Abstract
Glutaredoxins (GRXs) are small oxidoreductases that can modify target protein activities through control of the redox (reduction/oxidation) state by reducing or glutathionylating disulfide bridges. Although CC-type GRXs are plant specific and play important roles in many processes, the mechanisms by which they modulate the activity of target proteins in vivo are unknown. In this study, we show that a maize CC-type GRX, MALE STERILE CONVERTED ANTHER1 (MSCA1), acts redundantly with two paralogues, ZmGRX2 and ZmGRX5, to modify the redox state and the activity of its putative target, the TGA transcription factor FASCIATED EAR4 (FEA4) that acts as a negative regulator of inflorescence meristem development. We used CRISPR–Cas9 to create a GRX triple knockout, resulting in severe suppression of meristem, ear and tassel growth and reduced plant height. We further show that GRXs regulate the redox state, DNA accessibility and transcriptional activities of FEA4, which acts downstream of MSCA1 and its paralogues to control inflorescence development. Our findings reveal the function of GRXs in meristem development, and also provide direct evidence for GRX-mediated redox modification of target proteins in plants.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
All sequencing data have been deposited in the NCBI Sequence Read Archive with the accession code PRJNA639431. Source data are provided with this paper. All other data that support the findings of this study are available from the corresponding authors on request.
References
Mhamdi, A. & Van Breusegem, F. Reactive oxygen species in plant development. Development 145, dev164376 (2018).
Zeng, J., Dong, Z., Wu, H., Tian, Z. & Zhao, Z. Redox regulation of plant stem cell fate. EMBO J. 36, 2844–2855 (2017).
Weits, D. A. et al. An apical hypoxic niche sets the pace of shoot meristem activity. Nature 569, 714–717 (2019).
Martins, L., Trujillo-Hernandez, J. A. & Reichheld, J.-P. Thiol based redox signaling in plant nucleus. Front. Plant Sci. 9, 705 (2018).
Després, C. et al. The Arabidopsis NPR1 disease resistance protein is a novel cofactor that confers redox regulation of DNA binding activity to the basic domain/leucine zipper transcription factor TGA1. Plant Cell 15, 2181–2191 (2003).
Budimir, J., Treffon, K., Nair, A., Thurow, C. & Gatz, C. Redox‐active cysteines in TGACG‐BINDING FACTOR 1 (TGA1) do not play a role in salicylic acid or pathogen‐induced expression of TGA1‐regulated target genes in Arabidopsis thaliana. New Phytol. 230, 2420–2432 (2021).
Li, S. et al. Nuclear activity of ROXY1, a glutaredoxin interacting with TGA factors, is required for petal development in Arabidopsis thaliana. Plant Cell 21, 429–441 (2009).
Gutsche, N. & Zachgo, S. The N-terminus of the floral Arabidopsis TGA transcription factor PERIANTHIA mediates redox-sensitive DNA-binding. PLoS ONE 11, e0153810 (2016).
Xing, S. & Zachgo, S. ROXY1 and ROXY2, two Arabidopsis glutaredoxin genes, are required for anther development. Plant J. 53, 790–801 (2008).
Zander, M., Chen, S., Imkampe, J., Thurow, C. & Gatz, C. Repression of the Arabidopsis thaliana jasmonic acid/ethylene-induced defense pathway by TGA-interacting glutaredoxins depends on their C-terminal ALWL motif. Mol. Plant 5, 831–840 (2012).
Quon, T., Lampugnani, E. R. & Smyth, D. R. PETAL LOSS and ROXY1 interact to limit growth within and between sepals but to promote petal initiation in Arabidopsis thaliana. Front. Plant Sci. 8, 152 (2017).
Li, N. et al. TGACG-BINDING FACTORs (TGAs) and TGA-interacting CC-type glutaredoxins modulate hyponastic growth in Arabidopsis thaliana. N. Phytol. 221, 1906–1918 (2019).
Pautler, M. et al. FASCIATED EAR4 encodes a bZIP transcription factor that regulates shoot meristem size in maize. Plant Cell 27, 104–120 (2015).
Yang, F. et al. A maize glutaredoxin gene, Abphyl2, regulates shoot meristem size and phyllotaxy. Plant Cell 27, 121–131 (2015).
Chaubal, R. et al. The transformation of anthers in the msca1 mutant of maize. Planta 216, 778–788 (2003).
