Abstract
Some plants can ‘remember’ past environmental experience to become adapted to a given environment. For instance, after experiencing prolonged low-temperature exposure in winter (winter cold), vernalization-responsive plants remember past cold experience when temperature rises in spring, to acquire competence to flower at a later season favourable for seed production1,2. In Arabidopsis thaliana, prolonged cold induces silencing of the potent floral repressor FLOWERING LOCUS C (FLC) by Polycomb group (PcG) chromatin modifiers. This Polycomb-repressed chromatin state is epigenetically maintained and thus ‘memorized’ in subsequent growth and development upon return to warmth1,3. ‘Memory of winter cold’ has been viewed as being mitotically stable but meiotically unstable3,4,5, and thus not to be transmitted intergenerationally. In general, whether and how chromatin-mediated environmental memories are transmitted across generations are unknown in plants. Here, we show that the cold-induced Polycomb-repressed chromatin state at FLC or memory of winter cold is maintained in the egg cell, that is meiotically stable in the process of female gamete formation, and provide evidence that this Polycomb-mediated memory is not maintained in the sperm cell. Moreover, we show that this cold memory is inherited maternally but not paternally to the zygote and early embryos. Our study demonstrates and further provides mechanistic insights into intergenerational transmission of chromatin state-mediated environmental memories in plants.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The data supporting the findings in this study are included in this article and its extended data and Supplementary Information or available from the corresponding authors on request. Source data are provided with this paper.
References
Kim, D. H., Doyle, M. R., Sung, S. & Amasino, R. M. Vernalization: winter and the timing of flowering in plants. Annu. Rev. Cell Dev. Biol. 25, 277–299 (2009).
Andres, F. & Coupland, G. The genetic basis of flowering responses to seasonal cues. Nat. Rev. Genet. 13, 627–639 (2012).
Mylne, J. S. et al. LHP1, the Arabidopsis homologue of HETEROCHROMATIN PROTEIN1, is required for epigenetic silencing of FLC. Proc. Natl Acad. Sci. USA 103, 5012–5017 (2006).
Schiessl, S. V., Quezada-Martinez, D., Tebartz, E., Snowdon, R. J. & Qian, L. W. The vernalisation regulator FLOWERING LOCUS C is differentially expressed in biennial and annual Brassica napus. Sci. Rep. 9, 14911 (2019).
Finnegan, E. J., Genger, R. K., Kovac, K., Peacock, W. J. & Dennis, E. S. DNA methylation and the promotion of flowering by vernalization. Proc. Natl Acad. Sci. USA 95, 5824–5829 (1998).
Lee, I., Michaels, S. D., Masshardt, A. S. & Amasino, R. M. The late-flowering phenotype of FRIGIDA and luminidependens is suppressed in the Landsberg erecta strain of Arabidopsis. Plant J. 6, 903–909 (1994).
Angel, A., Song, J., Dean, C. & Howard, M. A Polycomb-based switch underlying quantitative epigenetic memory. Nature 476, 105–108 (2011).
Yuan, W. et al. A cis cold memory element and a trans epigenome reader mediate Polycomb silencing of FLC by vernalization in Arabidopsis. Nat. Genet. 48, 1527–1534 (2016).
Heo, J. B. & Sung, S. Vernalization-mediated epigenetic silencing by a long intronic noncoding RNA. Science 331, 76–79 (2011).
Yang, H. et al. Distinct phases of Polycomb silencing to hold epigenetic memory of cold in. Arabidopsis Sci. 357, 1142–1145 (2017).
Jiang, D. & Berger, F. DNA replication-coupled histone modification maintains Polycomb gene silencing in plants. Science 357, 1146–1149 (2017).
Tao, Z. et al. Embryonic epigenetic reprogramming by a pioneer transcription factor in plants. Nature 551, 124–128 (2017).
