Abstract
In randomized controlled trials of influenza vaccination, 550 children received trivalent-inactivated influenza vaccine, permitting us to explore relationship between vaccine response and host single nucleotide polymorphisms (SNPs) in 23 candidate genes with adjustment of multiple testing. For host SNPs in TLR7–1817G/T (rs5741880), genotype GT was associated with lower odds (OR: 0.22, 95% CI: 0.09, 0.53) of have post-vaccination hemagglutination-inhibiting (HAI) titers ≥40, compared with genotype GG and TT combined under the over-dominant model. For host SNPs in TLR8–129G/C (rs3764879), genotype GT was associated with lower odds (OR: 0.47; 95% CI: 0.28, 0.80) of have post vaccination HAI titers ≥40 compared with genotype GG and AA combined under the over-dominant model. Our results could contribute to the development of better vaccines that may offer improved protection to all recipients.
Similar content being viewed by others
Main
Influenza vaccination is one of the most effective strategies to control infection and transmission in community. However, there are some vaccinated people, such as elderly, and vary widely among others, with only limited immune responses and hence protection1,2,3. Comparing to other better-known determinants such as the degree of matching and various viral and host factors, host genetic make-up as a potential factor affecting immune responses after vaccination is less explored4,5,6,7,8. Currently, the field of vaccinology is still empirical in several aspects, and hence it remains difficult to understand poor vaccine immunogenicity in different pathological and physiological conditions9. Similar to other healthcare fields, a personalized approach is proposed to the practice of vaccinology9,10,11. Determining the host factors of immune response induced by influenza vaccine could contribute to development of new personalized vaccines, and new patient-oriented vaccination strategies6,12. Enhancing the understanding on genetic determinants of vaccine responses may help to identify individuals with potential of poor vaccine response for guiding more personalized vaccine development and targeted vaccination strategy.
In particular, there is scarcity of data about this topic in the pediatric population13,14,15. It is necessary to direct research toward the production of evidence related to vaccine response in the pediatric age, also in light of the important economic and social burden linked to influenza in this target population16.
Host polymorphisms has shown to be associated with immune response after vaccination17, vaccine-related adverse events17, and disease severity18 of various infectious diseases. Major examples included the association of polymorphisms in mannose-binding lectin (MBL)–2 gene encodes a calcium-dependent protein which is important for innate immunity, and associated with increased susceptibility to several infections19,20. Several polymorphisms in promoter regions in Interleukin (IL)–10 is associated with the regulation of cellular immune responses21,22, Toll-like receptor (TLR) gene with innate immune responses trigging23,24 and disease severity25. Polymorphism of genes involved in membrane trafficking and antigen processing, and was reported to have significant impact on human response to influenza vaccination26.
Here, we analyze the data of immune response and adverse response in two randomized placebo-controlled trials in influenza vaccination in children in Hong Kong2,3, to explore the relationship between host single nucleotide polymorphisms (SNPs) and immune responses, measured by post-vaccination hemagglutination-inhibiting (HAI) titers.
In these two trials, there were 550 children were recruited and randomized to receive TIV. After excluding 15–18 children with missing vaccine response with different definitions, and 50–82 children with missing genotype information (depending on the gene), 450–485 children were included in each analysis. Among those vaccinated children, 376/535 (70.3%) of them were classified as responder based on having post-vaccination titer ≥40 for the three vaccine strains, 181/535 (33.8%) of them were classified as responder based on ≥4-fold rise comparing pre- and post-vaccination titer for the three vaccine strains. The average increase of logarithm of GMT for the three vaccine strains after vaccination was 3.44 (95% confidence interval (CI): 3.28, 3.61) log2 GMT.
The frequency of genotypes of the 23 host SNPs were summarized (Supplementary Table 1). Five host SNPs were excluded from further analysis since >99% of participants had the same genotype. Also, two TLR8 SNPs (rs3764880 and rs3764879) had almost the same distribution among our participants, therefore we focused on rs3764880 in the analysis.