Diss, G., Ascencio, D., DeLuna, A. & Landry, C. R. Molecular mechanisms of paralogous compensation and the robustness of cellular networks. J. Exp. Zool. B 322B, 488–499 (2014).
Rodriguez-Leal, D. et al. Evolution of buffering in a genetic circuit controlling plant stem cell proliferation. Nat. Genet. 51, 786–792 (2019).
Kim, J.-R., Yoon, H. W., Kwon, K.-S., Lee, S.-R. & Rhee, S. G. Identification of proteins containing cysteine residues that are sensitive to oxidation by hydrogen peroxide at neutral pH. Anal. Biochem. 283, 214–221 (2000).
Tian, Y. et al. Hydrogen peroxide positively regulates brassinosteroid signaling through oxidation of the BRASSINAZOLE-RESISTANT1 transcription factor. Nat. Commun. 9, 1063 (2018).
Hill, B. G., Reily, C., Oh, J.-Y., Johnson, M. S. & Landar, A. Methods for the determination and quantification of the reactive thiol proteome. Free Radic. Biol. Med. 47, 675–683 (2009).
Zhang, Y. et al. Model-based analysis of ChIP-seq (MACS). Genome Biol. 9, R137 (2008).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Rouhier, N., Cerveau, D., Couturier, J., Reichheld, J.-P. & Rey, P. Involvement of thiol-based mechanisms in plant development. Biochim. Biophys. Acta Gen. Subj. 1850, 1479–1496 (2015).
Phillips, K. A. et al. vanishing tassel2 Encodes a grass-specific tryptophan aminotransferase required for vegetative and reproductive development in maize. Plant Cell 23, 550–566 (2011).
McSteen, P. et al. barren inflorescence2 Encodes a co-ortholog of the PINOID serine/threonine kinase and is required for organogenesis during inflorescence and vegetative development in maize. Plant Physiol. 144, 1000–1011 (2007).
Gallavotti, A. et al. sparse inflorescence1 Encodes a monocot-specific YUCCA-like gene required for vegetative and reproductive development in maize. Proc. Natl Acad. Sci. USA 105, 15196–15201 (2008).
Galli, M. et al. Auxin signaling modules regulate maize inflorescence architecture. Proc. Natl Acad. Sci. USA 112, 13372–13377 (2015).
Rouhier, N., Gelhaye, E. & Jacquot, J.-P. Plant glutaredoxins: still mysterious reducing systems. Cell. Mol. Life Sci. 61, 1266–1277 (2004).
Hong, L. et al. Somatic and reproductive cell development in rice anther is regulated by a putative glutaredoxin. Plant Cell 24, 577–588 (2012).
Díaz, M. G. et al. Redox regulation of PEP activity during seedling establishment in Arabidopsis thaliana. Nat. Commun. 9, 50 (2018).
Tsukagoshi, H., Busch, W. & Benfey, P. N. Transcriptional regulation of ROS controls transition from proliferation to differentiation in the root. Cell 143, 606–616 (2010).
Viola, I. L., Guttlein, L. N. & Gonzalez, D. H. Redox modulation of plant developmental regulators from the class I TCP transcription factor family. Plant Physiol. 162, 1434–1447 (2013).
Schippers, J. H., Foyer, C. H. & van Dongen, J. T. Redox regulation in shoot growth, SAM maintenance and flowering. Curr. Opin. Plant Biol. 29, 121–128 (2016).
Tognetti, V. B., Bielach, A. & Hrtyan, M. Redox regulation at the site of primary growth: auxin, cytokinin and ROS crosstalk: apical meristems plasticity in response to stress. Plant Cell Environ. 40, 2586–2605 (2017).
Yu, Q. et al. A P-loop NTPase regulates quiescent center cell division and distal stem cell identity through the regulation of ROS homeostasis in Arabidopsis root. PLoS Genet. 12, e1006175 (2016).
Huang, L. et al. Autophagy regulates glucose-mediated root meristem activity by modulating ROS production in Arabidopsis. Autophagy 15, 407–422 (2019).
Bashandy, T. et al. Interplay between the NADP-linked thioredoxin and glutathione systems in Arabidopsis auxin signaling. Plant Cell 22, 376–391 (2010).
Leiboff, S. et al. Genetic control of morphometric diversity in the maize shoot apical meristem. Nat. Commun. 6, 8974 (2015).
Greb, T. et al. Molecular analysis of the LATERAL SUPPRESSOR gene in Arabidopsis reveals a conserved control mechanism for axillary meristem formation. Genes Dev. 17, 1175–1187 (2003).
