor cells that divide several additional times and gradually differentiate. Paneth cells produce antimicrobial peptides and multiple signalling factors, such as epidermal growth factor (EGF), Wnt3, Delta-like ligands and cyclic ADP ribose (cADPR)2,12, which regulate stemness and function of the neighbouring ISCs. To more specifically address the separate roles of stem cells and their niche in age-associated intestinal decline, we used Lgr5-eGFP-IRES-creERT2 reporter mice1, which enable identification and isolation of Paneth cells, Lgr5–eGFPhi ISCs, and transit-amplifying cells that can be further divided into immediate eGFPmed and late eGFPlo progenitors.
The aged mouse crypts did not present gross histological alterations, and the fraction of ISCs and transit-amplifying cells that were 5-ethynyl-2′-deoxyuridine (EdU) + or Ki67+ was unchanged in old mouse and human samples (Extended Data Fig. 1f–h). However, flow cytometry of crypts from old mice revealed a significant drop in frequency of Lgr5hi ISCs (Fig. 1c), whereas Paneth cell frequency was significantly increased in old mice and humans (Fig. 1c, Extended Data Fig. 1i, j). As the Lgr5-eGFP-IRES-creERT2 mouse model exhibits mosaic expression of the eGFP-containing construct1, the alterations in cellular frequencies were also validated by immunohistochemical analyses of olfactomedin 4 and lysozyme as markers of ISCs and Paneth cells, respectively (Extended Data Fig. 1k). The reduction in ISC number, together with the unchanged EdU+ frequency suggested that old crypts may have a lower output of cells, which possibly contributes to villus blunting and slower intestinal turnover during ageing13. As Paneth cells positively regulate the number and function of Lgr5hi stem cells in young mice2,12, the decoupling of the Lgr5hi:Paneth cell ratio in old mice (Extended Data Fig. 1l) raised the possibility that interactions between these two cell types change during ageing. To address this, we investigated the organoid-forming capacity of co-cultured Lgr5hi and Paneth cells isolated from young and old mice (Fig. 1d). Old Paneth and Lgr5hi cells both showed cell-type-specific age-induced effects, and initiated organoids with reduced efficiency. Consistent with previous work, neither cell type formed organoids efficiently by themselves2,12 but, when co-cultured with Paneth cells, Lgr5hi cells from young mice formed organoids at a higher rate than old Lgr5hi cells. Of note, the age-induced stem cell defect was partially rescued by co-culture with young Paneth cells, whereas old Paneth cells did not fully support organoid formation by young Lgr5hi cells. These data indicate that both stem-cell-intrinsic and -extrinsic epithelial factors reduce the regenerative potential during intestinal ageing.
Decline in fatty acid oxidation was recently shown to intrinsically reduce the function of aged intestinal stem cells14. Old Paneth cells extrinsically decreased clonogenic growth of young Lgr5hi cells even in a long-term co-culture (Extended Data Fig. 1m, n). While the original Paneth cells lasted at least 14 days in such co-cultures (Extended Data Fig. 1o), new Paneth cells were continuously produced by the stem cells, which suggests that exposure to old Paneth cells had long-term effects on ISCs and their progeny. To understand the mechanics of how age-induced changes in niche–stem cell communication may influence stem cells, we performed RNA sequencing on both cell types (Supplementary Table 1). Old Paneth cells showed specific deregulation of genes that encode secreted or plasma-membrane-associated proteins (Extended Data Fig. 2a, b). Among the key stemness-regulating factors, we noted no alterations in expression of Wnt3 or Egf, whereas expression of cADPR-producing Bst1 was reduced (Extended Data Fig. 2c). However, targeting of Bst1 did not mimic the effects of ageing on ad libitum feeding (Extended Data Fig. 2d, e).
As the aged mouse intestine has also recently been reported to contain reduced Wnt activity13, we next focused on the extracellular Wnt inhibitor Notum, which was significantly upregulated in old Paneth cells (Fig. 2a, Extended Data Fig. 2c). Notum is a secreted Wnt deacylase that disengages Wnt ligands from Lrp5, Lrp6 and Frizzled receptors and reduces Wnt activity locally during development15,16. In the intestine, Wnts are produced by the mesenchymal cells lining the crypt17,18 and by Paneth cells2,19. Wnt ligands produced by the niche adhere to ISC plasma membrane and form a reservoir of stemness-maintaining factors until they become diluted owing to divisions outside the Wnt-producing niche20. Of note, the increase in Notum expression was strictly restricted to old Paneth cells (Fig. 2b), in which its secretion could counter the stemness-maintaining function of Wnt ligands. Correspondingly, expression of Wnt-responsive genes was reduced in old Lgr5hi cells (Fig. 2c, Extended Data Fig. 2f). Similarly, in the human intestine, NOTUM expression was restricted to Paneth cells, and its expression correlated with age, whereas LGR5 expression and age showed an inverse correlation (Extended Data Fig. 2g, h).
a, Relative Notum mRNA expression from isolated Paneth cells (n = 4 old, n = 3 young mice). b, In situ analysis and quantification of Notum mRNA expression in mouse jejunum (n = 5 mice per age group). Data are mean ± s.d. Scale bar, 10 μm. c, Expression of Wnt-responsive genes in isolated old Lgr5hi stem cells relative to young stem cells. Number of mice analysed is shown. d, Clonogenic capacity of isolated Lgr5+ cells from young and old mice cultured with or without 1 μg ml−1 recombinant Notum (n = 3 mice per age group). Representative images from day seven of culture. Data are mean ± s.d. Scale bar, 100 μm. e, Immunoblot and quantification of lysates from isolated young and old Paneth cells (n = 3 mice per age group). Tub, tubulin. Data are mean ± s.d. f, Relative Notum mRNA expression in small-intestinal organoids of young mice treated for 48 h with 5 μM of the PPAR-α inhibitor GW6471 (n = 4 biologically independent samples). g, Regenerative growth of small-intestinal organoids at day six. Organoids were treated with DMSO or GW6471 and/or 100 ng ml−1 Wnt3A for the first two days (n = 4 biologically independent samples). Student’s paired t-test. Other than in box plots, data are mean ± s.e.m.; two-tailed unpaired Student’s t-test; P values shown in the corresponding panels. P <0.05 is considered significant. For gel source data see Supplementary Fig. 3.
To test whether Notum indeed affects stemness, we cultured isolated Lgr5hi cells in the absence of Paneth cells and exogenous Wnt ligands. Under these conditions, single Lgr5hi cells form clonal spheroids, whereas more-differentiated cells do not (Fig. 2d). When cells from young mice were treated with biologically active recombinant Notum immediately after isolation to inactivate the membrane-bound Wnts that they were exposed to in vivo, their colony-forming efficacy and the size of formed spheroids were markedly reduced (Fig. 2d, Extended Data Fig. 3a, b). By contrast, colony formation by untreated cells from old mice was already reduced and Notum treatment did not have further effects. Correspondingly, exogenous Wnt ligands increased organoid-forming capacity and long-term regenerative growth specifically in the old crypts (Extended Data Fig. 3c, d). However, exogenously administered Notum had no effect on isolated crypts with tightly connected Paneth and stem cells (Extended Data Fig. 3e–g), which suggests that recombinant Notum could not access the Wnt ligands produced by the Paneth cells. Demonstrating the role of epithelial Wnt secretion, inhibition of Porcupine20 reduced clonogenic growth and the Lgr5hi:Paneth cell ratio of young organoids similarly to ageing (Extended Data Fig. 3h–j). In sum, these data highlight the consequences of reduced Wnt activity, and suggested that Notum expressed by Paneth cells could represent a mechanism for reducing Wnt activity in the old intestinal epithelium.
Notum is regulated by the canonical Wnt pathway, forming a negative-feedback loop16. However, unlike in Lgr5hi ISCs, expression of Wnt-responsive genes was not significantly altered in old Paneth cells (Supplementary Table 1). To find other candidate pathways that regulate Notum in Paneth cells, we performed gene set enrichment analysis (GSEA) and found that transcripts associated with activity of mTORC1 were significantly increased in old Paneth cells (Extended Data Fig. 4a). mTOR signalling is linked with ageing3, and in the intestine, mTORC1 modulates ISC activity via the Paneth cell niche in response to calorie intake12. We detected higher levels of phosphorylation of ribosomal protein S6 (pS6), a downstream effector of mTORC1, in Paneth cells of old mice (Fig. 2e, Extended Data Fig. 4b–d); this increase in pS6 phosphorylation was also reflected in whole-crypt preparations (Extended Data Fig. 4e). However, the frequencies of pS6+ Paneth cells (Extended Data Fig. 4c) or pS6+ crypts (data not shown) were not changed, supporting an increase in mTORC1 activity at the level of single Paneth cells. By contrast, pS6 levels in ISCs were unchanged (Extended Data Fig. 4f, g) but—as reported for the liver3—the age-induced mTORC1 activity in Paneth cells was associated with increased body mass (Extended Data Fig. 4h), potentially contributing to increased mTORC1 activity in old Paneth cells.
