Abstract
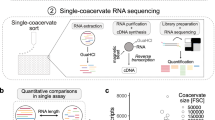
Phase separation inside mammalian cells regulates the formation of the biomolecular condensates that are related to gene expression, signalling, development and disease. However, a large population of endogenous condensates and their candidate phase-separating proteins have yet to be discovered in a quantitative and high-throughput manner. Here we demonstrate that endogenously expressed biomolecular condensates can be identified across a cell’s proteome by sorting proteins across varying oligomeric states. We employ volumetric compression to modulate the concentrations of intracellular proteins and the degree of crowdedness, which are physical regulators of cellular biomolecular condensates. The changes in degree of the partition of proteins into condensates or phase separation led to varying oligomeric states of the proteins, which can be detected by coupling density gradient ultracentrifugation and quantitative mass spectrometry. In total, we identified 1,518 endogenous condensate proteins, of which 538 have not been reported before. Furthermore, we demonstrate that our strategy can identify condensate proteins that respond to specific biological processes.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE114 partner repository with the dataset identifier PXD048218. Due to the size and lack of available condensate imaging databases, raw imaging data are available upon request to the corresponding author. All primary data are included in the source data associated with each figure accompanying this paper. Source data are provided with this paper.
References
Lyon, A. S., Peeples, W. B. & Rosen, M. K. A framework for understanding the functions of biomolecular condensates across scales. Nat. Rev. Mol. Cell Biol. 22, 215–235 (2021).
Alberti, S. & Hyman, A. A. Biomolecular condensates at the nexus of cellular stress, protein aggregation disease and ageing. Nat. Rev. Mol. Cell Biol. 22, 196–213 (2021).
Delarue, M. et al. mTORC1 controls phase separation and the biophysical properties of the cytoplasm by tuning crowding. Cell 174, 338–349 (2018).
Li, Y., Tang, W. & Guo, M. The cell as matter: connecting molecular biology to cellular functions. Matter 4, 1863–1891 (2021).
Walter, H. & Brooks, D. E. Phase separation in cytoplasm, due to macromolecular crowding, is the basis for microcompartmentation. FEBS Lett. 361, 135–139 (1995).
Fujioka, Y. et al. Phase separation organizes the site of autophagosome formation. Nature 578, 301–305 (2020).
Yasuda, S. et al. Stress- and ubiquitylation-dependent phase separation of the proteasome. Nature 578, 296–300 (2020).
Wei, M.-T. et al. Phase behaviour of disordered proteins underlying low density and high permeability of liquid organelles. Nat. Chem. 9, 1118–1125 (2017).
Frottin, F. et al. The nucleolus functions as a phase-separated protein quality control compartment. Science 365, 342–347 (2019).
Klosin, A. et al. Phase separation provides a mechanism to reduce noise in cells. Science 367, 464–468 (2020).
Liu, Q. et al. Glycogen accumulation and phase separation drives liver tumor initiation. Cell 184, 5559–5576 (2021).
Mathieu, C., Pappu, R. V. & Taylor, J. P. Beyond aggregation: pathological phase transitions in neurodegenerative disease. Science 370, 56–60 (2020).
Michaels, T. C. et al. Dynamics of oligomer populations formed during the aggregation of Alzheimer’s Aβ42 peptide. Nat. Chem. 12, 445–451 (2020).
Bremer, A. et al. Deciphering how naturally occurring sequence features impact the phase behaviours of disordered prion-like domains. Nat. Chem. 14, 196–207 (2022).
Lafontaine, D. L., Riback, J. A., Bascetin, R. & Brangwynne, C. P. The nucleolus as a multiphase liquid condensate. Nat. Rev. Mol. Cell Biol. 22, 165–182 (2021).
Lee, D. S., Wingreen, N. S. & Brangwynne, C. P. Chromatin mechanics dictates subdiffusion and coarsening dynamics of embedded condensates. Nat. Phys. 17, 531–538 (2021).
Riback, J. A. et al. Composition-dependent thermodynamics of intracellular phase separation. Nature 581, 209–214 (2020).
