Abstract
Maturation of iron-sulfur proteins in eukaryotes is initiated in mitochondria by the core iron-sulfur cluster assembly (ISC) complex, consisting of the cysteine desulfurase sub-complex NFS1-ISD11-ACP1, the scaffold protein ISCU2, the electron donor ferredoxin FDX2, and frataxin, a protein dysfunctional in Friedreich’s ataxia. The core ISC complex synthesizes [2Fe-2S] clusters de novo from Fe and a persulfide (SSH) bound at conserved cluster assembly site residues. Here, we elucidate the poorly understood Fe-dependent mechanism of persulfide transfer from cysteine desulfurase NFS1 to ISCU2. High-resolution cryo-EM structures obtained from anaerobically prepared samples provide snapshots that both visualize different stages of persulfide transfer from Cys381NFS1 to Cys138ISCU2 and clarify the molecular role of frataxin in optimally positioning assembly site residues for fast sulfur transfer. Biochemical analyses assign ISCU2 residues essential for sulfur transfer, and reveal that Cys138ISCU2 rapidly receives the persulfide without a detectable intermediate. Mössbauer spectroscopy assessing the Fe coordination of various sulfur transfer intermediates shows a dynamic equilibrium between pre- and post-sulfur-transfer states shifted by frataxin. Collectively, our study defines crucial mechanistic stages of physiological [2Fe-2S] cluster assembly and clarifies frataxin’s molecular role in this fundamental process.
Similar content being viewed by others
Introduction
Iron-sulfur (Fe/S) clusters are ancient inorganic protein cofactors present in almost all organisms, and are involved in numerous biological processes including respiration, metabolism, DNA replication and repair, translation, regulation, and antiviral defense1,2,3,4,5,6. Despite the chemical simplicity of Fe/S clusters, their biosynthesis and insertion into apoproteins requires complex proteinaceous machineries7,8,9,10. In eukaryotes, the mitochondrial Fe/S cluster assembly (ISC) system synthesizes Fe/S clusters de novo and inserts them into mitochondrial target apoproteins. Genetic mutations in the human ISC genes impair this process and cause severe metabolic, neurological and hematological diseases, often with fatal outcomes, the most prevalent being Friedreich’s ataxia with mutations in the frataxin (FXN) gene9,10,11,12,13.
Mitochondrial Fe/S protein biogenesis begins with the de novo assembly of a [2Fe-2S] cluster from Fe2+ and Cys-derived persulfide (SSH) on the scaffold protein ISCU214. Assembly requires the multi-protein ‘core ISC complex’ consisting of ISCU2, the cysteine desulfurase sub-complex NFS1-ISD11-ACP1, FXN, and the ferredoxin FDX215,16,17,18,19,20 (Fig. 1a). The assembled [2Fe-2S] cluster is then released from ISCU2 by a dedicated HSP70 chaperone system and transferred via numerous ISC trafficking and targeting factors toward the mitochondrial client Fe/S apoproteins8,21,22,23,24.
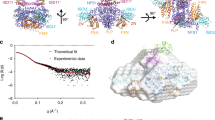
a Model for the synthesis of a [2Fe-2S] cluster on the ISCU2 scaffold protein by the core ISC complex in various steps. Free ISCU2 (Fe-U) with Fe2+ (red circle) bound at its assembly site residues associates with the heterodimeric cysteine desulfurase complex NFS1-ISD11-ACP1 ((NIA)2; only one half of the dimer is shown) to form the (Fe-NIAU)2 complex. Addition of Cys enables rapid persulfidation of NFS1 forming the persulfide-containing (Fe-NSIAU)2 complex and Ala. FXN (X) addition stimulates sulfur transfer from NFS1 to a Cys residue at the ISCU2 assembly site creating the (Fe-NIAUSX)2 intermediate. FDX2-mediated persulfide reduction and interaction of two ISCU2 of different (Fe-NIAUSX)2 complexes leads to the formation of holo-ISCU2 with a bound [2Fe-2S] cluster. b 2.4 Å consensus cryo-EM map (C2 symmetry applied) of the (Fe-NIAUX)2 complex during persulfide transfer, segmented and colored by subunit. Unless indicated otherwise, subunit coloring is consistent throughout the manuscript. Symmetry expansion and subsequent focused 3D classification allowed further separation of the (Fe-NIAU)2 (c), (Fe-NIAUSX)2 (d), and (Fe-NSIAUSX)2 (e) reaction intermediates (left). Middle: the cartoon representation shows a detailed view of the NFS1-ISCU2 interface with the different locations of the Cys loop of NFS1, either in an outward position with Cys381NFS1 coordinating the Fe2+ at the ISCU2 assembly site (c, e), or in an inward orientation (d) with Cys381NFS1 close to the PLP cofactor of NFS1. Right: the close-up of the respective ISCU2 assembly site regions (with or without Cys381NFS1) depicts the different modes of Fe2+ coordination in these sulfur transfer intermediates. The cryo-EM maps reveal clear densities for a persulfide (CSS) on Cys381NFS1 and/or Cys138ISCU2 (d, e).
Despite a number of biochemical and structural studies of the core ISC complex, the biochemical mechanism of de novo [2Fe-2S] cluster assembly on ISCU2 is poorly understood at the molecular level. So far, structural insights are restricted to complexes with either no metal or inhibitory Zn bound to ISCU2 rather than the physiological Fe. No structures of reaction intermediates are available to date. Biochemical studies revealed that initially the NFS1-ISD11-ACP1 heterodimer (hereafter termed (NIA)2) uses free Cys to generate alanine and a persulfide bound to the conserved Cys381NFS1 25,26 (Fig. 1a). This residue is located on a flexible loop (Cys loop) that moves toward the Fe2+ binding site of ISCU2 (denoted as U)19, thereby forming the (Fe-NSIAU)2 complex (S indicates persulfidation of the preceding ISC component). The (NIAU)2 complex was shown to have increased affinity for FXN (X)16,27,28,29. FXN binding to generate the intermediate (Fe-NSIAUX)2 complex strongly increases the rate of persulfide transfer from NFS1 to ISCU2 (yielding (Fe-NIAUSX)2) (Fig. 1a), implying a dedicated physiological role of FXN in this step, but FXN’s precise mechanistic function remains unclear17,25,26,30. Studies performed aerobically suggested Cys69ISCU2 and Cys138ISCU2 as potential primary residues accepting the sulfur from NFS1, and Cys138ISCU2 as the final acceptor25. In the human (Zn-NIAU)2 X-ray structure15, the latter residue is furthest from the sulfur-donating Cys381NFS1 (distances for ligand-metal coordination and for Cys-Cys pairs in various ISC complex structures are compiled in Table 1). In the (Zn-NIAUX)2 electron cryo-microscopy (cryo-EM) structure17, a rather different arrangement was observed at the metal site, with Cys381NFS1 being far away from the ISCU2-bound Zn and its coordinating residues, hence providing no clues on the mechanism of sulfur transfer. Clearly, structural information for the Fe-containing ISC complex intermediates is needed. Notably, in the related Bacillus subtilis SufS-SufU sulfur transfer system, Cys41SufU (corresponding to Cys69ISCU2) serves as the sulfur-accepting residue indicating a conspicuous difference for the ISC and SUF systems31.
Following sulfur transfer, the persulfide bound to ISCU2 in the (Fe-NIAUSX)2 complex is reduced to form sulfide (S2‒) by FDX2, which receives electrons from the NADPH-dependent ferredoxin reductase (FDXR)14,20,30,32. The reduction step generates a transient [1Fe-1S] intermediate that is finally fused to a [2Fe-2S] cluster on ISCU2 by dimerization facilitated by the hydrophobic interaction of the conserved N-terminal Tyr35 residues of two [1Fe-1S]-ISCU2 units18.
While Fe2+ binding to ISCU2 is essential for the persulfidation, reduction and cluster synthesis steps, the specific ISCU2 assembly site residues required for Fe coordination have been defined only for chemically reconstituted mouse Fe-ISCU (corresponding to human residues Cys69ISCU2, Asp71ISCU2, Cys95ISCU2, and His137ISCU2)19,30 (Table 1). Fe-loaded ISCU2 can bind to (NIA)2 (Fig. 1a), yet the structure of this complex is currently unknown. In structures of related Zn-loaded complexes, metal coordination by Cys69ISCU2 was found to be replaced by Cys381NFS1 in human (Zn-NIAU)215 and by Cys138ISCU2 in (Zn-NIAUX)217 (Table 1). Despite the fact that both structures do not represent physiological reaction intermediates, their apparent differences in metal coordination may suggest a structural plasticity of the ISCU2 assembly site. Clearly, this view needs to be verified by elucidation of the functionally relevant Fe coordination during the various biosynthetic stages of persulfide transfer and reduction within the active core ISC complex. Moreover, it is unclear how Fe and persulfide are handled simultaneously by the assembly site residues to facilitate sulfur transfer.
Here, we use a combination of structural, biochemical and spectroscopic approaches to address the molecular mechanism of sulfur transfer during mitochondrial de novo [2Fe-2S] cluster synthesis. We sought to (1) structurally and biochemically identify persulfide intermediates before and after sulfur transfer, (2) understand the molecular role of FXN, (3) identify the ISCU2 assembly site residues involved in sulfur transfer, and (4) define the dynamic changes in Fe coordination during the various reactions. Our current work provides a molecular understanding of both the mechanism and structural dynamics of sulfur transfer within the core ISC complex. Further, it clarifies the molecular role of FXN in this process, which may be relevant to the development of treatments for Friedreich’s ataxia.
