Abstract
RNA localization is highly regulated, with subcellular organization driving context-dependent cell physiology. Although proximity-based labelling technologies that use highly reactive radicals or carbenes provide a powerful method for unbiased mapping of protein organization within a cell, methods for unbiased RNA mapping are scarce and comparably less robust. Here we develop α-alkoxy thioenol and chloroenol esters that function as potent acylating agents upon controlled ester unmasking. We pair these probes with subcellular-localized expression of a bioorthogonal esterase to establish a platform for spatial analysis of RNA: bioorthogonal acylating agents for proximity labelling and sequencing (BAP-seq). We demonstrate that, by selectively unmasking the enol probe in a locale of interest, we can map RNA distribution in membrane-bound and membrane-less organelles. The controlled-release acylating agent chemistry and corresponding BAP-seq method expand the scope of proximity labelling technologies and provide a powerful approach to interrogate the cellular organization of RNAs.

Similar content being viewed by others
Main
The distribution of biomolecules in mammalian systems is tightly regulated, with compartmentalization in membrane-bound organelles as the most traditionally understood mechanism to dictate biomolecular interactions and localized regulation. Noncovalent interactions between molecules, such as protein–protein and protein–RNA interactions1,2, can also regulate the organization of molecules in a cell, in some cases through ‘membrane-less compartments’, such as stress granules and the nucleolus. This is especially true for RNA, as localization and local translation enable spatiotemporal regulation critical for processes such as embryonic patterning and cell motility and polarization. Regulatory non-coding RNAs often serve as scaffolds, mediating interactions between other RNAs and proteins within sub-organellar compartments, such as the nucleolus within the nucleus3,4. Thus, the location-based regulation of RNA function is critical to protein homoeostasis and cellular physiology5,6. Despite the criticality of RNA localization for function, few methods exist for transcriptome-wide unbiased mapping of RNAs in cells. While organelles can be isolated and their RNA content characterized in some cases, such methods are time-consuming and prone to contamination or do not exist for RNAs localized to membrane-free or sub-organellar compartments7.
Proximity labelling technologies employ the localized expression of an engineered protein or the delivery of a small-molecule catalyst to generate reactive molecules that covalently modify biomolecules within a local vicinity, enabling subsequent isolation and analysis8,9. For example, APEX2 is an engineered peroxidase that produces biotin–phenoxy radicals from biotin tyramide in the presence of hydrogen peroxide10,11, primarily labelling electron-rich residues, such as tyrosine, on nearby proteins. The APEX system has revolutionized spatial proteomics by providing maps of the mitochondrial12, macromolecular complexes13, the endoplasmic reticulum (ER14) and the synaptic cleft15 proteomes, among others. Various other peroxidase-based proximity biotinylation methods have likewise been developed for proximity labelling16,17. More recently, the photoPPI18 and µMap platforms have enabled the use of small-molecule iridium photocatalysis to generate carbenes from diazirines for proximity-dependent protein labelling19,20. Finally, acylation-based proximity labelling technologies for protein, such as BioID21,22 and TurboID23, generate biotinoyl-5′-AMP without the use of potentially confounding hydrogen peroxide or light.
In contrast to the robust and diverse technologies for mapping proteins, proximity labelling technologies for RNA remain nascent. APEX-RIP combines peroxidase-catalysed protein biotinylation with RNA–protein chemical crosslinking to isolate RNA polymers24. The discovery that APEX-derived biotin–phenoxy radicals could directly modify RNAs for direct proximity-based labelling resulted in the development of APEX-sequencing (APEX-seq)25,26 and the first reported high-resolution maps of RNA distribution in live cells. Since then, APEX-derived biotin–phenoxy radicals have been tuned for higher reactivity towards RNA27. Light-activated RNA-targeting proximity labelling technologies have also been developed, including CAP-sequencing28 and Halo-sequencing29, which use light-activated enzyme (miniSOG) or ligand (dibromofluorescein) activation to achieve reactive oxygen species (ROS)-mediated RNA proximity-based labelling. While these methods show promise, all rely on radical or ROS reactivity—species that primordial RNA-like polymers probably evolved into to minimize reactivity with. Thus, current RNA labelling methods require increased reagent concentration and/or labelling time to overcome poor RNA reactivity. This raises the possibility of incomplete labelling, biased by substrate abundance or structure and reflecting stress, not homoeostatic, subcellular RNA localizations. Interestingly, RNA is a well-known nucleophile through the reaction of its 2′-hydroxyl group with acylating agents and other electrophilic species30,31,32,33,34,35,36,37. However, the non-radical, acylation-based proximity reagents available—such as the anhydride-based TurboID biotin-5′-AMP molecules—are not known to be reactive enough to label RNA. Therefore, we set out to develop a proximity labelling platform that generates highly reactive and tunable acylating agents to label RNA in live cells efficiently.
In this Article, we describe the development of bioorthogonal acylating agents for proximity labelling and sequencing (BAP-seq), a proximity labelling method that combines a subcellularly targeted esterase from Bacillus subtilis (BS2) with masked enol-ester probes to generate highly reactive acylating agents that label nearby RNAs in live cells. Using previously unreported chemistry, we synthesize both a panel of masked aryl thioester probes with a spectrum of electrophilicity and a panel of masked acid chloride (AC) probes with diverse functional handles. Following in vitro and in cell analyses, acid chloride probe-2 (AC-2) emerged as the leading probe. AC-2 generates a highly reactive acid chloride acylating agent upon ester hydrolysis by BS2, labelling nearby nucleophiles with an alkyne for subsequent imaging or enrichment after click chemistry. Confocal imaging reveals that the AC-2 probe labels subcellular locales in a BS2 proximity-dependent manner, in both membrane-bound (for example, the nucleus) and membrane-less (for example, the nucleolus) compartments. We then demonstrate RNA labelling by the BS2–AC-2 system in vitro and in cells. Finally, we adapt BS2–AC-2 labelling and enrichment with RNA-sequencing (RNA-seq) to map the distribution of mitochondrial, nuclear and nucleolar RNAs. These results establish BAP-seq as a valuable addition to the repertoire of proximity labelling approaches.
Results
Developing bioorthogonal enol-ester-masked acylating agents
In designing acylation-based proximity labelling agents, we sought to develop a strategy in which an exogenously supplied enzyme could convert an inert probe to a reactive acylating agent. We envisioned that a bioorthogonal ester-masked enol with an α-leaving group would be unreactive towards biomolecules, but controlled ester hydrolysis would generate an enol that would rapidly tautomerize to the reactive acylating agent (Fig. 1). For the ester mask, we chose 1-methylcyclopropyl (mCP) ester, which is inefficiently cleaved by endogenous esterases in human and murine cells38 but can be hydrolysed quickly by exogenously expressed esterases, such as porcine liver esterase38 and BS239. To confirm that BS2 is compatible with subcellular expression, we performed cell viability experiments, which revealed no obvious cytotoxicity (Supplementary Fig. 1). To assess the kinetics of BS2 on the mCP ester mask, we synthesized mCP-masked 7-hydroxycoumarin (mCP-coumarin) as a profluorophore40 that generates the fluorescent 7-hydroxycoumarin upon BS2 activity. The kinetic measurements of the mCP-coumarin substrate revealed that BS2 can process mCP ester mask with a turnover number (Kcat) of 260 s−1 and Michaelis constant (Km) of 417 µM (Extended Data Fig. 1). We further validated that the esterase activity of BS2 on mCP-coumarin is inhibited by a known serine protease inhibitor, phenylmethylsulfonyl fluoride (PMSF; Supplementary Fig. 2). Owing to the established bioorthogonality of the mCP ester, the low cytotoxicity of BS2 and the rapid unmasking kinetics of the mCP ester by BS2, we selected the BS2 esterase–mCP ester pair as the genetically encoded release agent in our proposed proximity labelling platform.
A masked acylation agent is unreactive towards biomolecules. Selective unmasking by BS2 esterase through cleavage of a bioorthogonal mCP ester mask releases an enol, which rapidly tautomerizes to a carbonyl, the acylating agent. Nucleophilic biomolecules react with the acylating agent and form a covalent bond, which installs a visualization or affinity handle onto the target biomolecule. NBD, nitrobenzoxadiazole.
We next synthesized thioester-based (TE) probes, for which the resultant acylating agent’s electrophilicity can be predicted based on the pKa of the thiolate leaving group41. This necessitated development of methods to synthesize α-alkoxy thioenol esters, as this functional group has rarely been documented42 and existing methods were not compatible with key functional handle installation. Attempts to use various silver-based catalysts previously reported for trans-addition of carboxylic acids to ynol ethers43 on thioynol ether substrates yielded more than 90% of the 1,2-addition product, with less than 5% of the desired 1,1-addition product (α-alkoxy thioenol esters), probably owing to the reversed polarity of the triple bond caused by the electronegativity difference between the sulfur and oxygen atom44,45 (Extended Data Fig. 2). After systematically screening catalysts, ligands and solvent conditions on a model compound 1 (Extended Data Fig. 2 and Supplementary Fig. 3), we discovered [Ir(COD)OMe]2 could catalyse the addition of carboxylic acids to thioynol ethers to selectively generate the desired α-alkoxy thioenol esters (Fig. 2a). Preliminary investigation on the substrate scope revealed the necessity of an α-position heteroatom, possibly to serve as a σ-donor to coordinate with iridium. Despite the success in the reaction development, the synthesis of a minimalist probe failed in the conversion of alcohol 3 to the azide (Supplementary Fig. 4a,b). We reasoned that side reactions may arise from the close proximity of the α-alkoxy thioenol ester group to the azidation site. Therefore, we synthesized an alcohol substrate with an additional polyethylene glycol linker, which was then cleanly converted to the azide to install the click handle via a Mitsunobu azidation reaction46 (Fig. 2a). Having established a robust synthetic route, we synthesized four TE probes with a range of acylation strengths of the unmasked product by tuning the leaving group thiolate: phenyl thiol (TE-1, pKa 6.6), difluorophenyl thiol (TE-2, pKa 6.2), tetrafluorophenyl thiol (TE-3, pKa 3.8) and 4-nitrophenyl thiol (TE-4, pKa 3.9) (Supplementary Fig. 4c)41.