Sheldon, C. C. et al. Resetting of FLOWERING LOCUS C expression after epigenetic repression by vernalization. Proc. Natl Acad. Sci. USA 105, 2214–2219 (2008).
Choi, J. et al. Resetting and regulation of FLOWERING LOCUS C expression during Arabidopsis reproductive development. Plant J. 57, 918–931 (2009).
Santos-Mendoza, M. et al. Deciphering gene regulatory networks that control seed development and maturation in Arabidopsis. Plant J. 54, 608–620 (2008).
Li, Z., Fu, X., Wang, Y., Liu, R. & He, Y. Polycomb-mediated gene silencing by the BAH-EMF1 complex in plants. Nat. Genet. 50, 1254–1261 (2018).
Mozgova, I. & Hennig, L. The Polycomb group protein regulatory network. Annu. Rev. Plant Biol. 66, 269–296 (2015).
Belmonte, M. F. et al. Comprehensive developmental profiles of gene activity in regions and subregions of the Arabidopsis seed. Proc. Natl Acad. Sci. USA 110, E435–E444 (2013).
Tao, Z. et al. Embryonic resetting of the parental vernalized state by two B3 domain transcription factors in Arabidopsis. Nat. Plants 5, 424–435 (2019).
Sung, S. et al. Epigenetic maintenance of the vernalized state in Arabidopsis thaliana requires LIKE HETEROCHROMATIN PROTEIN 1. Nat. Genet. 38, 706–710 (2006).
Zhao, P. et al. Two-step maternal-to-zygotic transition with two-phase parental genome contributions. Dev. Cell 49, 882–893 (2019).
Nodine, M. D. & Bartel, D. P. Maternal and paternal genomes contribute equally to the transcriptome of early plant embryos. Nature 482, 94–97 (2012).
Borges, F. et al. Comparative transcriptomics of Arabidopsis sperm cells. Plant Physiol. 148, 1168–1181 (2008).
Wuest, S. E. et al. Arabidopsis female gametophyte gene expression map reveals similarities between plant and animal gametes. Curr. Biol. 20, 506–512 (2010).
Lafos, M. et al. Dynamic regulation of H3K27 trimethylation during Arabidopsis differentiation. PLoS Genet. 7, e1002040 (2011).
Gu, X., Xu, T. & He, Y. A histone H3 lysine-27 methyltransferase complex represses lateral root formation in Arabidopsis thaliana. Mol. Plant 7, 977–988 (2014).
Zhang, X. et al. The Arabidopsis LHP1 protein colocalizes with histone H3 Lys27 trimethylation. Nat. Struct. Mol. Biol. 14, 869–871 (2007).
Baroux, C., Gagliardini, V., Page, D. R. & Grossniklaus, U. Dynamic regulatory interactions of Polycomb group genes: MEDEA autoregulation is required for imprinted gene expression in Arabidopsis. Genes Dev. 20, 1081–1086 (2006).
Steffen, J. G., Kang, I. H., Macfarlane, J. & Drews, G. N. Identification of genes expressed in the Arabidopsis female gametophyte. Plant J. 51, 281–292 (2007).
Gaydos, L. J., Wang, W. & Strome, S. H3K27me and PRC2 transmit a memory of repression across generations and during development. Science 345, 1515–1518 (2014).
Zheng, H. et al. Resetting epigenetic memory by reprogramming of histone modifications in mammals. Mol. Cell 63, 1066–1079 (2016).
Borg, M. et al. Targeted reprogramming of H3K27me3 resets epigenetic memory in plant paternal chromatin. Nat. Cell Biol. 22, 621–629 (2020).
Chen, M. & Penfield, S. Feedback regulation of COOLAIR expression controls seed dormancy and flowering time. Science 360, 1014–1017 (2018).
Shindo, C. et al. Role of FRIGIDA and FLOWERING LOCUS C in determining variation in flowering time of Arabidopsis. Plant Physiol. 138, 1163–1173 (2005).