Results
Cryo-EM structures of the core ISC complex reveal heterogeneity at the FXN binding site
To gain structural insights into the persulfide transfer process from Cys381NFS1 to ISCU2, we prepared a (Fe-NIAUX)2 complex that is able to perform the persulfidation reaction, but is unable to generate the [2Fe-2S] cluster because FDX2 was omitted14,18,30 (Fig. 1a). To this end, the purified (NIAU)2 subcomplex was mixed under anaerobic conditions with Fe2+ and FXN (stoichiometry 1:5:1). Cys was added to initiate persulfidation, and the complex was rapidly vitrified under anaerobic conditions for cryo-EM single particle analysis. A structure determined from this sample reached a global resolution of 2.4 Å for the consensus refinement (C2 symmetry applied; Table 2 and Supplementary Figs. 1–4), and showed a typical (NIAUX)2 complex with an extensive dimeric interface in which the two NFS1 subunits and FXN interacted with ISCU2 via conserved residues17 (Fig. 1b). In our structure, the density for FXN was weaker in comparison to the (NIAU)2 components, indicating only partial FXN occupancy. Symmetry expansion and subsequent local 3D classification allowed separation of FXN-bound (77%) (Fe-NIAUX)2 and FXN-lacking (23%) (Fe-NIAU)2 states, which were locally refined to resolutions of 2.4 Å and 2.7 Å, respectively (Supplementary Figs. 1 and 2). The partial occupancy of the FXN binding site is best explained by the relatively weak affinity of FXN for the core ISC complex16,30,33. This fact was also evident from size exclusion chromatography (SEC) of the (Fe-NIAUX)2 complex. FXN showed partial dissociation from the complex, leading to sub-stoichiometric FXN occupancy (Supplementary Fig. 5).
Structure of the Fe-coordinated core ISC complex in the absence of FXN
The overall architecture of the (Fe-NIAU)2 complex was similar to that of the published Zn-containing complex15, yet showed important differences with potential functional implications (Table 2). The ISCU2-bound Fe ion was coordinated by residues Cys381NFS1, Asp71ISCU2, Cys95ISCU2, and likely a water (W1) molecule (Fig. 1c and Table 1). Although persulfidation of NFS1 in the absence of FXN occurred quickly (see below), we did not observe density for a Cys381NFS1-bound persulfide at the ISCU2 assembly site. Apparently, the presence of a metal (Fe or Zn) positions the active Cys381NFS1 residue at the ISCU2 assembly site, whereas without a bound metal the Cys loop of NFS1 is located close to the PLP site15,18. Active-site residue His137ISCU2, although oriented toward the ISCU2 assembly site, is too distant to coordinate the metal (His-Nε—Fe distance of 4.8 Å), distinguishing it from the X-ray structure of (Zn-NIAU)2 (Supplementary Fig. 6 and Table 1). Similarly, Cys69ISCU2 and Cys138ISCU2 are too far for Fe coordination. The fact that these three residues are not participating in Fe ligation may well explain why the densities for Cys69ISCU2 and His137ISCU2 show lower local resolution, indicating higher flexibility.
The Fe-containing conformation raised the important functional question of which Cys residue of ISCU2 may serve as the sulfur acceptor from the Cys loop Cys381NFS1. The latter is closest to the sulfur atoms of Cys95ISCU2 (3.8 Å) and Cys69ISCU2 (4.3 Å), yet rather distant from Cys138ISCU2 (6.5 Å; Table 1, bottom), which might suggest that Cys95ISCU2 or Cys69ISCU2 would be the most likely candidates for persulfide transfer from Cys381NFS1. However, previous biochemical experiments performed under aerobic conditions have suggested Cys69ISCU2 and Cys138ISCU2 as potential persulfide acceptors25. Intriguingly, His137ISCU2, which is hydrogen-bonded to the water ligand of Fe between Cys381NFS1 and Cys138ISCU2, appears to prevent a closer approach of the two Cys residues and to hinder direct sulfur transfer (Supplementary Fig. 6). Therefore, either a substantial conformational change is needed to make Cys138ISCU2 available as a direct sulfur acceptor, or one of the other Cys residues may serve as the intermediate or even final sulfur acceptor. Clearly, additional structural and biochemical data under anaerobic conditions are required to resolve this mechanistic problem.
Structural snapshots of pre- and post-sulfur-transfer states in (Fe-NIAUX)2 complexes
For the 3D class showing clear density for FXN, 3D refinement gave rise to a map with density for two alternate conformations of the catalytic Cys loop of NFS1, i.e., an “inward” conformation in which Cys381NFS1 was oriented toward the PLP cofactor of NFS1 and an “outward” conformation in which Cys381NFS1 faced the ISCU2 assembly site. This arrangement differed from the (Zn-NIAUX)2 cryo-EM structure17, where the Cys loop of NFS1 was positioned in an “intermediate” conformation. Using a multi-reference 3D classification approach (see “Methods”), followed by local refinements, our alternate conformations were separated and resolved at 2.5 Å (inward) and 2.6 Å (outward) (Fig. 1d, e, Table 2 and Supplementary Fig. 1). Most importantly, the inward conformation, henceforth referred to as (Fe-NIAUSX)2, contained additional density at Cys138ISCU2 that could be confidently modeled as a persulfide (Fig. 1d, right). This likely represents the post-sulfur-transfer state, in which the persulfide sulfur of Cys138ISCU2 (CSS138) ligates the Fe center together with Asp71ISCU2, Cys95ISCU2 and likely a water (W1) molecule (Table 1). How Cys138ISCU2 may have become available to accept the sulfur from Cys381NFS1, despite their large distance in the (Fe-NIAU)2 structure (see above), will be presented below. Apparently, after sulfur transfer, the Cys loop of NFS1 has moved away from the ISCU2 assembly site, thereby positioning Cys381NFS1 again close to the PLP site.
In the outward state (hereafter termed (Fe-NSIAUSX)2), we also observed additional densities that could be modeled as persulfides at both Cys381NFS1 (CSS381) and Cys138ISCU2 (CSS138) (Fig. 1e, right). Both persulfides are within Fe coordination distance and thus bind the metal together with Asp71ISCU2 and Cys95ISCU2 (Table 1). The presence of two persulfides was unforeseen. This arrangement could either represent both pre- and post-sulfur-transfer states (due to a second round of persulfidation), or be explained by a mixture of particles in either pre- or post-sulfur transfer states. Generally, the detection of a pre-sulfur-transfer state was unexpected due to its potentially short-lived character, and may indicate an incomplete sulfur transfer reaction or an equilibrium between the two states as suggested by findings below. Collectively, these structural snapshots of intermediate states before and after sulfur transfer from NFS1 to ISCU2 provide a view on the possible mechanism of sulfur trafficking (see below), yet do not readily explain why Cys138ISCU2 is the preferred persulfide acceptor.
FXN binding rearranges the ISCU2 assembly site to facilitate persulfide transfer
Comparing our (Fe-NIAU)2 and the two (Fe-NIAUX)2 structures allowed insights into the dedicated mechanistic role of FXN. Upon FXN binding, the overall architecture of the (Fe-NIAU)2 complex remained virtually unchanged, with the exception of significant local conformational alterations at the ISCU2 assembly site. Both Cys69ISCU2 and His137ISCU2 are shifted substantially away from the ISCU2-bound Fe, as previously observed in the (Zn-NIAUX)2 complex17, and now form contacts with FXN in both the outward- and inward-facing loop structures (Fig. 2a and Table 1). This makes sulfur acceptance by Cys69ISCU2 unlikely. Particularly, His137ISCU2, which appears to block close contact of Cys381NFS1 and Cys138ISCU2 in our (Fe-NIAU)2 structure, becomes inserted into a pocket formed by FXN residues Trp155, Ser157, and Pro163, thereby removing the steric separation of Cys381NFS1 and Cys138ISCU2 (Fig. 2b and Supplementary Movie 1). As a consequence, in the (Fe-NSIAUSX)2 structure, CSS138 is located close to both the Fe (Fe-S distance 2.3 Å) and CSS381 (S-S distance 3.6 Å) (Table 1). Even in this arrangement the distance between the pre-sulfur-transfer persulfide CSS381 and the Cys95ISCU2 sulfur (3.4 Å) is slightly shorter than to CSS138, raising the question of whether Cys95ISCU2 may serve as an intermediate sulfur-accepting moiety before sulfur transfer to Cys138ISCU2. In any case, the structural changes triggered by FXN binding move persulfide CSS381 into a position allowing sulfur transfer to ISCU2. The relatively open coordination sphere at the assembly site of the (Fe-NSIAUSX)2 complex may provide enough space for the sulfur transfer reaction by nucleophilic attack by either Cys95ISCU2 or Cys138ISCU2 sulfurs (Fig. 2c). The observed effects of FXN binding in moving the sulfur-donating and -accepting Cys residues into optimal positions for possible sulfur transfer may explain previous biochemical observations that FXN substantially accelerates the rate of persulfidation within the (NIAU)2 complex25,26.
a Overlay of the (Fe-NIAU)2 and (Fe-NSIAUSX)2 structures shows the structural changes upon FXN binding (Cys381NFS1 not shown for figure clarity). While the constantly Fe2+-coordinating Cys95ISCU2 and Asp71ISCU2 hardly change positions, Cys138ISCU2 moves toward the Fe2+ and now coordinates the metal. Concomitantly, His137ISCU2 and Cys69ISCU2 are shifted away from the ISCU2 assembly site, and His137ISCU2 is now accommodated in a pocket formed by residues including Trp155FXN, Ser157FDX and Pro163FDX (b). c The pre-sulfur transfer state of the (Fe-NSIAUSX)2 structure could conceivably allow for sulfur delivery from Cys381NFS1 to both Cys95ISCU2 (3.4 Å distance) and Cys138ISCU2 (3.6 Å distance).
Only ISCU2 assembly site residues Asp71, Cys95, and Cys138 are essential for sulfur transfer
To answer the question of whether persulfide transfer to Cys138ISCU2 in the (Fe-NIAUSX)2 complex occurs directly or indirectly via Cys95ISCU2, we first analyzed the relative importance of the five ISCU2 assembly site residues for the sulfur transfer reaction by employing a strictly anaerobic Cys alkylation-based gel-shift assay (see reaction scheme in Supplementary Fig. 7a)18. Persulfidation of wild-type (WT) and various assembly-site ISCU2 mutant proteins was initiated by Cys and FXN addition to (Fe-NIAU)2, and the reaction was quenched at different time points by consecutive addition of maleimide-polyethylene glycol11-biotin (MPB, for labeling of native and persulfidated Cys residues) and the denaturing agent SDS. Reductive cleavage of disulfides by TCEP reducing agent decreased the mass of persulfidated ISCU2 as detected by SDS-PAGE. Quantitation showed that already after 10 s, 75% of maximal mono-persulfidation (1x SSH; maximum of 86% reached after 2 min) had occurred (Supplementary Fig. 7b, c). At a much slower rate, a steady increase of doubly persulfidated (2x SSH) ISCU2 was seen, which likely represented a non-physiological side reaction. To avoid this unspecific persulfidation, further reactions were incubated for only 10 s.