Shimobayashi, S. F., Ronceray, P., Sanders, D. W., Haataja, M. P. & Brangwynne, C. P. Nucleation landscape of biomolecular condensates. Nature 599, 503–506 (2021).
Shin, Y. & Brangwynne, C. P. Liquid phase condensation in cell physiology and disease. Science 357, eaaf4382 (2017).
Garaizar, A. et al. Aging can transform single-component protein condensates into multiphase architectures. Proc. Natl Acad. Sci. USA 119, e2119800119 (2022).
Krainer, G. et al. Reentrant liquid condensate phase of proteins is stabilized by hydrophobic and non-ionic interactions. Nat. Commun. 12, 1085 (2021).
Shen, Y. et al. Biomolecular condensates undergo a generic shear-mediated liquid-to-solid transition. Nat. Nanotechnol. 15, 841–847 (2020).
Toprakcioglu, Z. et al. Adsorption free energy predicts amyloid protein nucleation rates. Proc. Natl Acad. Sci. USA 119, e2109718119 (2022).
Choi, S., Meyer, M. O., Bevilacqua, P. C. & Keating, C. D. Phase-specific RNA accumulation and duplex thermodynamics in multiphase coacervate models for membraneless organelles. Nat. Chem. 14, 1110–1117 (2022).
Dzuricky, M., Rogers, B. A., Shahid, A., Cremer, P. S. & Chilkoti, A. De novo engineering of intracellular condensates using artificial disordered proteins. Nat. Chem. 12, 814–825 (2020).
Lv, P. et al. O-GlcNAcylation modulates liquid–liquid phase separation of SynGAP/PSD-95. Nat. Chem. 14, 831–840 (2022).
You, K. et al. PhaSepDB: a database of liquid-liquid phase separation related proteins. Nucleic Acids Res. 48, D354–D359 (2020).
Hardenberg, M., Horvath, A., Ambrus, V., Fuxreiter, M. & Vendruscolo, M. Widespread occurrence of the droplet state of proteins in the human proteome. Proc. Natl Acad. Sci. USA 117, 33254–33262 (2020).
Saar, K. L. et al. Learning the molecular grammar of protein condensates from sequence determinants and embeddings. Proc. Natl Acad. Sci. USA 118, e2019053118 (2021).
Alberti, S., Gladfelter, A. & Mittag, T. Considerations and challenges in studying liquid-liquid phase separation and biomolecular condensates. Cell 176, 419–434 (2019).
Li, P. et al. Phase transitions in the assembly of multivalent signalling proteins. Nature 483, 336–340 (2012).
Nott, T. J. et al. Phase transition of a disordered nuage protein generates environmentally responsive membraneless organelles. Mol. Cell 57, 936–947 (2015).
Franzmann, T. M. & Alberti, S. Prion-like low-complexity sequences: key regulators of protein solubility and phase behavior. J. Biol. Chem. 294, 7128–7136 (2019).
Dong, R.-Y. & Granick, S. Reincarnations of the phase separation problem. Nat. Commun. 12, 911 (2021).
Han, T. W. et al. Cell-free formation of RNA granules: bound RNAs identify features and components of cellular assemblies. Cell 149, 768–779 (2012).
Kato, M. et al. Cell-free formation of RNA granules: low complexity sequence domains form dynamic fibers within hydrogels. Cell 149, 753–767 (2012).
Sanchez de Groot, N. et al. RNA structure drives interaction with proteins. Nat. Commun. 10, 3246 (2019).
Shi, M. et al. Quantifying the phase separation property of chromatin-associated proteins under physiological conditions using an anti-1,6-hexanediol index. Genome Biol. 22, 229 (2021).
Jalihal, A. P. et al. Multivalent proteins rapidly and reversibly phase-separate upon osmotic cell volume change. Mol. Cell 79, 978–990 (2020).
Markmiller, S. et al. Context-dependent and disease-specific diversity in protein interactions within stress granules. Cell 172, 590–604 (2018).
Esposito, M. et al. TGF-β-induced DACT1 biomolecular condensates repress Wnt signalling to promote bone metastasis. Nat. Cell Biol. 23, 257–267 (2021).