Inhibition of mTORC1 with rapamycin or by caloric restriction extends lifespan by inducing multisystemic effects21,22,23. When old crypts were transiently treated with rapamycin, regenerative function was restored (Extended Data Fig. 5a, b). Moreover, two-week-long in vivo treatment of old mice with rapamycin resulted in a marked rejuvenation of intestinal regenerative capacity that was attributable to effects on both Paneth cells and ISCs (Extended Data Fig. 5c–h). However, in contrast to caloric restriction12, systemic rapamycin treatment induced broad changes in expression of intestinal Wnt ligands, including the stromally produced Wnt4, which regulates Notum expression in the developing ovary24 (Extended Data Fig. 5i). Correspondingly, Notum expression was increased in crypts from rapamycin-treated mice (Extended Data Fig. 5j), possibly also reflecting an increase in the number of Paneth cells induced by rapamycin12. To address the role of mTORC1 activity in the intestinal epithelium in vivo without the rapamycin-induced systemic and stromal effects, we activated mTORC1 specifically in the intestinal epithelium of mice by Villin–Cre-mediated deletion of tuberculosis sclerosis complex 1 (Tsc1). Tsc1 deletion induced mTORC1 activation and Notum expression in Paneth cells, and reduced organoid-forming capacity (Extended Data Fig. 5k–m). In sum, these results indicate that increased cell-autonomous mTORC1 activity in Paneth cells contributes to the regenerative decline of the old intestinal epithelium.
As mTORC1 does not directly regulate transcription, we searched for factors that mediate Notum expression downstream of mTORC1 activation. To that end, GSEA analysis of old Paneth cells also indicated a significant reduction in expression of genes that are regulated by PPAR-α and PPAR-δ (Extended Data Fig. 6a). mTORC1 activity inhibits PPAR-α3, and we identified a putative binding site for PPAR-α in the Notum gene (Extended Data Fig. 6b). To test whether downregulation of PPAR-α may contribute to the observed ageing phenotypes, we treated young organoid cultures with the PPAR-α antagonist GW6471. GW6471 increased expression of Notum, reduced regenerative growth and decreased the Lgr5hi:Paneth cell ratio (Fig. 2f, g, Extended Data Fig. 6c). Moreover, the ageing-mimicking effects of GW6471 were abrogated by Wnt supplementation (Fig. 2g, Extended Data Fig. 6d). These data indicate that age-associated change in the mTOR–PPAR-α axis modifies Notum expression and the intestinal regenerative capacity in a Wnt-dependent fashion.
Finally, to investigate whether endogenous Notum expression is functionally relevant for the regenerative function, we targeted Notum in organoids to knock out gene function (Extended Data Fig. 7a). Notum-knockout organoids showed increased regenerative capacity in vitro and higher growth rate when orthotopically transplanted to recipient mouse submucosa (Fig. 3a, Extended Data Fig. 7b, c). Moreover, regenerative function of old organoids improved significantly after Notum deletion (Fig. 3b, Extended Data Fig. 7d). Conversely, activation of endogenous Notum expression by CRISPR activation decreased Wnt signalling and colony-forming capacity of CD24medSSClo cells containing the ISCs of the targeted organoids (Extended Data Fig. 7e–g). Finally, to test whether the regenerative capacity of old intestines can be increased via the intestinal stem cell niche, we used ABC9925, a small-molecule inhibitor of Notum. ABC99 blunted the effects of exogenous Notum and increased the frequency of Lgr5hi cells in vitro (Extended Data Fig. 8a, b), and in vivo treatment of mice by intraperitoneal injection with 10 mg per kg (body weight) ABC99 had no noticeable adverse effects (Extended Data Fig. 8c). Of note, the Lgr5hi cells that were isolated from old mice after seven days of in vivo treatment with ABC99 demonstrated colony-forming capacity comparable to cells from untreated young mice (Fig. 3c). Moreover, stem-cell-supporting function of Paneth cells was also restored, which suggests autocrine regulation (Extended Data Fig. 8d). To address whether Notum modulates Wnt activity of ISCs in vivo, we next compared the nuclear β-catenin levels of ISCs between Paneth cells to those of more-differentiated transit-amplifying cells that are not in contact with Notum-producing Paneth cells (Extended Data Fig. 8e). As expected, untreated old ISCs had reduced nuclear β-catenin levels (Fig. 3d). ABC99 increased the nuclear β-catenin levels of ISCs specifically in old mice (Fig. 3d, Extended Data Fig. 8f). This increased Wnt activity in old stem cells was reflected in increased proliferation, specifically in Olfm4+ stem and progenitor cells, in comparison to more differentiated transit-amplifying cells (Fig. 3e). To formally test whether Notum inhibition promotes regeneration of old intestine, we analysed how advance Notum inhibition affects recovery from 5-fluorouracil (5-FU) chemotherapy-induced mucositis26,27 that results in loss of body weight owing to compromised water retention and nutrient intake27. We treated mice with 100 mg per kg (body weight) 5-FU, as the weight of young mice recovered fully within 5 days of this dose, but old mice did not recover (Extended Data Fig. 9a). When Notum activity was inhibited with ABC99 for eight days before 5-FU treatment, weight loss in old mice was significantly reduced (Fig. 3f, Extended Data Fig. 9b). Moreover, the density of differentiated cells in the villi was restored to a level similar to that in the young mice, indicating increased regeneration by old stem cells (Fig. 3g). These data demonstrate that Notum produced by Paneth cells attenuates the regenerative capacity of aged intestinal epithelium in vivo by reducing Wnt activity specifically in stem cells.
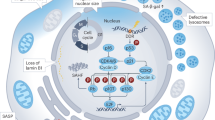
a, Growth of CRISPR-targeted organoids after orthotopic transplantation. Notum-knockout (KO) organoids were co-transplanted with competing scramble-targeted controls (Scr). (n = 8 mice transplanted). Scale bar, 1 mm. WT, wild type. b, Regenerative growth of CRISPR-targeted small-intestinal organoids from young and old mice (n = 4 mice per group). Student’s paired t-test. c, Clonogenic growth of Lgr5hi stem cells from ABC99-treated mice. Control mice received an equal amount of the inactive analogue ABC101 (yellow circles) or vehicle (number of analysed mice shown). d, Quantification of relative nuclear β-catenin intensity of crypt base columnar cells (number of analysed mice shown). e, Quantification of EdU+ cellular frequencies within the crypt, indicating Olfm4+ cells (number of analysed mice shown). f, Average weight change after 5-FU treatment for five days (number of analysed mice shown). g, Left, representative images of haematoxylin and eosin (H&E) staining of villi after 5-FU treatment for five days. Scale bar, 10 μm. Right, quantification of cellular density in ileal villi (cells per micrometre; number of analysed mice shown). h, Schematic of the model for stem cell maintenance by Paneth cells in the aged niche. Bold weighting of arrows, lines and text indicates a higher strength of the biological signal. In these experiments, old mice were more than 21 months old. Other than in box plots, data are mean ± s.d.; two-tailed unpaired Student’s t-test; P values shown in corresponding panels. P <0.05 is considered significant.
Appropriate Wnt levels are crucial for many stem cell compartments and alterations are seen in multiple pathologies28. Stromal Wnt signals have previously been shown to maintain the epithelial stem cell pool in the absence of Paneth cells under normal tissue homeostasis17,18,29. However, recent studies underline the importance of epithelial Wnt signalling in regeneration following injury30. Here we find that, during ageing, increased Notum expression in Paneth cells of the ISC niche in mouse and human inhibits Wnt signalling and reduces stem cell maintenance and regeneration. Simultaneously, reversing observed changes in mTORC1–PPAR-α signalling restored epithelial regeneration. Our findings underscore the importance of niche-regulated Wnt signals in promoting stemness and demonstrate a link between ageing-associated metabolic changes and tissue maintenance. Such effects could be missed by studies that demonstrate unaltered clonal dynamics of crypts during ageing31. Because Wnt–β-catenin signalling can modulate fatty acid oxidation32, increasing Wnt activity in old stem cells could also help to restore the age-induced decline in fatty acid oxidation14. However, further studies are required to address whether the mechanisms described here also affect tumour risk in the old intestine33. In any case, niche–stem cell interactions could provide safer strategies to target tissue renewal and age-related decline than strategies directly targeting stem cells. Activation of PPAR-α or PPAR-δ signalling is not an attractive option in this regard, as PPAR-δ was recently demonstrated to confer tumour-initiating capacity to non-stem cells in the intestine34. Notum inhibition with selective inhibitors, such as the ABC99 used here, may represent a safer way to treat gastrointestinal complications and reduce harmful side-effects of chemotherapeutic agents that pose a particular challenge for older individuals35.
Methods
Isolation of mouse small-intestinal crypts
Mouse small-intestinal crypts were isolated as previously described12. In brief, mouse small intestines were flushed with cold PBS and opened, and mucus was removed. The intestine was cut into small fragments and incubated with several changes of 10 mM EDTA in PBS on ice for 2 h. Epithelium was detached by vigorous shaking. To enrich crypts, tissue suspension was filtered through 70-μm nylon mesh. Enriched crypts were washed once with cold PBS and plated into 60% Matrigel (BD Biosciences) with ENR medium. Y-27632 (10 μM) was added to the medium for the first two days.