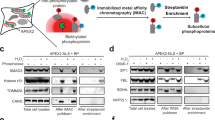
a, Key synthetic steps for the synthesis of TE probes. b, The unmasking reaction of TE-1 by BS2 yields the predicted thioester product, as confirmed by LC–MS. c, The synthetic scheme for the AC probes. d, The structure of all the AC probes. The leaving groups are highlighted in green, the bioorthogonal BS2 recognition site is highlighted in blue and the functional handle is highlighted in purple. e, The unmasking reaction of AC-5 by BS2 in the presence of a nucleophilic substrate (Fmoc–Lys–OMe) yields the predicted amide product. The newly formed bond is highlighted in red. f, The LC–MS trace of the reaction shown in e. mAu, milli-absorbance units; n-BuLi, n-butyllithium; TMEDA, tetramethylethylenediamine; DCE, 1,2-dichloroethane; DIAD, diisopropyl azodicarboxylate; DPPA, diphenylphosphoryl azide; THF, tetrahydrofuran; PhSSPh, phenyl disulfide; PhSH, thiophenol; DIBAL-H, diisobutylaluminum hydride; EDCI, N-ethyl-N′-(3-dimethylaminopropyl)carbodiimide hydrochloride; DMAP, 4-(dimethylamino)pyridine; DMF, N,N-dimethylformamide.
To confirm that the TE probes are substrates of BS2 and can generate the proposed reactive thioesters, we assessed the least reactive probe (TE-1) in vitro, which yields a stable thioester product that can be detected before hydrolysis47. While we observed minimal degradation of TE-1 (1 mM) after 3 h in aqueous solution (Supplementary Fig. 5a), the addition of BS2 esterase resulted in the complete consumption of TE-1 in 5 min and the formation of a mixture containing the predicted S-phenyl thioester (6), the thiophenol from thioester hydrolysis and the corresponding disulfide (Fig. 2b and Supplementary Fig. 5b–d). These data corroborate the hypothesis that mCP-masked TE probes are substrates of BS2 and that their BS2-mediated unmasking generates S-aryl thioesters, which are known acylating agents48.
After validating the proposed mCP-masked enol mechanism, we next aimed to generate probes that produce extremely reactive acylating agents with short half-lives due to competitive hydrolysis, resulting in intermediates that can covalently tag biomolecules in a proximity-dependent manner. Since acid chlorides are among the most reactive acylating agents known, with half-lives in water not reported owing to their high reactivity, we targeted the synthesis of masked AC probes. We reasoned that the functional warhead α-chloroenol ester could be synthesized by chromium(II) chloride-assisted transformation from trichloromethylcarbinol49 (Fig. 2c and Supplementary Fig. 6). We then introduced different handles, including alkynes with various linkers (AC-1, AC-2 and AC-3), an azide (AC-4) and a nitrobenzoxadiazole fluorophore for direct imaging (AC-5; Fig. 2d). Importantly, the (Z)-α-chloroenol ester is also stable in aqueous solution. We found no detectable decomposition of AC-5 in phosphate-buffered saline (PBS) after 1 h of incubation (Supplementary Fig. 7a,b). However, the addition of BS2 esterase resulted in rapid (<2 min) formation of the corresponding carboxylic acid, the expected hydrolysis product from acid chloride (Supplementary Fig. 7c,d). To verify the formation of the acid chloride, we incubated AC-5 with BS2 in the presence of an amine nucleophile (Fmoc–Lys ester) to ‘trap’ the acid chloride and observed the formation of the predicted amide product (Fig. 2e,f and Supplementary Fig. 7e,f). Further thin-layer chromatography (TLC) tests confirmed that all the TE and AC probes can be efficiently processed by BS2 (Supplementary Fig. 8). Together, these results indicate that both the TE and AC series of probes are stable in aqueous conditions and can rapidly generate acylating agents by BS2 esterase in vitro.
Proximity labelling using AC and TE probes in live cells
With the TE and AC probes in hand, we next sought to assess their bioorthogonality and ability to label biomolecules in live mammalian cells in a BS2-dependent manner. To do this, we first evaluated the cellular stability and labelling of each probe in cells overexpressing nuclear-localized BS2. After confirming nuclear localization of BS2 in transiently transfected HEK293T cells via immunostaining, we treated the cells with 25 µM of each probe for 10 min. The cells were then fixed, derivatized with Alexa-488-azide/alkyne via click chemistry (except AC-5, which was already conjugated to a fluorophore) and visualized by epi-fluorescence microscopy. As expected, TE-1 (Supplementary Fig. 9) and TE-2 (Supplementary Fig. 10), which generate the acylating agents with the lowest reactivity, showed no BS2-dependent labelling, which aligned with the low reactivity of their thioester products. In contrast, TE-3, TE-4 and all the AC probes showed a clear BS2-dependent nuclear-localized signal (Supplementary Figs. 11–17). We then compared the relative intensity of the nuclear signal of each probe to determine the optimal probe that features the highest signal-to-noise ratio (fluorescence signal in the presence versus absence of BS2). We found that the AC probes label more efficiently than TE probes (Fig. 3a), with AC-2 generating a 50-fold increase of signal in nuclear labelling with respect to dimethyl sulfoxide (DMSO; Fig. 3b) and an 8-fold signal-to-noise ratio (Supplementary Fig. 14). Examining the signal localization from AC-2 revealed potential nuclear staining (Fig. 3c). Therefore, we selected AC-2 as our lead probe for further characterization of labelling proximity.
a, Representative fluorescence images of HEK293T cells transiently transfected with a vector expressing an NLS–BS2 esterase construct, then treated with 25 µM of the indicated probes for 10 min. Probe localization, corresponding to labelled sites, is shown in green (via click-chemistry installation of Alexa-488), BS2 is shown in red (via immunofluorescence) and nuclear DAPI staining is shown in blue. Scale bars, 10 µm. b, Quantification of probe labelling from images in a. The data are shown as the mean ± standard deviaition from n = 4 images per probe. c, Detail of AC-2-treated cells, showing the inset outlined in a.
To further validate the proximity-dependent labelling of the BS2–AC-2 pair, we used confocal microscopy to visualize the co-localization of BS2 with AC-2 signal in cells expressing BS2 across a series of cellular compartments. Cells expressing BS2 localized to the nucleus (via a nuclear localization signal (NLS); Extended Data Fig. 3), the cytosol (via a nuclear export signal; Extended Data Fig. 4), ER (via a cytosol-facing outer ER membrane (ERM) localization tag; Extended Data Fig. 5), mitochondria (via a mitochondrial matrix localizing tag; Extended Data Fig. 6), nucleolus (via NIK3x fusion; Extended Data Fig. 7) or nuclear pore (via SENP2 fusion; Extended Data Fig. 8) were treated with 25 µM AC-2 for 10 min, fixed, derivatized with Alexa-488-azide via click chemistry and visualized by confocal microscopy. We detected robust BS2-dependent AC-2 labelling across all compartments (Fig. 4), including non-membrane-bound organelles such as the nucleolus and nuclear pore. Further analysis revealed that the AC-2 labelling signal tightly co-localized with the BS2 localization signal in the nucleus (Supplementary Fig. 18), nucleolus (Supplementary Fig. 19) and nuclear pore (Supplementary Fig. 20). These data confirm the suitability of the BS2–AC-2 system for proximity labelling. Given the high BS2-dependent labelling observed in subcellular compartments, we next sought to assess whether RNA acylation by BS2-unmasked AC-2 occurs in vitro and in cells.
Representative confocal images of HEK293T cells expressing BS2 in the cytosol, nucleus, ERM, mitochondrial matrix, nuclear pore or nucleolus, as indicated in the schematics on the right. All cells were treated with 25 µM of AC-2 for 10 min. Probe localization, corresponding to sites labelled via click-chemistry installation of Alexa-488, is shown in green. BS2 is shown in red (via immunofluorescence), and nuclear DAPI staining is shown in blue. Scale bars, 10 µm.
RNA labelling by BS2–AC-2
To assess whether the BS2–AC-2 probe system can label RNA, we performed an in vitro labelling reaction and observed BS2-dependent labelling of an RNA oligo by AC-5, the fluorophore-containing AC probe (Fig. 5a). To test RNA labelling in cells, we incubated cells expressing BS2 in the cytosol, nucleus or mitochondria with our lead probe, AC-2 (25 µM), for 10 min. We then lysed the cells, extracted and purified total cellular RNA and performed click chemistry with biotin azide to install biotin on modified RNAs. We assessed the degree of RNA labelling using a dot blot and observed robust BS2-dependent RNA labelling across all organelles (Fig. 5b). RNase treatment eliminated this signal, while proteinase K treatment did not, confirming that the signals were indeed from RNA. Importantly, we detected no labelling in cells without BS2 expression, indicating the low background and further demonstrating the bioorthogonality of AC-2. These data confirm that the BS2–AC-2 pair labels RNAs in cells, and as such, we sought to analyse the RNA distribution across several compartments.
a, A total of 2.5 µM of a 74-mer RNA was treated with 500 µM AC-5 and 600 nM BS2 for 5 min; then RNA labelling was analysed via fluorescent gel electrophoresis. SYBR Gold shows the amount of RNA loaded across all samples. The experiment was repeated twice independently, showing similar results. b, Dot blot showing BS2-dependent RNA labelling by AC-2 when BS2 is expressed in the cytosol, nucleus and mitochondria. HEK293T cells were transfected with a control vector or a vector expressing BS2 in the cytoplasm, nucleus or mitochondria, then treated with 25 µM AC-2 for 10 min, followed by lysis, RNA isolation and click chemistry to install biotin on labelled RNAs. RNA labelling was analysed via dot blot with 500 ng of isolated RNA using a streptavidin-conjugated horseradish peroxidase antibody. The methylene blue blot shows the amount of RNA loaded across all samples. For RNase- or protease-treated samples, RNA was treated with 85 µg ml−1 of RNase A or 85 µg ml−1 of proteinase K for 15 min, purified and then blotted on the same membrane.
