Abstract
Intercellular communication orchestrates a multitude of physiologic and pathologic conditions. Algorithms to infer cell–cell communication and predict downstream signalling and regulatory networks are needed to illuminate mechanisms of stem cell differentiation and tissue development. Here, to fill this gap, we developed and applied CellComm to investigate how the aorta–gonad–mesonephros microenvironment dictates haematopoietic stem and progenitor cell emergence. We identified key microenvironmental signals and transcriptional networks that regulate haematopoietic development, including Stat3, Nr0b2, Ybx1 and App, and confirmed their roles using zebrafish, mouse and human models. Notably, CellComm revealed extensive crosstalk among signalling pathways and convergence on common transcriptional regulators, indicating a resilient developmental programme that ensures dynamic adaptation to changes in the embryonic environment. Our work provides an algorithm and data resource for the scientific community.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The processed and raw scRNA-seq data supporting the findings of this study have been deposited in the Gene Expression Omnibus under accession code GSE160526. We re-analysed publicly available datasets from the following accession numbers: GSE137117 and GSE144240. Reference genome data for scRNA-seq analyses were downloaded from https://support.10xgenomics.com/single-cell-gene-expression/software/downloads/2.2. All other data supporting the findings of this study are available from the corresponding authors on reasonable request. Source data are provided with this paper.
Code availability
CellComm and CellRouter are available through the Framework for Unified Single-Cell Analysis (FUSCA) R package available at https://github.com/edroaldo/fusca. We also provide a web resource to run CellComm and other algorithms at http://hematopoieticniches.com.
References
Baron, C. S. et al. Single-cell transcriptomics reveal the dynamic of haematopoietic stem cell production in the aorta. Nat. Commun. 9, 2517 (2018).
Zhou, F. et al. Tracing haematopoietic stem cell formation at single-cell resolution. Nature 533, 487–492 (2016).
Hou, S. et al. Embryonic endothelial evolution towards first hematopoietic stem cells revealed by single-cell transcriptomic and functional analyses. Cell Res. 30, 376–392 (2020).
Gao, P. et al. Transcriptional regulatory network controlling the ontogeny of hematopoietic stem cells. Genes Dev. 34, 950–964 (2020).
Zhu, Q. et al. Developmental trajectory of pre-hematopoietic stem cell formation from endothelium. Blood https://doi.org/10.1182/blood.2020004801 (2020).
Taoudi, S. et al. Extensive hematopoietic stem cell generation in the AGM region via maturation of VE-cadherin+CD45+ pre-definitive HSCs. Cell Stem Cell 3, 99–108 (2008).
Rybtsov, S. et al. Hierarchical organization and early hematopoietic specification of the developing HSC lineage in the AGM region. J. Exp. Med. 208, 1305–1315 (2011).
Lummertz da Rocha, E. et al. Reconstruction of complex single-cell trajectories using CellRouter. Nat. Commun. 9, 892 (2018).
Nguyen, P. D. et al. Haematopoietic stem cell induction by somite-derived endothelial cells controlled by meox1. Nature 512, 314–318 (2014).
McGarvey, A. C. et al. A molecular roadmap of the AGM region reveals BMPER as a novel regulator of HSC maturation. J. Exp. Med. 214, 3731–3751 (2017).
Oatley, M. et al. Single-cell transcriptomics identifies CD44 as a marker and regulator of endothelial to haematopoietic transition. Nat. Commun. 11, 586 (2020).
Khurana, S. et al. Outside-in integrin signalling regulates haematopoietic stem cell function via Periostin–Itgav axis. Nat. Commun. 7, 13500 (2016).
Ransom, D. G. et al. Characterization of zebrafish mutants with defects in embryonic hematopoiesis. Development 123, 311–319 (1996).
North, T. E. et al. Prostaglandin E2 regulates vertebrate haematopoietic stem cell homeostasis. Nature 447, 1007–1011 (2007).
Huang, H.-T. et al. A network of epigenetic regulators guides developmental haematopoiesis in vivo. Nat. Cell Biol. 15, 1516–1525 (2013).
Rho, S.-S. et al. Rap1b promotes Notch-signal-mediated hematopoietic stem cell development by enhancing integrin-mediated cell adhesion. Dev. Cell 49, 681–696.e6 (2019).
Ditadi, A. et al. Human definitive haemogenic endothelium and arterial vascular endothelium represent distinct lineages. Nat. Cell Biol. 17, 580–591 (2015).