ISCU2 variants C69S, H137A (both assembly-site residues) and C130S (a non-conserved Cys residue) yielded 70–101% persulfide formation compared to ISCU2 WT (Fig. 3a and Supplementary Fig. 8a–c), clearly showing that these residues are not essential for persulfidation. Since Fe was essential for this reaction (Fig. 3a), Cys69ISCU2 and His137ISCU2 appeared not to serve as Fe ligands during sulfur transfer, despite the fact that they are involved in the initial Fe binding to ISCU2 (Supplementary Fig. 9)19. This finding is consistent with our (Fe-NIAU)2 structure, in which neither Cys69ISCU2 nor His137ISCU2 coordinate the Fe. A previous study reported that the mouse ISCU-C69S variant abrogates persulfidation, and a function of this residue in sulfur transfer was suggested25. Based on our structural data the effect is more likely explained by the Fe coordination function of this ISCU2 residue (Table 1). For free ISCU2, Fe (or Zn) binding strictly requires Cys69 as a ligand (Supplementary Fig. 9)30, as opposed to NFS1-bound ISCU2 (Fig. 1c–e). Because the previous study had performed ISCU-C69S persulfidation with as-purified, non-metal reconstituted protein25, distorted metal binding may have disabled persulfidation.
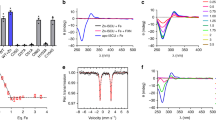
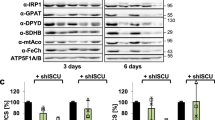
a In vitro persulfidation of ISCU2 by the cysteine desulfurase NFS1 was investigated via a MPB alkylation-based gel-shift assay (Supplementary Fig. 7). In reactions containing 20 µM ISCU2, 20 µM (NIA)2, 40 µM FXN, as well as excess Fe2+ and ascorbate, persulfide formation was initiated by addition of excess Cys. Reactions performed at room temperature (rt) were quenched after 10 s by addition of MPB and SDS. Persulfidation of ISCU2 mutant proteins and control reactions with wild-type (WT) ISCU2 lacking Fe or FXN was quantified by densitometry of the corresponding bands in SDS-PAGE gels (Supplementary Fig. 8), and normalized to control reactions with WT ISCU2 (dotted line). “WT/ no Fe” reactions contained no Fe2+ and ascorbate but 1 mM DTPA to chelate Fe. Error bars indicate the SEM (for WT/ no Fe, WT/ no FXN, C69S, H137A: n = 5; for D71A, C95S, C130S, C138S: n = 4; for others: n = 3; independent experiments). b–e Persulfidation of individual Cys residues of WT ISCU2 was investigated via an iodoTMT-based isobaric alkylation approach (Supplementary Fig. 10). Reactions containing 20 µM ISCU2, 20–40 µM of (NIA)2 and optionally FXN, and either 100 µM Fe2+ (b, d, e) or 500 µM DTPA (no Fe) (c) were quenched with excess iodoTMT (from stocks containing SDS and EDTA) at the indicated time points after addition of 100 µM Cys. Reactions were performed at room temperature (b–d) or at 0 °C (e). Following TCEP-mediated persulfide cleavage, samples were labeled with a second iodoTMT reagent. Trypsin-digested samples were analyzed by nanoHPLC-MS2, and iodoTMT reporter fragment signals of both labels were integrated to quantitate the relative amount of persulfidation of individual Cys residues. Error bars indicate the SEM (n = 3; independent experiments). Source data are provided as a Source Data file.
In contrast to the ISCU2 variants described above, the ISCU2-D71A, -C95S, and -C138S assembly-site variants and Cys double mutant proteins produced only background levels of persulfide (Fig. 3a and Supplementary Fig. 8b, d, e). Since residues Asp71ISCU2 and Cys95ISCU2 serve as metal ligands in all metal-containing (NIAU)2 and (NIAUX)2 structures including those presented here (Table 1), the sulfur transfer defect of these variants can satisfactorily be explained by a lack of Fe coordination during persulfidation. The failure of the ISCU2-C138S variant could be a dual effect, first by lack of metal binding, and second by its proposed role as a persulfide-receiving residue. Collectively, only three of the five essential ISCU2 assembly site residues are required for the sulfur transfer step.
Rapid, specific and direct persulfidation of ISCU2 Cys138 by NFS1
Previous mass spectrometric (MS) analyses showing that Cys69ISCU2 and Cys138ISCU2 may act as persulfide acceptors25 were carried out under aerobic conditions using as-purified, Zn-bound mouse ISCU and reaction times on the order of 3 min (which favors unspecific persulfidation; cf. Supplementary Fig. 7c). Here, we developed a rapid MS-based persulfide detection approach using the Fe-loaded (Fe-NIAUX)2 complex under anaerobic conditions. Isobaric thiol labeling reagents of different masses allowed quantitation of the persulfide levels on each individual ISCU2 Cys residue on a time scale of seconds. Thereby, the assay can also determine transient persulfidation (see scheme in Supplementary Fig. 10a). Briefly, Cys-dependent persulfide formation on (Fe-NIAUX)2 was quenched with a mixture of denaturing SDS detergent, the metal chelator EDTA, and the iodoacetyl tandem mass tag (iodoTMT) labeling reagent. As a control, CD spectroscopy was used to verify that Fe dissociated rapidly and quantitatively from Fe-loaded ISCU2, thereby documenting the efficiency of terminating persulfidation (Supplementary Fig. 10b, c). Excess label was quenched by Cys addition, and after reductive cleavage of persulfides, a second iodoTMT reagent with different mass was used to label the cleavage products (Supplementary Fig. 10a). Persulfide adducts at each Cys-containing peptide were quantified by Nano HPLC-MS2 analysis of trypsin-digested samples. We first analyzed persulfidation of NFS1, which occurred at a high rate and showed specific labeling of only the active-site Cys381NFS1 independently of Fe2+ (Supplementary Fig. 11a, b).
Similarly, ISCU2 was persulfidated with high rate and specificity at Cys138ISCU2, yet unlike NFS1 this happened in a completely Fe-dependent manner (Fig. 3b, c). In the absence of Fe, all Cys residues were labeled only after longer incubation times and rather inefficiently, clearly indicating a non-physiological effect. The rapid persulfidation of Cys138ISCU2 was in good agreement with our MPB-based persulfidation assay (cf. Supplementary Fig. 7b, c). The lower amount of Cys138ISCU2 persulfide that was formed in the iodoTMT assay (ca. 50%) compared to the MPB assay (up to 90%) likely corresponds to the lower stoichiometry of (NIA)2 and FXN over ISCU2 (1:1:1, as opposed to 2:2:1 in the MPB assay), which was chosen to minimize the amount of costly iodoTMT reagent. Interestingly, specific persulfidation of Cys138ISCU2 was also observed in samples lacking FXN, yet at much slower rates, while NFS1 was modified at a normal high rate (Fig. 3d and Supplementary Fig. 11c). This observation fits well to the slow FXN-independent [2Fe-2S] cluster formation in vitro (detected by the CD-based Fe/S cluster biosynthesis assay14,18) indicating that FXN is not essential for [2Fe-2S] cluster assembly, yet substantially increases its rate. Moreover, the non-essential character of FXN is evident in S. cerevisiae, where YFH1 is the only gene of the core ISC system that can be deleted without losing cell viability34,35.
No sulfur transfer intermediate on Cys95ISCU2 was observed, despite the fact that in the (Fe-NSIAUSX)2 cryo-EM structure this residue was as close to the sulfur-donating Cys381NFS1 as Cys138ISCU2 (Figs. 1e and 2c and Table 1). To accumulate a potential Cys95ISCU2 intermediate, we used the ISCU2-C138S variant and performed the persulfidation reaction. No labeling of Cys95ISCU2 (or other ISCU2 Cys residues) above background was detectable (Fig. 3a and Supplementary Fig. 11e). As an internal control, Cys381NFS1 was persulfidated as in wild-type controls. We finally examined whether the potential Cys95ISCU2 intermediate was converted too rapidly for the Cys138ISCU2 persulfide to be detectable by our experimental setup. We therefore performed the iodoTMT assay at 0 °C to slow down the reaction, and we measured the first data point already after <1 s of persulfidation ((NIA)2:FXN:ISCU2 = 2:2:1). Under these conditions, persulfidation of Cys138ISCU2 was much slower and reached the levels obtained at room temperature only after 5 min, while Cys381NFS1 persulfidation was still virtually complete after <1 s (Fig. 3e and Supplementary Fig. 11d). Clearly, no other ISCU2 Cys residues were modified above background, making persulfidation intermediates including that of Cys95ISCU2 unlikely. We conclude that Cys138ISCU2 is the rapid, specific, and direct sulfur acceptor from Cys381NFS1 upon substantial FXN-facilitated structural changes of the ISCU2 assembly site to properly position the sulfur-donating and -accepting residues around the Fe ion.
Mössbauer spectroscopy identifies a FXN- and persulfidation-shifted equilibrium of two ISCU2-bound Fe2+ species
The findings above provide evidence for a highly dynamic ISCU2 assembly site that undergoes multiple conformational changes during Cys381NFS1 persulfide binding to ISCU2-bound Fe, FXN-mediated (Fe-NIAU)2 rearrangement, and subsequent sulfur transfer to Cys138ISCU2. We used Mössbauer spectroscopy to further elucidate the dynamic alterations in Fe coordination at the ISCU2 assembly site. With the exception of Fe binding to free murine ISCU19, the ISCU2 Fe coordination and its dynamics during the intermediate steps of de novo [2Fe-2S] cluster formation have not been analyzed. We recorded Mössbauer spectra for different biochemically generated reaction intermediates, in particular the 57Fe-loaded complexes (Fe-NIAU)2, (Fe-NIAUX)2 (no Cys added), and persulfidated (Fe-NIAUSX)2 (Fig. 4 and Table 3). As controls we used 57Fe-loaded ISCU2 (Fe-U)19 (Fig. 4a), (NIAX)2 plus added 57Fe (lacking ISCU2; Fig. 4e), and the final product, i.e., mature [2Fe-2S] cluster-loaded ISCU2 (Fig. 4f).