Isolation of human colonic crypts
Crypts from human colonic biopsies were isolated by vigorous shaking after 1-h incubation in ice cold PBS with 10 mM EDTA. To enrich crypts, tissue suspension was filtered through 70-μm nylon mesh. Enriched crypts were washed once with cold PBS and plated into 60% Matrigel (BD Biosciences) and cultured as previously described36.
Organoid culture
Crypts were plated (50–200 crypts per 20 μl drop of 60% Matrigel) and overlaid with ENR medium (Advanced DMEM/F12 (Gibco), 1× Glutamax (Gibco), 100 U/ml penicillin and streptomycin, 10 mM Hepes, 1× B-27 (Gibco), 1× N-2 (Gibco), 50 ng/ml of mouse EGF (RnD), 100 ng/ml noggin (Peprotech), 500 ng/ml of R-spondin-1 (RnD), 1 μM N-acetyl-l-cysteine (Sigma-Aldrich)). Y-27632 (10 μM) was added for the first two days of culture. Organoid starting frequency was counted after two days of culture unless otherwise stated in the figure legend. Primary organoids were cultured for five-to-nine days, after which regenerative growth (number of de novo crypt domains per organoid) was quantified and organoids were subcultured. Subculturing was performed by mechanically disrupting organoids to single-crypt fragments, which were replated (1:3) with fresh Matrigel. Secondary cultures were confirmed to start from single crypt domains by inspection, and their survival and de novo crypt number were quantified two days after replating. When indicated, ENR medium was supplemented with rapamycin (CST), GW6417 (Tocris), CHIR99021 (BioVision), IWP-237 (Sigma) or Wnt3A (RnD). Equal amounts of vehicle (ethanol or DMSO) was used in controls. ENR supplemented with 10 nM gastrin (Sigma-Aldrich), 100 ng/ml Wnt3A (RnD), 10 mM nicotinamide (Sigma-Aldrich), 500 nM A-83-01 (Sigma-Aldrich) and 10 μM SB202190 (Sigma-Aldrich) was used for isolated human colonic crypts36. Colonic organoid starting frequency was counted on day seven.
Single-cell sorting and analysis
To isolate single cells, isolated crypts or grown organoids were dissociated in TrypLE Express (Gibco) with 1,000 U/ml of DnaseI (Roche) at 32 °C (90 s for crypts, 5 min at 37 °C for cultured organoids). Cells were washed and stained with the following antibodies: CD31–PE (Biolegend, Mec13.3), CD45–PE (eBioscience, 30-F11), Ter119–PE (Biolegend, Ter119), EpCAM–APC (eBioscience, G8.8) and CD24–Pacific Blue (Biolegend, M1/69), all at 1:500. Finally, cells were resuspended in SMEM medium (Sigma) supplemented with 7-AAD (Life) (2 μg/ml) for live-cell separation. Cells were sorted using a FACSAria II or FACSAria Fusion (BD Biosciences). Sorting strategies: intestinal stem cells, Lgr5–eGFPhiEpcam+CD24med(or −)CD31−Ter119 −CD45−7-AAD−; Paneth cells, CD24hiSideScatterhiLgr5–eGFP−Epcam+CD31−Ter119−CD45−7-AAD−; enteroendocrine cells, CD24hiSideScatterloLgr5–eGFP−Epcam+CD31−Ter119−CD45−7-AAD−. When analysing organoids, eGFP gates were applied directly on the Epcam+CD31−Ter119−CD45−7-AAD− population. Equal numbers of Lgr5hi and Paneth cells were co-cultured with ENR medium supplemented with additional 500 μg/ml of R-spondin-1 (to yield a final concentration of 1 μg/ml), 100 ng/ml Wnt3A and 10 μM of Jagged-1 peptide (Anaspec) for the first six days. Y-27632 (10 μM) was added to the medium for the first four days. Single-cell starting frequency and clonogenic growth of primary organoids were analysed at days 5–9. Long-term organoid-forming capacity was quantified from twice-subcultured organoids 21 days after isolation. Culture of isolated Lgr5hi or CD24medSidescatterlo cells without Paneth cells and Wnt ligands was performed in ENR supplemented with 10 μM Chir99021, 10 μM Y-27632 and 1 μg/ml recombinant human Notum (RnD) and/or 50–500 nM ABC9925 when indicated for the first 5 days followed by culture in regular ENR medium. Colony-forming capacity was quantified on day 5 or day 7 as indicated in the figure legends. Cross-sectional area of colonies was quantified from bright-field images taken with an inverted cell culture microscope (Nikon TS100 Eclipse, DS-Qi1Mc camera) on day 7. Paneth cells from mT/mG mouse-derived organoids were isolated as CD24hiSideScatterhiTomato+Epcam+7-AAD− and co-cultured with freshly isolated Lgr5hi stem cells. Cell population analysis was performed with FlowJo software.
CRISPR–Cas9 gene editing of intestinal organoids
Guide RNAs for the target-gene knockout38 were designed with the CRISPR design tool (http://crispr.mit.edu). Guides were cloned into lentiCRISPR v2 vector. Lentiviral vectors were produced in 293fT cells (ThermoFisher, R70007) and concentrated with Lenti-X concentrator (Clontech). The 293fT cell line was not authenticated in the laboratory, but tested negative for mycoplasma. Cultured intestinal organoids were exposed to 1 mM nicotinamide for 2 days before they were processed for transduction. Organoids were mechanically disrupted and dissociated to small fragments with TrypLE Express supplemented with 1,000 U/ml DnaseI for 5 min at 32 °C. Fragments were washed once with SMEM medium and resuspended to transduction medium (ENR medium supplemented with 8 μg/ml polybrene (Sigma-Aldrich), 1 mM of nicotinamide, 10 μM Y-27632) and mixed with concentrated virus. Samples were spinoculated 1 h at 600g 32 °C followed by 2–4 h incubation at 37 °C, after which they were collected and plated on 60% Matrigel overlaid with transduction medium without polybrene. Two to three days after transduction, infected clones were selected by adding 2 μg/ml of puromycin (Sigma-Aldrich) to the medium. Four days after selection, clones that survived were expanded in normal ENR medium and clonogenic growth was assessed. Knockout was confirmed by three-primer PCR around the gRNA-target site. In experiments comparing young and old gene-edited organoids, organoids were for a maximum of seven days before transduction. LentiCRISPR v2 was a gift from F. Zhang (Addgene plasmid 52961)39.
Oligonucleotides used for generation of gRNAs: Notum (1), CACCGGGCGGGGCTGCCGTCATTGC, AAACGCAATGACGGCAGCCCCGCCC; Notum (2), CACCGTCGGCGGTGGTTACTCTTTC, AAACGAAAGAGTAACCACCGCCGAC; Bst-1, CACCGTTCTGGGGGCAAGAGCGCGG, AAACCCGCGCTCTTGCCCCCAGAAC; Scramble (1), CACCGCTAAAACTGCGGATACAATC, AAACGATTGTATCCGCAGTTTTAGC; Scramble (2), CACCGAAAACTGCGGATACAATCAG, AAACCTGATTGTATCCGCAGTTTTC. Oligonucleotides used for confirming gene-editing: Notum (1), TATGGCGCAAGTCAAGAGCC, CACGTCGGTGACCTGCAATG, CAAGCCAGGTTGACGGCCT; Notum (2), CGGTTTGGGGATGAGGGTAG, GTCGGCGGTGGTTACTCTTT, GCCAGTCTTTGGAGCTCAT; Bst-1, CCACGGGCTAGAGGAATCAA, GCAAGAGCGCGGTGGAC, CTCAGCAGCGTGGTGTACT.
CRISPR–Cas9 gene activation of intestinal organoids
Lenti-SAM-Cre vector40 was constructed by assembling five DNA fragments with overlapping ends using Gibson Assembly. In brief, fragments containing sequences corresponding to U6-sgRNA-MS2 (PCR amplified from lenti-sgRNA(MS2)-zeo, Addgene plasmid 61427), the PGK promoter, MS2-p65-HSF1-T2A (PCR amplified from lenti-MS2-P65-HSF1-Hygro, Addgene plasmid 61426), and T2A-Cre were assembled by Gibson assembly into a lentiviral backbone following the manufacturer’s guidelines. For short guide RNA (sgRNA) cloning, the Lenti-SAM-Cre vector was digested with BsmBI and ligated with BsmBI-compatible annealed oligonucleotides. sgRNAs were designed using the Cas9 activator tool (http://sam.genome-engineering.org). At least five nucleotides were removed from the 5′ end of candidate sgRNAs to enable use of the SAM system with nuclease-active Cas941. If the first nucleotide in the truncated sgRNA sequence was not a G, an additional nucleotide was removed and replaced with a G to enable efficient expression of the sgRNA from the U6 promoter. Sequence against tdTomato was used as a control42. LSL-Cas9-eGFP mouse-derived small-intestinal organoids were infected with Lenti-SAM-Cre-derived virus. Cells with successfully integrated constructs were selected by sorting GFP+ cells from organoid cultures. Organoids were grown in ENR containing 3 μM CHIR99021 to avoid selection against Notum expression. Activation of Notum expression was confirmed by quantitative PCR with reverse transcription (RT–qPCR) analysis from whole organoids cultured for two days in ENR without CHIR99021. For assessing the effect of endogenous Notum on stem cells, CD24medSidescatterlo cells were sorted from organoids cultured for 4–5 days in ENR without CHIR99021.
sgRNA sequences used for generating Lenti-SAM-Cre vectors: Notum (dANotum), GCTGGCCGCGGAGAA; tdTomato (dATom), CGAGTTCGAGATCGA.