Umemoto, T. et al. CD61 enriches long-term repopulating hematopoietic stem cells. Biochem. Biophys. Res. Commun. 365, 176–182 (2008).
Huang, K. et al. Generation and analysis of GATA2 human ESCs reveal ITGB3/CD61 as a reliable marker for defining hemogenic endothelial cells during hematopoiesis. Stem Cell Rep. 7, 854–868 (2016).
Mariani, S. A. et al. Pro-inflammatory aorta-associated macrophages are involved in embryonic development of hematopoietic stem cells. Immunity 50, 1439–1452 (2019).
Frame, J. M. et al. Metabolic regulation of inflammasome activity controls embryonic hematopoietic stem and progenitor cell production. Dev. Cell https://doi.org/10.1016/j.devcel.2020.07.015 (2020).
Clements, W. K. et al. A somitic Wnt16/Notch pathway specifies haematopoietic stem cells. Nature 474, 220–224 (2011).
Souilhol, C. et al. Inductive interactions mediated by interplay of asymmetric signalling underlie development of adult haematopoietic stem cells. Nat. Commun. 7, 10784 (2016).
Francis, K. & Palsson, B. O. Effective intercellular communication distances are determined by the relative time constants for cyto/chemokine secretion and diffusion. Proc. Natl Acad. Sci. USA. 94, 12258–12262 (1997).
Linnekin, D. Early signaling pathways activated by c-Kit in hematopoietic cells. Int. J. Biochem. Cell Biol. 31, 1053–1074 (1999).
Shu, R. et al. APP intracellular domain acts as a transcriptional regulator of miR-663 suppressing neuronal differentiation. Cell Death Dis. 6, e1651 (2015).
Stehling-Sun, S., Dade, J., Nutt, S. L., DeKoter, R. P. & Camargo, F. D. Regulation of lymphoid versus myeloid fate ‘choice’ by the transcription factor Mef2c. Nat. Immunol. 10, 289–296 (2009).
Rosenbauer, F. & Tenen, D. G. Transcription factors in myeloid development: balancing differentiation with transformation. Nat. Rev. Immunol. 7, 105–117 (2007).
Liang, J. et al. The C-kit receptor-mediated signal transduction and tumor-related diseases. Int. J. Biol. Sci. 9, 435–443 (2013).
Xu, K. et al. Integrating ChIP-sequencing and digital gene expression profiling to identify BRD7 downstream genes and construct their regulating network. Mol. Cell. Biochem. 411, 57–71 (2016).
Lumsden, A. L. et al. Dysregulation of neuronal iron homeostasis as an alternative unifying effect of mutations causing familial Alzheimer’s disease. Front. Neurosci. 12, 533 (2018).
Muto, Y., Nishiyama, M., Nita, A., Moroishi, T. & Nakayama, K. I. Essential role of FBXL5-mediated cellular iron homeostasis in maintenance of hematopoietic stem cells. Nat. Commun. 8, 16114 (2017).
Cahan, P. et al. CellNet: network biology applied to stem cell engineering. Cell 158, 903–915 (2014).
Kinney, M. A. et al. A systems biology pipeline identifies regulatory networks for stem cell engineering. Nat. Biotechnol. 37, 810–818 (2019).
Ng, A. H. M. et al. A comprehensive library of human transcription factors for cell fate engineering. Nat. Biotechnol. https://doi.org/10.1038/s41587-020-0742-6 (2020).
Morris, S. A. et al. Dissecting engineered cell types and enhancing cell fate conversion via CellNet. Cell 158, 889–902 (2014).
Chen, M. J. et al. Transcriptome dynamics of hematopoietic stem cell formation revealed using a combinatorial Runx1 and Ly6a reporter system. Stem Cell Rep. 14(5), 956–971 (2020).
Lummertz da Rocha, E. et al. Prediction of intercellular communication networks using CellComm. Protoc. Exch. https://doi.org/10.21203/rs.3.pex-1843/v1 (2022).
Faith, J. J. et al. Large-scale mapping and validation of Escherichia coli transcriptional regulation from a compendium of expression profiles. PLoS Biol. 5, e8 (2007).
Sturgeon, C. M., Ditadi, A., Awong, G., Kennedy, M. & Keller, G. Wnt signaling controls the specification of definitive and primitive hematopoiesis from human pluripotent stem cells. Nat. Biotechnol. 32, 554–561 (2014).