Spectra of the indicated complexes reconstituted with 1 eq. (NH4)257Fe(SO4)2 per ISCU2. For the detailed sample composition, see Supplementary Table 2a. U: ISCU2; X: FXN; F: FDX2. a Mössbauer spectrum recorded for 57Fe-loaded ISCU2 (Fe-U). b–d Mössbauer spectra of samples containing (NIA)2 and/or FXN in excess over ISCU2. Cysteine was added as the last component in (d) to initiate ISCU2 persulfidation. e Mössbauer spectrum in the presence of (NIAX)2, lacking ISCU2. f Mössbauer spectrum of enzymatically reconstituted [2Fe-2S]-ISCU2 from a reaction containing (NIA)2, FXN, FDX2 and FDXR in catalytic amounts, as well as Cys and NADPH. Each Mössbauer spectrum was recorded at 77 K and zero applied external field. The red lines display the best fit of the data using the components shown as black lines with parameters given in Table 3. Mössbauer data points are presented as relative transmission per velocity channel derived from detector counts (see “Methods”). Error bars indicate ±SD. g Scheme depicting the shift in equilibrium of ISCU2-bound Fe-species (components 1 and 2) by FXN binding and persulfidation (+Cys) during consecutive stages of [2Fe-2S] cluster biosynthesis on ISCU2 (Fe components 3 and 4) identified by Mössbauer spectroscopy.
Various 57Fe species were identified by Mössbauer spectroscopy across the different samples, and were grouped according to their isomer shifts δ. Two distinct Fe2+ species (components 1 and 2) were not present in the 57Fe + (NIAX)2 sample, hence being specific for ISCU2 (Fig. 4a–e and Table 3). Conversely, the Fe3+/Fe2+ species observed in (NIAX)2 (represented by component A with δ = 0.30 mms−1 and component B with δ = 1.24 mms−1) were not related to ISCU2. Component A (but not B) was present in the various (NIAU)2 complexes, yet could be minimized by optimizing the ionic conditions for assembly of these complexes (compare Fig. 4 and Table 3 with Supplementary Fig. 12 and Supplementary Table 1; conditions in Supplementary Table 2a, b). As a further control, we analyzed ISCU2 after enzymatic [2Fe-2S] cluster reconstitution with catalytic amounts of (NIA)2, FXN, and the electron transfer chain NADPH-FDX2-FDXR14. Two distinct 57Fe/S cluster-related components (3 and 4, Fig. 4f and Table 3) were detected with parameters comparable to those published for mouse [2Fe-2S]-ISCU30, yet were clearly distinct from the ISCU2-related components 1 and 2, showing the specificity of the latter Fe species for (NIAU)2 samples.
The ISCU2-specific component 1 (isomer shift δ = 0.89, quadrupole splitting ΔEQ = 3.4–3.6 mms−1; Table 3) was the only species present in Fe-U and indicated high-spin Fe2+ ligated by 1‒2 S and 2‒3 N/O ligands. This finding is consistent with the established tetrahedral metal coordination by Cys69ISCU2, Asp71ISCU2, Cys95ISCU2, and His137ISCU2 in the NMR structure of Zn-bound murine ISCU (PDB 1WFZ) (Fig. 4a and Supplementary Fig. 9)19,30. Strikingly, the amount of component 1 in the various (NIAU)2 complexes successively decreased along the Fe/S biosynthetic pathway. Concomitantly, component 2 (isomer shift δ = 1.10–1.14, quadrupole splitting ΔEQ = 3.15–3.70 mms−1; Table 3) amounts increased, particularly upon (NIA)2 binding to Fe-U from 0 to 38%, and further upon FXN addition ((Fe-NIAUX)2) from 38 to 66% (Fig. 4a–d, g and Table 3). After persulfidation upon Cys addition ((Fe-NIAUSX)2 intermediate), component 2 further increased from 66 to 83%. Notably, the isomer shift slightly decreased to δ = 1.10 mms−1, and the quadrupole splitting significantly increased to 3.70 mms−1. Whether the latter increase is due to a rearrangement of the Fe ligand environment (for instance, dissociation of Cys381NFS1 from the Fe and binding of Cys138ISCU2) or an effect induced by persulfidation remains unclear. These findings suggested that (NIAU)2 complex formation, FXN binding and persulfidation successively led to a transition toward component 2, thereby indicating substantial changes in Fe coordination at the ISCU2 assembly site during biosynthesis, with (at least) two distinct Fe-coordination modes (Fig. 4g and Table 3).
Component 1 represented the major ISCU2-specific Fe species (55%) in (Fe-NIAU)2 (Table 3). When compared to the Fe coordination in Fe-U, our (Fe-NIAU)2 cryo-EM structure showed Cys69ISCU2 being replaced by Cys381NFS1, and His137ISCU2 being exchanged for a water molecule (Fig. 1c). This substantial rearrangement upon Fe-U binding to (NIA)2 may, however, not be detectable by Mössbauer spectroscopy, because the respective numbers of S- and N/O-ligands remained unchanged. Interestingly, the movement of His137ISCU2 away from the Fe coordination site may partially mimic the alterations observed upon FXN binding, and generally shows the high structural dynamics of the ISCU2 assembly site. Its plasticity is also evident from comparison to the crystal structure of (Zn-NIAU)2, where metal ligation via His137ISCU2 was maintained, in addition to coordination by Cys381NFS1, Asp71ISCU2, Cys95ISCU2, i.e., similar as in Fe-U but with an exchange of Cys69ISCU2 for Cys381NFS1 (Table 1 and Supplementary Fig. 6).
Component 2 with δ = 1.10–1.14 mms−1 constituted the major fraction of ISCU2-specific Fe-species in both FXN-containing (Fe-NIAUX)2 and (Fe-NIAUSX)2 complexes (66% and 83%, respectively; Fig. 4c, d and Table 3). The tetrahedral (2S-1N-1O) coordinated Fe2+ site of Fe-U represented by component 1 shows δ = 0.89 mms−1. A possible bidentate binding of Asp71 increases the coordination number by one. This would lead to an increase in the isomer shift by ~0.1 mms−1 36,37 leading to ~1 mms−1 for a (2S-1N-2O) coordination. Changing His137ISCU2 to a water ligand as observed by our cryo-EM data would not affect the isomer shift much. Thus, Mössbauer spectroscopy indicates a bidentate binding of Asp71 which could not be assigned with certainty from the obtained resolution of the cryo-EM structures (Fig. 1). Moreover, an octahedral (2S-4O) coordination of Fe2+ with a further N/O ligand (e.g., water) would lead to a further increase in the isomer shift by ~0.1 mms−1 resulting in 1.1 mms−1, close to the values experimentally observed for component 2 in the ISC complexes, in particular for (Fe-NIAUSX)2. However, we note that an octahedral coordination was not observed in the (Fe-NIAUSX)2 cryo-EM structure, where bidentate Asp71ISCU2, Cys95ISCU2, CSS138ISCU2, and one water (i.e., 2S-3O) have been assigned as Fe ligands (Fig. 1d). Notably, the line width Γ of component 2 in Mössbauer analyses (Table 3) is significantly decreased (by 40%) in (Fe-NIAUX)2 and (Fe-NIAUSX)2 compared to (Fe-NIAU)2 samples, suggesting that FXN binding structurally rigidifies this Fe species.
The presence of a substantial amount (31%) of component 1 identified by our Mössbauer analysis of (Fe-NIAUX)2 may be related to the dynamic association and dissociation of FXN. Such behavior is suggested by our work (Supplementary Fig. 5), NMR data showing only 65% of FXN being bound to (Zn-NIAU)2 under stoichiometric conditions30, and recent biochemical studies33. Hence, in our (Fe-NIAUX)2 Mössbauer sample, the fraction of non-FXN-bound (Fe-NIAU)2 complex likely accounts for the Mössbauer parameters of component 1. Collectively, these findings suggest an equilibrium between (at least) two distinct core ISC complex states with different Fe coordination (Fig. 4g and Table 3). The first state (component 1) predominates in the (Fe-NIAU)2 sample, where Fe is ligated by Cys381NFS1, Asp71ISCU2, Cys95ISCU2 and a water molecule (as in our (Fe-NIAU)2 structure) in a tetrahedral fashion (stage 2 in Fig. 5). In contrast, component 2 prevails upon FXN binding, e.g., in (Fe-NIAUSX)2. The fact that the isomer shift of component 2 by Mössbauer data is not fully consistent with the observed Fe coordination in our (Fe-NIAUSX)2 structure may be due to a malleable ISCU2 assembly site, the presence of additional water molecules not resolved by cryo-EM, and/or the drastically different experimental conditions required for cryo-EM and Mössbauer experiments including pH differences (see “Methods”). At any rate, the equilibrium between the two different Fe binding states can be shifted by FXN binding and persulfidation suggesting physiological relevance.
Top: Cartoon of the ISC complexes analyzed in this study. Bottom: Coordination of Fe by the five residues of the ISCU2 assembly site, Cys381NFS1 (orange), and water (blue) at different stages of the biosynthetic pathway. (1) Free ISCU2 (U) tetrahedrally coordinates Fe2+ via its residues Cys69, Asp71, Cys95 and His137. (2) Binding of Fe-U to (NIA)2 leads to an Fe coordination in which Cys69ISCU2 is replaced by Cys381NFS1 and His137 by water. (3) Cys addition creates the Cys381NFS1 persulfide which also can coordinate the Fe. In this state, steric hindrance by His137ISCU2 and the distal location of Cys138ISCU2 preclude persulfide transfer from Cys381NFS1 to Cys138ISCU2. (4) Dynamic binding of FXN to (Fe-NSIAU)2 promotes His137ISCU2 to move away from the assembly site, while Cys138ISCU2 moves closer to the Fe and Cys381-SNFS1. (5) Thereby, Cys138ISCU2 coordinates the Fe as a prerequisite for direct persulfide acceptance from Cys381NFS1 to form the (Fe-NIAUSX)2 intermediate. Despite the fact that Cys95ISCU2 is within a similar distance from the Cys381NFS1 persulfide as Cys138ISCU2, no persulfidation is observed. (A) As a rather unlikely alternative (see Discussion), the preformed (Fe-NIAUX)2 complex (similar to the (Zn-NIAUX)2 structure17) may coordinate the Fe by Asp71ISCU2, Cys95ISCU2, Cys138ISCU2, and a water and after Cys addition and Cys381NFS1 persulfidation is ready for sulfur transfer. Finally, FDX2-dependent reduction of the persulfide to inorganic sulfide and transient dimerization of two [1Fe-1S]-bound ISCU2 molecules enables [2Fe-2S] cluster formation18.