BAP-seq
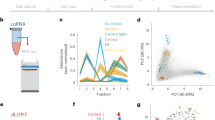
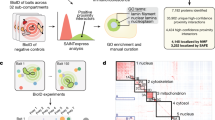
We next performed proximity-dependent labelling of RNA by BS2–AC-2 in multiple compartments, including the cytosol, nucleus, mitochondrial matrix and nucleolus, and analysed enriched RNAs by high-throughput sequencing, the resulting pipeline which we named BAP-seq (Fig. 6a). Before proceeding to BAP-seq, we tested a gradient of AC-2 concentrations and treatment times to optimize the treatment conditions for the best labelling output. Interestingly, we observed BS2-dependent proximity labelling in all tested concentrations (25–100 µM) and treatment times (1–10 min), further establishing the robustness of the technology. In general, longer treatment times and higher concentrations increased labelling and background signal (non-BS2-dependent signal measured in non-transfected cells) (Supplementary Figs. 21 and 22). As such, 3 min of treatment with 50 µM AC-2 resulted in the best signal-to-noise ratio for biomolecule labelling, being comparable to that of the APEX system (Supplementary Fig. 23).
a, The schematic workflow of BAP-seq. b, A volcano plot depicting the enrichment of mitochondrial transcripts via BAP-seq in cells expressing mitochondrial BS2, compared with cells expressing BS2 in the cytoplasm with cut-offs of log2(fold change) >0.75 and adjusted P value <0.05. c, A volcano plot depicting the enrichment of mitochondrial transcripts via BAP-seq in cells expressing nuclear BS2, as compared with cells expressing BS2 in the cytoplasm with cut-offs of log2(fold change) >0.75 and adjusted P value <0.05. d, A volcano plot depicting the enrichment of mitochondrial transcripts via BAP-seq in cells expressing BS2 in the nucleolus, as compared with the nucleus with cut-offs of log2(fold change) >0.75 and adjusted P value <0.05. In b–d, the horizontal dashed line indicates adjusted P value = 0.05. The vertical dashed line indicates log2(fold change) = 0.75 or −0.75. The enrichment values were obtained from DEG analysis using Limma. FDR, false discovery rate.
We then treated HEK293T cells with AC-2 under the optimized conditions after transiently expressing BS2 in each compartment of interest. We lysed the cells and isolated total RNA. Our initial trial enrichment was performed with 25 µg of RNA per sample. The labelled RNA was isolated by clicking on a biotin tag and subsequent enrichment with streptavidin magnetic beads. We detected a significant amount of enriched RNA that is both BS2 expression and AC-2 dependent (Extended Data Fig. 9a), in line with the dot blots. A subsequent enrichment with 50 µg of input RNA followed by analysis by high-resolution automated electrophoresis revealed a strong de-enrichment of ribosomal RNAs in the mitochondria and nucleus and an enrichment of rRNAs in the nucleolus, as expected (Supplementary Fig. 24). Given these results, we next subjected the enriched RNAs to polyA capture and analysis by RNA-seq.
To test whether BS2 transfection led to any aberration changes to the transcriptome, we compared the counts per million (CPM) reads of non-transfected input samples (a fraction of total cellular RNA) to the inputs of nuclear-, mitochondrial- and cytosolic BS2-transfected samples (Supplementary Fig. 25), which revealed no significant changes in the transcriptome of cells. Principal component analysis of the enriched samples of the RNA-seq experiments showed distinct populations for cytosol, mitochondria, nucleus and nucleolus, suggesting that different subsets of RNA were enriched for each (Extended Data Fig. 9b). For these analyses, we compared RNA-seq results between pairs of AC-2 treated biological samples expressing BS2 in separate compartments. The enrichment values were obtained from differentially expressed gene (DEG) analysis using Limma with the cut-offs of log2fold change of 0.75 and P value of 0.05, based on similar pipelines25,26. Our analysis pipeline, which compared the relative enrichment between samples expressing BS2 in different compartments, identifies transcripts that are differentially localized in one locale versus another.
In a simple assessment, we compared enriched transcripts when BS2 was localized to the mitochondrial matrix versus when BS2 was localized to the cytoplasm. As expected, we observed several mitochondrial encoded RNAs—all 13 mitochondrial messenger RNAs (mt-mRNAs), both mt-rRNAs (MT-RNR1 and MT-RNR2) and some mt-transfer RNAs (MT-TL1, MT-TE and MT-TA)—were enriched in these datasets (Fig. 6b). Using the standard cut-off of P < 0.05 (refs. 25,29), a total of 190 RNAs were enriched in the mitochondria. A more stringent cut-off of P < 0.01 identified 70 RNAs enriched in mitochondria, including all the mt-mRNAs and mt-rRNAs. We then compared the enrichment of mitochondrial encoded transcripts when BS2 was expressed in the mitochondrial matrix versus other compartments and found that samples expressing mitochondrial BS2 have more normalized counts for mt-mRNAs than any other samples. These data demonstrate that mt-RNAs are consistently enriched in mitochondria samples, regardless of which compartment is used for comparison (Extended Data Fig. 9c), as expected, and indicates the robustness of BAP-seq. In addition to the known mitochondrial transcripts, we found long non-coding RNAs (lncRNAs), such as MALAT1 and NEAT1, enriched in our mitochondrial datasets relative to the cytosol (Supplementary Fig. 26). These have recently been discovered to partially localize to mitochondria50,51. As such comparisons of enriched datasets are prone to sub-sampling bias, we tested mitochondrial enrichment with respect to cytosol post input normalization. The resultant volcano plot shows better enrichment of bona fide mitochondrial-encoded transcripts (Supplementary Fig. 27a), and the top enriched transcripts are common to both analyses (Supplementary Fig. 27b), indicating that the enriched datasets using both methods are comparable.
Given BAP-seq’s ability to faithfully enrich known mitochondrial RNAs, we next analysed the samples expressing nuclear BS2. We expected and found an enrichment of intronic sequences in the nucleus datasets when compared across all compartments (Extended Data Fig. 9d) despite performing polyA enrichment, which may not capture some intronic RNA species. Next, we compared RNAs enriched in our nuclear dataset to the cytosol, where we found 2,375 selectively enriched RNAs in the nucleus, including over 1,000 lncRNAs, when using the conventional P value cut-off of 0.05 (Fig. 6c). With the more stringent P value cut-off of 0.01, we found 1,473 RNAs enriched in our dataset, including several known nuclear lncRNAs. While the majority of the identified lncRNAs have not been characterized, our dataset identified several well-studied nuclear lncRNAs, such as MALAT1, XIST and NEAT1, as well as other lesser-studied lncRNAs that are known to reside in the nucleus, such as INE1, MIAT, TSIX, GAS5, PURPL, RAD51-AS1, MIR503HG and LINC00839 (Supplementary Fig. 28a).
Finally, we compared transcripts enriched with nucleolar BS2 versus nuclear BS2—which also includes the nucleolus—a very stringent comparison as both BS2 proteins are present in the same membrane-bound compartment (the nucleus) but in different sub-compartments (nucleoplasm versus nucleolus). In accordance with the highly differentiated labelling obtained following nucleolar BS2 expression compared with nuclear BS2 expression (Fig. 3), we observed distinct enrichment of transcripts in the two datasets. We found enrichment of validated nucleolar-residing mRNA, IGF2 (ref. 52) and several small nucleolar RNAs known to reside in the nucleolus, such as SNORA22, SNORD3B-2 and SNORA14B53, and corresponding de-enrichment of known nucleoplasm-residing lncRNAs, such as XIST, NEAT1 and MIAT54,55 (Fig. 6d). We sought to validate two of the disease associated lncRNAs identified in our nucleolar datasets—HELLPAR (associated with HELLP syndrome56) and OSTM1-AS1 (a known oncogenic factor57) by RNA fluorescence in situ hybridization (FISH) imaging. FISH analysis of the two lncRNAs confirmed preferential nucleolus localization of both transcripts over nucleoplasm (Supplementary Figs. 29–31). Taken together, these data demonstrate that BAP-seq can identify transcripts selectively enriched between compartments (that is, mitochondria versus cytoplasm versus nucleus), as well as between sub-compartments within a single membrane-bound region of the cell (nucleolus versus nucleus).
Discussion
Since its debut in 2004, the biotin ligase-generated biotin–AMP anhydride system has been developed into the most widely used non-radical proximity-dependent protein labelling platform. However, despite significant advances in radical and carbene systems, non-radical based labelling chemistries have been limited. The synthetic strategies shown here introduce a class of tunable, enzyme-activable, acylating agents that can be applied for proximity labelling of RNA and probably further applications in proteins, glycans, metabolites and other biomolecules with known propensities to react with acylating agents. Our method could also be adapted to unmask natural bioactive electrophiles, a critical class of spatially regulated signalling molecules58. To bypass the transfection or protein localization challenges in certain model systems, RNA proximity labelling experiments can be performed in fixed cells, as recently demonstrated with the oligonucleotide-directed proximity-interactome mapping method59. Given the activity of purified BS2 in vitro experiments, BAP-seq is probably amenable to adaptation to a fixed cell protocol, which we are currently pursuing. While we utilized a suite of thioesters and acid chloride-based acylating agents in this study, the concept presented here should, in principle, be adaptable to a wide range of acylating agents, opening opportunities to tune chemistries for a desired reactivity, tuning reactivity for specific classes of biomolecules and adjusting the radius of labelling.
Intriguingly, by imaging co-localization data, we observed excellent micrometre (µm)-scale labelling of membrane-bound compartments, such as the nucleus, and smaller membrane-free compartments, including the nucleolus (hundreds of nanometres) and nuclear pores (~100 nm). These data indicate a tight labelling radius around BS2 expression, which is expected on the basis of the high reactivity of acid chlorides in aqueous biological conditions. While immunofluorescence experiments are a quick way to assess the labelling proximity to BS2 expression, it should be noted that immunofluorescence shows all labelled biomolecules, not just RNA, and the acid chloride can probably react with nucleophilic sites on other classes of biomolecules. Thus, BAP-seq datasets are a better measure of RNA labelling specificity. Though BAP-seq offers good subcellular resolution, it is limited by the localization specificity of the expressed BS2. As overexpression of proteins risks mislocalization, BS2 expression, and proper subcellular localization, requires empirical tuning and validation by immunostaining for any given experiment. However, the fact that BAP-seq works even with transient transfection adds more flexibility to future applications. Notably, the high activity level of BS2, because of fast enzyme kinetics (Extended Data Fig. 1), bolsters the efficiency of the BS2–AC-2 system. Indeed, even in cells expressing very little BS2—a common consequence due to transfection heterogeneity—we still observed a robust AC-2 signal. This indicates that probe concentration, rather than enzyme (BS2) concentration, limits labelling, which is an advantage in such a system. However, this also means that any mislocalized BS2, which is inevitable, will lead to background in the experiment.
The BS2–AC-2 pair and resultant BAP-seq do not require light or oxidants, which provides a potentially less perturbative method compared with existing RNA proximity labelling approaches and significantly simplifies the experimental workflow. The high temporal resolution of our method (3 min) and low probe concentration (50 µM) are also probably less perturbative than other methods (Extended Data Table 1). In the analysis of BAP-seq datasets, we performed a stringent comparison by comparing two different cellular compartments to identify preferential RNA localization. Alternatively, as is done in other methods24, one can also compare the enriched transcripts of a given compartment to that of input samples (total cellular RNA) to identify bona fide residents of that compartment. While polyA capture was used in this proof-of-principle BAP-seq analysis, alternate library preparation strategies, such as rRNA depletion or whole transcriptome sequencing, should be considered to capture the non-polyadenylated RNAs of interest.