Mandegar, M. A. et al. CRISPR interference efficiently induces specific and reversible gene silencing in human iPSCs. Cell Stem Cell 18, 541–553 (2016).
Acknowledgements
This work was supported by the following grants from the National Institutes of Health: RC2DK120535 (G.Q.D. and T.E.N.), U01HL134812 (G.Q.D. and T.E.N.), R24DK092760 (G.Q.D. and T.E.N.), R01DK098241 (T.E.N.), F32DK122715 and K01DK129409 (W.W.S.). E.L.d.R. thanks Coordination for the Improvement of Higher Education Personnel (CAPES), Brazil. M.F. is supported by a PhD fellowship from the Brazilian National Council for Scientific and Technological Development (CNPq). C.K. was supported by a postdoctoral fellowship from the German Research Foundation (DFG; grant KU 3580/1-1).
Author information
Authors and Affiliations
Contributions
E.L.d.R., C.K. and G.Q.D. conceived and designed the project. E.L.d.R. designed and developed CellComm. E.L.d.R. and C.K. designed experiments to sequence the AGM niche. C.K. and M.A.N. generated the scRNA-seq dataset. E.L.d.R. processed the raw scRNA-seq and analysed all data described. C.K. designed and performed mouse and iPSC experiments, with A.M. W.W.S. designed and performed all zebrafish experiments, with Z.C.L. R.J. performed some T-cell differentiation experiments. R.d.S.P. created the R package with CellComm and the updated version of CellRouter. M.F. wrote the protocol for Protocol Exchange. E.L.d.R., C.K. and W.W.S. wrote the paper. J.J.C. provided critical feedback on the manuscript. G.Q.D. and T.E.N. oversaw the project, critiqued experiments and revised the paper.
Corresponding authors
Ethics declarations
Competing interests
G.Q.D. and Boston Children’s Hospital have filed provisional patent applications (US11261430B2, US20220049221A1) covering derivation of blood cells from pluripotent stem cell sources. J.J.C. is an academic co-founder and board member of Cellarity. The other authors declare no competing interests.
Peer review
Peer review information
Nature Cell Biology thanks Fuchou Tang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Gating strategy and Identification of the hemogenic endothelium population.
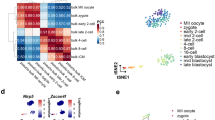
a, Single, live, non-red blood cells (7-AADnegativeTER119negative) were sampled. b, Gating strategy for isolation of endothelial cells (EC) and type 1 preHSCs (T1preHSCs). Cells gated in A were further gated on VE-Cadherin+CD31+CD45negativeCD41negativeKITlow/+ for EC and VE-Cadherin+CD31+CD45negative CD41lowKITlow/+ for T1-preHSC. c, tSNE analysis of the entire AGM ecosystem data color-coded by sample replicates. d, tSNE analysis of the entire AGM ecosystem data color-coded by transcriptional clusters. e, Proportion of replicates in each transcriptional cluster. f, tSNE analysis of sorted endothelial cells (ECs) and immunophenotypic T1preHSCs colored-coded by transcriptional clusters. g, Proportion of sorted cells (ECs, T1reHSCs) across transcriptional clusters. h, Manual cluster annotation based on transcriptional clusters and the sample of origin. i, Gene expression distribution of selected EC, HE (hemogenic endothelium), and T1preHSC genes. The middle line in the box plot indicates the median. The lower and upper hinges correspond to the 25th and 75th percentiles. Whiskers show min to max. Data beyond the whiskers are ‘outlying’ points and are plotted individually. Source numerical data are available in source data.
Extended Data Fig. 2 Experimental validation of ligands and receptors.