We estimated the association between host SNPs and vaccine response by logistic regressions (Supplementary Table 2), under dominant model (Supplementary Table 3), recessive model (Supplementary Table 4), over-dominant model (Supplementary Table 5) and multiplicative model (Supplementary Table 6) for the 18 host SNPs. After adjusting for multiple testing, we identified that SNPs in TLR7 (rs5741880) and TLR8 (rs3764880) were associated with vaccine response. For host SNPs in TLR7 (Table 1), genotype GT was associated with lower odds (OR: 0.22, 95% CI: 0.09, 0.53) of have post-vaccination titers ≥40, and 72% (95% CI: 37%, 87%) lower average increase of logarithm of GMT after vaccination for the three vaccine strains, compared with genotype GG and TT combined under the over-dominant model, after adjustment of multiple testing. Consistently, differences in vaccine response among genotype, measured by vaccination titers ≥40, or lower average increase of logarithm of GMT after vaccination for the three vaccine strains, were also detected in dominant model and multiplicative model, although these differences cannot reach statistical significance after multiple testing.
For host SNPs in TLR8 (Table 2), genotype GA was associated with lower odds (OR: 0.47; 95% CI: 0.28, 0.80) of have post vaccination titers ≥40 compared with genotype GG and AA combined under the over-dominant model, after adjustment of multiple testing. Differences in vaccine response for these two definitions were also detected for dominant model, but could not reach statistical significance after adjusting for multiple testing.
No serious post-vaccination response including anaphylaxis or shock was reported by any recipient. We found no association between host SNPs and vaccine adverse responses, defined as presence of more than two symptoms, after adjusting for multiple testing (Supplementary Table 7). Before adjusting for multiple testing, we found that host SNPs in IL-6 (rs1818879) was associated with vaccine adverse response under the dominant and over-dominant model. Host SNPs in CCL1 (rs2282691) were associated with vaccine adverse response under the over-dominant model.
Immune response of influenza vaccination is showed to be heterogeneous, despite constant vaccine formulation, possibility due to both host factors like vaccine history27, and viral factors like mutation rate of influenza virus28. Here, we examined the relationship between variation in the host genetic make-up and the immune response of influenza vaccination, based on a sample of 550 individuals that participated in two large-scale clinical trials of influenza vaccination. We found a significant association between host SNPs in TLR7 and TLR 8 gene on immune response to vaccination.
The TLR family is important for activation of innate immunity and pathogen recognition29. Various TLRs exhibit different patterns of expression. Animal studies show that TLR7 is associated with pathology of influenza A virus infection30. TLR7 is showed to be a vital component of antiviral immunity31 and play an important role of triggering the immune response of COVID-1932. TLR8 is showed to madidate reversal of CD4 + regulatory T cell function33 and linked with the susceptibility of pulmonary tuberculosis34. TLR7–8 recognizes single-stranded RNA virus including influenza35,36 and HIV. TLR7–8 agonists can enhance activation of innate immune cells such as CD8 + T cell responses23,24, and are suggested to be vaccine adjuvants37. Specifically, TLR7 encodes pattern-recognition receptors that regulate immune responses acting as viral RNA sensors, is strongly activated only in symptomatic subjects. This unique transcriptional signature manifests 36 h before peak symptoms and is predictive of disease severity38. TLR7 and TLR8 involving in viral sensing play a central role in the vaccine response to trivalent influenza vaccine (TIV) in adults within 24 h after immunization39. While TLR polymorphism has been reported to be associated with increase influenza virus A replication, its pathogenicity, and fatality40, its association with influenza vaccine response has not been reported in previous studies17,40. Our results illustrate the potential role of TLR7–8 gene as key regulators in immunogenicity of seasonal influenza vaccine.