Aside from the technological advances in this work, we also generated datasets identifying RNA species present in various locales using BAP-seq. The datasets yielded plenty of known targets, validating the method, as well as many targets not previously reported but that warrant further investigation. When comparing our nuclear-enriched dataset with that of APEX-seq, we found that, out of our top 25 enriched RNAs with the lowest P value, 23 overlap with the APEX-seq dataset, and out of the top 50, 37 are present in the APEX-seq enriched nuclear datasets (Supplementary Fig. 32). We listed the top 100 lncRNAs enriched in the nuclear dataset over cytosol, which consists of known nuclear lncRNAs, such as MALAT1, XIST and NEAT1, and several understudied lncRNAs with no validation of subcellular localization in HEK293T cells (Supplementary Fig. 28b). In the nucleolus, even when using a very stringent cut-off of log2fold change greater than 1.5 and an adjusted P value of 0.01, we identified 459 RNAs enriched in the nucleolus versus the nucleus, which includes IGF2. Among those, we also identified and validated two disease-relevant transcripts not previously known to localize to the nucleolus-HELLPAR and OSTM1-AS1. Collectively, these observations indicate that BAP-seq provides a robust hypothesis generation pipeline across multiple cellular locales.
Each RNA proximity labelling technology will have its own false positives and negatives, and in some cases, identified targets may be investigated and not yield expected outcomes, perhaps owing to low abundance or their ephemeral presence in non-canonical locales. Therefore, combining multiple approaches with distinct chemistries will likely improve data robustness. BAP-seq has some clear advantages, including avoidance of light or ROS, tunable chemistry based on probe acylation strength and rapid kinetics. More broadly, the chemistry established here opens up the possibilities to trigger labelling reagents based on new input signals or couple with reactivity-based RNA profiling strategies60, thus allowing mapping of RNA localization in more intricate ways. With newer and more refined tools, proximity-labelling chemistries will be vital to moving beyond the ‘parts list’ of biology, which is now quite established, to a better understanding of the dynamic organization and reorganization of biomolecules during physiological and pathological processes.
Methods
General methods
For chemical synthesis, reagents and dry solvents were purchased from commercial sources (Sigma-Aldrich, Combi-blocks and Thermo Fisher) and used without further purification. Silica gel P60 (SiliCycle, 40–63 μM, 230–400 mesh) was used for column chromatography. Analytical TLC was performed using SiliCycle 60 F254 silica gel (pre-coated sheets, 0.25-mm-thick) with detection at 214 nm. Low-resolution mass-spectral analysis and liquid chromatography analysis were carried out on an Advion Expression-L mass spectrometer with electron spray ionization in the positive mode, coupled to an Agilent 1220 Infinity LC System with an Agilent Poroshell 120 column. Automated flash column chromatography purification was carried out on a Biotage system Isolera One using SNAP Biotage columns. Nuclear magnetic resonance (NMR) spectra were recorded on the BRUKER Ascend 400 at the Department of Chemistry NMR Facility at the University of Chicago, for 1H 400 MHz and 13C 101 MHz measurements. Chemical shifts are given in parts per million (δ) referenced to tetramethylsilane (δ = 0.00 ppm 1H-NMR and 13C-NMR). The coupling constants are given in hertz. The high-resolution mass spectra measurements were performed on an Agilent 6224 TOF using a combination of atmospheric-pressure chemical ionization and electrospray ionization at the Department of Chemistry Mass Spectrometry Facility at the University of Chicago.
Synthesis methods
The detailed synthetic methods for all the probes are provided in the Supplementary Information.
Cloning
All the plasmids were constructed by Gibson Assembly with polymerase chain reaction products generated using Q5 or Phusion DNA polymerases (New England Biolabs). The plasmids were sequenced by the University of Chicago Comprehensive Cancer Center DNA Sequencing and Genotyping facility. The full vector sequences are included in Supplementary Fig. 33.
Mammalian cell culture and plasmid transfection
HEK293T (ATCC) cells were cultured in Dulbecco’s modified Eagle medium (DMEM; GlutaMAX, high glucose, sodium pyruvate and Phenol Red; Thermo Fisher) supplemented with 10% (v/v) fetal bovine serum (Gemini Benchmark) and 1% (v/v) penicillin–streptomycin (Gibco/Life Technologies). The cells were maintained in a water-saturated, 5% CO2-containing, 37 °C incubator. The cells used for experiments never exceeded passage number 25. Fresh HEK293T cells were obtained from ATCC and frozen at an early passage number (five) in individual aliquots. There was no testing for mycoplasma infection as a result. The transient transfections were performed using Lipofectamine 3000 (Thermo, L3000015; Invitrogen) following the manufacturer’s protocol.
Cell viability/MTS assay to test compatibility of BS2 expression in different subcellular compartments
HEK293T cells were plated on a six-well dish without antibiotics and at 70% confluency. Each well was transfected with 2,000 ng of empty vector or different BS2 vectors. At 24 h post transfection, they were trypsinized, and an equivalent number of cells for each condition were replated into 10 wells of a 96-well plate. The cells were allowed to grow for another 2 days before replacing the media with 20% 3-(4,5-dimethylthiazol-2-yl)-5-(3-carboxymethoxyphenyl)-2-(4-sulfophenyl)-2H-tetrazolium (MTS) reagent (6 mM MTS, Biovision and 0.33 mM phenosine methosulfate, Sigma-Aldrich) in DMEM. The cells were incubated with the MTS reagent for 4 h at 37 °C and 5% CO2 in the incubator. The totalformazan product formed from the MTS reagent was then measured by recording the absorbance at 490 nm using the Synergy Neo2 hybrid multi-mode reader (BioTek Instruments). The background absorbance from the empty wells containing the MTS reagent was subtracted from all the samples, followed by normalization to that of the empty vector.
Recombinant protein expression and purification
The pET–BS2 plasmid (Supplementary Fig. 33) was transformed into chemically competent Escherichia coli BL21 cells. A total of 500 ml cultures were grown at 37 °C in lysogeny broth (40 μg ml−1 Kan) in a 2 litre flask to mid-log phase (measured by absorbance OD600 of 0.65). The cultures were induced with 0.2 mM isopropyl β-d-1-thiogalactopyranoside and incubated at 30 °C overnight. The cells were collected by 4 °C centrifugation (4,000g for 15 min), the supernatants were discarded and the pellets were resuspended in 30 ml of lysis buffer (50 mM Tris (pH 7.5), 1 M NaCl, 10 mM tris(2-carboxyethyl)phosphine hydrochloride (TCEP) and 20% glycerol) and protease inhibitor cocktail consisting of bestatin (10 μM), pepstatin A (1 μM) and E-64 (20 μM), avoiding serine protease inhibitors that could inhibit BS2 enzymatic function. The mixtures were then sonicated (total on time: 10 min, with 10 s on and 20 s off; amplitude 30%) and spun down at 12,000 rpm for 40 min. The BS2 was purified from the supernatants by nickel affinity chromatography. After discarding the cell debris, the supernatant was incubated with 1 ml Takara His60 Ni Superflow Resin with gentle rotation at 4 °C for 1 h. The His-tagged proteins were purified using a standard protocol of washing and eluting the resin with wash buffer 1 (20 mM imidazole, 50 mM Tris, 1 M NaCl, 20% glycerol and 10 mM TCEP; pH 7.5), wash buffer 2 (40 mM imidazole, 50 mM Tris, 1 M NaCl, 20% glycerol and 10 mM TCEP; pH 7.5) and elution buffer (300 mM imidazole, 50 mM Tris, 1 M NaCl, 20% glycerol and 10 mM TCEP; pH 7.5). The purified proteins were then desalted on GE Disposable PD-10 desalting columns and stored in the protein storage buffer (50 mM Tris, 100 mM NaCl, 2 mM TCEP and 50% glycerol; pH 7.5). The final BS2 concentrations were measured using a standard bicinchoninic acid assay.
Measurement of BS2 kinetics on mCP-coumarin substrate
A varying concentration of mCP-coumarin (50–800 µM) was added to 5 nM BS2 in PBS containing 10% DMSO by volume at room temperature. Higher concentration (greater than 800 µM) of mCP-coumarin crashed out in PBS and was, thus, avoided for the assay. For completely unmasked mCP-coumarin control, 0.5 M NaOH was added to 10× stocks of different concentrations of mCP-coumarin (0.5–8 mM) in DMSO for 5 min, followed by the addition of 0.5 M HCl to neutralize. They were diluted with PBS to get the final concentrations of completely unmasked mCP-coumarin (50–800 µM). All the reactions were plated in triplicates in a 96-well plate, and the kinetic measurement of BS2 unmasking mCP-coumarin was recorded using the Synergy Neo2 hybrid multi-mode reader (BioTek Instruments). The fluorescence measurements (λex = 320/20 nm and λem = 450/20 nm) were recorded every 30 s for 20 min immediately after the addition of mCP-coumarin to BS2. The fluorescence signal of the NaOH-treated samples corresponded to the maximum fluorescence signal and was used to convert random fluorescence units to corresponding concentrations of 7-hydroxycoumarin. Using the initial few minutes of kinetic measurements for each concentration of mCP-coumarin, the linear rates of BS2 unmasking/7-hydroxycoumarin generation were obtained. The linear rates of unmasking at each concentration were then plotted using GraphPad Prism and fit with a Michaelis–Menten curve to obtain maximum velocity (Vmax) and Km. Kcat was obtained by dividing Vmax by the BS2 concentration used in the assay (5 nM).
For the PMSF inhibition assay, 100 nM of BS2 was incubated with 1 mM PMSF or DMSO control for 5 min before the addition of 100 µM of mCP-coumarin in PBS. For completely unmasked mCP-coumarin control, 0.5 M NaOH was added to 1 mM mCP-coumarin in DMSO, followed by 0.5 M HCl and was finally diluted to 100 µM of unmasked mCP-coumarin with PBS. Immediately after adding the mCP-coumarin to BS2, kinetic fluorescence measurements were carried out, as described above, in 30 s intervals for 20 min.