Geladaki, A. et al. Combining LOPIT with differential ultracentrifugation for high-resolution spatial proteomics. Nat. Commun. 10, 331 (2019).
Mitrea, D. M. et al. Methods for physical characterization of phase-separated bodies and membrane-less organelles. J. Mol. Biol. 430, 4773–4805 (2018).
Zhou, M. et al. Phase-separated condensate-aided enrichment of biomolecular interactions for high-throughput drug screening in test tubes. J. Biol. Chem. 295, 11420–11434 (2020).
Caudron-Herger, M. et al. R-DeeP: proteome-wide and quantitative identification of RNA-dependent proteins by density gradient ultracentrifugation. Mol. Cell 75, 184–199 (2019).
Sridharan, S. et al. Systematic discovery of biomolecular condensate-specific protein phosphorylation. Nat. Chem. Biol. 18, 1104–1114 (2022).
Hernández, A. R., Klein, A. M. & Kirschner, M. W. Kinetic responses of β-catenin specify the sites of Wnt control. Science 338, 1337–1340 (2012).
Kim, S.-E. et al. Wnt stabilization of β-catenin reveals principles for morphogen receptor-scaffold assemblies. Science 340, 867–870 (2013).
Ma, W. et al. Single-molecule dynamics of dishevelled at the plasma membrane and Wnt pathway activation. Proc. Natl Acad. Sci. USA 117, 16690–16701 (2020).
Sasagawa, S., Ozaki, Y.-i, Fujita, K. & Kuroda, S. Prediction and validation of the distinct dynamics of transient and sustained ERK activation. Nat. Cell Biol. 7, 365–373 (2005).
Brangwynne, C. P., Tompa, P. & Pappu, R. V. Polymer physics of intracellular phase transitions. Nat. Phys. 11, 899–904 (2015).
Elbaum-Garfinkle, S. et al. The disordered P granule protein LAF-1 drives phase separation into droplets with tunable viscosity and dynamics. Proc. Natl Acad. Sci. USA 112, 7189–7194 (2015).
Reichheld, S. E., Muiznieks, L. D., Keeley, F. W. & Sharpe, S. Direct observation of structure and dynamics during phase separation of an elastomeric protein. Proc. Natl Acad. Sci. USA 114, E4408–E4415 (2017).
Ribeiro, S. S., Samanta, N., Ebbinghaus, S. & Marcos, J. C. The synergic effect of water and biomolecules in intracellular phase separation. Nat. Rev. Chem. 3, 552–561 (2019).
Stradner, A. et al. Equilibrium cluster formation in concentrated protein solutions and colloids. Nature 432, 492–495 (2004).
Bracha, D., Walls, M. T. & Brangwynne, C. P. Probing and engineering liquid-phase organelles. Nat. Biotechnol. 37, 1435–1445 (2019).
Banani, S. F., Lee, H. O., Hyman, A. A. & Rosen, M. K. Biomolecular condensates: organizers of cellular biochemistry. Nat. Rev. Mol. Cell Biol. 18, 285–298 (2017).
Davis, R. B., Moosa, M. M. & Banerjee, P. R. Ectopic biomolecular phase transitions: fusion proteins in cancer pathologies. Trends Cell Biol. 32, 681–695 (2022).
Tiwary, A. K. & Zheng, Y. Protein phase separation in mitosis. Curr. Opin. Cell Biol. 60, 92–98 (2019).
Bracha, D. et al. Mapping local and global liquid phase behavior in living cells using photo-oligomerizable seeds. Cell 175, 1467–1480 (2018).
Cai, D. et al. Phase separation of YAP reorganizes genome topology for long-term YAP target gene expression. Nat. Cell Biol. 21, 1578–1589 (2019).
Li, Y. et al. Volumetric compression induces intracellular crowding to control intestinal organoid growth via Wnt/β-catenin signaling. Cell Stem Cell 28, 63–78 (2021).
Cox, B. & Emili, A. Tissue subcellular fractionation and protein extraction for use in mass-spectrometry-based proteomics. Nat. Protoc. 1, 1872–1878 (2006).