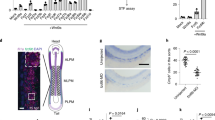
a, Whole-mount in situ hybridization (WISH) for runx1/cmyb in control and itgb3b morpholino-injected embryos at 36 hpf. Scale bar, 100 µm, N = 23 embryos (control) and N = 26 embryos (itgb3b morpholino). b, Representative flow plots for tested surface receptors on hemogenic endothelial cells from hiPSCs (HE, CD34+CD45−, day 8) and Hematopoietic cells (CD34+CD45+, day 8 + 7), pre gated on live, single cells. c, qPCR confirming significant reduction of CD93 transcript 48 h after siRNA transfection. Bars represent mean + /- SD. N = 4 independent EHT cultures. Paired t-test, **p = 0.0065. d, Flow cytometric analysis of CD93 surface expression 48 h after siRNA transduction. Bars represent mean + /- SD. N = 3 independent differentiation experiments. Paired t-test, one-tailed, *p = 0.0261. e, CFU assay from d7 floating cells after hiPSC-derived EHT with and without CD93 siRNA KD. Bars represent mean + /- SD of N = 6 assays. f, WISH timecourse for zebrafish mfap2 across the window of EHT. Scale bar 500 µm. g, Maximum intensity projection of a confocal z-stack of double fluorescent in situs for zebrafish mfap2 and runx1 mRNA at 24 hpf. Vasculature is visualized with GFP antibody staining in transgenic embryos. Yellow arrowhead indicates position of cross section. Scale bars 50 µm, 25 µm and 10 µm. h, Viability and quantification of floating, hematopoietic cells at d8 + 7 of endothelial to hemogenic transition at increasing concentrations of recombinant human MFAP2. Mean + /- SD is plotted, N = 6 independent differentiation experiments. i, Representative flow plots (left) and violin plots (median and quartiles) for CD34 and CD45 staining of floating hematopoietic cells after hiPSC derived EHT culture in the presence of increasing recombinant human MFAP2 concentrations (N = 6 independent experiments). RM One-way ANOVA, p-values have been adjusted for multiple comparison using Dunnett’s test, **p = 0.0029. Source numerical data are available in source data.
Extended Data Fig. 3 Application of CellComm to other cell-cell interactions.
a, UMAP analysis of scRNA-seq of E + HE + IAC sorted populations from E10.5 mouse embryos. b, Inferred intercellular communication network. c, Top ligand-receptor pairs across selected cell types prioritized by CellComm. d, Reconstructed signaling pathways connecting cell surface receptors to TRs across reported cell-cell interactions. e, Predicted downstream signaling pathways from cell surface receptors leading to transcriptional regulators in neuron subtypes upon interaction with macrophages. f, Gene ontology enrichment analysis of the regulons of selected transcriptional regulators identified in e. The color and size scales both represent the -log10 (P value) of the pathway enrichment result. AE: arterial endothelial cell. VE: ventral endothelial cell. Source numerical data are available in source data.
Extended Data Fig. 4 Further prioritization of transcriptional regulators.
a, The endothelial-to-hematopoietic transition trajectory reconstructed by CellRouter using sorted ECs and T1preHSCs. b, Transcriptional regulators predicted to be important for the cell state transitions reported in the x-axis, for example EC.HE means the differentiation trajectory from EC to HE. c, Gene expression distribution of genes selected for experimental validation. The middle line in the box plot indicates the median. The lower and upper hinges correspond to the 25th and 75th percentiles. Whiskers show min to max. Data beyond the whiskers are ‘outlying’ points and are plotted individually. d, Kinetic profiles of selected genes along the CellRouter trajectory from endothelial cells (ECs) to the T1preHSC10 state. e, Kinetic profiles of Gfi1b and Gfi1 along the EC to HE differentiation trajectory. f, Expression dynamics by qPCR of hematopoietic transcripts RUNX1C and GFI1 as well as CellComm predicted candidate regulators APP and YBX1. N = 4 (d1), N = 6 (d4), N = 5 (d7) independent EHT cultures, N = 2 independent CB CD34 + donor samples. Bar graphs represent mean. g, Kinetic profiles of selected genes along the CellRouter predicted EHT trajectory using the dataset generated by1. h, Increase in APP and YBX1 expression upon ITGB3 inhibition in hiPSC-derived HSPCs. N = 6, from 3 independent cultures. Bar graphs represent mean + /- SD. i, Cell cycle signature scores along the CellRouter trajectory from EC to the T1preHSC10 state. j, Clustered kinetic profiles, which group genes with similar gene expression trends along the reported CellRouter trajectory. k, Gene ontology enrichment of gene sets in each kinetic cluster identified in l. The color and size scales both represent the -log10 (P value) of the pathway enrichment result. Hypergeometric test. The p-values were adjusted for multiple comparisons using the Benjamini-Hochberg method. Source numerical data are available in source data.