RT–qPCR
RNA from crypts, single cells and cultured organoids was isolated by Trizol purification according to the manufacturer’s instructions (Life Technologies) using glycogen as co-precipitant (Life Technologies). Full tissue samples were shredded with ceramic beads (Precellys) in RLT buffer and RNA was isolated by RNAeasy+ kit (Qiagen) according to the manufacturer’s instructions. Isolated RNA was transcribed with cDNA synthesis kit using OligodT primers (Molecular probes). qPCR amplification was detected by the SYBRGreen (2× SYBRGreen mix, Applied Biosciences) method. Samples were run as triplicates and genes of interest were normalized to Gapdh or Actb. Primers used for qPCR: Actb, CCTCTATGCCAACACAGTGC, CCTGCTTGCTGATCCACATC; Gapdh, ATGGTGAAGGTCGGTGTGAA, TGGAAGATGGTGATGGGCTT; Notum, CTGCGTGGTACACTCAAGGA, CCGTCCAATAGCTCCGTATG; Bst1, ACCCCATTCCTAGGGACAAG, GCCTCCAATCTGTCTTCCAG; Cd44, GCACTGTGACTCATGGATCC, TTCTGGAATCTGAGGTCTCC; Myc, CAAATCCTGTACCTCGTCCGATTC, CTTCTTGCTCTTCTTCAGAGTCGC; Ascl2, CTACTCGTCGGAGGAAAG, ACTAGACAGCATGGGTAAG; Lgr5, ACCCGCCAGTCTCCTACATC, GCATCTAGGCGCAGGGATTG; Axin2, AGTGCAAACTCTCACCCACC, TCGCTGGATAACTCGCTGTC; Wnt2b, CGTGTAGACACGTCCTGGTG, GTAGCGTTGACACAACTGCC; Wnt3, TGGAACTGTACCACCATAGATGAC, ACACCAGCCGAGGCGATG; Wnt4, GTACCTGGCCAAGCTGTCAT, CTTGTCACTGCAAAGGCCAC; Wnt5a, ATGAAGCAGGCCGTAGGAC, CTTCTCCTTGAGGGCATCG; Rnf43, CACCATAGCAGACCGGATCC, TATAGCCAGGGGTCCACACA; Sox9, GAGCCGGATCTGAAGAGGGA, GCTTGACGTGTGGCTTGTTC.
RNA sequencing and data processing
Total RNA from sorted Paneth (four young and five old biological replicates) and Lgr5hi (three young and three old biological replicates) cells were isolated by Trizol purification. Samples were first treated with HL-dsDNase (Articzymes 80200-050) to remove residual DNA. An Ovation Universal RNA-Seq System kit was used for Illumina library preparations (NuGEN Technologies). Purified total RNA (100 ng) was used and primers for ribosomal removal were designed and used as outlined in the kit manual. Libraries were purified with AMPure XP beads (Beckman Coulter), quantified and run on a NextSeq 500 sequencer using 75b single-read kits (Illumina). Adaptor sequences and low-quality reads were removed from the data using cutadapt43. The data were mapped to Mus musculus genome GRCm38.p4 using STAR44. Count data were processed using GenomicFeatures and GenomicAlignments45, and the differential expression analysis was carried out using DESeq246 in R. PreRanked GSEA analysis (http://software.broadinstitute.org/gsea/index.jsp) was performed for fold-change ranked genes with 1,000 permutations47. Hallmark, Biocarta and Kyoto Encyclopedia of Genes and Genomes (KEGG) gene sets are available via GSEA Molecular Signatures Database. The ‘PPARd’ gene set was adopted from a previous publication48. Gene Ontology enrichment analysis was done for the significantly (adjusted P <0.1) altered genes with Gene Ontology Consortium Enrichment analysis (http://geneontology.org/page/go-enrichment-analysis), using Fisher’s exact test not corrected for multiple testing. Subcellular localization for significantly altered genes was taken from the Uniprot database (http://www.uniprot.org/). Putative transcription factor binding sites for PPAR-α on mouse and human NOTUM genes were found by using the DECODE database (SABiosciences) and confirmed for mouse using the JASPAR database using PPRE motif (PPARG; RXRA) (http://jaspar.genereg.net). RNA sequencing data from human terminal ileal samples were obtained from the GTEx Portal at the Human Proteome Atlas (https://proteinatlas.org) on 1 June 2018. Sex-matched samples (51 males) were divided into three age groups (20–39, 40–59, >60 years old) and proportions of expression presented. The data are publicly available through ArrayExpress under accession code: E-MTAB-7916 (http://www.ebi.ac.uk/arrayexpress/experiments/E-MTAB-7916).
Immunoblotting
Isolated crypts and cells were lysed in RIPA buffer with 1× Halt Protease inhibitor cocktail (ThermoFisher Scientific) and 1× PhosStop (Roche) phosphatase inhibitors. Protein concentrations of cleared lysates were measured by BCA Protein Kit (ThermoFisher Scientific). For sorted cells equal loading was adjusted by sorting the same number of cells. Samples were run on 4–12% Bis-Tris protein gels (Life Technologies) and blotted on nitrocellulose membranes. Membranes were incubated with primary antibodies: pS6 (Ser240/244, CST,5364 for Fig. 2h and Extended Data Figs. 4e, 5d; 1:1,000), pS6 (Ser235/236, CST, 4858; 1:500 for Extended Data Fig. 5k), S6 (CST, 2217; 1:500), H3 (CST, 4499; 1:1,000), β-actin (CST, 4967; 1:2,000), α-tubulin (CST, 2144; 1:1,000) and pS6K (ImmunoWay,YP0886; 1:500) at 4 °C overnight, and HRP-conjugated anti-rabbit (Sigma-Aldrich; 1:5,000) or anti-mouse (CST; 1:1,000) for 1 h at room temperature. Signal was detected using ECL reagent Supersignal West Femto (ThermoFisher Scientific). Densitometry was performed with ImageJ, normalizing to β-actin or α-tubulin.
Immunohistochemistry and immunofluorescence
Tissues were fixed in 4% PFA, paraffin-embedded and sectioned. Antigen retrieval was performed by boiling in pH 6 citrate buffer (Sigma-Aldrich) for 5 min. Antibodies: lysozyme (DAKO, EC3.2.1.17; 1:750), Ki67 (Abcam, ab15580; 1:300), pS6 (Ser240/244) (CST, 5364; 1:1,000), Olfm4 (clone PP7, a gift from CST (Extended Data Fig. 1k), CST, 39141 (Extended Data Fig. 8g); 1:300), β-catenin (BD, 610153; 1:300), E-cadherin (BD, 610181; 1:500). Antigen retrieval was followed by permeabilization with 0.5% Triton-X100 (Sigma) and, in the case of analysis of EdU incorporation, was followed by EdU Click-IT chemistry according to manufacturer’s instructions (ThermoFisher Scientific). Primary antibodies were detected with biotin-conjugated secondary antibodies and DAB substrate on peroxidase-based system (Vectastain Elite ABC, Vector Labs). For immunofluorescence, Alexa Fluor 488-, Alexa Fluor 594-, Alexa Fluor 633- and Alexa Fluor 647-conjugated anti-rabbit or anti-mouse secondary antibodies (Life Technologies, all 1:500) were used. Nuclei were co-stained with DAPI (Life Technologies, 1 μg/ml) or Hoechst 33342 (Life Technologies, 1 μg/ml).
Immunocytochemistry
Sorted cell populations were treated as previously described12. In brief, they were either centrifuged onto charged microscope slides with a Shandon Cytospin 4 (ThermoFisher) for 3 min at 800 r.p.m. or allowed to settle on poly-l-lysine-coated coverglass-bottomed MatTek dishes for 15 min at 37 °C followed by fixation with 4% PFA and immunostaining. Antibodies: lysozyme (DAKO, EC3.2.1.17; 1:500), Muc2 (SantaCruz,H-300; 1:50), counterstains: Hoechst 33342 (Life Technologies, 1 μg/ml), Phalloidin-647 (Life Technologies; 1:50).
Quantification of nuclear β-catenin localization
Three-micrometre-thick confocal sections of β-catenin-stained ileal segments were captured with a Leica SP5IIHCS confocal microscope and 63× water-immersion objective and 12-bit image colour depth. β-catenin mean fluorescence intensity was measured by a blinded investigator from three nuclear regions of interest of cells in the transit-amplifying zone (cell position +6 and above relative to crypt bottom) followed by measurement of intensities in the nuclei of crypt base columnar cells (CBCs) and Paneth cells (identified by nuclear morphology and cellular shape). Nuclear intensities of CBCs and Paneth cells were always normalized to transit-amplifying cells from the same image.