Discussion
Fe/S protein biogenesis in eukaryotes starts with the de novo synthesis of [2Fe-2S] clusters inside mitochondria, and involves several steps: (1) the sulfur release from free Cys by the cysteine desulfurase complex NFS1-ISD11-ACP1, (2) FXN-stimulated transfer of the NFS1-bound persulfide to the scaffold protein ISCU2 which has Fe2+ bound, (3) the FDX2-dependent reduction of the persulfide to sulfide, and finally, (4) the formation of the [2Fe-2S] cluster by interaction of two [1Fe-1S]-containing ISCU2 on different core ISC complexes14,15,18,25,26,30. To date, many of the molecular mechanistic details of how Fe and sulfur are handled by the ISC proteins generating the [2Fe-2S] cluster are poorly understood. We have used a combination of structural biology, biochemistry, and Mössbauer spectroscopy under anaerobic conditions to structurally and biochemically characterize various intermediates of this multi-stage process including those containing the transient persulfides. Thereby, we could define the pathway of the persulfide sulfur from Cys381NFS1 to the receiving Cys138ISCU2 and elucidate the conformational rearrangements of the five essential ISCU2 active-site residues in this sulfur-trafficking process and in ISCU2-bound Fe coordination.
The model depicted in Fig. 5 presents the various conformational states of the core ISC complex defined in this work. The entire process starts with the assembly of Fe-bound ISCU2 and (NIA)2 to form the (Fe-NIAU)2 complex (Fig. 5, stages 1 and 2). According to Mössbauer- and CD-spectroscopic analyses, free ISCU2 binds ferrous Fe in a tetrahedral coordination via Cys69, Asp71, Cys95 and His137 similar to mouse Fe-U30 (PDB 1WFZ). Mössbauer data suggest that a similar Fe species is present in a major fraction of the (Fe-NIAU)2 sample, yet structural data for both (Fe-NIAU)2 (this work) and (Zn-NIAU)215 indicate that in these complexes Cys69ISCU2 is replaced by Cys381NFS1 and His137ISCU2 by a water (Fig. 5, stage 2). Such a conformational change is, however, Mössbauer-silent (component 1 in Table 3). The Mössbauer technique additionally revealed the presence of another (minor) species with distinct parameters (component 2). Addition of both Cys and FXN substantially increased the amount of this component 2 indicating an equilibrium between (at least) two states (Fig. 5, stages 3 and 4, 5). As FXN does not stably and quantitatively bind to (NIAU)2 due to low affinity, the residual amount of component 1 likely represents (Fe-NIAU)2 lacking FXN, while the FXN-containing complexes (i.e., (Fe-NSIAUX)2 sulfur transfer intermediate of stage 4 and (Fe-NIAUSX)2 of stage 5) may feature component 2. In vivo, it is well possible that the (NIA)2 complex already contains a persulfidated NFS1 when binding to Fe-ISCU2, thus creating stage 3 without a transient stage 2. Nevertheless, our experiments suggest that also stage 2, upon addition of Cys, can be readily converted to stage 3, possibly due to the high dynamics of the Cys-loop of NFS1.
The cryo-EM structures suggest a highly conformationally dynamic ISCU2 assembly site upon transition from (Fe-NIAU)2 to persulfidated and FXN-bound states involving dissociation of Cys381NFS1 from the Fe and binding of Cys138ISCU2 (Fig. 5, stages 3–5). Since our persulfide-containing cryo-EM structures (Fe-NSIAUSX)2 and (Fe-NIAUSX)2 likely represent pre- and post-sulfur transfer states, the conformational flexibility at the ISCU2 assembly site may be central for bringing the sulfur-donating Cys381NFS1 and sulfur-accepting Cys138ISCU2 in close vicinity (from 6.5 Å in (NIAU)2 to 3.6 Å in (Fe-NSIAUSX)2; Table 1) to facilitate sulfur transfer. This distance may be sufficient for direct transfer, considering the S–S bond length of a persulfide of 2.0 Å38. FXN, even though not essential, apparently shifts the equilibrium of these states and thereby accelerates the structural transitions toward the configuration that is optimal for sulfur transfer. This action may be represented by the (Fe-NSIAUSX)2 structure. It is not clear whether the unexpected detection of two persulfide sulfurs in this structure represents a true reaction intermediate or is explained by the presence of particles in both pre- and post-sulfur transfer states that were averaged during cryo-EM analysis. However, the vicinity of Cys381NFS1 and Cys138ISCU2 at the coordinating Fe provides a chemically favorable orientation for facile sulfur exchange by nucleophilic attack.
Since only the conformation stabilized by FXN but not that of (Fe-NIAU)2 (or component 1) sterically allows the persulfide transfer, this structural transition likely defines the molecular mechanistic role of FXN in the process. Precisely, FXN interaction with persulfidated (Fe-NSIAU)2 leads to movements of (1) His137ISCU2 into a (hydrophobic) pocket formed by Pro163FXN and Trp155FXN and (2) Cys69ISCU2 further away from the assembly site, thereby (3) closing ranks between Fe and Cys138ISCU2 (Fig. 2a and Supplementary Movie 1). The resulting closer vicinity of sulfur-donating Cys381NFS1 and sulfur-accepting Cys138ISCU2 and the exit of sterically hindering His137ISCU2 from the assembly site facilitates the sulfur transfer reaction. Interestingly, this conformational transition can also occur, yet at much slower rate, without FXN, as seen in our persulfidation experiments (Fig. 3d), suggesting that FXN simply shifts the equilibrium between these states. Structurally, a similar transition is observed in the persulfide-free and Zn-containing (NIAUX)2 complex, despite the presence of non-physiological Zn which supports sulfur transfer but not [2Fe-2S] cluster formation15,17,30. To finish the biosynthetic process, persulfide reduction by FDX2 enables the formation of mature holo-ISCU2 (Fig. 5; [2Fe-2S]-U)18, 20.
For detailed analysis of the persulfidation reaction, we have developed an assay employing iodoTMT labels with different masses to quantitatively determine persulfidation of all involved Cys residues in parallel under anaerobic conditions. Using this assay, we show a direct, rapid, and specific persulfide transfer from Cys381NFS1 to Cys138ISCU2. By slowing down the reaction at 0 °C and by using short reaction times (1 s), we could make other Cys persulfides as intermediates unlikely, even though sterically Cys95ISCU2 could serve as a sulfur acceptor (see below). Our persulfidation experiments further resolved the potential roles of the other active-site residues in sulfur transfer. While Asp71ISCU2 and Cys95ISCU2 are essential for persulfidation because of their continuous Fe coordination, surprisingly both Cys69ISCU2 and His137ISCU2 were not. Since the latter two residues coordinate the Fe in free ISCU2, Fe must be able to enter the process also at a later stage, e.g., at the (NIAU)2 complex, thus making a free Fe-U complex dispensable. The non-essential role of Cys69ISCU2 for persulfidation is fully consistent with its structural distance to the Cys381NFS1 persulfide after (Fe-NSIAU)2 formation and later steps. Together, these results suggest a non-essential role of Cys69ISCU2 and His137ISCU2 during persulfide transfer.
Concerning the mechanism of persulfidation, our structural data showing similar distances of Cys381NFS1 persulfide to both Cys95ISCU2 and Cys138ISCU2 (Table 1) did not readily exclude a sulfur relay to Cys95ISCU2. With a CSS bond angle of around 103° 39, both Cys95ISCU2 and Cys138ISCU2 would be similarly positioned to accept the sulfur from persulfidated Cys381NFS1. To detect such a possible (intermediate) species, we adapted our assay conditions so that the fast time course of Cys138ISCU2 persulfidation could be temporally resolved. In these experiments at 0 °C and short reaction times, exclusive persulfidation of Cys138ISCU2 was observed, making a Cys95ISCU2 intermediate unlikely (Fig. 3e). Consistently, no persulfidation of other Cys residues was detectable for the ISCU2-C138S variant. These results raise the interesting question of why Cys138ISCU2 is preferred over Cys95ISCU2. One possible explanation could be that Cys95ISCU2, together with Asp71ISCU2, continuously coordinates the Fe throughout the pathway, and thus may not be flexible enough to accept an extra atom. In contrast, Cys138ISCU2 shows comparatively high conformational flexibility, e.g., during the movement toward the Fe upon the FXN-accelerated conformational change of ISCU2 (Supplementary Movie 1). Interestingly, persulfide transfer from NFS1 to ISCU2 observed here (Fig. 3) appears to be much faster than previously assumed based on measurements from 10 s to several hours25,26,30. This high rate was even exceeded by NFS1 persulfidation which at 0 °C could not kinetically be resolved (Supplementary Fig. 11d) and may occur at similar fast rates as the formation of the PLP-intermediates26. Hence, in vivo, (NIA)2 may be persulfidated before binding its partner proteins, and persulfidation of Cys381NFS1 only after (Fe-NIAUX)2 complex assembly seems kinetically unlikely (Fig. 5, stage A). Overall, our study suggests that the persulfidation steps are not rate-limiting for the overall process of [2Fe-2S] cluster synthesis.
Our work delineates a potential mechanism of persulfide transfer within the core ISC complex and defines the molecular role of FXN in this process. As indicated by our study, the molecular transitions facilitated by FXN occur at low efficiency in the absence of this protein, and may serve as the basis for the development of small molecules mimicking FXN function for therapeutic regimes of Friedreich’s ataxia. A further important objective for future investigations is the structural characterization of intermediates upon and after FDX2 reduction of (Fe-NIAUSX)2 to yield a putatively [1Fe-1S]-containing complex. The presumably short-lived nature of such intermediates may render their characterization rather challenging.