Extended Data Fig. 5 Experimental validation of transcriptional regulators.
a, Stat3 inhibitor dose-response curve on runx1/cmyb expression in zebrafish embryos treated from 14-36hpf. b, Representative flow plots for CD34 and CD45 staining of floating hematopoietic cells after hiPSC derived EHT culture in the presence of increasing STAT3 inhibitor concentrations. Bar graphs represent mean + /- SD. Ordinary One-way ANOVA, Dunnett’s multiple comparisons test, N = 5, from 4 independent cultures. c, Gating strategy on positive control samples (CB MNC) and representative flow plots of anti-CD5 and anti-CD7 stained cells, 14 days in T cell differentiation culture in the presence of indicated STAT3 inhibitor concentrations. d, No decrease in viability (DAPI negativity) in hiPSC-derived EHT cultures treated with a STAT3 inhibitor. Mean + /- SD is depicted. N = 4, from 3 independent differentiation experiments. e, Phenotypic distribution plots of runx1/cmyb + staining in 36hpf zebrafish embryos after control and experimental morpholino injection. N = 42 and 46 (ctrl MO and cebpa MO) and N = 51 and 53 (ctrl MO and cebpb MO) from two independent experiments each. Of note, ybx1 morphants displayed severe embryonic toxicity at doses as low as 1 ng, which prevented further analysis in zebrafish. f, qRT-PCR verifying YBX1 knockdown after 72 h in dox inducible hiPSC derived HE carrying a dCAS9-KRAB fusion protein and sgRNAs against YBX1. Bars represent mean + /- SD. N = 3 independent cultures, unpaired t-test, p = 0.0115. g, CRISPRi of YBX1 does not affect the percentage of CD34 + CD45 + cells. N = 2 independent sgRNAs. Data and mean is plotted. h, Reduction of CFU potential upon CRISPRi-mediated reduction of YBX1 expression in hiPSC derived HSPCs. Bar graphs represent mean. N = 3 sgRNAs for NTC, N = 2 sgRNAs for YBX1. i, Reduced lymphoid differentiation potential (N = 4 independent cultures) upon CRISPRi-mediated reduction of YBX1 expression in hiPSC derived HSPCs. ****p < 0.0001, ***p = 0.0001. Bar graphs represent mean + /- SD. j, Representative flow cytometric plots of nascent preHSCs (VE-CAD + CD45 + ) in DMSO or APPi-treated E9.5 explant cultures. k, Quantification of phenotypic T1- and T2-preHSCs in control E10.5 AGM explant cultures (N = 9) or cultures treated with APP inhibitor (N = 8). Bars represent mean + /- SD. Unpaired t-test, two-tailed, T1-preHSCs p = 0.0722, T2-preHSCs p = 0.0258. l, Representative flow cytometric plots of E14.5 fetal liver samples from APP wild type (wt), heterozygous (het) and knockout (ko) embryos pregated on single, live, lineage negative cells. Source numerical data are available in source data.
Extended Data Fig. 6 CellComm infers cell communication from spatial transcriptomics data.
a, Transcriptional clusters overlaid on the tissue image. Only spots are shown. c, Probabilistic transfer of annotations from a reference cSCC cell atlas to the spatial transcriptomics data, c, Probabilistic scores for a Tumor Specific Keratinocyte (TSK) population overlaid on the tissue image. d, Spots color-coded by niche identity overlaid on the tissue. Only spots are shown. e, left panel: number of ligand-receptor interactions co-expressed between reported cell types; middle panel: spatial proximity between niche centroids in the tissue image; right panel: interaction score taking into account the number of ligand-receptor interactions and the co-localization of niche centroids in the tissue image. f, Selected niches with higher interaction scores overlaid on the tissue image. g, Ligand-receptor pairs mediating crosstalk between the tumor microenvironment (TME) and the tumor cells. h, Spatially resolved expression of selected ligand-receptor pairs identified by CellComm. Source numerical data are available in source data.
Extended Data Fig. 7 CellComm analysis at spatial subcluster resolution.
a, Subclustering of niche identities based on spatial information, showing spatial subclusters of tumor-specific keratinocytes (TSK), the fibrovascular niche identified, as well as spots classified as lymphoid cells. b, Number of ligand-receptor pairs co-expressed between spatial subclusters (left panel); spatial proximity between subcluster centroids of reported niches (middle) and interaction score taking into account the number of ligand-receptor pairs and spatial distance of subclusters. Source numerical data are available in source data.