RNA in situ hybridization
RNA in situ hybridization was performed with RNAScope 2.5HD Assay–Brown according to the manufacturer’s protocol (RNAScope ACDBio). Probes used: mouse Notum: Mm-Notum 428981; mouse (positive control) Lgr5: Mm-Lgr5 312178; Human NOTUM: Hs-NOTUM 430311; human UBC (positive control): Hs-UBC 310041; Dapb (negative control): Probe-DapB 310043.
Of note, human NOTUM was only detected from samples freshly fixed in 4% PFA followed by paraffin-embedding. Pathological samples fixed in 10% NBF were not compatible with this probe.
Statistical analysis
No statistical method was used to calculate the sample size. For analysis of in vitro organoid cultures, investigators were blinded when possible, but owing to features of co-culture experiments this was not always possible. Blinded investigators performed all histological quantification. Microsoft Excel v.16.16.8 and Graphpad Prism v.8.0.0 were used for statistical analysis and visualization of data. All data were analysed by two-tailed Student’s t-test, except RNA sequencing data (see ‘RNA sequencing and data processing’), exact P values are presented in the corresponding figures. A paired t-test was applied if the day of organoid growth quantification varied between pairs (samples processed the same day were paired) or phenotype after treatment was compared to the control from the same mouse (samples from the same mouse were paired). Whether a test was paired or unpaired is noted in the figure legends. P <0.05 was considered significant.
Human biopsy samples
Human ileal and colon tissue biopsies were obtained from 24 healthy subjects who were undergoing a routine colonoscopy. Human jejunal samples were obtained from patients undergoing Roux-en-Y gastric bypass surgery and fixed in 4% PFA before a routine paraffin-embedding protocol. The specimens used for organoid functional assay were stored in normal saline on ice until analysis. Exclusion criteria included any history of malignancy, chronic liver disease, history suggesting a malabsorption disorder, previous intestinal surgery, renal disease, bleeding disorder that would preclude biopsy, active infection or systemic inflammatory disorder. The study regarding relevant samples and associated ethical regulations were approved by the institutional review board of Massachusetts General Hospital and Helsinki University Hospital. Written and informed consent was obtained before enrolment.
Mice
Lgr5-eGFP-IRES-creERT2 mice1 were maintained in a C57BL/6J background. Rosa26mTmG (JAX 007576), Tsc1fl/fl (ref. 49) (Tsc1tm1Djk/J, JAX 005680), Rosa26LSL-ZsGreen (JAX 007906), Rosa26LSL-TdTomato (JAX 007909), Rag2−/− (B6(Cg)-Rag2tm1.1Cgn/J, JAX 008449) and Rosa26LSL-Cas9-eGFP (JAX 024857) mice were obtained from Jackson Laboratories and were on mixed background. Villin-creERT2 mice were a gift from S. Robine and have previously been described50. All animal housing and experiments were done under local institutional regulations. Mice were allocated to experimental groups randomly, but without proper randomization. Investigators were not blinded owing to the apparent phenotype of aged mice. For in vivo proliferation analysis, 10 mg/kg of EdU (Sigma) in PBS was injected intraperitoneally 2 h before mice were euthanized. For in vivo Tsc1 deletion, Villin-creERT2;Tsc1fl/fl mice were given 5 intraperitoneal injections of 100 mg/kg tamoxifen (Sigma) on alternate days. Rapamycin treatment was performed as previously described12. ABC99 was produced as previously described25: 33.3 mg/ml stock solution in ethanol was prepared freshly and further mixed 1:1:1:17 into Tween-80 (Sigma), PEG-400 (HamiltonResearch) and 0.9% NaCl. Mice were injected intraperitoneally with 10 mg/kg ABC99 daily with a last dose together with 10 mg/kg EdU 2 h before they were euthanized. Control mice were treated with vehicle or an equal amount of the inactive control compound ABC10125. 5-FU (Sigma) was reconstituted in DMSO at 100 mg/ml and a single intraperitoneal injection was given to mice with a dose of 100–200 mg/kg (as described in the figure legends). Mice over 24 months of age were considered old, and mice between 3 and 9 months of age were considered young (denoted ‘O’ and ‘Y’, respectively, throughout the figure legends), with the exception of Fig. 3f, g, Extended Data Fig. 9a, b and Supplementary Fig. 2, in which old mice were 20–22 months of age. Both sexes were used in all experiments. All animal experiments were approved and carried out in accordance with the guidelines of the Finnish National Animal Experimentation Board and the Committee on Animal Care at MIT.
Organoid transplantation
Notum wild-type and Notum knockout intestinal organoids were generated using Notum (2) guide RNAs, as described above, in Villin-creERT2;Rosa26LSL-ZsGreen and Villin-creERT2;Rosa26LSL-tdTomato intestinal organoids cultured with 4-OHT to generate zsGreen+ wild-type, zsGreen+ knockout, tdTomato+ wild-type and tdTomato+ knockout organoids. Organoids were grown in Matrigel and cultured with crypt medium. Before transplantation, ZsGreen+ knockout and tdTomato+ wild-type (and, in parallel, ZsGreen+ wild-type and tdTomato+ knockout) organoids were chemically dissociated using Cell Recovery Solution (Corning, 354253), and then resuspended in a 1:1 ratio in 90% crypt medium and 10% Matrigel at a concentration of 25 organoids per μl. Organoids were orthotopically transplanted into the colonic submucosa of Rag2−/− recipient mice, as previously described51,52. The average volume of each injection was 60 μl. Eight weeks later, engrafted organoids were assessed using fluorescence colonoscopy followed by fluorescence microscopy using GFP and tdTomato filters. Tissues were then fixed in 4% paraformaldehyde for 4–6 h, cryopreserved with 30% sucrose in PBS overnight, and then frozen in OCT. Frozen tissue sections were stained with DAPI to visualize nuclei, and then imaged for tdTomato and GFP. The total number of tdTomato+ and GFP+ cells per mouse was then counted using Fiji.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this paper.
Data availability
RNA sequencing data are publicly available through ArrayExpress with accession code E-MTAB-7916. Source Data for Fig. 1–3 and Extended Data Figs. 1–9 are available with the online version of the paper. All other data are available from the corresponding author upon reasonable request.
References
Barker, N. et al. Identification of stem cells in small intestine and colon by marker gene Lgr5. Nature 449, 1003–1007 (2007).
Sato, T. et al. Paneth cells constitute the niche for Lgr5 stem cells in intestinal crypts. Nature 469, 415–418 (2011).
Sengupta, S., Peterson, T. R., Laplante, M., Oh, S. & Sabatini, D. M. mTORC1 controls fasting-induced ketogenesis and its modulation by ageing. Nature 468, 1100–1104 (2010).
Molofsky, A. V. et al. Increasing p16INK4a expression decreases forebrain progenitors and neurogenesis during ageing. Nature 443, 448–452 (2016).
Rossi, D. J. et al. Deficiencies in DNA damage repair limit the function of haematopoietic stem cells with age. Nature 447, 725–729 (2007).
Conboy, I. M. & Rando, T. A. Heterochronic parabiosis for the study of the effects of aging on stem cells and their niches. Cell Cycle 11, 2260–2267 (2012).
Sato, T. et al. Single Lgr5 stem cells build crypt–villus structures in vitro without a mesenchymal niche. Nature 459, 262–265 (2009).
Warren, P. M., Pepperman, M. A. & Montgomery, R. D. Age changes in small-intestinal mucosa. Lancet 312, 849–850 (1978).
Feibusch, J. M. & Holt, P. R. Impaired absorptive capacity for carbohydrate in the aging human. Dig. Dis. Sci. 27, 1095–1100 (1982).
Feldman, M., Cryer, B., McArthur, K. E., Huet, B. A. & Lee, E. Effects of aging and gastritis on gastric acid and pepsin secretion in humans: a prospective study. Gastroenterology 110, 1043–1052 (1996).
Potten, C. S., Martin, K. & Kirkwood, T. B. Ageing of murine small intestinal stem cells. Novartis Found Symp. 235, 66–79 (2001).
Yilmaz, O. H. et al. mTORC1 in the Paneth cell niche couples intestinal stem-cell function to calorie intake. Nature 486, 490–495 (2012).
Nalapareddy, K. et al. Canonical Wnt signaling ameliorates aging of intestinal stem cells. Cell Rep. 18, 2608–2621 (2017).
Mihaylova, M. M. et al. Fasting activates fatty acid oxidation to enhance intestinal stem cell function during homeostasis and aging. Cell Stem Cell 22, 769–778 (2018).
Giráldez, A. J., Copley, R. R. & Cohen, S. M. HSPG modification by the secreted enzyme Notum shapes the Wingless morphogen gradient. Dev. Cell 2, 667–676 (2002).
Kakugawa, S. et al. Notum deacylates Wnt proteins to suppress signalling activity. Nature 519, 187–192 (2015).
Shoshkes-Carmel, M. et al. Subepithelial telocytes are an important source of Wnts that supports intestinal crypts. Nature 557, 242–246 (2018).
Degirmenci, B., Valenta, T., Dimitrieva, S., Hausmann, G. & Basler, K. GLI1-expressing mesenchymal cells form the essential Wnt-secreting niche for colon stem cells. Nature 558, 449–453 (2018).