There are limitations in our study. First, the host SNPs but not the entire genome was assessed. Second, the sample size was insufficient to properly account for multiple testing and may only allow detection of large effect associated with host SNPs. Third, further exploration to include other SNPs is needed, such as the SNPs recently reported to affect disease severity (rs12252-C IFITM3)41; rs1755609, rs2438409 GLDC)41,42 and influenza vaccination (rs12252-C IFITM3; rs743811 HO-1, rs2160567 HO-2; rs10220412 IGHV1–69; rs8099917 IL-28B; rs17793951, rs1175540, rs2972164 PPARG, rs2071045 LEP, rs876537 CRP; HLA gene polymorphism)7,43,44,45,46,47. Interplay between different gene polymorphism and humoral response48, the immunogenetics of different influenza vaccines49, and the induced immune response against evolving influenza virus, and the mechanism of influenza-host genetic interactions may be explored in future studies. Fourth, our sample size may be underpowered in some genetic models (Supplementary Table 8), and hence there could be some SNPs that could have effect on influenza vaccine response, but unidentified. Finally, there could be difference in vaccine response among strains (Supplementary Table 9), and such heterogeneity may reduce the power to detect association.
In conclusion, the result from this study has demonstrated the importance of host genetic variation in affecting the response to influenza vaccination. Our findings may help to explain the great variability in the protection achieved by influenza vaccination in different individuals. The identification of genetic variations associated with poor response and adverse effect on receiving influenza vaccination also enhanced our understanding in the area and could contribute to the development of better vaccines that may offer improved protection to all recipients. Our study could help to overcome barriers in the field of vaccinology and the response of vaccines, particularly for pediatric population. Our study also provides a framework how influenza vaccines can be optimized by considering immunogenetics in its design, including the exploration on adjuvants that target the proteins encoded by these TLR genes to circumvent immunogenetic restrictions.
Methods
Study design
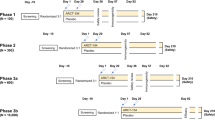
Data were collected in two community-based randomized controlled trials of influenza vaccination conducted in 2008–2009 (pilot study) and 2009–2010 (main study) in Hong Kong2,3. In these trials, children (6–15 y in pilot study and 6–17 y in main study) were randomly allocated to receive either a single dose of trivalent-inactivated influenza vaccine (TIV, Sanofi Pasteur) or saline placebo. Serum specimens were collected at the enrollment to the study and 1 month after vaccination.
Ethics
Proxy written consent from parents or legal guardians was obtained for participants who were <18 years old, with additional written assent from those ages 8–17 years. The study protocol was approved by the Institutional Review Board of the University of Hong Kong.
Laboratory methods
Serum specimens were tested against the vaccine strains A/Brisbane/59/2007(H1N1) and B/Brisbane/60/2008-like (Victoria lineage), the prevalent seasonal strain A/Perth/16/2009-like(H3N2), in parallel by hemagglutination inhibition assays in serial doubling dilutions from an initial dilution of 1:10 using standard methods50.
Whole blood samples were collected for genetic analysis in this study. DNA was extracted and genotyped for SNPs for IL-1B -511G > A (rs16944), IL-6–5843A/G (rs1818879), IL-8–251T/A (rs4073), IL-10–1082T/C (rs1800896), -819A/G (rs1800871), -592T/G (rs1800872), MBL-2–5232C > T, (rs1800451). -221C/G (rs7096206), -34G > A (rs5030737). -550G > C (rs11003125), MxA-88G/T (rs2071430), OAS1–347A/G (rs2660), RIG1 G/C (rs9695310), TLR3–1377T/G (rs3775290), -7C/A (rs3775296), TLR4 G/A (rs5030718), Asp299Gly (rs4986790), TLR7 Gln11Leu (rs179008), 1817G/T (rs5741880), TLR8–129G/C (rs3764879), Met1Val (rs3764880), and (rs11003131)G/T. These SNPs were selected based on a candidate gene approach, selected based on previous literatures17,18,19,20,21,22,23,24,25.