Schonrock, N. et al. Polycomb-group proteins repress the floral activator AGL19 in the FLC-independent vernalization pathway. Genes Dev. 20, 1667–1678 (2006).
Michaels, S. D. & Amasino, R. M. FLOWERING LOCUS C encodes a novel MADS domain protein that acts as a repressor of flowering. Plant Cell 11, 949–956 (1999).
Blazquez, M. Quantitative GUS activity assay of plant extracts. CSH Protoc. 8, 4690 (2007).
Wang, Y., Gu, X., Yuan, W., Schmitz, R. J. & He, Y. Photoperiodic control of the floral transition through a distinct Polycomb repressive complex. Dev. Cell 28, 727–736 (2014).
Tofanelli, R., Vijayan, A., Scholz, S. & Schneitz, K. Protocol for rapid clearing and staining of fixed Arabidopsis ovules for improved imaging by confocal laser scanning microscopy. Plant Methods 15, 120 (2019).
Wu, Y. et al. Arabidopsis FIMBRIN5, an actin bundling factor, is required for pollen germination and pollen tube growth. Plant Cell 22, 3745–3763 (2010).
Acknowledgements
We are grateful to R. M. Amasino for kindly providing the FRI-Col seeds and I. E. Somssich for the pJawohl8-RNAi plasmid. We thank Y. Li for assistance with constructing the FRI clf and FRI lhp1 lines and Z. Gao for assistance with genetic crossing. This work was supported by the National Natural Science Foundation of China (grant nos. 31721001 and 31830049 to Y.H.) and the Chinese Academy of Sciences (grant no. XDB27030202 to Y.H.).
Author information
Authors and Affiliations
Contributions
Y.H., X.L., Y.O. and R.L. conceived and designed the experiments. X.L., Y.O. and R.L. performed the experiments. All authors took part in data analysis. Y.H. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Plants thanks Daniel Woods and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Analysis of embryonic FLC reactivation following parental vernalization.
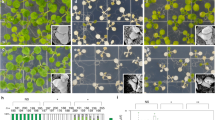
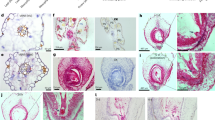
a, Embryonic FLC::GUS reactivation in the plants vernalized for 35 d (V35). Shown are representative FLC::GUS expression patterns in the indicated F1 developing seeds. Number of each staining out of all scored seeds is indicated at bottom left of each image. Scale bars, 100 μm. b, Levels of FLC mRNA in the developing seeds of FRI-Col (WT) vernalized for 35 d or 45 d. Transcript levels were normalized directly to PP2A. Values were are means ± s.d. of three biological replicates. Lower-case letters denote statistically distinct differences (two-way ANOVA, p < 0.05).
Extended Data Fig. 2 Analyses of fertilization and de novo FLC transcription in early developing seeds.
a, Ovules from WT plants (vernalized for 45 d) are fertilized, 6 h after pollination. Pollen tubes were stained by aniline blue, and shown is a representative micrograph of two independent experiments with similar results. Scale bar, 100 μm. b, A DNA gel image to show that residue genomic DNA was completely removed by a DNase treatment in the indicated samples. cDNAs are reverse transcribed from DNase-treated total RNAs, extracted from the F1 developing seeds of NV x V35. The experiment was repeated independently twice with similar results. c, Levels of primary FLC transcripts (unspliced) in the indicated F1 developing seeds. Transcript levels were normalized directly to PP2A. Values are means ± s.d. of three biological replicates.
Extended Data Fig. 3 Temporal and spatial expression patterns of LEC1 from gametophytes through the heart-stage embryo.
A LEC1pro-LEC1:GUS line12 that reflects the endogenous LEC1 expression was examined. Each shown micrograph is a representative from over 26 examined samples; scale bars, 100 μm.