Farin, H. F., Van Es, J. H. & Clevers, H. Redundant sources of Wnt regulate intestinal stem cells and promote formation of Paneth cells. Gastroenterology 143, 1518–1529 (2012).
Farin, H. F. et al. Visualization of a short-range Wnt gradient in the intestinal stem-cell niche. Nature 530, 340–343 (2016).
McCay, C. M., Maynard, L. A., Sperling, G. & Barnes, L. L. Retarded growth, life span, ultimate body size and age changes in the albino rat after feeding diets restricted in calories. Nutr. Rev. 33, 241–243 (1975).
Harrison, D. E. et al. Rapamycin fed late in life extends lifespan in genetically heterogeneous mice. Nature 460, 392–395 (2009).
Lamming, D. W. et al. Rapamycin-induced insulin resistance is mediated by mTORC2 loss and uncoupled from longevity. Science 335, 1638–1643 (2012).
Naillat, F. et al. Identification of the genes regulated by Wnt-4, a critical signal for commitment of the ovary. Exp. Cell Res. 332, 163–178 (2015).
Suciu, R. M., Cognetta, A. B., III, Potter, Z. E. & Cravatt, B. F. Selective irreversible inhibitors of the Wnt-deacylating enzyme NOTUM developed by activity-based protein profiling. ACS Med. Chem. Lett. 9, 563–568 (2018).
Longley, D. B., Harkin, D. P. & Johnston, P. G. 5-fluorouracil: mechanisms of action and clinical strategies. Nat. Rev. Cancer 3, 330–338 (2003).
Song, M. K., Park, M. Y. & Sung, M. K. 5-fluorouracil-induced changes of intestinal integrity biomarkers in BALB/c mice. J. Cancer Prev. 18, 322–329 (2013).
Nusse, R. & Clevers, H. Wnt/β-catenin signaling, disease, and emerging therapeutic modalities. Cell 169, 985–999 (2017).
Kim, T. H., Escudero, S. & Shivdasani, R. A. Intact function of Lgr5 receptor-expressing intestinal stem cells in the absence of Paneth cells. Proc. Natl Acad. Sci. USA 109, 3932–3937 (2012).
Zou, W. Y. et al. Epithelial WNT ligands are essential drivers of intestinal stem cell activation. Cell Rep. 22, 1003–1015 (2018).
Kozar, S. et al. Continuous clonal labeling reveals small numbers of functional stem cells in intestinal crypts and adenomas. Cell Stem Cell 13, 626–633 (2013).
Frey, J. L. et al. Wnt–Lrp5 signaling regulates fatty acid metabolism in the osteoblast. Mol. Cell. Biol. 35, 1979–1991 (2015).
Huels, D. J. et al. Wnt ligands influence tumour initiation by controlling the number of intestinal stem cells. Nat. Commun. 9, 1132 (2018).
Beyaz, S. et al. High-fat diet enhances stemness and tumorigenicity of intestinal progenitors. Nature 531, 53–58 (2016).
Chang, S., Goldstein, N. E. & Dharmarajan, K. V. Managing an older adult with cancer: considerations for radiation oncologists. BioMed Res. Int. 2017, 1695101 (2017).
Sato, T. et al. Long-term expansion of epithelial organoids from human colon, adenoma, adenocarcinoma, and Barrett’s epithelium. Gastroenterology 141, 1762–1772 (2011).
Chen, B. et al. Small molecule-mediated disruption of Wnt-dependent signaling in tissue regeneration and cancer. Nat. Chem. Biol. 5, 100–107 (2009).
Shalem, O. et al. Genome-scale CRISPR–Cas9 knockout screening in human cells. Science 343, 84–87 (2014).
Sanjana, N. E., Shalem, O. & Zhang, F. Improved vectors and genome-wide libraries for CRISPR screening. Nat. Methods 11, 783–784 (2014).
Konermann, S. et al. Genome-scale transcriptional activation by an engineered CRISPR–Cas9 complex. Nature 517, 583–588 (2015).
Dahlman, J. E. et al. Orthogonal gene knockout and activation with a catalytically active Cas9 nuclease. Nat. Biotechnol. 33, 1159–1161 (2015).
Sánchez-Rivera, F. J. et al. Rapid modelling of cooperating genetic events in cancer through somatic genome editing. Nature 516, 428–431 (2014).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.journal 17, 17.1.2001 (2011).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Lawrence, M. et al. Software for computing and annotating genomic ranges. PLOS Comput. Biol. 9, e1003118 (2013).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Adhikary, T. et al. Genomewide analyses define different modes of transcriptional regulation by peroxisome proliferator-activated receptor-β/δ (PPARβ/δ). PLoS ONE 6, e16344 (2011).
Kwiatkowski, D. J. et al. A mouse model of TSC1 reveals sex-dependent lethality from liver hemangiomas, and up-regulation of p70S6 kinase activity in Tsc1 null cells. Hum. Mol. Genet. 11, 525–534 (2002).
el Marjou, F. et al. Tissue-specific and inducible Cre-mediated recombination in the gut epithelium. Genesis 39, 186–193 (2004).
Roper, J. et al. Colonoscopy-based colorectal cancer modeling in mice with CRISPR-Cas9 genome editing and organoid transplantation. Nat. Protoc. 13, 217–234 (2018).
Roper, J. et al. In vivo genome editing and organoid transplantation models of colorectal cancer and metastasis. Nat. Biotechnol. 35, 569–576 (2017).
Acknowledgements
This study was supported by the Academy of Finland (Research Fellow and Centre of Excellence, MetaStem), Marie Curie CIG (618774), ERC-STG (677809), Swedish Research Council 2018-03078, Sigrid Juselius Foundation, Center for Innovative Medicine, and Wallenberg Academy Fellows program to P.K. N.P. was supported by the Integrative Life Science Doctoral program and by the Research foundation of University of Helsinki. B.F.C. was supported by grant R35 CA231991. We thank the personnel of the DNA sequencing and genomics laboratory for performing the RNA sequencing assays. We thank J. Bärlund, A. Sola-Carvajal, M. Simula and A. Kegel for technical assistance.
Author information
Authors and Affiliations
Contributions
N.P. and P.K. designed and interpreted the results of all experiments. N.P., S.I., M.M., S.A., A.B.C. III, R.M.S., K.L., E.M., S.G., S.N. and T.T. performed all experiments and analysed the results. J.R. performed and analysed organoid transplantations. O.-P.S. processed and analysed the RNA sequencing data with the help of P.A., P.K. and N.P. N.P., M.M., K.L., E.M., S.A. and S.G. performed and analysed the immunohistochemistry, immunofluorescence and RNA in situ hybridizations. N.G., T.S., A.J., K.P. and A.R. provided the human biopsy material. Ö.H.Y., D.M.S., T.T., T.J. and B.F.C. participated in the design and interpretation of experiments. P.K. and N.P. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Characterization of aged intestine.
a, Organoid-forming capacity of crypts from young and old mice (n = 4 mice per group). Student’s paired t-test. b, Frequency of organoids unable to form new crypts (fission deficiency) in young and old mice (n = 6 mice per group) analysed 5–9 days after isolation. Student’s paired t-test. c, Distribution of regenerative growth capacity of primary organoids from young and old mice (n = 6 mice per group). d, Regenerative growth of subcultured secondary mouse organoids (n = 6 mice per group). Student’s paired t-test. e, Distribution of regenerative growth capacity of subcultured secondary organoids from old and young mice (n = 6 mice per group). f, Representative H&E staining of mouse jejunal sections from young and old mice (four mice analysed per group). g, Quantification of EdU+ cells in jejunal crypts 2 h after administration. Only cells next to lysozyme+ Paneth cells were quantified as CBC stem cells. Crypt cells that were not touching lysozyme+ cells were quantified as transit-amplifying (TA) cells (n = 5 mice per group). Representative image of crypt stained for EdU (cyan), DAPI (nuclei, blue), lysozyme (white) and E-cadherin (red). Scale bar, 20 μm. h, Quantification of Ki67+ cells in human ileal biopsies. Cells at the crypt bottom with elongated nuclei next to postmitotic Paneth cells were counted as CBCs. Cells not at the crypt base were considered transit-amplifying cells. (n = 6 for 20–25-year-old donors, n = 10 for ≥ 75-year-old donors). i, Representative gating of Lgr5hi, Lgr5med, Lgr5lo, Paneth and enteroendocrine cells (in relation to Fig. 1c). Quantification of enteroendocrine cells (n = 30 young, n = 26 old). For FACS gating strategy, see Supplementary Fig. 1. j, Analysis of human ileal biopsy material for lysozyme+ Paneth cells (n values for analysed samples shown). k, Immunostaining and quantification of Olfm4+ stem and progenitor cells (green background, n = 75 crypts from young and old. Five individuals per age group) and lysozyme+ Paneth cells in jejunal crypts (red background, n = 115 crypts from young and n = 117 crypts from old mice, five individuals per age group). Whiskers plotted according to Tukey’s method. Scale bars, 10 μm. l, Ratio of Lgr5hi stem cells and Lgr5med progenitor cells and ratio of Lgr5hi stem cells and Paneth cells analysed by flow cytometry from isolated crypts (n = 30 young, n = 26 old). Whiskers plotted according to Tukey’s method. m, Regenerative growth of young Lgr5hi stem cells co-cultured with young or old Paneth cells. Quantification at day 8–11 (n = 6). Representative images are from day 8. Scale bar, 100 μm. Student’s paired t-test. n, Long-term clonogenicity of young and old Lgr5hi stem cells co-cultured with young and old Paneth cells. Serially passaged organoids were quantified 21 days after initial plating (n = 14 mice per age group). Combinations compared to average of young Lgr5hi cells co-cultured with young and old Paneth cells. o, Fourteen-day co-culture of Paneth cells from tdTomato-expressing mouse (R26-mTmG) with Lgr5hi stem cells from Lgr5-eGFP-IRES-creERT2 mouse show long-term niche interactions in organoid culture. Scale bar, 100 μm. Similar results were seen in three replicate wells from co-cultures of the same mice. Y, mice between 3 and 9 months of age; O, mice over 24 months of age in all experiments. In box plots, unless otherwise indicated, the line represents median, the box shows interquartile range and whiskers represent the range. All other data are mean ± s.d.; two-tailed unpaired Student’s t-test; exact P values shown in corresponding panels.