Statistical analysis
We measured the response to influenza vaccine by (1) post-vaccination titers ≥1:40 for the three vaccine strains, (2) ≥4-fold rise after vaccination for the three vaccine strains and (3) the average increase of logarithm of geometric mean titers (GMT) for the three vaccine strains. To evaluate the relationship between host SNPs with vaccine response, we tested four genetic models, including dominant model, recessive model, over-dominant model and multiplicative models51. Therefore, for each host SNPs, genotypes were combined under different genetic models in the statistical analysis. In each genetic model, we used logistic regression to estimate the odds ratio of antibody response to vaccination among different genotype. We used the Benjamini-Hochberg Procedure to adjust the p value for multiple testing52. The same procedure was repeated to explore the relationship between adverse vaccine response, defined as presence of two or more symptoms out of ten following symptoms within 4 days after vaccination: fever, chills, fatigue, headache, cough, muscle pain, swell, redness, bruising and injection pain. All statistical analyses were conducted using R version 3.6.1 (R Foundation for Statistical Computing, Vienna, Austria).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The data that support the findings of this study are available on request from the corresponding author T.K.T. The data are not publicly available due to containing information that could compromise research participant privacy.
References
Bellei, N. C., Carraro, E., Castelo, A. & Granato, C. F. Risk factors for poor immune response to influenza vaccination in elderly people. Braz. J. Infect. Dis. 10, 269–273 (2006).
Cowling, B. J. et al. Protective efficacy against pandemic influenza of seasonal influenza vaccination in children in Hong Kong: a randomized controlled trial. Clin. Infect. Dis. 55, 695–702 (2012).
Cowling, B. J. et al. Protective efficacy of seasonal influenza vaccination against seasonal and pandemic influenza virus infection during 2009 in Hong Kong. Clin. Infect. Dis. 51, 1370–1379 (2010).
Poland, G. A., Ovsyannikova, I. G. & Jacobson, R. M. Immunogenetics of seasonal influenza vaccine response. Vaccine 26, D35–D40 (2008).
Linnik, J. E. & Egli, A. Impact of host genetic polymorphisms on vaccine induced antibody response. Hum. Vaccin Immunother. 12, 907–915 (2016).
Domnich, A., Manini, I., Calabro, G. E., Waure, C. & Montomoli, E. Mapping Host-Related Correlates of Influenza Vaccine-Induced Immune Response: An Umbrella Review of the Available Systematic Reviews and Meta-Analyses. Vaccines (Basel) 7, 215 (2019).
Posteraro, B. et al. The link between genetic variation and variability in vaccine responses: systematic review and meta-analyses. Vaccine 32, 1661–1669 (2014).
Gelder, C. M. et al. Associations between human leukocyte antigens and nonresponsiveness to influenza vaccine. J. Infect. Dis. 185, 114–117 (2002).
Poland, G. A., Ovsyannikova, I. G. & Kennedy, R. B. Personalized vaccinology: a review. Vaccine 36, 5350–5357 (2018).
Poland, G. A., Kennedy, R. B. & Ovsyannikova, I. G. Vaccinomics and personalized vaccinology: is science leading us toward a new path of directed vaccine development and discovery. PLoS Pathog. 7, e1002344 (2011).
Traversi, D. et al. Precision Medicine and Public Health: New Challenges for Effective and Sustainable Health. J. Pers. Med. 11, 135 (2021).
Calabro, G. E. et al. The Value(s) of Vaccination: Building the Scientific Evidence According to a Value-Based Healthcare Approach. Front. Public Health 10, 786662 (2022).
Newport, M. J. The genetic regulation of infant immune responses to vaccination. Front. Immunol. 6, 18 (2015).
Zhu, W. et al. A whole genome transcriptional analysis of the early immune response induced by live attenuated and inactivated influenza vaccines in young children. Vaccine 28, 2865–2876 (2010).
Alcorn, J. F. et al. Differential gene expression in peripheral blood mononuclear cells from children immunized with inactivated influenza vaccine. Hum. Vaccin Immunother. 16, 1782–1790 (2020).
Villani, L., D’Ambrosio, F., Ricciardi, R., de Waure, C. & Calabro, G. E. Seasonal influenza in children: Costs for the health system and society in Europe. Influenza Other Respir. Viruses 16, 820–831 (2022).
Tang, Y. W., Li, H., Wu, H., Shyr, Y. & Edwards, K. M. Host single-nucleotide polymorphisms and altered responses to inactivated influenza vaccine. J. Infect. Dis. 196, 1021–1025 (2007).
MacLeod, H. & Wetzler, L. M. T cell activation by TLRs: a role for TLRs in the adaptive immune response. Sci. STKE 2007, pe48 (2007).