LC–MS detection of TE-1 stability and BS2 activity in PBS
A total of 0.7 mM of TE-1 was incubated in 500 µl PBS containing 15% DMSO by volume at room temperature in the presence of a reference compound (1,3,5-trimethoxybenxzene, 621-23-8; Sigma-Aldrich). At regular intervals (1 h), 20 µl of the solution was removed, diluted with 20 µl of methanol and assayed by liquid chromatography–mass spectrometry (LC–MS). The absorbance maxima, normalized to that of the reference peak, were plotted as a function of time. To test the activity of BS2 on TE-1, 0.85 mM TE-1 was dissolved in 350 µl PBS containing 15% DMSO by volume. A total of 30 µl of the solution was taken out at this stage, diluted with an equal volume (30 µl) of methanol and assayed by LC–MS. The BS2 stock solution was added to the TE-1 solution in PBS to achieve 385 nM final BS2 concentration and incubated at 37 °C with shaking at 900 rpm. After 1, 2 and 5 min, 30 µl of the solution was removed, quenched with an equal volume of methanol and assayed by LC–MS.
LC–MS detection of AC-5 stability, BS2 activity and trapping of acid chloride with Fmoc–Lys in PBS
A total of 0.5 mM AC-5 was dissolved in 500 µl PBS containing 30% DMSO by volume at room temperature. After 0, 30 and 60 min, 50 µl aliquots of the solution were removed and assayed by LC–MS. To test BS2 activity on AC-5, 0.5 mM AC-5 was dissolved in 100 µl of PBS containing 30% DMSO by volume. After recording LC–MS spectra for the no-BS2 control, 1 µl of BS2 stock solution was added to the solution at room temperature to achieve 500 nM final BS2 concentration, which was then assayed by LC–MS immediately. To trap the active acylating agent, 0.5 mM of Fmoc–l-Lys–methyl ester and 0.5 mM AC-5 was dissolved in PBS containing 30% DMSO by volume and assayed by LC–MS before and after adding BS2 at a final concentration of 500 nM.
TLC assay to detect BS2 activity on TE and AC probes
A total of 200 nM BS2 was added to each of the AC and TE probes in PBS containing 10% DMSO and incubated at 37 °C for 1 min and then quenched by an equal volume of methanol. An equal volume of this reaction mixture and all the controls (no-probe and no-BS2) was then dotted onto a TLC plate, developed in hexane/ethyl acetate solvent mixture and visualized by ultraviolet (UV) light, iodine or KMnO4 staining.
Fluorescence microscopy
For epi-fluorescence microscopy, an inverted epi-fluorescence microscope (Leica DMi8) equipped with a Hamamatsu Orca-Flash 4.0 camera, a 63× oil objective (numerical aperture 1.4) and a 300 W xenon light source (Sutter Lamda XL) was used. Leica LASX software was used to obtain images for Alexa-488 (490/20×, ET Quad-S, ET 525/36 m), Hoechst 33342 (ET 402/15×, Quad-S, ET 455/50 m), Alexa-555 (ET 555/25×, Quad-S, ET 605/52 m) and bright field. The acquisition time ranged from 30 to 800 ms. The acquisition time and fluorescence intensity manager were kept constant for a given channel across all samples in an experiment that was compared. For confocal microscopy, the slides were imaged on a Leica Stellaris 8 laser scanning confocal microscope (DMI8-CS) equipped with near-UV and white light laser (WLL). The 4′,6-diamidino-2-phenylindole (DAPI; HyD S1 detector, 405 nm laser excitation and 430/500 emission), Alexa-488 (HyD X2, 499 nm laser excitation and 510/560 nm emission), Alexa-555 (HyD S3 detector, WLL 80 mHz, 558 nm laser excitation and 590/650 nm emission) and bright field (Trans PMT and WLL 80 mHz) images were taken using a 63×/1.4 UV oil objective at 3× magnification with eight-line averaging on a single focal plane without Z stacking.
The HEK293T cells were plated without antibiotics on glass coverslips (Neuvitro Corporation) pre-treated with 0.1 mg ml−1 poly-d-lysine (for at least 2 h at room temperature) and contained within a 24-well dish. A total of 18–24 h post plating, the cells were transfected with 500 ng of the respective BS2 plasmids or no-BS2-containing dummy plasmid at 60% confluency. A total of 24 h post transfection, the relevant concentration of the corresponding probe or DMSO in pre-warmed DMEM was added for varying incubation times at 37 °C. The cells were then incubated with 1 mM PMSF in Dulbecco’s phosphate-buffered saline (DPBS) for 1 min to stop labelling by quenching BS2, followed by 2× washes with DPBS. The cells were then fixed with 4% paraformaldehyde (15710; Electron Microscopy Science) solution in PBS for 15 min, washed twice with PBS, permeabilized with ice-cold MeOH for 8 min at −20 °C and washed twice with PBS. The samples were then incubated in a CuAAC reaction solution in PBS with the following final concentrations: 2 mM 2-(4-((bis((1-(tert-butyl)-1H-1,2,3-triazol-4-yl)methyl)amino)methyl)-1H-1,2,3-triazol-1-yl)acetic acid (BTTAA; Click Chemistry Tools, 1236-100), 1 mM CuSO4, 10 μM of 488-PEG4-alkyne (Sigma, 761621) for azide-handle probes or AzDye 488 azide (Click Chemistry Tools, 1275-5) for alkyne handle probes and 10 mM sodium ascorbate (prepared fresh). The CuSO4 was omitted for the no-click control. The BTTAA, CuSO4 and 488 alkyne/azide were pre-mixed, and the sodium ascorbate was added last, immediately before cell bathing. The samples were incubated in the CuAAC solution for 1 h at room temperature in the dark, followed by three washes with PBS for 5 min each. The samples were then blocked with 3% bis(trimethylsilyl)acetamide (BSA) in PBS for 1 h at room temperature, followed by overnight incubation with 1:1,000 diluted anti-V5 antibody (Invitrogen, R960-25) in 3% BSA in PBS at 4 °C. The following day, after three washes with PBS, 1:1,000 diluted secondary antibody-anti mouse Alexa-555 (Invitrogen, A21427) in 3% BSA in PBS was added for 1 h at room temperature. The samples were washed twice with PBS and incubated with 1 μM of DAPI in PBS for 5 min, followed by two more washes with PBS. After removing the last PBS wash, the cell-containing coverslips were then removed from the 24-well plate, inverted and transferred to the fixative (ProLong diamond antifade mountant, ThermoFisher; P36961), pre-dotted on a glass slide. The slides were dried for at least 5 h in the dark before imaging. The samples were imaged on the inverted epi-fluorescence microscope or Leica Stellaris 8 laser scanning confocal microscope.
Image analysis
All image analyses were performed in ImageJ. Measurements of four different images/frames of the same slide were taken for each condition. For Fig. 3b, the signal intensity for the probe channel (Alexa-488) was measured with thresholding from the BS2 channel (Alexa-555) to count the signal from transfected cells only. An auto-threshold was used to choose BS2-transfected nuclei in the AF-555 channel of each image, resulting in the loss of some low-expressing cells, which were excluded from the quantification. All signals were normalized to the fluorescence signal from the nucleus of non-transfected DMSO-treated cells, which was measured by DAPI thresholding.
To compare the signal-to-noise ratio for each probe in Supplementary Figs. 9–17, the signal intensity for the probe channel (Alexa-488) was measured with thresholding from the DAPI channel to select the nuclear signal, which included both transfected and non-transfected nuclei. This signal was normalized to the signal intensity of the probe/Alexa-488 channel with DAPI thresholding of non-transfected samples treated with the respective probes.
All fluorescent images were analysed as collected without additional filtering or background subtraction. The brightness and contrast were adjusted for better visualization of the probe signal, and the optimized settings were applied to all images of the probe channel compared in each panel.
For co-localization analysis, on confocal images, a straight line was drawn across the object of interest (nuclei, nuclear pores or nucleoli) in the raw images of the BS2 channel (Alexa-555) spanning no apparent signal on both ends and a strong signal at the centre. Using the region-of-interest manager in ImageJ, the straight line was transferred to the probe channel, and the plot profile was measured along the line in both channels. The normalized plot profiles were then plotted for each object.
In vitro RNA labelling
A total of 3 µg of a 74-mer RNA oligo (a gift from the Szostak laboratory) (UCUGUGUGGGGUAGGUGGUGCUGGGUAAUGGGACUGCAUUGCAUAUCCCUAGGCACCUAUUGAGAUUUCUCUG) in PBS was boiled at 92 °C for 1 min and immediately cooled to 4 °C for 5 min. Purified BS2 and AC-5 were added to the folded RNA solution to get a final concentration of 600 nM BS2 and 500 µM AC-5 in a reaction volume of 50 µl in PBS. The reaction mixture was incubated at 37 °C at 900 rpm for 5 min. The labelled RNA was then purified using the RNA Clean and Concentrator Kit (Zymo, RCC-5) and eluted in 10 µl of water. For RNase treatment, purified labelled RNA was incubated with 85 µg ml−1 of RNase A at 37 °C for 15 min, re-purified using the RCC-5 kit and eluted in 10 µl of water. The entire RNA product for all the treatment conditions was loaded onto a 10% urea polyacrylamide gel electrophoresis gel with 2× RNA loading dye (New England Biolabs) and run at 80 V for 2 h. AC-5 labelling was observed by fluorescence imaging using the Amersham Typhoon laser scanner in the Cy2 channel. After fluorescence imaging, the gel was incubated in SYBR Gold for 15 min and imaged under UV for RNA loading control.
In cell AC-2 RNA labelling and detection by streptavidin dot blot
The HEK293T cells were plated in a 12-well plate without antibiotics and transfected with 800 ng of cyto-BS2, NLS-BS2, mito-BS2 or no-BS2-dummy plasmid at 60% confluency. A total of 24 h after transfection, all the wells were treated with 25 µM of AC-2 or DMSO in pre-warmed DMEM for 10 min at 37 °C. The labelling reaction was quenched by treating the cells with 1 mM PMSF in DPBS for 1 min, followed by two rounds of DPBS washing. The cells were lysed immediately, and the RNA was extracted using the QIAGEN kit. Eluted RNA was then clicked with biotin at 25 °C for 30 min by adding 5× click mixture to get the final concentration of 0.4 mM CuSO4, 2 mM biotin picolyl azide (Sigma-Aldrich, 900912), 1 mM tris-hydroxypropyltriazolylmethylamine (THPTA; Click Chemistry Tools, 760952-88-3) and 10 mM sodium ascorbate along with 1 mM SUPERase•In RNase inhibitor (Thermo Fisher, AM2694) in PBS. The click reaction was quenched with 10 mM EDTA, and the RNA was purified using the Zymo RCC-25 kit. For every sample, 1.5 μg of biotinylated RNA was treated with either DMSO, RNase A (85 μg ml−1), or proteinase K (85 μg ml−1) at 25 °C and 900 rpm for 30 min, followed by Zymo RCC-5 clean up. RNA was eluted in 12 µl of water, split equally and blotted in two Hybond membranes (Cytiva, RPN1210B) presoaked in 2× saline–sodium citrate (SSC) buffer (Thermo Fisher, BP1325-1) and air dried. The blotted RNA on the membrane was allowed to dry at room temperature for 15 min before UV crosslinking at 2,400 mJ cm−2 (254 nm wavelength; UV Stratalinker 2,400). Both the membranes were washed twice with Tris-buffered saline with 0.1% Tween-20 detergent, and one of the blots was used for methylene blue staining for 5 min, washed several times with water and imaged for RNA loading control. The other blot was blocked with 3% BSA solution in phosphate-buffered saline with 0.1% Tween-20 detergent (PBST) at room temperature for 1 h, followed by overnight incubation with 1:2,000 streptavidin–horseradish peroxidase antibody (Sigma-Aldrich, S2438) in 3% BSA solution in PBST. The next day, the blot was washed thrice with PBST and imaged for chemiluminescence using SuperSignal West Pico PLUS chemiluminescent substrate (Fisher Scientific, 34577) in an Azure c300 imager.