Methods
Expression and purification of recombinant proteins
An overview of the plasmid constructs used for recombinant protein expression is given in Supplementary Table 3. FDXR22 and all other proteins20 were overproduced and purified at 4 °C or on ice as described hereinafter. His-tagged constructs were purified by His-affinity chromatography and nontagged constructs by anion exchange chromatography (AEC). Pellets were resuspended in His or AEC buffers (respective protein-specific buffer composition given in Supplementary Table 4). Protease inhibitor (cOmplete Protease Inhibitor Cocktail), lysozyme and DNase I (and PLP for NIA purification) were added to the suspension. Cells were lysed by sonication and the lysate cleared by centrifugation. The cell extract was loaded onto either a His-binding (Ni-NTA-Agarose) or an anion exchange column (Source 30Q) pre-equilibrated with His or AEC buffers. His-binding columns were washed using His buffer plus 10 mM ((NIA)2) or 70 mM (ISCU2, FXN) imidazole and proteins eluted in His buffer plus 250 mM imidazole. Anion exchange columns were washed with AEC buffer, and proteins were eluted using a gradient of 0–100% AEC elution buffer (Supplementary Table 4). Eluates were concentrated to a volume of ca. 1.5 ml. FXN was treated with recombinant TEV protease to remove the N-terminal His-tag. Proteins were transferred to an anaerobic chamber and (NIA)2, ISCU2 and FXN incubated for 60 min with additives (Supplementary Table 5) to remove metals and/or polysulfanes/sulfides. Protein samples were subjected to anaerobic size exclusion chromatography using HiLoad 16/600 Superdex columns pre-equilibrated with degassed SEC buffer (Supplementary Table 4). Desired fractions were combined, concentrated to roughly 1 ml, aliquoted in air-tight glass vials and checked for purity by SDS–PAGE. Protein concentration was determined via the Bradford assay. A correction factor for the resulting concentrations was determined for (NIA)2, ISCU2, FXN, and FDX2 by quantitative amino acid analysis (Leibniz-Institut für Analytische Wissenschaften).
Purification of (NIA)2 typically consisted of His affinity chromatography followed by SEC. For cryo-EM sample preparation, an AEC step was added after His purification to achieve the highest possible purity. ISCU2-D71A mutant protein showed aggregate formation and Fe/S cluster binding, when purified under the same conditions employed for wild-type ISCU2 and other ISCU2 variants. To maintain ISCU2-D71A in a monomeric apoform, buffer and additive compositions were optimized as detailed in Supplementary Tables 4 and 5.
Cryo-EM sample preparation
Protein preparations of high purity were used for cryo-EM experiments (85% purity of (NIA)2 and 92–99% purity of all other proteins, based on densitometry of SDS-PAGE bands). Cryo-EM sample preparation was performed at room temperature under oxygen-free conditions in an anaerobic chamber (Coy Laboratories) using anaerobic buffers. (NIA)2 and ISCU2 were mixed in a 1:2 molar ratio and incubated for 5 min. The formed (NIAU)2 complex was purified by SEC using a Superdex 200 increase 10/300 GL column (GE Healthcare) equilibrated with cryo-EM buffer (35 mM Tris/HCl pH 7.4, 150 mM NaCl) and concentrated using a 100 kDa centrifugal filter (10 min, 4000 × g).
UltrAuFoil 0.6/1 300 mesh gold grids (Quantifoil) were made hydrophilic by plasma cleaning using a NanoClean Model 1070 (Fischione Instruments) running for 2 min at 40% power with an Ar/O2 gas mixture (90:10 ratio) prior to sample vitrification. 27.5 µM (NIA)2, 275 µM Na-ascorbate, 275 µM FeCl2 and 55 µM FXN were mixed in cryo-EM buffer and the persulfidation reaction was initiated by addition of 300 µM cysteine. The sample (2.7 µl) was mixed with 0.3 µl fluorinated fos-choline-8 (Anatrace; final concentration 1.5 mM), and applied to the freshly plasma-cleaned grid. The sample was vitrified in liquid ethane using a Vitrobot Mark IV (Thermo Scientific) inside the anaerobic chamber. The grid was blotted for 4 s with a nominal blot force of 4 at 4 °C and 100% humidity using grade 595 filter paper (Whatman Products). The time between adding cysteine and sample vitrification was less than 1 min.
Data acquisition and image processing
Cryo-EM data were acquired in Energy-Filtered Transmission Electron Microscopy (EFTEM) mode using a Titan Krios G3i electron microscope (Thermo Scientific), equipped with a BioQuantum energy filter (Gatan) and operated at 300 kV. Electron-optical alignments were adjusted with EPU (Thermo Scientific). In total, 5410 dose-fractionated movies were recorded with a K3 direct electron detector (Gatan) in electron counting mode, using aberration-free image shift (AFIS) automation strategies of EPU, with a total dose of ~62 e‒/Å2 spread equally over 60 frames at a nominal magnification of ×105,000, corresponding to a calibrated pixel size of 0.837 Å.
Data quality was monitored during data acquisition using cryoSPARC live40. In total, 4298 micrographs with a CTF estimation better than 5 Å and astigmatism less than 500 Å were used for initial data processing in cryoSPARC as described hereafter (Supplementary Fig. 1a). Particles were picked with the blob-picker implementation and subjected to 2D classification. 2D classes showing the (NIAUX)2 complex were selected and used for template picking. Initially, 2,189,680 particles were picked, extracted with a box size of 256 × 256 pixels and classified in 2D. Based on the appearance of 2D class averages, a subset of 1,075,763 particles was subjected to a homogeneous refinement in cryoSPARC resulting in a global resolution of 2.9 Å. Particle coordinates from this refinement were then converted into a star file using Pyem41 for further data processing in RELION-3.142.
Movies were sorted into optics groups according to their EPU AFIS metadata (https://github.com/DustinMorado/EPU_group_AFIS). Full-frame motion correction was performed with MotionCor243 using 3 × 3 patches and the contrast transfer function (CTF) was calculated using CTFFind4.144. Only micrographs with a CTF estimated better than 5 Å were used (4508 micrographs). Particle coordinates initially obtained in cryoSPARC live were extracted in RELION-3.1 with a box size of 256 × 256 pixels and 4x down sampling (1,003,002 particles) and subjected to 2D classification. A subset of 483,881 particles was selected on the basis of high-quality 2D classes and used to generate an initial model. A 3D classification (25 iterations, T = 4) of all 1,003,002 particles with this initial model as 3D reference (60 Å low pass filtered) resulted in a major class containing 696,662 particles, which were used for further 3D refinements. After re-extraction of the refined particles using 1.2x down sampling, a 3D refinement applying C2 symmetry gave a global resolution of 2.86 Å (0.143 FSC criterion). Two rounds of CTF refinement, correcting for higher order aberrations, anisotropic magnification and per-particle defocus, with every optics group being refined independently45, followed by Bayesian particle polishing, further improved the overall resolution to 2.41 Å. Particles were C2 symmetry expanded and local 3D classification without alignment (T = 4, 50 iterations) was performed with a mask focusing on the FXN-binding region (Supplementary Fig. 1a). This separated the dataset into 2 classes—FXN-bound (77.1%) and non-FXN-bound (22.9%), which were refined by local 3D refinements using a solvent mask yielding resolutions of 2.41 Å and 2.71 Å, respectively.
The non-FXN-bound subset was further separated by a focused 3D classification (40 iterations, T = 16, without alignment) applying a mask on ISCU2. The major class contained 269,101 particles, which were subjected to a local 3D refinement (2.75 Å final resolution). Map interpretability could be further improved by density modification46 and local anisotropic sharpening47 using the Phenix software package48.
The FXN-bound map showed density consistent with a persulfide on Cys138ISCU2 and revealed two alternate conformations of the flexible Cys loop of NFS1. To separate these loop conformations a multi-reference 3D classification approach was used. References were generated from a model based on PDB 6NZU, which was real-space refined against the FXN-bound cryo-EM map followed by manual fitting of two versions of the Cys loop of NFS1 (residues 377-389) into density corresponding to the two alternate conformations using Coot-0.949 (Supplementary Fig. 1a). 3D volumes, low-pass filtered to 8 Å, were generated from these models using the molmap command in ChimeraX50. Particle subtraction with a mask focusing on the Cys loop region was performed and multi-reference 3D classification (T = 64, 25 iterations, without alignment) was carried out using 10 Å low-pass filtered 3D references to separate the alternate loop conformations. Particles from both classes were subjected to another round of multi-reference classification using the same parameters and references. Particles assigned to the major classes were selected, reverted to the original (non-signal subtracted) particles and subjected to a local 3D refinement using the initial model as a 3D reference. Conformation 1 (inward), resolved to a global resolution of 2.49 Å, shows Cys381NFS1 near the PLP cofactor, while in conformation 2 (outward), resolved to 2.58 Å, Cys381NFS1 is close to the ISCU2-bound Fe. Density modification46 and local anisotropic sharpening47 in Phenix48 further improved map clarity.
For the electron dose analysis (Supplementary Fig. 4), movie frames 1–2 (dose 2.07 e− Å−2), 1–3 (dose 3.1 e− Å−2), 1–5 (dose 5.17 e− Å−2), 1–10 (dose 10.33 e− Å−2), 1–40 (dose 41.33 e− Å−2) and 1–60 (dose 62 e− Å−2) were combined during Bayesian particle polishing in RELION-3.1. Half-maps were reconstructed with relion_reconstruct and post-processed using the relion_postprocess command with automatically estimated B-factor.
Model building and refinement
The cryo-EM structure of human (Zn-NIAUX)2 (PDB 6NZU)17 was used as a template for model building and refinement. The model was rigid-body fitted into the respective cryo-EM map in ChimeraX followed by manual refinement in Coot-0.949. Persulfide sulfur (S-mercaptocysteine; CSS) and Fe2+ (FE2) were added to the model in Coot-0.9. Metal restraints were generated from the manually refined model using ReadySet! in Phenix, and adjusted according to published high-resolution structures. Water molecules were built manually in agreement with the density and the hydrogen-bonding distances. The final models were real-space refined against the respective map using phenix.real_space_refine and validated by phenix.validation_cryoem within Phenix48 (Table 2). Figures and the movie of models and maps were prepared with ChimeraX50.
Enzyme assays, CD spectroscopy and analytical SEC
All persulfide transfer and reconstitution assays, analytical SEC (aSEC) as well as sample preparation for circular-dichroism (CD) spectroscopy were performed under oxygen-free conditions in an anaerobic chamber (Coy Laboratory products) using anaerobic buffers. aSEC was performed on an UltiMate 3000 UHPLC system (Thermo Fisher Diagnostics) using a Superdex 200 increase 10/300 GL column (GE Healthcare) equilibrated with anaerobic SEC buffer (Supplementary Table 4). CD spectroscopy was performed anaerobically at room temperature in a quartz cuvette with 4 mm path length sealed with an air-tight rubber capping on a Jasco J-815 (JASCO Inc.).