Extended Data Fig. 2 Characterization of gene expression in old Paneth and ISCs.
a, Venn diagram of gene-expression changes in old Paneth cells. (n = 5 mice in old, n = 4 mice in young) b, List of Gene Ontology (GO) terms with the highest enrichment among genes deregulated in old Paneth cells. Fisher’s exact test, no correction for multiple testing. c, Expression of stem-cell-maintaining factors Wnt3 and Egf, and of Notum and Bst1 in old Paneth cells (RNA sequencing (RNA-seq)). Values show fold change in comparison to young Paneth cells. (n = 5 mice in old, n = 4 mice in young). d, Gene editing of Bst1 confirmed by PCR strategy with primers flanking the editing site (191-bp product) and hitting the edited site (89-bp product). Representative agarose gel image is shown. Experiment repeated once to validate the organoid line used in e. e, Regenerative growth of Bst-1 knockout intestinal organoids. Organoids were quantified two days after subculturing (n = 5 repeated experiments with the same organoid line). f, Venn diagram of gene-expression changes in old Lgr5hi stem cells. GSEA preranked analysis of old versus young Lgr5hi stem cells for the gene list ‘KEGG WNT signalling pathway’. Nominal P value is shown (n = 3 mice per age group). g, RNA-scope for NOTUM mRNA (brown) in human jejunal section. Expression seen exclusively in Paneth cells (arrowheads and inset). Experiment repeated twice with similar results in independent samples. h, Expression of human NOTUM and LGR5 from terminal ileal samples of GTEx Consortium (n = 51 sex-matched samples). Expression range is divided into three equal-sized tertiles. In box plots, unless otherwise indicated, the line represents median, the box shows interquartile range and whiskers represent the range. All other data are mean ± s.d.; two-tailed unpaired Student’s t-test; exact P values shown in corresponding panels. For gel source data, see Supplementary Fig. 3.
Extended Data Fig. 3 Wnt ligands increase regenerative capacity of ISCs.
a, Distribution of organoid size on day 5 in ENR + 100 ng ml−1 Wnt3A ± 1μg ml−1 recombinant Notum (n = 50 organoids for Notum-treated (red), n = 38 organoids for untreated (black)). b, Area of colonies from sorted Lgr5hi stem cells from young and old mice (n = 3 mice per age group). Area quantified at day 7. c, Organoid-forming capacity of crypts from young and old mice treated with 100 ng ml−1 Wnt3A. Starting frequency was quantified on day 2 and is represented relative to untreated control (n = 10 mice per age group). d, Primary and secondary regenerative growth of young and old organoids treated with or without 100 ng ml−1 Wnt3A for the first 2 days of culture. Primary organoids were quantified at day 6 and secondary organoids two days after subculturing. Data are represented relative to untreated control (n = 9 mice per age group). e, Organoid-forming capacity of isolated crypts from young mice treated with or without 1 μg ml−1 recombinant Notum (n = 3 mice). f, Primary regenerative growth of organoids from young mice treated with or without 1 μg ml−1 recombinant Notum, quantified on day 6 (n = 3 mice). g, Secondary regenerative growth of organoids from young mice treated with or without 1 μg ml−1 recombinant Notum, quantified on day 2 after subculture (n = 3 mice). h, Organoid-forming capacity of isolated crypts at day 2 from young mice treated with Porcupine inhibitor IWP-237 (n = 3 mice). i, Primary regenerative growth of organoids treated with IWP-2 for the first two days of culture. Organoids were quantified on day 6 (n = 3 mice). j, Flow cytometry analysis of cellular frequencies from Lgr5–eGFP organoids two days after treatment with IWP-2 (n = 4 mice). Y, mice between 3 and 9 months of age; O, mice over 24 months of age in all experiments. In box plots, unless otherwise indicated, the line represents median, the box shows interquartile range and whiskers represent the range. All other data are mean ± s.d.; two-tailed unpaired Student’s t-test; exact P values shown in corresponding panels.
Extended Data Fig. 4 Increased mTORC1 activity in old Paneth cells but not in ISCs.
a, GSEA analysis for gene list ‘Hallmark mTORC1’ (for statistics, see ‘RNA sequencing and data processing’ in Methods). Nominal P value is shown (n = 5 mice in old, n = 4 mice in young). b, Immunohistochemical staining of pS6 (Ser240/244) at mouse jejunal crypt. pS6+ Paneth cells at the crypt bottom are separated by pS6− CBCs. Scale bar, 25 μm. Experiment was repeated for 14 mice with similar results. c, Quantification of pS6+ cells in jejunal crypts (n = 7 mice per age group). d, Isolated Paneth cells from young and old mice, stained with pS6 antibody (red), DAPI (nuclei, blue). Scale bar, 10 μm. Representative image from two independent experiments. e, Left, immunofluorescent image of isolated crypt stained with pS6 antibody (red), Lgr5–eGFP (green) and DAPI (nuclei, blue). Scale bar, 10 μm. Representative of two independent experiments. Right, immunoblots of pS6 and pS6K from isolated crypts of young and old mice, and densitometric quantification (ratio to actin) (n = 14 mice per age group). An outlier (red) deviating >2 s.d. was removed from the analysis. f, Isolated Lgr5hi stem cells (eGFP, green) from young and old mice, stained with pS6 antibody (red), phalloidin (F-actin, white), DAPI (nuclei, blue). Cells were distributed to pS6hi (cells with higher than mean pS6 intensity) or pS6lo (lower than mean pS6 intensity) categories. n = 3 independent experiments. g, Distribution of pS6 intensity in isolated Lgr5hi cells from young and old mice (n = 3 mice per age group, number of cells analysed shown above the corresponding box and whisker plots). h, Mouse weights (n = 25 for young female, n = 26 for old female, n = 20 for young male, n = 19 for old male). Whiskers plotted according to Tukey’s method. Y, mice between 3 and 9 months of age; O, mice over 24 months of age in all experiments. In box plots, unless otherwise indicated, the line represents median, the box shows interquartile range and whiskers represent the range. All other data are mean ± s.d.; two-tailed unpaired Student’s t-test; exact P values shown in corresponding panels. For gel source data, see Supplementary Fig. 3.
Extended Data Fig. 5 Inhibiting mTORC1 activity in old mice restores intestinal regenerative capacity.
a, Organoid-forming capacity and survival of subcultured intestinal organoids treated with rapamycin. Crypts were either treated continuously for four days (2 nM) or with a two-day pulse (2 nM pulse, 10 nM pulse) followed by two days in normal medium before subculturing and quantification (n = 3). b, Regenerative growth of organoids from young and old mice treated with 2 nM rapamycin for 2 days ex vivo. Crypt number was scored six to seven days after treatment from secondary subcultures (two days after passage) (n = 5 mice per age group). Student’s paired t-test. Representative images are from subcultures on day 2. Scale bar, 100 μm. c, Weight of mice receiving daily injections of rapamycin (4 mg kg−1) or vehicle (n = 5 mice per group). Daily data points represent median (circles) and interquartile range (dashed line). d, Immunoblots of pS6 from isolated crypts of vehicle (V)- or rapamycin (R)-treated young and old mice t (n = 4 mice per group). e, Organoid-forming capacity of isolated crypts from old mice treated with vehicle or rapamycin (n = 4 mice per group). f, Primary regenerative growth of organoids from old mice treated with vehicle or rapamycin (n = 4 mice per group). g, Organoid-forming capacity of young Lgr5hi stem cells co-cultured with Paneth cells isolated from young or old mice treated with vehicle or rapamycin (n = 4 mice per group). Combinations compared to average of co-cultures with young vehicle- and old rapamycin-treated Paneth cells. h, Clonogenic growth of Lgr5hi stem cells from young or old mice treated with vehicle or rapamycin (n = 4 mice per group); colonies quantified at day 7. i, RT–qPCR analysis of relative Wnt2b, Wnt5a, Wnt4, Wnt3 and Lgr5 expression from full jejunal samples of old mice treated with rapamycin. Values show fold change (expressed in log2) in comparison to old vehicle treated (n values of mice analysed shown). Data are mean ± s.e.m. j, RT–qPCR analysis of relative Notum and Bst1 expression from crypts of old mice treated with rapamycin. Values show fold change (expressed in log2) in comparison to old vehicle-treated (n = 3 mice per group). Data are mean ± s.e.m. k, Immunoblots of pS6, S6 and H3 from isolated Epcam+ cells of wild type (Tsc1WT) and Tsc1 knockout (Tsc1Δ) epithelium (n = 3 mice per group). l, Quantification of RNA-scope for Notum mRNA in wild type (Tsc1(WT)) and Tsc1 knockout (Tsc1(Δ)) ileal crypts (n = 6 mice for Tsc1(WT) and 5 mice for Tsc1(Δ)). An outlier (red) deviating >2 s.d was removed from the analysis. Representative images of crypts used in quantifications with Notum mRNA (brown) in Paneth cells (inset). m, Organoid-forming capacity of isolated crypts from Tsc1(WT and Tsc1(Δ) epithelium. Quantification was done on day 8. Y, mice between 3 and 9 months of age; O, mice over 24 months of age in all experiments. For gel source data, see Supplementary Fig. 3.