Zhang, Z., Yang, J. & Wu, Y. Transcriptional regulation of zein gene expression in maize through the additive and synergistic action of opaque2, prolamine-box binding factor, and O2 heterodimerizing proteins. Plant Cell 27, 1162–1172 (2015).
Botello-Morte, L. et al. Cysteine mutational studies provide insight into a thiol-based redox switch mechanism of metal and DNA binding in FurA from Anabaena sp. PCC 7120. Antioxid. Redox Signal. 24, 173–185 (2016).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Thorvaldsdóttir, H., Robinson, J. T. & Mesirov, J. P. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief. Bioinform. 14, 178–192 (2013).
Tian, T. et al. agriGO v2.0: a GO analysis toolkit for the agricultural community, 2017 update. Nucleic Acids Res. 45, W122–W129 (2017).
Acknowledgements
We thank M. Bai (Shandong University) for technical assistance and critical discussions. We thank L. Qin (Core Facility for Bioimaging, Huazhong Agricultural University) for SEM imaging assistance. This work was supported by the National Natural Science Foundation of China (nos. 31871633 to F.Y., R.S.Y., Y.M.W., W.S.Z. and H.J.S.); the Fundamental Research Funds for the Central Universities (no. ZK201908 and 2021ZKPY010 to F.Y., L.D., Y.N.S. and T.J.T.); the 111 Project (no. B20051 to F.Y.); the Next-Generation BioGreen 21 Program SSAC Project (no. PJ01322602) from the Rural Development Administration, Republic of Korea (to D.J.); the National Science Foundation Plant Genome Research Program (nos. IOS-1546837 and 2129189 to D.J.); the Taishan Scholars of Shandong Province (no. tsqn201812018 to F.X.); and the Human Frontier Science Program Long-Term Fellowship (no. LT000227/2016 to F.X.).
Author information
Authors and Affiliations
Contributions
F.Y. and D.J. conceived and designed experiments. R.Y., F.X., W.Z., Y.W., T.T. and H.S. performed experiments. L.D. and Y.S. analysed data. F.Y., F.X., R.Y. and D.J. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Plants thanks the anonymous reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 The agronomic traits of msca1 and its paralog mutants.
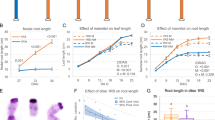
a, The knock-out mutations (red asterisks on the gene model) for ZmGRX2 and ZmGRX5 were caused by CRISPR/Cas9 gene editing. zmgrx2-1 and zmgrx2-2 contain 55 bp and 70 bp deletions before the conserved CCMC motif, respectively. zmgrx5-1 contains 13 bp deletion within the CCMC motif, and zmgrx5-2 harbors 5 bp deletion before the motif. b, Mutations in MSCA1 and its paralogs have effects on the plant architecture, with the msca1;zmgrx5-1 double and grx triple-1 (msca1;zmgrx2-1;zmgrx5-1) mutants being obviously shorter. c-f, The plant height (c), kernel row number (d), tassel branch number (e) and spikelet density (f) of msca1;zmgrx5 and grx triple were significantly decreased in comparison to wild type (WT). Data are shown as mean ± SD, dots show data distribution. (n > 10 biologically independent samples, P values calculated using two-tailed t-test). g-i, grx triple-2 (msca1;zmgrx2-2;zmgrx5-2) mutants were shorter (g), with less tassel branches (h), and had smaller ears with less kernels (i) in comparison to wild type plants. Scale bars, 30 cm in b and g, 5 cm in h and i.
Extended Data Fig. 2 Compensation expression of three GRX genes.
All three GRX genes show a complementary expression upon its paralog mutations. Expression was measured by real-time quantitative RT-PCR using the tips of 3-5 mm ears. The experiments were repeated at least three times with similar results. Data are shown as mean ± SD, dots show data distribution. Error bars show the standard deviation (SD) from three independent experiments. The statistical significance was estimated using two-tailed t test with **P < 0.01, ***P < 0.001.
Extended Data Fig. 3 Agronomic traits of the mutants containing fea4 and grx mutations.
a-c, Combination of fea4 and grx triple mutations mimics to the grx triple in plant height (a, b) and tassel branch number (c). Data are shown as mean ± SD, dots show data distribution. (n > 10 biologically independent samples, P values calculated using two-tailed t-test). Scale bar, 30 cm in a.