Comparison of RNA labelling by APEX2 versus BS2–AC-2 labelling by dot blot
The HEK293T cells were transfected with 2,000 ng of nuclear BS2, nuclear-APEX2 or dummy plasmid at 60% confluency on a six-well plate. A total of 24 h post transfection, the labelling reaction was carried out. For APEX-seq samples, the cells were incubated with 500 µM of biotin tyramide in full media without antibiotics for 30 min at 37 °C in the incubator. H2O2 was then added to the samples to achieve a final concentration of 1 mM, and the plate was gently agitated for 1 min. The reaction was quenched by replacing the media with quenching solution consisting of 5 mM Trolox, 10 mM sodium ascorbate and 10 mM sodium azide in DPBS. The cells were washed twice with the quenching solution and once with DPBS. For BS2–AC-2 samples, 24 h post transfection, cells were treated with 50 µM of AC-2 probe in pre-warmed DMEM for 3 min at 37 °C. The media was then replaced with 1 mM PMSF in DPBS, incubated for 1 min and washed with DPBS. After APEX and AC-2 labelling, the cells were lysed, and the RNA was extracted using the Qiagen kit. The RNA samples from BS2–AC-2 labelling was then clicked with biotin for 30 min at room temperature using the click reaction mixture of 0.4 mM CuSO4, 2 mM biotin picolyl azide, 1 mM THPTA and 10 mM sodium ascorbate along with 1 mM SUPERase•In RNase inhibitor in PBS. The click reaction was quenched with 10 mM EDTA, and the biotinylated RNA was purified using the Zymo RNA Clean and Concentrator Kit-25. The RNA concentration of all the samples was measured by nanodrop. A total of 5 µg of RNA for each sample was then blotted onto each of the two Hybond membranes, and the dot blot assay was performed as described above.
Sample preparation for BAP-seq
HEK293T cells were transfected at 60% confluency with DNA vectors encoding cyto-BS2, nuclear BS2, mitochondrial BS2, NIK3x-BS2 (nucleolus targeting) or no-BS2-dummy plasmid, with each plasmid having two to three biological replicates. A total of 24 h post transfection, the cells were treated with 50 µM of AC-2 probe in pre-warmed DMEM for 3 min at 37 °C. The labelling reaction was quenched by incubating with 1 mM PMSF in DPBS for 1 min, followed by a quick DPBS wash. The cells were then lysed, and the RNA was extracted using the Qiagen kit. For each replicate of a BS2 construct, 50 µg of RNA (two wells of a six-well plate) was used for the next steps. The labelled RNA was clicked with biotin using the click reaction cocktail of 0.4 mM CuSO4, 2 mM biotin picolyl azide, 1 mM THPTA and 10 mM sodium ascorbate along with 1 mM SUPERase•In RNase inhibitor in PBS for 30 min at room temperature. The click reaction was quenched with 10 mM EDTA, and the biotinylated RNA was purified using the Zymo RNA Clean and Concentrator Kit-25. The biotinylated RNAs were enriched using the APEX-seq enrichment method24, with some modification. Briefly, 50 μl Pierce streptavidin magnetic beads (Thermo, 88816) were utilized per 50 μg RNA sample. The beads were washed three times on/off the magnet with 500 μl binding and wash buffer (5 mM Tris-HCl, pH 7.5, 0.5 mM EDTA, 1 M NaCl and 0.1% Tween-20) per sample, followed by two 500 μl washes with solution A (100 mM NaOH and 50 mM NaCl) per sample, one 500 μl wash with solution B (100 mM NaCl) and a final resuspension of the beads in 250 μl solution B containing 1 μl SUPERase•In RNase inhibitor per sample. The resuspended beads were mixed thoroughly with the 250 μl of eluted, biotin-labelled RNA from above and incubated at 4 °C for 2 h on a rotator. After this incubation, the samples were washed five times with binding and wash buffer as described above, and the enriched RNAs were eluted with Trizol45. For elution of enriched RNAs off the beads, they were first resuspended in 100 μl of PBS, followed by the addition of 300 μl of Trizol (Invitrogen, 16096040), and were incubated at 37 °C for 10 min to dissociate the RNAs. The beads were placed on the magnet, and the dissociated RNAs in the supernatant were pipetted out. Finally, the enriched RNAs were recovered using the Zymo Direct-zol RNA Microprep kit.
PolyA enrichment and sequencing
The enriched RNAs for all the samples were subjected to polyA capture. Subsequently, the DNA library with dual-index adaptors was generated and amplified using the mRNA Hyper Prep Kit (Kapa Biosystems, KK8581) following the manufacturer’s protocol. After amplification, the quality check of the library was performed with tapestation analysis. The libraries were sequenced on Next-seq 550 using a High Output kit at ~13 million single 75-cycle reads per library.
Analysis of BAP-seq data
The RNA-seq library generated was mapped to the Homo sapiens GRCh38 reference genome using STAR61 version 2.7.9, and the aligned reads were quantified using Salmon62 version 1.4.0. After read mapping with Salmon63, the method described in ref. 63 was used to read Salmon outputs into the R environment. Annotation data from Gencode version 34 were used to summarize data from the transcript level to the gene level. Filtering was performed to remove particular gene biotypes or genes with low expression. We first selected the gene biotype of interest: protein coding, lncRNA, Mt_rRNA, Mt_tRNA, rRNA, snRNA, scRNA and small nucleolar RNA. This reduced the number of genes from 60,728 to 39,823. The genes were then filtered for low expression, keeping only genes with greater than 1 CPM. This reduced the number of genes from 39,823 to 24,878. To identify DEGs, precision weights were applied to trimmed mean of M-values-normalized gene counts based on within-group sample-level variance and gene-level mean-variance trends using VOOM64. The count data were fitted to a gene-wise linear model with group status as a coefficient, and the contrasts were specified for comparisons of interest. The Limma65 empirical Bayes method was used to estimate the posterior odds of differential expression (|log(fold change)| >0) after adjusting for gene-level posterior residual standard deviations. Statistically significant DEGs were decided with |log2fold change| > 0.75 and a false discovery rate of 0.05. The results of differential expression testing are visualized by volcano plots comparing samples with BS2 expression in two different locales (cytoplasm versus nucleus, cytoplasm versus mitochondria and nucleus versus nucleolus).
For input-normalized analyses of mitochondrial enrichments, the CPM reads of mitochondrial- and cytoplasmic-enriched samples were first normalized to that of the grouped input reads consisting of inputs from non-transfected, cytoplasmic, mitochondrial and nuclear samples. These normalized counts were then transformed using VOOM and subjected to DEG analysis as described above.
FISH of HELLPAR and OSTM1-AS1 lncRNAs
Using the Stellaris Probe Designer from Biosearch, 37 FISH probes for HELLPAR and 28 FISH probes for OSTM1-AS1 were designed with a masking level of 5, oligo length of 20 and a minimum spacing length of two nucleotides in between two FISH probes. The oligo sequences for all the FISH probes used are provided in Supplementary Fig. 34. To make the FISH probes, 3′ amine modified oligos were first obtained from Integrated DNA Technologies and then conjugated with AlexaFluor 647 N-hydroxysuccinimide ester (Thermo Fisher, A37573) following a reported protocol66. For the fluorophore coupling, 100 µM of the pooled FISH probes for each target was added to 40 mM of AlexaFluor N-hydroxysuccinimide ester in 0.1 M NaHCO3 solution with a pH of 8.6 and was incubated overnight at 37 °C. To precipitate the oligos, at first one-ninth by volume, 3 M NaOAc (pH 5) was added, followed by 2.5 times by volume of absolute ethanol. It was allowed to precipitate overnight and was centrifuged, and the supernatant was removed. The pellet was rewashed with absolute ethanol to remove any unreacted fluorophore. The pellet was redissolved in 30–40 μl water and purified using P-6 columns (Bio-Rad, 7326221), and the final concentration of the probes was measured with nanodrop.
For the RNA FISH assay, HEK293T cells were plated on an eight-well glass chambered dish (Cellvis, C8-1.5H-N) and allowed to grow up to 70% confluency and fixed with 4% paraformaldehyde in PBS for 10 min at room temperature. After three rounds of washes with PBS, the cells were permeabilized on ice with 0.5% Triton in PBS containing 2 mM vanadyl ribonucleoside complex (Sigma-Aldrich). Post permeabilization, the cells were washed twice with PBS and twice again with 2× SSC buffer. Cells were then washed with wash buffer consisting of 10% formamide in 2× SSC buffer. The FISH probe solution with the corresponding final concentration (5 nM multiplied by the number of FISH probes for each target) was prepared by diluting the probes with 10% dextran sulfate in the wash buffer in the presence of 10 mM DTT. The cells were incubated with the FISH probe solution overnight at 37 °C. The next day, the samples were washed with wash buffer for 30 min followed by 2× washes with PBS and fixed again with 4% paraformaldehyde in PBS for 10 min at room temperature. After three rounds of washes with PBS, the cells were blocked with 5% BSA in PBS at room temperature for 1 h. A 1:100 dilution of anti-NPM1 antibody (Thermo Fisher; PA517742) in 1% BSA in PBS was added to the samples and incubated for 1 h, followed by three rounds of washes. A 1:1,000 dilution of anti-rabbit AlexaFluor-488 antibody (Abcam, ab150077) was further added for 1 h, followed by three washes with PBS. A total of 1 μM of DAPI was then added for 10 min and washed twice before proceeding to confocal microscopy.