Extended Data Fig. 8 Signaling processes in the tumor microenvironment.
a, Ligand-receptor pairs mediating interactions between TSK spatial subclusters and the fibrovascular niche (left) as well as the downstream transcription factors in the fibrovascular niche identified by CellComm (right). b, Ligand-receptor pairs mediating interactions between the fibrovascular niche and the TSK spatial subclusters (left) as well as the downstream transcriptional regulators in tumor cells identified by CellComm. c, Gene ontology analysis of the regulons predicted to be controlled by the reported transcriptional regulators. Hypergeometric test. The p-values were adjusted for multiple comparisons using the Benjamini-Hochberg method. d, Signature scores for TSKs, demonstrating spatial localization of this tumor-specific population, as well as signature scores calculated from the regulons of the reported transcriptional regulators. Source numerical data are available in source data.
Extended Data Fig. 9 Conceptual and practical comparison of CellComm to related algorithms.
Conceptual design of CellPhoneDB, NicheNet and CellComm. a, CellPhoneDB infers intercellular communication based on ligand-receptor pairs. b, NicheNet uses prior knowledge of signaling pathways and regulatory interactions to build a model to predict which ligands regulate expression of target genes, such as differentially expressed genes or any other gene set of interest. c, CellComm infers intercellular communication by exploring co-expression patterns of ligand-receptor pairs across cell types. Then, CellComm weights a large-scale protein interaction network based on cell type-specific co-expression measurements derived from the scRNA-seq data. CellComm implements an optimization algorithm to search for paths connecting cell surface receptors to downstream transcriptional regulators and prioritizes signaling pathways and transcriptional regulators based on the statistical enrichment of the regulons of each transcriptional regulator on cell type-specific signatures. The predicted regulons are identified by gene regulatory network reconstruction from the scRNA-seq data. TR = Transcriptional Regulator. d, CellComm analysis showing predicted signaling pathways connecting cell surface receptors in the T1preHSC1 subset to downstream transcriptional regulators. e, NicheNet analysis showing which ligands are predicted to regulate the expression of T1preHSC1 signature genes (predicted target genes). f, The Venn diagram of the downstream transcriptional regulators identified by CellComm and target genes identified by NicheNet shows no overlap, demonstrating the orthogonal approach to cell-cell communication implemented in CellComm. g, Ligands and receptors identified by NicheNet. h, Overlap between ligands and receptors identified by CellComm and NicheNet. Genes in bold were experimentally validated in our study (hypergeometric test). Genes not in bold are known genes involved in hematopoiesis. Source numerical data are available in source data.
Extended Data Fig. 10 Comparison of CellComm to CytoTalk and CellPhoneDB.
a, Ligand-receptor interactions predicted by CellComm for the UGR1-T1preHSC1 interaction. b, Ligand-receptor interactions predicted by CytoTalk for the UGR1-T1preHSC1 interaction. c, Overlap of ligands/receptors identified by CellComm and CytoTalk. Genes in bold were experimentally validated in our study. d, Overlap of downstream genes predicted by CellComm and CytoTalk. e, Overlap of transcriptional regulators predicted by CellComm and transcriptional regulators in the downstream CytoTalk subnetwork (hypergeometric test). f, Mean expression of ligands and receptors identified by CellComm and CytoTalk. g, Cell-cell interaction network inferred by CellPhoneDB using the AGM scRNA-seq data generated in this study. h, Overlap of ligands/receptors identified by CellComm and CellPhoneDB (hypergeometric test). Genes in bold were experimentally validated in our study. Genes not in bold are known genes involved in hematopoiesis. i, Ligand-receptor pairs identified by CellPhoneDB in the AGM scRNA-seq data. Source numerical data are available in source data.
Supplementary information
Supplementary Information
Supplementary Figs. 1 and 2, and Notes 1–6.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 9
Statistical source data.
Source Data Extended Data Fig. 10
Statistical source data.
Rights and permissions
About this article
Cite this article
Lummertz da Rocha, E., Kubaczka, C., Sugden, W.W. et al. CellComm infers cellular crosstalk that drives haematopoietic stem and progenitor cell development. Nat Cell Biol 24, 579–589 (2022). https://doi.org/10.1038/s41556-022-00884-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-022-00884-1
This article is cited by
-
The diversification of methods for studying cell–cell interactions and communication
Nature Reviews Genetics (2024)
-
Comparative transcriptomics coupled to developmental grading via transgenic zebrafish reporter strains identifies conserved features in neutrophil maturation
Nature Communications (2024)
-
Zebrafish: a convenient tool for myelopoiesis research
Cell Regeneration (2023)
-
Non-cell-autonomous cancer progression from chromosomal instability
Nature (2023)