Extended Data Fig. 4 Molecular characterization of the FLC locus in the Se-0 accession.
a, Levels of FLC transcripts in the indicated seedlings. Total RNAs were extracted from non-vernalized (NV) WT (FRI-Col) and Se-0 seedlings, as well as vernalized seedlings (V45) that were grown for 3 d at normal temperature (22 °C) following a 45-d cold exposure. Transcript levels were normalized to PP2A. Values are means ± s.d. of three biological replicates; relative expression to WT (NV) is presented. b, Levels of H3K27me3 on FLC chromatin in the indicated seedlings. NV, non-vernalized; V45, 5 d at normal temperature following 45-d cold. Shown are relative levels of H3K27me3 in the examined two FLC regions (as illustrated in Fig. 1d), following a normalization to the reference gene SHOOTMERISTEMLESS (STM)10. Values are means ± s.d. of three biological replicates. c, d, Genomic polymorphisms of FLC (c) and UBC21 (d) between WT (FRI-Col) and Se-0. The genotype-specific forward primers used to amplify genomic and cDNAs of FLC or UBC21 are denoted. Black boxes for exons, and A of ATG as +1.
Extended Data Fig. 5 Expression of LHP1 and CLF proteins in the egg cell, sperm cell and/or early embryo cells.
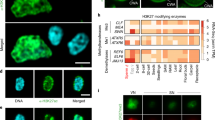
a, LHP1:GFP in the nuclei of mature ovule cells. ECN, egg-cell nucleus; CCN, central-cell nucleus; SYN, synergid cell nucleus. Shown is a representative micrograph from over 50 examined ovules; scale bars, 10 μm. b, c, LHP1:GFP protein is absent in mature pollen grains (b) and germinating pollen grains (c). Nuclei were stained with Hoechst. SN, sperm cell nucleus; VN, vegetative cell nucleus. Scale bars, 10 μm. Note that there is green autofluorescence from the pollen grain wall. d, LHP1:GFP and CLF:GFP expression in 2-DAP F1 embryos of the indicated crosses. Scale bars, 10 μm. b–d, Experiments were repeated independently once with similar results.
Extended Data Fig. 6 Analysis of egg-cell or central-cell knockdown of FIE, CLF, SWN and/or MEA expression.
a, b, Levels of FIE, CLF, SWN and MEA transcripts in mature ovules (a) and 2-DAP developing seeds (b) of the indicated lines. Transcript levels were normalized to PP2A, and values are means ± s.d. of three biological replicates. Statistically significant differences are indicated by lower-case letters (one-way ANOVA, p < 0.001) or p values (two tailed t-test); n.s., not significant. c, FLC::GUS expression patterns in the indicated F1 developing seeds (plants were vernalized for 35 d). Number of each staining out of all scored seeds is indicated at bottom left of each image. Scale bars, 100 μm. d, Normal seed development in the indicated lines. Shown are siliques at around 7 DAP.
Supplementary information
Supplementary Information
Supplementary Table 1.
Source data
Source Data Extended Data Fig. 2
Uncropped DNA gel image.
Rights and permissions
About this article
Cite this article
Luo, X., Ou, Y., Li, R. et al. Maternal transmission of the epigenetic ‘memory of winter cold’ in Arabidopsis. Nat. Plants 6, 1211–1218 (2020). https://doi.org/10.1038/s41477-020-00774-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-020-00774-0
This article is cited by
-
A molecular mechanism for embryonic resetting of winter memory and restoration of winter annual growth habit in wheat
Nature Plants (2024)
-
Distinct chromatin signatures in the Arabidopsis male gametophyte
Nature Genetics (2023)
-
The biological concept of stress revisited: relations of stress and memory of plants as a matter of space–time
Theoretical and Experimental Plant Physiology (2022)
-
Epigenetic repression and resetting of a floral repressor, FLC, in the life cycle of winter-annual Arabidopsis
Plant Biotechnology Reports (2022)
-
Wheat genomic study for genetic improvement of traits in China
Science China Life Sciences (2022)