Extended Data Fig. 6 Decreased PPAR activity in aged intestine.
a, GSEA analysis for ‘Biocarta PPARa’ and ‘PPARd’ gene sets (for statistics, see ‘RNA sequencing and data processing’ in Methods). Nominal P value is shown (n = 5 mice in old, n = 4 mice in young). b, Schematic of the putative PPARα-binding site on the mouse and human NOTUM genes found with DECODE. Mouse sequence shown. Bottom, score for the discovered site using JASPAR matrix models for mouse PPAR-response element (PPRE). PPARG; RXRA motif was used. c, FACS analysis of cell populations in primary organoids treated for three days with DMSO, CHIR99021 or GW6471 (n = 6 mice for DMSO, n = 5 mice for CHIR99021 and GW6471). Ratios of Lgr5hi to Paneth cells, and Lgr5hi to Lgr5lo cells from the same analysis. Mean CD24 expression of live Epcam+ cells are also shown. d, Representative images of mouse intestinal organoids treated for 4 days with DMSO, 5 μM GW6471 or 5 μM GW6471 + 100 ng ml−1 Wnt3A. Arrowheads indicate surviving, and red asterisks collapsed, organoids. Scale bar, 100 μm. Experiment was repeated four times with similar results. Unless otherwise indicated, data are mean ± s.d.; two-tailed unpaired Student’s t-test; exact P values shown in corresponding panels.
Extended Data Fig. 7 Notum regulates intestinal stem cell function.
a, Notum gene targeting. Schematic represents sites of genome editing. Gene editing was confirmed by PCR with primers flanking the editing site (174 bp product for Notum KO1 and 294 bp product for Notum KO2) and hitting the edited site (84 bp product for Notum KO1 and 188 bp product for Notum KO2). Representative agarose gel images from two independent experiments with similar results are shown. b, Regenerative growth of Notum knockout organoids. De novo crypt domains were quantified two days after subculture (n = 5 repeated experiments with the same organoid lines). Representative images of organoids two days after subculture are shown. Scale bar, 100 μm. c, Schematic presenting in vivo competition assay of gene-edited organoid growth by orthotopic transplantation to immunodeficient Rag2−/− mice. Representative colonoscopy, necroscopy and histology images used for assay quantification (n = 8 mice transplanted). Scale bars, 1 mm for necroscopy and 200 μm for histology. d, Representative images of CRISPR-targeted young and old organoids two days after subculturing (n = 4 mice per group). Scale bar, 100 μm. e, Relative Notum expression in organoids with synergistic activator mediator complex (SAM) targeted to Notum promoter (dA Notum) grown for two days in ENR medium. Three independent experiments; relative to control (dA Tom). f, Quantification and representative images of day-5 colonies formed by isolated CD24medSSClo cells from Notum activator (dA Notum) and control (dA Tom) organoids. Scale bar, 100 μm. n = 4 repeated experiments with the same organoid line. g, RT–qPCR analysis of relative Axin2 and Lgr5 expression in CD24medSSClo cells sorted from Notum activator (dA Notum) organoids. Values show fold change (expressed in log2) in comparison to control (dATom) (n = 3 replicate wells per organoid line). In box plots, unless otherwise indicated, the line represents median, the box shows interquartile range and whiskers represent the range. All other data are mean ± s.e.m.; two-tailed unpaired Student’s t-test; exact P values shown in corresponding panels. For gel source data see Supplementary Fig. 3.
Extended Data Fig. 8 Notum inhibitor ABC99 prevents Wnt inactivation.
a, Flow cytometry analysis of cell populations in primary organoids treated for eight days with 500 nM ABC99 (n = 3 mice) relative to DMSO control. Student’s paired t-test. b, Clonogenic growth of Lgr5hi stem cells on day 5 treated with or without 50 nM ABC99 and/or 500 ng ml−1 recombinant Notum (two independent experiments with similar results, one experiment with three replicate wells shown). c, Relative weight of mice treated with daily injections of ABC99 (10 mg per kg (body weight)) or control (vehicle or ABC101 10 mg per kg (body weight)) (n = 10 mice for young control and young ABC99, n = 8 mice for old control and n = 9 mice for old ABC99). Daily data points represent median (circles) and interquartile range (dashed line). d, Clonogenic growth of young Lgr5hi stem cells co-cultured with young or old Paneth cells from mice treated with ABC99 or control (n values for analysed mice shown). Combinations compared to average of co-cultures with young control (−) and old ABC-treated (+) Paneth cells. Control mice received an equal amount of the inactive analogue ABC101 (yellow circles) or vehicle. e, Representative image of immunofluorescent staining of ileal crypts used for quantification of nuclear β-catenin (white) intensity. Paneth cells (red arrowheads) and CBCs (green arrowheads) were identified by cellular and nuclear (DAPI, blue) morphology. Their nuclear β-catenin levels were compared to transit-amplifying cells (white arrowheads). Scale bar, 20 μm. Experiment was repeated twice with a total of 26 mice all showing strongest nuclear β-catenin at the crypt bottom. f, Immunofluorescent staining of histological sections from old ileum. β-catenin (white), lysozyme (red) and DAPI (nuclei, blue). Scale bar, 10 μm. Quantification of relative nuclear β-catenin intensity of Paneth cells (red arrowhead) (n values for analysed mice shown). For quantification of CBCs (green arrowhead) see Fig. 3d. g, Immunofluorescent staining of histological sections from old ileum. Olfm4, green; EdU, red; DAPI (nuclei), blue. Scale bar, 20 μm. Quantification of EdU+ cellular frequencies within the crypt (n values for analysed mice shown). Y, mice between 3 and 9 months of age; O, mice over 24 months of age in all experiments. In box plots, unless otherwise indicated, the line represents median, the box shows interquartile range and whiskers represent the range. All other data are mean ± s.d.; two-tailed unpaired Student’s t-test; exact P values shown in corresponding panels.
Extended Data Fig. 9 Old intestine recovers poorly from 5-FU-induced damage.
a, Body weights of young and old mice following single injection of 5-FU (100–200 mg kg−1). Two mice per group, body weight relative to day of injection (day 0). b, Relative body weight of young and old mice treated for one week with or without ABC99 followed by single 5-FU (100 mg kg−1) injection (n = 8 mice for young vehicle, old vehicle and young ABC99-treated, n = 10 mice for old ABC99-treated). Daily data points represent median (circles) and interquartile range (dashed line). Daily weight of old ABC99-treated mice were compared to old controls with two-tailed unpaired Student’s t-test; exact P values shown under the corresponding daily weight. Young mice between 3 and 4 months of age; old mice over 20 months of age.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-3 which include the FACS gating strategy (Supplementary Fig. 1), cell purity assessment (Supplementary Fig. 2), uncropped scans of immunoblots and gel electrophoresis (Supplementary Fig. 3) and a full description for Supplementary Table 1.
Supplementary Table 1
Analysis of RNA-sequencing results for young and old Paneth and Lgr5+ ISCs – see Supplementary Information document for full description.
Source data
Rights and permissions
About this article
Cite this article
Pentinmikko, N., Iqbal, S., Mana, M. et al. Notum produced by Paneth cells attenuates regeneration of aged intestinal epithelium. Nature 571, 398–402 (2019). https://doi.org/10.1038/s41586-019-1383-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1383-0
This article is cited by
-
Intestinal stem cells: guardians of homeostasis in health and aging amid environmental challenges
Experimental & Molecular Medicine (2024)
-
NAD+ dependent UPRmt activation underlies intestinal aging caused by mitochondrial DNA mutations
Nature Communications (2024)
-
The secreted protein Amuc_1409 from Akkermansia muciniphila improves gut health through intestinal stem cell regulation
Nature Communications (2024)
-
A stromal lineage maintains crypt structure and villus homeostasis in the intestinal stem cell niche
BMC Biology (2023)
-
Early weaning causes small intestinal atrophy by inhibiting the activity of intestinal stem cells: involvement of Wnt/β-catenin signaling
Stem Cell Research & Therapy (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.