Money, N. P. Osmotic pressure of aqueous polyethylene glycols: relationship between molecular weight and vapor pressure deficit. Plant Physiol. 91, 766–769 (1989).
Qian, D. et al. Tie-line analysis reveals interactions driving heteromolecular condensate formation. Phys. Rev. X 12, 041038 (2022).
Akabayov, B., Akabayov, S. R., Lee, S.-J., Wagner, G. & Richardson, C. C. Impact of macromolecular crowding on DNA replication. Nat. Commun. 4, 1615 (2013).
Alghoul, E., Basbous, J. & Constantinou, A. An optogenetic proximity labeling approach to probe the composition of inducible biomolecular condensates in cultured cells. STAR Protoc. 2, 100677 (2021).
An, H., Ordureau, A., Körner, M., Paulo, J. A. & Harper, J. W. Systematic quantitative analysis of ribosome inventory during nutrient stress. Nature 583, 303–309 (2020).
Garcia-Jove Navarro, M. et al. RNA is a critical element for the sizing and the composition of phase-separated RNA–protein condensates. Nat. Commun. 10, 3230 (2019).
Ku, W. L. et al. Single-cell chromatin immunocleavage sequencing (scChIC-seq) to profile histone modification. Nat. Methods 16, 323–325 (2019).
Meszaros, B., Erdos, G. & Dosztanyi, Z. IUPred2A: context-dependent prediction of protein disorder as a function of redox state and protein binding. Nucleic Acids Res. 46, W329–W337 (2018).
Hou, C. et al. PhaSepDB in 2022: annotating phase separation-related proteins with droplet states, co-phase separation partners and other experimental information. Nucleic Acids Res. 51, D460–D465 (2023).
Zeng, W. J. et al. Initiation of stress granule assembly by rapid clustering of IGF2BP proteins upon osmotic shock. Biochim. Biophys. Acta Mol. Cell Res. 1867, 118795 (2020).
Jung, J. U., Taylor, C. A. T. & Cobb, M. H. Crank up the volume: osmotic stress induces WNK1 phase separation. Cell Res. 33, 265–266 (2023).
Majumder, S. & Jain, A. Osmotic stress triggers phase separation. Mol. Cell 79, 876–877 (2020).
Kundinger, S. R. et al. Phosphorylation regulates arginine-rich RNA-binding protein solubility and oligomerization. J. Biol. Chem. 297, 101306 (2021).
Liu, X.-M., Ma, L. & Schekman, R. Selective sorting of microRNAs into exosomes by phase-separated YBX1 condensates. eLife 10, e71982 (2021).
Lu, X. et al. Copy number amplification and SP1-activated lncRNA MELTF-AS1 regulates tumorigenesis by driving phase separation of YBX1 to activate ANXA8 in non-small cell lung cancer. Oncogene 41, 3222–3238 (2022).
Guo, M. et al. Cell volume change through water efflux impacts cell stiffness and stem cell fate. Proc. Natl Acad. Sci. USA 114, E8618–E8627 (2017).
Li, Y. et al. Helical nanofiber yarn enabling highly stretchable engineered microtissue. Proc. Natl Acad. Sci. USA 116, 9245–9250 (2019).
Li, Y. et al. Compression-induced dedifferentiation of adipocytes promotes tumor progression. Sci. Adv. 6, eaax5611 (2020).
Bergmann, J. E. & Lodish, H. F. A kinetic model of protein synthesis. Application to hemoglobin synthesis and translational control. J. Biol. Chem. 254, 11927–11937 (1979).
Besse, F. & Ephrussi, A. Translational control of localized mRNAs: restricting protein synthesis in space and time. Nat. Rev. Mol. Cell Biol. 9, 971–980 (2008).
Brittis, P. A., Lu, Q. & Flanagan, J. G. Axonal protein synthesis provides a mechanism for localized regulation at an intermediate target. Cell 110, 223–235 (2002).
Haschemeyer, A. E. Kinetics of protein synthesis in higher organisms in vivo. Trends Biochem. Sci. 1, 133–136 (1976).
Rannels, D. E., Wartell, S. A. & Watkins, C. A. The measurement of protein synthesis in biological systems. Life Sci. 30, 1679–1690 (1982).