All the FISH images shown are maximum projection of 6–35 Z stack images. For the FISH probe channel (Alexa-647), averaging over two to four lines was carried out, and the images were filtered using the median filter with a rolling ball radius of 0.8 pixels in ImageJ. After filtering, the FISH images were converted to 16-colour images using lookup tables, and the foci with at least a single pixel of yellow or higher intensity was considered a true FISH signal. For each image, the localization of each FISH foci was manually assigned to the nucleolus when it co-localized with the NPM1 channel, while otherwise it was assigned as nucleoplasmic.
Statistics and reproducibility
All the AC and TE probe immunofluorescence imaging experiments were independently repeated twice, with similar results. In Fig. 3b, Supplementary Figs. 9–17 and 21–22 and Extended Data Figs. 3–8, four images with different field of views of a single set of samples prepared at the same time are used for representation and quantification. The fluorescence in situ hybridization experiments were independently repeated thrice, with similar outcomes. The images from two independent biological replicates were used in Supplementary Figs. 29–31. The RNA-seq experiment was performed once with two to three independent biological replicates per sample.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All BAP-seq data are available in the Sequence Read Archive through accession number GSE229451. All the raw images used for Supplementary Figs. 11–30 and Figs. 3 and 4 are provided with Mendeley Data (https://doi.org/10.17632/kzpvsdjrgy.1)67. All other data generated or analysed in this study are available within the main text and Supplementary Information. Links to vector maps are included in Supplementary Fig. 33. Source data are provided with this paper.
References
Keskin, O., Tuncbag, N. & Gursoy, A. Predicting protein–protein interactions from the molecular to the proteome level. Chem. Rev. 116, 4884–4909 (2016).
Hentze, M. W., Castello, A., Schwarzl, T. & Preiss, T. A brave new world of RNA-binding proteins. Nat. Rev. Mol. Cell Biol. 19, 327–341 (2018).
Engreitz, J. M., Ollikainen, N. & Guttman, M. Long non-coding RNAs: spatial amplifiers that control nuclear structure and gene expression. Nat. Rev. Mol. Cell Biol. 17, 756–770 (2016).
Rinn, J. & Guttman, M. RNA function. RNA and dynamic nuclear organization. Science 345, 1240–1241 (2014).
Tiruchinapalli, D. M. et al. Activity-dependent trafficking and dynamic localization of zipcode binding protein 1 and beta-actin mRNA in dendrites and spines of hippocampal neurons. J. Neurosci. 23, 3251–3261 (2003).
Huttelmaier, S. et al. Spatial regulation of beta-actin translation by Src-dependent phosphorylation of ZBP1. Nature 438, 512–515 (2005).
Satori, C. P., Kostal, V. & Arriaga, E. A. Review on recent advances in the analysis of isolated organelles. Anal. Chim. Acta 753, 8–18 (2012).
Qin, W., Cho, K. F., Cavanagh, P. E. & Ting, A. Y. Deciphering molecular interactions by proximity labeling. Nat. Methods 18, 133–143 (2021).
Seath, C. P., Trowbridge, A. D., Muir, T. W. & MacMillan, D. W. C. Reactive intermediates for interactome mapping. Chem. Soc. Rev. 50, 2911–2926 (2021).
Rhee, H. W. et al. Proteomic mapping of mitochondria in living cells via spatially restricted enzymatic tagging. Science 339, 1328–1331 (2013).
Lam, S. S. et al. Directed evolution of APEX2 for electron microscopy and proximity labeling. Nat. Methods 12, 51–54 (2015).
Hung, V. et al. Proteomic mapping of the human mitochondrial intermembrane space in live cells via ratiometric APEX tagging. Mol Cell 55, 332–341 (2014).
Han, S. et al. Proximity biotinylation as a method for mapping proteins associated with mtDNA in living cells. Cell Chem. Biol. 24, 404–414 (2017).
Hung, V. et al. Proteomic mapping of cytosol-facing outer mitochondrial and ER membranes in living human cells by proximity biotinylation. eLife 6, e24463 (2017).
Loh, K. H. et al. Proteomic analysis of unbounded cellular compartments: synaptic clefts. Cell 166, 1295–1307(2016).
Rees, J. S., Li, X. W., Perrett, S., Lilley, K. S. & Jackson, A. P. Selective Proteomic Proximity Labeling Assay Using Tyramide (SPPLAT): a quantitative method for the proteomic analysis of localized membrane-bound protein clusters. Curr. Protoc. Protein Sci. 80, 19.27.1–19.27.18 (2015).
Bar, D. Z. et al. Biotinylation by antibody recognition—a method for proximity labeling. Nat. Methods 15, 127–133 (2018).
McCutcheon, D. C., Lee, G., Carlos, A., Montgomery, J. E. & Moellering, R. E. Photoproximity profiling of protein–protein interactions in cells. J. Am. Chem. Soc. 142, 146–153 (2020).
Geri, J. B. et al. Microenvironment mapping via Dexter energy transfer on immune cells. Science 367, 1091–1097 (2020).
Oakley, J. V. et al. Radius measurement via super-resolution microscopy enables the development of a variable radii proximity labeling platform. Proc. Natl Acad. Sci. 119, e2203027119 (2022).
Choi-Rhee, E., Schulman, H. & Cronan, J. E. Promiscuous protein biotinylation by Escherichia coli biotin protein ligase. Protein Sci. 13, 3043–3050 (2004).
Roux, K. J., Kim, D. I., Raida, M. & Burke, B. A promiscuous biotin ligase fusion protein identifies proximal and interacting proteins in mammalian cells. J. Cell Biol. 196, 801–810 (2012).
Branon, T. C. et al. Efficient proximity labeling in living cells and organisms with TurboID. Nat. Biotechnol. 36, 880–887 (2018).
Kaewsapsak, P., Shechner, D. M., Mallard, W., Rinn, J. L. & Ting, A. Y. Live-cell mapping of organelle-associated RNAs via proximity biotinylation combined with protein–RNA crosslinking. eLife 6, e29224 (2017).
Fazal, F. M. et al. Atlas of subcellular RNA localization revealed by APEX-Seq. Cell 178, 473–490 e426 (2019).
Padron, A., Iwasaki, S. & Ingolia, N. T. Proximity RNA labeling by APEX-Seq reveals the organization of translation initiation complexes and repressive RNA granules. Mol. Cell 75, 875–887 e875 (2019).
Zhou, Y. et al. Expanding APEX2 substrates for proximity-dependent labeling of nucleic acids and proteins in living cells. Angew. Chem. Int. Ed. Engl. 58, 11763–11767 (2019).
Wang, P. et al. Mapping spatial transcriptome with light-activated proximity-dependent RNA labeling. Nat. Chem. Biol. 15, 1110–1119 (2019).
Engel, K. L. et al. Analysis of subcellular transcriptomes by RNA proximity labeling with Halo-seq. Nucleic Acids Res. 50, e24 (2022).
McGinnis, J. L., Dunkle, J. A., Cate, J. H. & Weeks, K. M. The mechanisms of RNA SHAPE chemistry. J. Am. Chem. Soc. 134, 6617–6624 (2012).
Lee, B. et al. Comparison of SHAPE reagents for mapping RNA structures inside living cells. RNA 23, 169–174 (2017).
Xiao, L., Fang, L., Chatterjee, S. & Kool, E. T. Diverse reagent scaffolds provide differential selectivity of 2′-OH acylation in RNA. J. Am. Chem. Soc. 145, 143–151 (2023).
Sun, W. et al. Genetically encoded chemical crosslinking of RNA in vivo. Nat. Chem. 15, 21–32 (2023).
Spitale, R. C. et al. Structural imprints in vivo decode RNA regulatory mechanisms. Nature 519, 486–490 (2015).
Xiao, L., Jun, Y. W. & Kool, E. T. DNA tiling enables precise acylation-based labeling and control of mRNA. Angew. Chem. Int. Ed. Engl. 60, 26798–26805 (2021).
Xiao, L., Habibian, M. & Kool, E. T. Site-selective RNA functionalization via DNA-induced structure. J. Am. Chem. Soc. 142, 16357–16363 (2020).
Fang, L., Xiao, L., Jun, Y. W., Onishi, Y. & Kool, E. T. Reversible 2′-OH acylation enhances RNA stability. Nat. Chem. 15, 1296–1305 (2023).
Tian, L. et al. Selective esterase–ester pair for targeting small molecules with cellular specificity. Proc. Natl Acad. Sci. USA 109, 4756–4761 (2012).
Jones, K. A. et al. Development of a split esterase for protein–protein interaction-dependent small-molecule activation. ACS Cent. Sci. 5, 1768–1776 (2019).
Franzini, R. M. & Kool, E. T. 7-Azidomethoxy-coumarins as profluorophores for templated nucleic acid detection. ChemBioChem 9, 2981–2988 (2008).
Zheng, Y. Y., Zheng, W. R., Zhu, D. F. & Chang, H. F. Theoretical modeling of pKa’s of thiol compounds in aqueous solution. New J. Chem. 43, 5239–5254 (2019).
Beddoes, R. L. et al. An example of a tin-pummerer rearrangement. Tetrahedron Lett. 33, 417–420 (1992).
Yin, J., Bai, Y., Mao, M. & Zhu, G. Silver-catalyzed regio- and stereoselective addition of carboxylic acids to ynol ethers. J. Org. Chem. 79, 9179–9185 (2014).
Stang, P. J. Ynol esters and alkynyl(phenyl)iodonium chemistry. Russ. Chem. Bull. 42, 12–23 (1993).
Gray, V. J. & Wilden, J. D. The chemistry of ynol and thioynol ethers. Org. Biomol. Chem. 14, 9695–9711 (2016).
Lal, B., Pramanik, B. N., Manhas, M. S. & Bose, A. K. Diphenylphosphoryl azide a novel reagent for the stereospecific synthesis of azides from alcohols. Tetrahedron Lett. 18, 1977–1980 (1977).
Bracher, P. J., Snyder, P. W., Bohall, B. R. & Whitesides, G. M. The relative rates of thiol-thioester exchange and hydrolysis for alkyl and aryl thioalkanoates in water. Orig. Life Evol. Biosph. 41, 399–412 (2011).
Dawson, P. E., Muir, T. W., Clark-Lewis, I. & Kent, S. B. Synthesis of proteins by native chemical ligation. Science 266, 776–779 (1994).
Bejot, R. et al. Stereoselective transformations of trihalomethylcarbinols induced by chromous chloride. Angew. Chem. Int. Ed. Engl. 44, 2008–2011 (2005).
Zhao, Y. et al. Nuclear-encoded lncRNA MALAT1 epigenetically controls metabolic reprogramming in HCC cells through the mitophagy pathway. Mol. Ther. Nucleic Acids 23, 264–276 (2021).
