Abstract
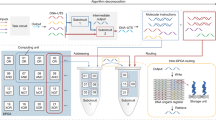
Molecular biology provides an inspiring proof-of-principle that chemical systems can store and process information to direct molecular activities such as the fabrication of complex structures from molecular components. To develop information-based chemistry as a technology for programming matter to function in ways not seen in biological systems, it is necessary to understand how molecular interactions can encode and execute algorithms. The self-assembly of relatively simple units into complex products1 is particularly well suited for such investigations. Theory that combines mathematical tiling and statistical–mechanical models of molecular crystallization has shown that algorithmic behaviour can be embedded within molecular self-assembly processes2,3, and this has been experimentally demonstrated using DNA nanotechnology4 with up to 22 tile types5,6,7,8,9,10,11. However, many information technologies exhibit a complexity threshold—such as the minimum transistor count needed for a general-purpose computer—beyond which the power of a reprogrammable system increases qualitatively, and it has been unclear whether the biophysics of DNA self-assembly allows that threshold to be exceeded. Here we report the design and experimental validation of a DNA tile set that contains 355 single-stranded tiles and can, through simple tile selection, be reprogrammed to implement a wide variety of 6-bit algorithms. We use this set to construct 21 circuits that execute algorithms including copying, sorting, recognizing palindromes and multiples of 3, random walking, obtaining an unbiased choice from a biased random source, electing a leader, simulating cellular automata, generating deterministic and randomized patterns, and counting to 63, with an overall per-tile error rate of less than 1 in 3,000. These findings suggest that molecular self-assembly could be a reliable algorithmic component within programmable chemical systems. The development of molecular machines that are reprogrammable—at a high level of abstraction and thus without requiring knowledge of the underlying physics—will establish a creative space in which molecular programmers can flourish.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The AFM data generated and analysed here are available from the authors’ website (http://www.dna.caltech.edu/SupplementaryMaterial/Algorithmic_SST/), as is the Python code for data analysis and DNA sequence design.
Change history
02 August 2019
An Amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Whitesides, G. M. & Grzybowski, B. Self-assembly at all scales. Science 295, 2418–2421 (2002).
Winfree, E. Simulations of Computing by Self-Assembly. Technical Report CaltechCSTR:1998.22 (California Institute of Technology, 1998).
Doty, D. Theory of algorithmic self-assembly. Commun. ACM 55, 78–88 (2012).
Seeman, N. C. & Sleiman, H. F. DNA nanotechnology. Nat. Rev. Mater. 3, 17068 (2017).
Mao, C., LaBean, T. H., Reif, J. H. & Seeman, N. C. Logical computation using algorithmic self-assembly of DNA triple-crossover molecules. Nature 407, 493–496 (2000); erratum 408, 750 (2000).
Rothemund, P. W. K., Papadakis, N. & Winfree, E. Algorithmic self-assembly of DNA Sierpinski triangles. PLoS Biol. 2, e424 (2004).
Schulman, R. & Winfree, E. Synthesis of crystals with a programmable kinetic barrier to nucleation. Proc. Natl Acad. Sci. USA 104, 15236–15241 (2007).
Barish, R. D., Schulman, R., Rothemund, P. W. K. & Winfree, E. An information-bearing seed for nucleating algorithmic self-assembly. Proc. Natl Acad. Sci. USA 106, 6054–6059 (2009).
Schulman, R., Yurke, B. & Winfree, E. Robust self-replication of combinatorial information via crystal growth and scission. Proc. Natl Acad. Sci. USA 109, 6405–6410 (2012).
Evans, C. G. Crystals that Count! Physical Principles and Experimental Investigations of DNA Tile Self-Assembly. PhD thesis, Caltech (2014).
Schulman, R., Wright, C. & Winfree, E. Increasing redundancy exponentially reduces error rates during algorithmic self-assembly. ACS Nano 9, 5760–5771 (2015).
Winfree, E., Liu, F., Wenzler, L. A. & Seeman, N. C. Design and self-assembly of two-dimensional DNA crystals. Nature 394, 539–544 (1998).
Yin, P. et al. Programming DNA tube circumferences. Science 321, 824–826 (2008).
Rothemund, P. W. K. Folding DNA to create nanoscale shapes and patterns. Nature 440, 297–302 (2006).
Wei, B., Dai, M. & Yin, P. Complex shapes self-assembled from single-stranded DNA tiles. Nature 485, 623–626 (2012).
Wang, W. et al. Self-assembly of fully addressable DNA nanostructures from double crossover tiles. Nucleic Acids Res. 44, 7989–7996 (2016).
Ong, L. L. et al. Programmable self-assembly of three-dimensional nanostructures from 10,000 unique components. Nature 552, 72–77 (2017).
Rothemund, P. W. K. & Winfree, E. The program-size complexity of self-assembled squares. In STOC: Proceedings of the 32nd Annual ACM Symposium on Theory of Computing (eds Yao, F. & Luks, E.) 459–468 (Association for Computing Machinery, 2000).
Soloveichik, D. & Winfree, E. Complexity of self-assembled shapes. SIAM J. Comput. 36, 1544–1569 (2007).
Moore, C. & Mertens, S. The Nature of Computation (Oxford Univ. Press, Oxford, 2011).
Cook, M. Universality in elementary cellular automata. Complex Syst. 15, 1–40 (2004).