Heinrich, R., Neel, B. G. & Rapoport, T. A. Mathematical models of protein kinase signal transduction. Mol. Cell 9, 957–970 (2002).
Schwanhäusser, B. et al. Global quantification of mammalian gene expression control. Nature 473, 337–342 (2011).
Su, Q., Mehta, S. & Zhang, J. Liquid–liquid phase separation: orchestrating cell signaling through time and space. Mol. Cell 81, 4137–4146 (2021).
Tripathi, S., Levine, H. & Jolly, M. K. The physics of cellular decision making during epithelial-mesenchymal transition. Annu. Rev. Biophys. 49, 1–18 (2020).
Wong, I. Y. et al. Collective and individual migration following the epithelial–mesenchymal transition. Nat. Mater. 13, 1063–1071 (2014).
Zhang, J. et al. TGF-β-induced epithelial-to-mesenchymal transition proceeds through stepwise activation of multiple feedback loops. Sci. Signal. 7, ra91 (2014).
Erdel, F. et al. Mouse heterochromatin adopts digital compaction states without showing hallmarks of HP1-driven liquid–liquid phase separation. Mol. Cell 78, 236–249 (2020).
Fu, H. et al. Poly(ADP-ribosylation) of P-TEFb by PARP1 disrupts phase separation to inhibit global transcription after DNA damage. Nat. Cell Biol. 24, 513–525 (2022).
Gardell, S. J. et al. Boosting NAD+ with a small molecule that activates NAMPT. Nat. Commun. 10, 3241 (2019).
Attisano, L. & Wrana, J. L. Signal integration in TGF-β, WNT and Hippo pathways. F1000Prime Rep. 5, 17 (2013).
Su, X. et al. Phase separation of signaling molecules promotes T cell receptor signal transduction. Science 352, 595–599 (2016).
Zbinden, A., Pérez-Berlanga, M., De Rossi, P. & Polymenidou, M. Phase separation and neurodegenerative diseases: a disturbance in the force. Dev. Cell 55, 45–68 (2020).
Li, Q. et al. LLPSDB: a database of proteins undergoing liquid-liquid phase separation in vitro. Nucleic Acids Res. 48, D320–D327 (2020).
Mészáros, B. et al. PhaSePro: the database of proteins driving liquid-liquid phase separation. Nucleic Acids Res. 48, D360–D367 (2020).
Guo, M. et al. Probing the stochastic, motor-driven properties of the cytoplasm using force spectrum microscopy. Cell 158, 822–832 (2014).
Gupta, S. K. & Guo, M. Equilibrium and out-of-equilibrium mechanics of living mammalian cytoplasm. J. Mech. Phys. Solids 107, 284–293 (2017).
Jawerth, L. et al. Protein condensates as aging Maxwell fluids. Science 370, 1317–1323 (2020).
Spruijt, E., Sokolova, E. & Huck, W. T. Complexity of molecular crowding in cell-free enzymatic reaction networks. Nat. Nanotechnol. 9, 406–407 (2014).
Zhang, Q. et al. Visualizing dynamics of cell signaling in vivo with a phase separation-based kinase reporter. Mol. Cell 69, 334–346 (2018).
Kanshin, E., Bergeron-Sandoval, L.-P., Isik, S. S., Thibault, P. & Michnick, S. W. A cell-signaling network temporally resolves specific versus promiscuous phosphorylation. Cell Rep. 10, 1202–1214 (2015).
Chakfe, Y. & Bourque, C. W. Excitatory peptides and osmotic pressure modulate mechanosensitive cation channels in concert. Nat. Neurosci. 3, 572–579 (2000).
Molloy, M. P. Two-dimensional electrophoresis of membrane proteins using immobilized pH gradients. Anal. Biochem. 280, 1–10 (2000).
Rimpilainen, M. A. & Righetti, P. G. Membrane protein analysis by isoelectric focusing in immobilized pH gradients. Electrophoresis 6, 419–422 (1985).
Zhang, L. et al. Proteomic analysis of mouse liver plasma membrane: use of differential extraction to enrich hydrophobic membrane proteins. Proteomics 5, 4510–4524 (2005).