Mohammad, G. & Kowluru, R. A. Nuclear genome-encoded long noncoding RNAs and mitochondrial damage in diabetic retinopathy. Cells 10, 3271 (2021).
Reyes-Gutierrez, P., Ritland Politz, J. C. & Pederson, T. A mRNA and cognate microRNAs localize in the nucleolus. Nucleus 5, 636–642 (2014).
Wang, H. et al. Long noncoding RNA miR503HG, a prognostic indicator, inhibits tumor metastasis by regulating the HNRNPA2B1/NF-kappaB pathway in hepatocellular carcinoma. Theranostics 8, 2814–2829 (2018).
Sone, M. et al. The mRNA-like noncoding RNA Gomafu constitutes a novel nuclear domain in a subset of neurons. J. Cell Sci. 120, 2498–2506 (2007).
Yu, B. & Shan, G. Functions of long noncoding RNAs in the nucleus. Nucleus 7, 155–166 (2016).
van Dijk, M. et al. Mutations within the LINC-HELLP non-coding RNA differentially bind ribosomal and RNA splicing complexes and negatively affect trophoblast differentiation. Hum. Mol. Genet. 24, 5475–5485 (2015).
Du, J. et al. LncRNA OSTM1-AS1 acts as an oncogenic factor in Wilms’ tumor by regulating the miR-514a-3p/MELK axis. Anticancer Drugs 33, 720–730 (2022).
Long, M. J. C., Huang, K.-T. & Aye, Y. The not so identical twins: (dis)similarities between reactive electrophile and oxidant sensing and signaling. Chem. Soc. Rev. 50, 12269–12291 (2021).
Tsue A. F., et al. Oligonucleotide-directed proximity-interactome mapping (O-MAP): a unified method for discovering RNA-interacting proteins, transcripts and genomic loci in situ. Preprint at bioRxiv https://doi.org/10.1101/2023.01.19.524825 (2023).
Fang, L. et al. Pervasive transcriptome interactions of protein-targeted drugs. Nat. Chem. 15, 1374–1383 (2023).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Patro, R., Duggal, G., Love, M. I., Irizarry, R. A. & Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 14, 417–419 (2017).
Soneson, C., Love, M. I. & Robinson, M. D. Differential analyses for RNA-seq: transcript-level estimates improve gene-level inferences. F1000Res 4, 1521 (2015).
Law, C. W., Chen, Y., Shi, W. & Smyth, G. K. voom: Precision weights unlock linear model analysis tools for RNA-seq read counts. Genome Biol 15, R29 (2014).
Ritchie, M. E. et al. Limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Fei, J. et al. RNA biochemistry. Determination of in vivo target search kinetics of regulatory noncoding RNA. Science 347, 1371–1374 (2015).
Dickinson, B. Raw images from “Bioorthogonal masked acylating agents for proximity-dependent RNA labeling”. Mendeley Data https://doi.org/10.17632/kzpvsdjrgy.1 (2023).
Acknowledgements
This work was supported by the National Institute of General Medical Sciences (GM119840), The National Institute of Mental Health (MH122142) and the National Human Genome Research Institute (RM1 HG008935) of the National Institutes of Health, as well as The G. Harold and Leila Y. Mathers Charitable Foundation. S.-A.A. was supported by the National Institute of Diabetes and Digestive and Kidney Diseases of the National Institutes of Health (F30 DK125088). We thank A. Radakovic from J.W. Szostak’s laboratory for supplying synthetic RNA oligos and C. He for helpful discussions. We thank E. Wu and the University of Chicago CRI Bioinformatics Core for assistance with data acquisition and analysis. We also thank the University of Chicago Integrated Light Microscopy Core (RRID: SCR_019197) and the Chicago Immunoengineering Innovation Center at the Pritzker School of Molecular Engineering for training and support with confocal microscopy and Next-seq sequencing facilities, respectively. We thank S. Ahmadiantehrani for assistance with preparing this paper.
Author information
Authors and Affiliations
Contributions
S.P., T.Q., K.K. and B.C.D. conceived the project. S.P. and T.Q. synthesized all compounds and performed experiments. S.-A.A. assisted with experiments and data analysis. S.P., T.Q. and B.C.D. wrote the paper with input from all authors.
Corresponding author
Ethics declarations
Competing interests
B.C.D. is a founder and holds equity in Tornado Bio, Inc. The other authors declare no competing interests.
Peer review
Peer review information
Nature Chemistry thanks Yimon Aye, Ryan Flynn and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Measurement of the kinetics of mCP-coumarin unmasked by 5 nM of BS2 esterase.
a, Schematic of synthesis of mCP-coumarin and the unmasking reaction by BS2. b, Linear range of mCP ester unmasking measured with different concentrations of mCP-coumarin incubation. Y-axis represents the normalized concentration of coumarin generated from mCP unmasking. Error bars represent the standard deviation of the mean, n = 3 independent in vitro reactions. c, Michaelis-Menten plot showing the rate of mCP unmasking as a function of the substrate (mCP-coumarin) concentration. Error bars represent the standard deviation of the mean n = 3 independent in vitro reactions, and the curve represents the best fit. d, Kinetic parameters obtained by Michaelis-Menten plot of BS2 unmasking mCP-coumarin. All plots and curve fit were made using GraphPad Prism. Error bars represent the standard deviation, n = 3 independent in vitro reactions.
Extended Data Fig. 2 Optimization of catalyst conditions for regioselective carboxylic acid addition to synthesize α-alkoxy thioenol esters.
Silver-based catalysts yielded more than 90% of the undesired 1,2-addition product (entry 1). The screening of various metal catalysts revealed the iridium catalyst could provide desired 1,1-addition product (entries 2–8), especially the [Ir(OMe)(COD)]2. Further screening of several ligands suggested the ligand has minimal effects on yield (entries 9–16).
Extended Data Fig. 3 Confocal microscopy images showing BS2 dependent proximity labelling by AC-2 probe in the nucleus.
BS2 with NLS and V5 tag was transiently expressed in HEK293T cells, and the expression was visualized with V5 antibody; labelling activity was visualized after click reaction with azide-Alexa488. DAPI is a nuclear marker. The scale bar on Brightfield images is 10 µm.
Extended Data Fig. 4 Confocal microscopy images showing BS2 dependent proximity labelling by AC-2 probe in the cytosol.
BS2 with nuclear export signal (NES) and V5 tag was transiently expressed in HEK293T cells, and the expression was visualized with V5 antibody; labelling activity was visualized after click reaction with azide-Alexa488. DAPI is a nuclear marker. The scale bar on Brightfield images is 10 µm.
Extended Data Fig. 5 Confocal microscopy images showing BS2 dependent proximity labelling by AC-2 probe in the ERM.
BS2 with outer ER membrane localization via ER transmembrane anchor facing the cytosol was transiently expressed in HEK293T cells, and the expression was visualized with V5 antibody; labelling activity was visualized after click reaction with azide-Alexa488. DAPI is a nuclear marker. The scale bar on Brightfield images is 10 µm.
Extended Data Fig. 6 Confocal microscopy images showing BS2 dependent proximity labelling by AC-2 probe in the mitochondria.
BS2 with MTS-matrix (Mitochondria Targeting Signal) and V5 tag was transiently expressed in HEK293T cells, and the expression was visualized with V5 antibody; labelling activity was visualized after click reaction with azide-Alexa488. DAPI is a nuclear marker. The scale bar on Brightfield images is 10 µm.
Extended Data Fig. 7 Confocal microscopy images showing BS2 dependent proximity labelling by AC-2 probe in the nucleolus.
BS2 with nucleolar localization via three tandem nucleolar targeting sequences from NF-κB-inducing kinase (NIK) and V5 tag was transiently expressed in HEK293T cells, and the expression was visualized with V5 antibody; labelling activity was visualized after click reaction with azide-Alexa488. DAPI is a nuclear marker. The scale bar on Brightfield images is 10 µm.
Extended Data Fig. 8 Confocal microscopy images showing BS2 dependent proximity labelling by AC-2 probe in the nuclear pore.
BS2 fused with SENP2 protein and V5 tag was transiently expressed in HEK293T cells, and the expression was visualized with V5 antibody; labelling activity was visualized after click reaction with azide-Alexa488. DAPI is a nuclear marker. The scale bar on Brightfield images is 10 µm.
Extended Data Fig. 9 Enrichment of BS2 proximal RNA post AC-2 labelling.
a, Bioanalyzer quantification of enriched RNAs post AC-2 labelling. 25 µg of input RNA was taken for each sample for the enrichment, and the experiment was performed with 2 biological replicates. b, Principal component analysis (PCA) analysis of gene expression values for different samples of BAP-seq, n = 2-3 biological replicates. c, Comparison of sequencing counts post differentially expressed gene (DEG) analysis for Mt-genes across different compartments, n = 2 biological replicates. d, Comparison of intronic reads across different samples Numbers are presented as a percentage of total reads that includes exons and intergenic regions. Error bars represent the standard deviation of mean, n = 2 biological replicates. P-value was determined by two-tailed unpaired t-tests with Welch’s correction. ** represents p-value < 0.01 (p = 0.0099) and *** represents p-value < 0.001 (p = 0.0006).
Supplementary information
Supplementary Information
Supplementary Figs. 1–34, synthesis methods and characterization, detailed BAP-seq protocol and information-only references.
Supplementary Data 1
BAP-seq CPM reads of all samples.
Source data
Source Data Fig. 2
LC trace used for Fig. 2b,f. Both Excel workbooks are provided in the same folder.
Source Data Fig. 3
The data points used for the bar graph in Fig. 3b. The raw images used for Fig. 3a,c are provided with Mendeley. The DOI is mentioned in the ‘Data availability’ section.
Source Data Fig. 4
The raw images used are provided with Mendeley.
Source Data Fig. 5
Unprocessed gels and blots used in Fig. 5a,b.
Source Data Fig. 5
Cy2-fluorescence image for Fig. 5a.
Source Data Fig. 5
Streptavidin chemiluminiscence blot for Fig. 5b.
Source Data Fig. 5
Methylene blue staining for Fig. 5b.
Source Data Fig. 6
DEG tables used for plots in Fig. 6b–d.
Source Data Extended Data Fig. 1
Data used for the plots in the figure.
Source Data Extended Data Fig. 9
Data used for plots in the figure.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Pani, S., Qiu, T., Kentala, K. et al. Bioorthogonal masked acylating agents for proximity-dependent RNA labelling. Nat. Chem. (2024). https://doi.org/10.1038/s41557-024-01493-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41557-024-01493-1