Tang, Y. W. et al. Analysis of candidate-host immunogenetic determinants in herpes simplex virus-associated Mollaret’s meningitis. Clin. Infect. Dis. 30, 176–178 (2000).
Ip, W. K. et al. Mannose-binding lectin in severe acute respiratory syndrome coronavirus infection. J. Infect. Dis. 191, 1697–1704 (2005).
McGuire, W., Hill, A. V., Allsopp, C. E., Greenwood, B. M. & Kwiatkowski, D. Variation in the TNF-alpha promoter region associated with susceptibility to cerebral malaria. Nature 371, 508–510 (1994).
Mira, J. P. et al. Association of TNF2, a TNF-alpha promoter polymorphism, with septic shock susceptibility and mortality: a multicenter study. JAMA 282, 561–568 (1999).
Wille-Reece, U. et al. HIV Gag protein conjugated to a Toll-like receptor 7/8 agonist improves the magnitude and quality of Th1 and CD8+ T cell responses in nonhuman primates. Proc. Natl Acad. Sci. USA 102, 15190–15194 (2005).
Craft, N. et al. The TLR7 agonist imiquimod enhances the anti-melanoma effects of a recombinant Listeria monocytogenes vaccine. J. Immunol. 175, 1983–1990 (2005).
Texereau, J. et al. The importance of Toll-like receptor 2 polymorphisms in severe infections. Clin. Infect. Dis. 41, S408–S415 (2005).
Franco, L. M. et al. Integrative genomic analysis of the human immune response to influenza vaccination. Elife 2, e00299 (2013).
Ng, S. et al. The effect of age and recent influenza vaccination history on the immunogenicity and efficacy of 2009-10 seasonal trivalent inactivated influenza vaccination in children. PLoS ONE 8, e59077 (2013).
Bedford, T. et al. Global circulation patterns of seasonal influenza viruses vary with antigenic drift. Nature 523, 217–220 (2015).
Bowie, A. G. Translational mini-review series on Toll-like receptors: recent advances in understanding the role of Toll-like receptors in anti-viral immunity. Clin. Exp. Immunol. 147, 217–226 (2007).
To, E. E. et al. Intranasal and epicutaneous administration of Toll-like receptor 7 (TLR7) agonists provides protection against influenza A virus-induced morbidity in mice. Sci. Rep. 9, 2366 (2019).
Ramirez-Ortiz, Z. G. et al. The receptor TREML4 amplifies TLR7-mediated signaling during antiviral responses and autoimmunity. Nat. Immunol. 16, 495–504 (2015).
van der Made, C. I. et al. Presence of Genetic Variants Among Young Men With Severe COVID-19. JAMA 324, 663–673 (2020).
Peng, G. et al. Toll-like receptor 8-mediated reversal of CD4+ regulatory T cell function. Science 309, 1380–1384 (2005).
Davila, S. et al. Genetic association and expression studies indicate a role of toll-like receptor 8 in pulmonary tuberculosis. PLoS Genet. 4, e1000218 (2008).
Diebold, S. S. et al. Innate antiviral responses by means of TLR7-mediated recognition of single-stranded RNA. Science 303, 1529–1531 (2004).
Lund, J. M. et al. Recognition of single-stranded RNA viruses by Toll-like receptor 7. Proc. Natl Acad. Sci. USA 101, 5598–5603 (2004).
Kumar, S., Sunagar, R. & Gosselin, E. Bacterial Protein Toll-Like-Receptor Agonists: A Novel Perspective on Vaccine Adjuvants. Front. Immunol. 10, 1144 (2019).
Huang, Y. et al. Temporal dynamics of host molecular responses differentiate symptomatic and asymptomatic influenza a infection. PLoS Genet. 7, e1002234 (2011).
Bucasas, K. L. et al. Early patterns of gene expression correlate with the humoral immune response to influenza vaccination in humans. J. Infect. Dis. 203, 921–929 (2011).
Nogales, A. & M, L. D. Host Single Nucleotide Polymorphisms Modulating Influenza A Virus Disease in Humans. Pathogens 8, 168 (2019).