Consortium, T. U. UniProt: the universal protein knowledgebase in 2021. Nucleic Acids Res. 49, D480–D489 (2020).
Cox, J. et al. Accurate proteome-wide label-free quantification by delayed normalization and maximal peptide ratio extraction, termed MaxLFQ. Mol. Cell. Proteomics 13, 2513–2526 (2014).
Conesa, A. et al. Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21, 3674–3676 (2005).
Perez-Riverol, Y. et al. The PRIDE database resources in 2022: a hub for mass spectrometry-based proteomics evidences. Nucleic Acids Res. 50, D543–D552 (2022).
Acknowledgements
We acknowledge financial support from the National Natural Science Foundation of China (grants 32171248 and 12102142 to Y.L., 22074047 and 21775049 to B.-F.L. and 31700746 to P.C.) and the Fundamental Research Funds for Central Universities (HUST no. 2021GCRC056 to Y.L.).
Author information
Authors and Affiliations
Contributions
Conceptualization was provided by Y.L. and B.-F.L., methodology by Y.L. and P.L., investigations by P.L., P.C., Y.L., H.X., L.L., M.L., X.R., W.W., W.Z., L.Z., X.X., Y.Z. and L.X., visualization by Y.L., P.L., P.C., F.Q., J.S., J.L., P.Z., Z.G., X.F., W.D. and X.L., funding acquisition by Y.L. and B.-F.L., project administration by Y.L. and supervision by Y.L. and B.-F.L. The original draft was written by Y.L. and P.L., and writing, review and editing by Y.L., P.L. and B.-F.L.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Chemistry thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 In situ imaging confirmed that the isolated condensates remained undissociated upon the treatment of the RIPA or NP-40 lysis buffer.
a, Cells were transfected YBX1-EGFP to form YBX1-EGFP condensates in microfluidic channel. The RIPA buffer or NP-40 buffer was introduced into the cells by microchannels. b, YBX1-EGFP condensates remained for at least 40 mins with the treatment of NP-40 buffer. However, NP-40 was observed to insufficiently dissolve cell membrane and be unable to disrupt the nuclear membrane. c, YBX1-EGFP condensates remained for at least 40 mins with the treatment of RIPA buffer, while cellular lipid membrane and nuclear structure were sufficiently dissolved by RIPA buffer. Representative results from three independent experiments.
Extended Data Fig. 2 In situ imaging confirmed that the condensates still retained upon the step of pre-clear centrifugation.
a, Cell lysate was centrifuged by 12000 g, and supernatant was introduced into the cells by microchannels. b, YBX1-EGFP condensates remained after pre-clear centrifugation step. c, YAP1-EGFP condensates remained after pre-clear centrifugation step. Representative results from two independent experiments.
Extended Data Fig. 3 Testing droplet-like behaviors of MRPL23 condensates and TOE1 condensates.
a, Time-dependent images showed the liquid-like fusion and splitting behaviors of the MRPL23-GFP condensates. b, Fluorescent image of MRPL23-GFP condensates. The dash line circle indicated the area of nucleus. c, The percentage of MRPL23-GFP in the condensed form as compared to the total cellular protein. n = 7 independent experiments. d-e, Quantification (d) and sequential images (e) of FRAP assays on MRPL23-GFP condensates showed the recovery of MRPL23-GFP after photo-bleaching. n = 6 independent experiments. f, Time-dependent images showed the liquid-like fusion and splitting behaviors of the TOE1-GFP condensates. g, Fluorescent image of TOE1-GFP condensates. The dash line circle indicated the area of nucleus. h, The percentage of TOE1-GFP in the condensed form as compared to the total cellular protein. n = 10 independent experiments. i-j, Quantification (i) and sequential images (j) of FRAP assays on TOE1-GFP condensates showed the recovery of TOE1-GFP after photo-bleaching. n = 6 independent experiments. In c, d, h, i, data are means ± SD and were analysed by two-sided unpaired student’s t test.