Neary, T. & Woods, D. P-completeness of cellular automaton Rule 110. In ICALP 2006: International Colloquium on Automata, Languages, and Programming (eds Bugliesi, M., Preneel, B., Sassone, V. & Wegener, I.) 132–143 (Springer, 2006).
Schulman, R. & Winfree, E. Programmable control of nucleation for algorithmic self-assembly. SIAM J. Comput. 39, 1581–1616 (2009).
Evans, C. G. & Winfree, E. Physical principles for DNA tile self-assembly. Chem. Soc. Rev. 46, 3808–3829 (2017).
Winfree, E. & Bekbolatov, R. Proofreading tile sets: error correction for algorithmic self-assembly. In DNA9: Proceedings of the 9th International Conference on DNA Computing (eds Chen, J. & Reif, J.) 126–144 (Springer, 2004).
Mohammed, A. M. & Schulman, R. Directing self-assembly of DNA nanotubes using programmable seeds. Nano Lett. 13, 4006–4013 (2013).
Evans, C. G. & Winfree, E. DNA sticky end design and assignment for robust algorithmic self-assembly. In DNA19: Proceedings of the 19th International Conference on DNA Computing and Molecular Programming (Eds Soloveichik, D. & Yurke, B.) 61–75 (Springer, 2013).
Zadeh, J. N. et al. NUPACK: analysis and design of nucleic acid systems. J. Comput. Chem. 32, 170–173 (2011).
Lorenz, R. et al. ViennaRNA Package 2.0. Algorithms Mol. Biol. 6, 26 (2011).
Wei, B., Luvena, L. L., Ong, J., Jaffe, A. S. & Yin, P. Complex reconfiguration of DNA nanostructures. Angew. Chem. Int. Ed. 126, 7605–7609 (2014).
Chalk, C., Fu, B., Martinez, E., Schweller, R. & Wylie, T. Concentration independent random number generation in tile self-assembly. Theor. Comput. Sci. 667, 1–15 (2017).
Jacobs, W. M., Reinhardt, A. & Frenkel, D. Rational design of self-assembly pathways for complex multicomponent structures. Proc. Natl Acad. Sci. USA 112, 6313–6318 (2015).
Hedges, L. O., Mannige, R. V. & Whitelam, S. Growth of equilibrium structures built from a large number of distinct component types. Soft Matter 10, 6404–6416 (2014).
Cairns-Smith, A. G. Genetic Takeover and the Mineral Origins of Life (Cambridge Univ. Press, Cambridge, 1982).
Acknowledgements
We thank C. Evans, A. Gopinath, B. Wei, C. Geary, R. Schulman, S. Woo, P. Rothemund and Y. Rondelez for experimental advice; R. Barish and R. Hariadi for contributing to preliminary designs for algorithmic self-assembly by SST; C. Moore, T. Stérin, C. Thachuk, P.-É. Meunier and C. Geary for discussions on theory; and L. Qian, G. Tikhomirov and P. Petersen for AFM usage. This work was supported by National Science Foundation (NSF) grants CCF-1162589 (to E.W., D.D. and D.W.), CCF-1162459 (to P.Y.), CCF-1219274 (to D.W. and D.D.), CCF-1619343 (to D.D.), CCF-0832824 and CCF-1317694 (Expeditions in Computing, to E.W.) and CCF-1317291 (Expeditions in Computing, to P.Y.), and by NASA grant NNX13AJ56G (to D.W.). C.M. was funded by the Fannie and John Hertz Foundation. F.Z. and J.H. received support from the Caltech Summer Undergraduate Research Fellowship program.
Author information
Authors and Affiliations
Contributions
D.W., D.D., E.W. and P.Y. conceived the study. D.W., D.D. and E.W. designed the circuits and wrote the manuscript. D.W. and D.D. carried out all data analysis and experiments reported except for the nanotube nucleation/melt experiments (which were performed by J.H. and D.W.) and the unzipping and other early experimental protocols (performed by F.Z., C.M. and D.D.).
Corresponding authors
Ethics declarations
Competing interests
D.W., D.D., J.H., F.Z. and E.W. declare that they have no competing interests. P.Y. and C.M. declare competing interests: they are both listed as inventors on pending and issued patents on single-stranded tiles; and P.Y. is a co-founder of Ultivue Inc. and NuProbe Global.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
This file contains Supplementary Information section A, which contains details about the IBC model, the abstract tile assembly model and proofreading, the SST binding domain schematics, DNA sequence design, data supporting design decisions, experimental methods, AFM image analysis, and the 21 implemented circuits
Supplementary Data
This file contains Supplementary Information section B, which contains sequences for the DNA systems explored in this paper
Supplementary Figures
This file contains Supplementary Information section C, four sample high-resolution AFM images of DNA nanostructures for several circuits
Rights and permissions
About this article
Cite this article
Woods, D., Doty, D., Myhrvold, C. et al. Diverse and robust molecular algorithms using reprogrammable DNA self-assembly. Nature 567, 366–372 (2019). https://doi.org/10.1038/s41586-019-1014-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1014-9
This article is cited by
-
DNA as a universal chemical substrate for computing and data storage
Nature Reviews Chemistry (2024)
-
Pattern recognition in the nucleation kinetics of non-equilibrium self-assembly
Nature (2024)
-
Self-assembling DNA recognizes patterns
Nature (2024)
-
The Impacts of Dimensionality, Diffusion, and Directedness on Intrinsic Cross-Model Simulation in Tile-Based Self-Assembly
Algorithmica (2024)
-
The Need for Seed (in the Abstract Tile Assembly Model)
Algorithmica (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.