Everitt, A. R. et al. IFITM3 restricts the morbidity and mortality associated with influenza. Nature 484, 519–523 (2012).
Zhou, J. et al. Identification and characterization of GLDC as host susceptibility gene to severe influenza. EMBO Mol. Med. 11, e9528 (2019).
Cummins, N. W. et al. Heme oxygenase-1 regulates the immune response to influenza virus infection and vaccination in aged mice. FASEB J. 26, 2911–2918 (2012).
Avnir, Y. et al. IGHV1-69 polymorphism modulates anti-influenza antibody repertoires, correlates with IGHV utilization shifts and varies by ethnicity. Sci. Rep. 6, 20842 (2016).
Egli, A. et al. IL-28B is a key regulator of B- and T-cell vaccine responses against influenza. PLoS Pathog. 10, e1004556 (2014).
Ovsyannikova, I. G. et al. Leptin and leptin-related gene polymorphisms, obesity, and influenza A/H1N1 vaccine-induced immune responses in older individuals. Vaccine 32, 881–887 (2014).
Lambkin, R., Novelli, P., Oxford, J. & Gelder, C. Human genetics and responses to influenza vaccination: clinical implications. Am. J. Pharmacogenom. 4, 293–298 (2004).
Verhein, K. C., Vellers, H. L. & Kleeberger, S. R. Inter-individual variation in health and disease associated with pulmonary infectious agents. Mamm. Genome 29, 38–47 (2018).
Cianci, R., Newton, E. E. & Pagliari, D. Efforts to Improve the Seasonal Influenza Vaccine. Vaccines (Basel) 8, 645 (2020).
Cowling, B. J. et al. Comparative epidemiology of pandemic and seasonal influenza A in households. N. Engl. J. Med. 362, 2175–2184 (2010).
Horita, N. & Kaneko, T. Genetic model selection for a case-control study and a meta-analysis. Meta Gene 5, 1–8 (2015).
Benjamini, Y., Drai, D., Elmer, G., Kafkafi, N. & Golani, I. Controlling the false discovery rate in behavior genetics research. Behav. Brain Res. 125, 279–284 (2001).
Acknowledgements
The authors thank Drs. SS Cherny, MM Garcia-Barcelo, Man So and T Zhu for facilitating the DNA extraction. The authors thank anonymous reviewers for their helpful and insightful comments and suggestions. The authors thank Xiaotong Huang for technical assistance. The authors thank Chan Kit Man, Kwok Hung Chan, Calvin Cheng, Lai-Ming Ho, Ho Yuk Ling, Nicole Huang, Lam Yiu Pong, Tom Lui, Edward Ma, Sophia Ng, Tong Hok Leung, Loretta Mak, Winnie Wai, Kevin Yau and Jenny Yuen for research support. This study was supported by the Research Fund for the Control of Infectious Diseases of the Health, Welfare and Food Bureau of the Hong Kong SAR Government (grant CHP-CE-03), the Theme-based Research Scheme project no. T11–712/19 N from the Hong Kong Government, the Health and Medical Research Fund, Food and Health Bureau, Government of the Hong Kong Special Administrative Region (grant no. HKU 767510 M),
Author information
Authors and Affiliations
Contributions
T.K.T. and D.K.M.I designed research. B.J.C., V.J.F., and R.A.P.M.P. conducted the cohort study. T.K.T. and C.W. analyzed data. T.K.T. wrote the paper. D.K.M.I., N.N.Y.T., and J.S.M.P. contributed to revision of the paper. All authors discussed the results and commented on the paper.
Corresponding authors
Ethics declarations
Competing interests
B.J.C. reports honoraria from AstraZeneca, Fosun Pharma, GlaxoSmithKline, Haleon, Moderna, Pfizer, Roche, and Sanofi Pasteur. The authors report no other potential competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tsang, T.K., Wang, C., Tsang, N.N.Y. et al. Impact of host genetic polymorphisms on response to inactivated influenza vaccine in children. npj Vaccines 8, 21 (2023). https://doi.org/10.1038/s41541-023-00621-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41541-023-00621-1