Reconstitution of Fe-ISCU2
Fifty µM ISCU2 was incubated in reconstitution buffer (35 mM Tris/HCl pH 8.0, 150 mM NaCl, 5% w/v glycerol, 2 mM TCEP) for 5 min at room temperature. CD spectra (250–350 nm) were recorded before and after addition of 2 eq. FeCl2 per ISCU2. The apo-ISCU2 spectrum was subtracted from the Fe-ISCU2 spectrum to remove signal contributions from aromatic amino acids51.
Persulfidation assay with MPB-based persulfide quantification
NFS1-dependent persulfidation of ISCU2 was analyzed based on Cys alkylation25. Standard reactions contained 100 µM FeCl2, 100 µM Na-ascorbate, 20 µM ISCU2, 40 µM FXN and 20 µM (NIA)2 in MPB buffer (35 mM Tris/HCl pH 7.4, 150 mM NaCl, 5% w/v glycerol). After incubation for 3 min at room temperature, 200 µM cysteine was added to initiate persulfide formation, and the reaction was quenched after 10 s by mixing with 2–8 eq. (per total thiol concentration) of maleimide-polyethylenglycol11-biotin (MPB) dissolved in DMSO. Directly afterwards, 1% w/v SDS was added, and samples were mixed vigorously and incubated for 10 min. Aliquots containing 1 µg ISCU2 were mixed with SDS-PAGE sample buffer containing 50 mM TCEP and incubated at room temperature for 15 min. The reduced samples were analyzed either by standard SDS-PAGE using 15% w/v acrylamide gels (for ISCU2 WT and variants with at least three Cys residues remaining) or by tricine SDS-PAGE using 16% w/v acrylamide gels (ISCU2 variants with fewer than three Cys residues). Gels were stained with InstantBlue (Expedeon), and ISCU2 bands were quantified by densitometry using Image Studio Lite version 5.2 to calculate the percentage of persulfidated ISCU2-species.
Persulfidation assay with iodoTMT-based persulfide quantification
Persulfidation of individual Cys residues of NFS1 and ISCU2 was quantified via mass spectrometric analysis of persulfide transfer reactions labeled with thiol-specific iodoacetyl Tandem Mass Tag™ (iodoTMT) reagents. Stocks of 22.1–30.0 mM iodoTMT dissolved in 50% v/v methanol, 5–8% w/v SDS, 3–5 mM Na-EDTA and 100 mM Tris pH 8.5 were prepared directly before the experiment. Reactions contained 100–150 µM FeCl2 (5 eq. over ISCU2), 20–30 µM ISCU2, 20–40 µM FXN and 15–40 µM NIA in iodoTMT buffer (35 mM Tris/HCl pH 8.0, 150 mM NaCl, 5% w/v glycerol). Following incubation for 3 min at room temperature, 75–200 µM cysteine (2.5 eq. over (NIA)2) was added to initiate persulfide formation. Reactions were quenched at time points up to 320 s with 4.4–15.0 mM iodoTMT (10.3–39.5 eq. over total thiols) and incubated for 10 min. Samples were incubated with 6.2–22.0 mM cysteine (1.5 eq. over remaining iodoTMT) for 10 min to quench the labeling reagent, and persulfides were cleaved by incubation with 5 mM TCEP for 10 min. Cys residues were subsequently labeled for 10 min with 4.4–15 mM (2.0 eq. over remaining cysteine) of a second iodoTMT reagent of different reporter fragment mass. Remaining labeling reagent was quenched with 6.2–15 mM cysteine for 10 min. Samples were digested by the addition of Sequencing Grade Modified Trypsin (Serva) and incubated at 37 °C overnight. Peptides were not reduced or carbamidomethylated because of iodoTMT-labels. After tryptic digestion, resulting peptides were desalted and concentrated using Chromabond C18WP spin columns (Macherey-Nagel, Part No. 730522). Finally, peptides were dissolved in 25 µl of water with 5% acetonitrile and 0.1% formic acid.
The mass spectrometric analysis of the samples was performed using either an Orbitrap Velos Pro mass spectrometer or an Orbitrap Exploris 480 (both Thermo Scientific). Both systems were connected to Ultimate nanoRSLC-HPLC systems (Thermo Scientific), equipped with custom end-fritted 50 cm × 75 µm C18 RP columns, filled with 2.4 µm beads (Dr. Maisch) and connected online to the mass spectrometers through nanospray sources. Two µl, corresponding to 2 µg, of the tryptic digests were injected onto a 300 µm ID × 1 cm C18 PepMap pre-concentration column (Thermo Scientific). Automated trapping and desalting of the sample were performed at a flowrate of 6 µl/min using water/0.05% formic acid as solvent. Separation of the tryptic peptides was achieved using a gradient of water/0.05% formic acid (solvent A) and 80% acetonitrile/0.045% formic acid (solvent B) at a flow rate of 300 nl/min, holding 4% B for 5 min, followed by a linear gradient to 45% B within 30 min and linear increase to 95% solvent B for further 5 min.
On the Orbitrap Velos Pro system, a survey scan with a resolution of 60,000 within the Orbitrap mass analyzer was combined with targeted FT-MS2 scans (resolution of 15,000) using HCD fragmentation (charge state adapted, normalized CE 35%). Ionization parameters were as follows: Capillary temperature 270 °C, source voltage 2.7 kV, S-lens RF-level 64.6. On the Orbitrap Exploris 480, a resolution of 60,000 was used for the survey scans and of 45,000 for targeted MS2 scans. A fixed scan range of 120–1200 m/z was used for the targeted MS2 scans to ensure that the reporter ions were properly measured. Ionization parameters were as follows: Temperature of ion transfer tube 275 °C, source voltage 2.3 kV. On both mass spectrometers, targeted MS2-spectra were recorded for the following m/z of the targets: ISCU2: 959.5085, 748.7231, 614.3708, 830.4421; NFS1: 485.2727, 447.7556, 815.4144, 848.1060. Extracted ion chromatograms corresponding to the reporter fragment ion masses were generated from MS2 data scans with 5 ppm (Orbitrap Velos Pro) or 10 ppm (Orbitrap Exploris 480) mass accuracy using the QualBrowser of the Xcalibur software package (Thermo Scientific). The resulting signals were manually integrated for quantification purposes (Supplementary Fig. 13). For each protein Cys residue, the percentage of persulfidation was calculated from the ratio of detected amounts of both iodoTMT labels.
Mössbauer spectroscopy
Samples were prepared under strictly anaerobic conditions in an anaerobic chamber (Coy). Two sets of Mössbauer samples were prepared. The first set (Supplementary Fig. 12 and Supplementary Table 1) was prepared employing 57Fe-enriched ferric ammonium citrate (FAC). Chemicals listed in Supplementary Table 2b were mixed in buffer 1 (50 mM Tris, 150 mM NaCl, 5% w/v glycerol, pH 8.0) in the given order. Subsequently, a buffer exchange into buffer E (buffer 1 plus 2 mM TCEP) was performed using ZebaSpin 7 K columns immediately before flash freezing. Resulting samples exhibited significant amounts of non-ISCU2-specific Fe species (components A and B) as impurities (Supplementary Table 1). The second set was prepared under optimized conditions (Supplementary Table 2a) that drastically decreased these Fe impurities (Fig. 4 and Table 3). In this case, (NH4)257Fe(SO4)2 was synthesized from elemental 57Fe and a mixture of (NH4)2SO4 and H2SO452. Chemicals listed in Supplementary Table 2a were mixed with buffer 1 (50 mM Tris, 150 mM NaCl, 70% w/v glycerol, pH 8.0) to achieve a final glycerol content of 30% w/v at a final volume of 360 µl. Buffers, ISCU2, (NH4)257FeSO4 and Na-ascorbate were mixed first, and samples were incubated for 5–10 min. Cysteine was added last to samples to initiate persulfide formation. Directly afterwards, 240 µl were pipetted into Mössbauer cups. Samples were flash-frozen in liquid nitrogen inside the anaerobic chamber. This resulted in an incubation time with Cys of ~2 min. The [2Fe-2S]-loaded ISCU2 sample was incubated for 40 min before flash freezing, and the formation of the [2Fe-2S] cluster was confirmed by analyzing an aliquot (50 µl) by CD spectroscopy.
Mössbauer transmission data of the prepared samples were recorded and processed using a conventional Mössbauer spectrometer in constant acceleration mode. The setup included a 512-channel analyzer (WissEl GmbH) in the time-scale mode. Calibration of spectra was performed at room temperature with α-Fe foil. Zero-field spectra were recorded upon sample cooling to 77 K via a continuous flow cryostat (Oxford Instruments) filled with liquid nitrogen. Data analysis was performed using the Microsoft Excel add-on VindaB53, employing least-squares fits with Lorentzian line shapes. The relative transmissions given as data points in Fig. 4 and Supplementary Fig. 12 were calculated by dividing the registered total counts of each channel by the spectral background.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Cryo-EM maps have been deposited in the Electron Microscopy Data Bank (EMDB) under the following accession codes: EMD-17731 ((Fe-NIAUX)2 consensus map), EMD-17732 ((Fe-NIAUSX)2 structure), EMD-17733 ((Fe-NSIAUSX)2 structure), and EMD-17734 ((Fe-NIAU)2 structure). Atomic models have been deposited in the Protein Data Bank (PDB) under the following accession codes: 8PK8 ((Fe-NIAUSX)2 structure), 8PK9 ((Fe-NSIAUSX)2 structure), and 8PKA, ((Fe-NIAU)2 structure). Previously published structures used in this study are available in the PDB with the accession codes 1WFZ (M. musculus IscU solution NMR structure), 5WLW (H. sapiens (Zn-NIAU)2 X-ray structure) and 6NZU (H. sapiens (Zn-NIAUX)2 cryo-EM structure). Any additional information required to reanalyze the data reported in this study is available from the corresponding authors. Source data are provided with this paper.
References
Beinert, H., Holm, R. H. & Münck, E. Iron-sulfur clusters: nature’s modular, multipurpose structures. Science 277, 653–659 (1997).