Extended Data Fig. 4 FEA4 also interact with ZmGRX2 and ZmGRX5, and the point mutations in FEA4 and MSCA1 for the interaction assays.
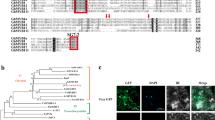
a, FEA4 interacts with ZmGRX2 and ZmGRX5 in yeast-two hybrid assay, SD-LT represents non-selective plates lacking leucine and tryptophan, and SD-AHLT represents selective plates lacking adenine, histidine, leucine and tryptophan. BD: GAL4 DNA binding domain; AD: GAL4 activation domain. b, Luciferase complementation image assay (LCI) confirmed that FEA4 interacts with ZmGRX2 and ZmGRX5. c, Amino acid sequence alignment of FEA4 with its orthologs shows Cys321 (red arrow) is the conserved cysteine residue. Cys321 and Cys209 (pink) were mutated to Serine (S). PAN (PERIANTHIA) and Os06g0265400 are the closest orthologs in Arabidopsis and rice (Oryza sativa), respectively. d, Amino acid sequence alignment of MSCA1 with its orthologs shows that the ‘CCMC’ motif (red solid line) and a functional site Valine 65 (V65) are conserved (red arrows). The conserved cysteines were mutated to serine (C72S;C75S), and V65 was mutated to Methionine (V65M). ROXY1&2 are the close orthologs in Arabidopsis.
Extended Data Fig. 5 Detection of the fusion proteins expression in the Y2H assays and LCI assays in Fig. 3 and Fig. 5.
a, Western blots of yeast extracts from the cells expressing GAL4 AD domain (AD) fused with different proteins using HA antibody, and GAL4 BD domain (BD) fused with different proteins using Myc antibody (the same number represents the same cell expressing a combination of AD and BD fusion proteins). Lanes 1-3 were for Fig. 3d, lanes 4-6 were for Fig. 3e, and lanes 7-10 were for Fig. 5a. All samples were shown to have proper proteins expressed. The Ponceau S staining was provided as the loading controls. The experiments were repeated at least three times with similar results. b-d, Western blots of the protein extracts from N. benthamiana leaves co-infiltrated with the combinations as labeling. The anti-luciferase antibodies were used to detect the N-terminus and C-terminus of the luciferase fused with different proteins, respectively, based on the size differences. (b) is for Fig. 3f, (c) is for Fig. 3g, and (d) is for Fig. 5b. All samples were shown to have proper proteins expressed. The Ponceau S staining is provided as the loading control. The experiments were repeated at least three times with similar results.
Extended Data Fig. 6 MSCA1 can be oxidized and reduced in vitro.
The AMS trapping assay detected the reduced His-SUMO-MSCA1 form is more abundant than the oxidized form. The reduced glutathione (GSH) could efficiently reduce the oxidized MSCA1 even at the presence of H2O2. The experiments were repeated at least three times with similar results.
Extended Data Fig. 7 The YFP-FEA4 protein accumulations in wild type and the grx triple mutant.
Total protein extracts of YFP-FEA4 transgene in two backgrounds were detected by western blotting. The actin was used as the loading control detected by anti-actin antibody. The experiments were repeated at least three times with similar results.
Extended Data Fig. 8 Comparison of FEA4 binding profiles and its target genes between wild type and grx triple mutant.
a, The replicate rates of FEA4 binding peaks identified by ChIP-seq using 3–5 mm ears were about 60% in both wild type (WT, left panel) and the grx triple mutant (triple, right panel). Each replicate was done using more than 30 ears. b, A large portion (>90%) of potential binding peaks (Left) and the corresponding genes (Right) identified for FEA4 were overlap between WT and the grx triple mutants. The number of FEA4 peaks was almost doubled in the grx triple mutants compared to WT. c, The distributions of the high-confident binding peaks of FEA4 in wild type (WT) and the grx triple mutants (triple). ‘1 kb upstream’ and ‘1–3 kb upstream’ represent the regions 1 kb and 1–3 kb upstream of the transcription start sites (TSS), respectively. ‘Downstream’ represents the regions 3 kb downstream of the transcriptional termination sites (TTS). ‘intergenic’ represents the intergenic regions outside the 3 kb upstream of TSS and 3 kb downstream of TTS. d, The TGA motif (TGACG) was enriched in the FEA4-bound promoter regions of both wild type (WT, 56%) and the grx triple mutant (triple, 58.2%). The promoter regions include the regions within 3 kb upstream of TSS and 3 kb downstream of TTS of annotated coding genes. In total 3,059 out of 5,459 and 7,094 out of 12,181 FEA4 binding sites in the promoter regions of WT and triple mutant, respectively, contain at least one TGA motif. e, The overlapping DEGs caused by fea4 mutation with FEA4-bound genes either in wild type (left panel) or grx triple mutant background (right panel). f, On the common FEA4-bound sites present in the 75 overlapping genes, the peaks captured in the grx triple (triple) mutants are significantly more abundant than those captured in wild type (WT). Pileup values representing the aligned reads with a given extension size were shown. In total, 91 binding peaks of FEA4-YFP captured in both wild type and the grx triple mutants are used for analysis. The box plot boundaries show the interquartile range, and the center line is the median, extended to minimum and maximum values, the whiskers represent 1.5-fold to the interquartile range from the lower and upper quartiles. n = 91, P = 3 ×10−4. P value calculated using two-sided Wilcoxon test.