Extended Data Fig. 4 Testing droplet-like behaviors of ASNS condensates and LTA4H condensates.
a, Time-dependent images showed the liquid-like fusion and splitting behaviors of the ASNS-GFP condensates. b, Fluorescent image of ASNS-GFP condensates. The dash line circle indicated the area of nucleus. c, The percentage of ASNS-GFP in the condensed form as compared to the total cellular protein. n = 7 independent experiments, OE represents overexpression. d-e, Quantification (d) and sequential images (e) of FRAP assays on ASNS-GFP condensates showed the recovery of ASNS-GFP after photo-bleaching. n = 6 independent experiments. f, Time-dependent images showed the liquid-like fusion and splitting behaviors of the LTA4H-GFP condensates. g, Fluorescent image of LTA4H-GFP condensates. The dash line circle indicated the area of nucleus. h, The percentage of LTA4H-GFP in the condensed form as compared to the total cellular protein. n = 7 independent experiments, OE represents overexpression. i-j, Quantification (i) and sequential images (j) of FRAP assays on LTA4H-GFP condensates showed the recovery of LTA4H-GFP after photo-bleaching. n = 6 independent experiments. k, Numbers of detected proteins of 8 types of known membraneless condensates in the detected condensate proteins upon the short-term treatment of TGF-β. In c, d, h, i, data are means ± SD and were analysed by two-sided unpaired student’s t test.
Extended Data Fig. 5 Testing droplet-like behaviors of DAP3 condensates and IDH1 condensates.
a, Time-dependent images showed the liquid-like fusion and splitting behaviors of the DAP3-GFP condensates. b, Fluorescent image of DAP3-GFP condensates. The dash line circle indicated the area of nucleus. c, The percentage of DAP3-GFP in the condensed form as compared to the total cellular protein. n = 7 independent experiments, OE represents overexpression. d-e, Quantification (d) and sequential images (e) of FRAP assays on DAP3-GFP condensates showed the recovery of ASNS-GFP after photo-bleaching. n = 6 independent experiments. f, Time-dependent images showed the liquid-like fusion and splitting behaviors of the IDH1-GFP condensates. g, Fluorescent image of IDH1-GFP condensates. The dash line circle indicated the area of nucleus. h, The percentage of IDH1-GFP in the condensed form as compared to the total cellular protein. n = 7 independent experiments, OE represents overexpression. i-j, Quantification (i) and sequential images (j) of FRAP assays on IDH1-GFP condensates showed the recovery of IDH1-GFP after photo-bleaching. n = 6 independent experiments. k, Fluorescence images of DAP3-EGFP and IDH1-EGFP transfected into H1975 cells without TGF-β induction (left) and with 2 days of TGF-β treatment (right). Representative results from three independent experiments. l, Numbers of detected proteins of 8 types of known membraneless condensates in the detected condensate proteins upon the long-term treatment of TGF-β. In c, d, h, i, data are means ± SD and were analysed by two-sided unpaired student’s t test.
Supplementary information
Supplementary Information
Resource table, materials and methods, discussions sections 1–8, Supplementary Figs. 1–30, Tables 1–5, references and raw data of uncropped gel scans.
Supplementary Table 1
Results of high-throughput identification of phase separated proteins.
Source data
Source Data Fig. 2
Statistical source data for Figs. 2c, 2d.
Source Data Fig. 3
Statistical source data for Figs. 3e, 3g, 3j, 3k, 3o, 3p.
Source Data Fig. 4
Statistical source data for Figs. 4e, 4g, 4j, 4k, 4o, 4p.
Source Data Fig. 5
Statistical source data for Figs. 5e, 5g, 5j, 5k, 5o, 5p.
Source Data Extended Data Fig. 3
Statistical source data Extended Data Figs. 3c, 3d, 3h, 3i.
Source Data Extended Data Fig. 4
Statistical source data Extended Data Figs. 4c, 4d, 4h, 4i.
Source Data Extended Data Fig. 5
Statistical source data Extended Data Figs. 5c, 5d, 5h, 5i.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, P., Chen, P., Qi, F. et al. High-throughput and proteome-wide discovery of endogenous biomolecular condensates. Nat. Chem. (2024). https://doi.org/10.1038/s41557-024-01485-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41557-024-01485-1