Extended Data Fig. 9 Redox state of FEA4 affects its DNA binding affinity.
a, Protein detection of the recombinant MBP-FEA4 proteins used for EMSA assays in Fig. 6g. MBP-FEA4C321 is present only as monomer. b, Compared to FEA4 under normal redox state, oxidized MBP-FEA4 showed a slightly stronger binding, whereas reduced MBP-FEA4 exhibited a slightly weaker binding to the probe. The upper asterisk points to probe bound by recombinant MBP-FEA4 proteins (shift), the lower asterisk indicates free probes. c, MBP-FEA4 forms more dimer with H2O2 and less dimer with DTT treatment, compared to the untreated MBP-FEA4. The experiments were repeated at least three times with similar results.
Extended Data Fig. 10 Gene Ontology functional enrichment analysis.
Gene Ontology functional enrichment analysis of the partial common regulated genes in the fea4 mutants with the grx triple mutants (a), and the partial differentially expressed genes between grx triple mutants and wild type (b).
Supplementary information
Supplementary Table
Supplementary Table 1. Significant binding peaks called from YFP-FEA4 ChIP–seq in both WT replicates.
Source data
Source Data Fig. 1
Source data for Fig. 1g,j. SAM and IM size measurements of each individual from different genotypes.
Source Data Fig. 3
Source data for Fig. 3c. IM size measurements of each individual from different genotypes.
Source Data Fig. 4
Source data for Fig. 4 b,d,e,g. Unprocessed immunoblots.
Source Data Fig. 5
Source data for Fig. 5 c,d,h,f,j. Unprocessed immunoblots.
Source Data Fig. 6
Source data for Fig. 6e,f. ChIP–qPCR and RT–qPCR results.
Source Data Fig. 6
Source data for Fig. 6g. Full scan of EMSA results.
Source Data Extended Data Fig. 1
Source data for Supplementary Fig. 1c–f. Agronomic trait survey data of each individual from different genotypes.
Source Data Extended Data Fig. 2
Source data for Supplementary Fig. 2. RT–qPCR results.
Source Data Extended Data Fig. 3
Source data for Supplementary Fig. 3b,c. Agronomic trait survey data of each individual from different genotypes.
Source Data Extended Data Fig. 5
Source data for Supplementary Fig. 5a–d. Unprocessed immunoblots.
Source Data Extended Data Fig. 6
Source data for Supplementary Fig. 6. Unprocessed immunoblots.
Source Data Extended Data Fig. 7
Source data for Supplementary Fig. 7. Unprocessed immunoblots.
Source Data Extended Data Fig. 9
Source data for Supplementary Fig. 9b. Full scan of EMSA results.
Rights and permissions
About this article
Cite this article
Yang, R.S., Xu, F., Wang, Y.M. et al. Glutaredoxins regulate maize inflorescence meristem development via redox control of TGA transcriptional activity. Nat. Plants 7, 1589–1601 (2021). https://doi.org/10.1038/s41477-021-01029-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-021-01029-2
This article is cited by
-
Complexation and immobilization of arsenic in maize using green synthesized silicon nanoparticles (SiNPs)
Scientific Reports (2024)
-
Asymmetric gene expression and cell-type-specific regulatory networks in the root of bread wheat revealed by single-cell multiomics analysis
Genome Biology (2023)
-
Novel insights into maize (Zea mays) development and organogenesis for agricultural optimization
Planta (2023)