Johnson, D. C., Dean, D. R., Smith, A. D. & Johnson, M. K. Structure, function and formation of biological iron-sulfur clusters. Ann. Rev. Biochem. 74, 247–281 (2005).
Upadhyay, A. S. et al. Cellular requirements for iron-sulfur cluster insertion into the antiviral radical SAM protein viperin. J. Biol. Chem. 292, 13879–13889 (2017).
Barton, J. K., Silva, R. M. B. & O’Brien, E. Redox chemistry in the genome: emergence of the [4Fe4S] cofactor in repair and replication. Annu. Rev. Biochem. 88, 163–190 (2019).
Braymer, J. J., Freibert, S. A., Rakwalska-Bange, M. & Lill, R. Mechanistic concepts of iron-sulfur protein biogenesis in Biology. Biochim. Biophys. Acta Mol. Cell Res. 1868, 118863 (2021).
Maio, N. et al. Fe-S cofactors in the SARS-CoV-2 RNA-dependent RNA polymerase are potential antiviral targets. Science 373, 236–241 (2021).
Py, B. & Barras, F. Genetic approaches of the Fe-S cluster biogenesis process in bacteria: historical account, methodological aspects and future challenges. Biochim. Biophys. Acta 1853, 1429–1435 (2015).
Ciofi-Baffoni, S., Nasta, V. & Banci, L. Protein networks in the maturation of human iron-sulfur proteins. Metallomics 10, 49–72 (2018).
Lill, R. & Freibert, S. A. Mechanisms of mitochondrial iron-sulfur protein biogenesis. Annu. Rev. Biochem. 89, 471–499 (2020).
Maio, N. & Rouault, T. A. Mammalian iron sulfur cluster biogenesis and human diseases. IUBMB Life 74, 705–714 (2022).
Beilschmidt, L. K. & Puccio, H. M. Mammalian Fe-S cluster biogenesis and its implication in disease. Biochimie 100, 48–60 (2014).
Frazier, A. E., Thorburn, D. R. & Compton, A. G. Mitochondrial energy generation disorders: genes, mechanisms, and clues to pathology. J. Biol. Chem. 294, 5386–5395 (2019).
Stehling, O., Wilbrecht, C. & Lill, R. Mitochondrial iron–sulfur protein biogenesis and human disease. Biochimie 100, 61–77 (2014).
Webert, H. et al. Functional reconstitution of mitochondrial Fe/S cluster synthesis on Isu1 reveals the involvement of ferredoxin. Nat. Commun. 5, 5013 (2014).
Boniecki, M. T., Freibert, S. A., Muhlenhoff, U., Lill, R. & Cygler, M. Structure and functional dynamics of the mitochondrial Fe/S cluster synthesis complex. Nat. Commun. 8, 1287 (2017).
Cai, K., Frederick, R. O., Dashti, H. & Markley, J. L. Architectural features of human mitochondrial cysteine desulfurase complexes from crosslinking mass spectrometry and small-angle X-ray scattering. Structure 26, 1127–1136.e4 (2018).
Fox, N. G. et al. Structure of the human frataxin-bound iron-sulfur cluster assembly complex provides insight into its activation mechanism. Nat. Commun. 10, 2210 (2019).
Freibert, S. A. et al. N-terminal tyrosine of ISCU2 triggers [2Fe-2S] cluster synthesis by ISCU2 dimerization. Nat. Commun. 12, 6902 (2021).
Srour, B. et al. Iron insertion at the assembly site of the ISCU scaffold protein is a conserved process initiating Fe-S cluster biosynthesis. J. Am. Chem. Soc. 144, 17496–17515 (2022).
Schulz, V. et al. Functional spectrum and specificity of mitochondrial ferredoxins FDX1 and FDX2. Nat. Chem. Biol. 19, 206–217 (2023).
Dutkiewicz, R. & Nowak, M. Molecular chaperones involved in mitochondrial iron-sulfur protein biogenesis. J. Biol. Inorg. Chem. 23, 569–579 (2018).
Weiler, B. D. et al. Mitochondrial [4Fe-4S] protein assembly involves reductive [2Fe-2S] cluster fusion on ISCA1-ISCA2 by electron flow from ferredoxin FDX2. Proc. Natl Acad. Sci. USA 117, 20555–20565 (2020).
Lill, R. From the discovery to molecular understanding of cellular iron-sulfur protein biogenesis. Biol. Chem. 401, 855–876 (2020).
Przybyla-Toscano, J., Christ, L., Keech, O. & Rouhier, N. Iron-sulfur proteins in plant mitochondria: roles and maturation. J. Exp. Bot. 72, 2014–2044 (2021).
Parent, A. et al. Mammalian frataxin directly enhances sulfur transfer of NFS1 persulfide to both ISCU and free thiols. Nat. Commun. 6, 5686 (2015).
Patra, S. & Barondeau, D. P. Mechanism of activation of the human cysteine desulfurase complex by frataxin. Proc. Natl Acad. Sci. USA 116, 19421–19430 (2019).
Schmucker, S. et al. Mammalian frataxin: an essential function for cellular viability through an interaction with a preformed ISCU/NFS1/ISD11 iron-sulfur assembly complex. PLoS ONE 6, e16199 (2011).
Tsai, C. L. & Barondeau, D. P. Human frataxin is an allosteric switch that activates the Fe-S cluster biosynthetic complex. Biochemistry 49, 9132–9139 (2010).
Manicki, M. et al. Overlapping binding sites of the frataxin homologue assembly factor and the heat shock protein 70 transfer factor on the Isu iron-sulfur cluster scaffold protein. J. Biol. Chem. 289, 30268–30278 (2014).
Gervason, S. et al. Physiologically relevant reconstitution of iron-sulfur cluster biosynthesis uncovers persulfide-processing functions of ferredoxin-2 and frataxin. Nat. Commun. 10, 3566 (2019).
Fujishiro, T. et al. Zinc-ligand swapping mediated complex formation and sulfur transfer between SufS and SufU for iron-sulfur cluster biogenesis in Bacillus subtilis. J. Am. Chem. Soc. 139, 18464–18467 (2017).
Sheftel, A. D. et al. Humans possess two mitochondrial ferredoxins, Fdx1 and Fdx2, with distinct roles in steroidogenesis, heme, and Fe/S cluster biosynthesis. Proc. Natl Acad. Sci. USA 107, 11775–11780 (2010).
Uzarska, M. A. et al. During FeS cluster biogenesis, ferredoxin and frataxin use overlapping binding sites on yeast cysteine desulfurase Nfs1. J. Biol. Chem. 298, 101570 (2022).
Babcock, M. et al. Regulation of mitochondrial iron accumulation by Yfh1p, a putative homolog of frataxin. Science 276, 1709–1712 (1997).
Mühlenhoff, U., Richhardt, N., Ristow, M., Kispal, G. & Lill, R. The yeast frataxin homolog Yfh1p plays a specific role in the maturation of cellular Fe/S proteins. Hum. Mol. Genet. 11, 2025–2036 (2002).
Bill, E. et al. Fe(II)-substituted horse liver alcohol dehydrogenase, a model for non-heme iron enzymes. Various states of iron-dioxygen interaction investigated by Mössbauer and EPR spectroscopy. Eur. J. Biochem. 180, 111–121 (1989).
Schünemann, V. et al. Iron coordination geometry in full-length, truncated, and dehydrated forms of human tyrosine hydroxylase studied by Mössbauer and X-ray absorption spectroscopy. J. Biol. Inorg. Chem. 4, 223–231 (1999).
Bailey, T. S., Zakharov, L. N. & Pluth, M. D. Understanding hydrogen sulfide storage: probing conditions for sulfide release from hydrodisulfides. J. Am. Chem. Soc. 136, 10573–10576 (2014).
Snyder, J. P. & Carlsen, L. Lone pair-lone pair interactions in unsymmetrical systems: RSSR vs. RSOR. J. Am. Chem. Soc. 99, 2931–2942 (1977).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Asarnow, D., Palovcak, E. & Cheng, Y. UCSF pyem v0.5. Zenodo. https://doi.org/10.5281/zenodo.3576630 (2019).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. Elife 7, e42166 (2018).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Zivanov, J., Nakane, T. & Scheres, S. H. W. Estimation of high-order aberrations and anisotropic magnification from cryo-EM data sets in RELION-3.1. IUCrJ 7, 253–267 (2020).
Terwilliger, T. C., Ludtke, S. J., Read, R. J., Adams, P. D. & Afonine, P. V. Improvement of cryo-EM maps by density modification. Nat. Methods 17, 923–927 (2020).
Terwilliger, T. C., Sobolev, O. V., Afonine, P. V. & Adams, P. D. Automated map sharpening by maximization of detail and connectivity. Acta Crystallogr. D Struct. Biol. 74, 545–559 (2018).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr. D Struct. Biol. 75, 861–877 (2019).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Kelly, S. M. & Price, N. C. The use of circular dichroism in the investigation of protein structure and function. Curr. Protein Pept. Sci. 1, 349–384 (2000).
Konz, T., Montes-Bayon, M. & Sanz-Medel, A. Incorporation of (57)Fe-isotopically enriched in apoferritin: formation and characterization of isotopically enriched Fe nanoparticles for metabolic studies. Analyst 139, 5451–5459 (2014).
Gunnlaugsson, H. P. Spreadsheet based analysis of Mössbauer spectra. Hyperfine Interact. 237, 79 (2016).
Acknowledgements
We acknowledge the contribution of the Core Facilities “Mass spectrometry and Elemental analysis”, “Protein Biochemistry and Spectroscopy” of the Philipps-Universität Marburg, and the “Central Electron Microscopy Facility” of the Max Planck Institute of Biophysics in Frankfurt. We thank Dr. Timo Glatter (MPI Marburg) for letting us use the Exploris 480 mass spectrometer. R.L. acknowledges generous financial support from Deutsche Forschungsgemeinschaft (Koselleck grant LI 415/6, and SPP 1927, LI 415/7), B.J.M. from the Max Planck Society, and Vo.S. from Deutsche Forschungsgemeinschaft (SPP 1927, SCHU 1251/17-2).
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
Conceptualization was by Vo.S., B.J.M., and R.L. V.S. and S.A.F. prepared the protein samples for cryo-EM, biochemistry, and Mössbauer spectroscopy. R.S. prepared the cryo-EM grids, acquired and processed cryo-EM data, and built the atomic models. S.W. aligned the electron microscope and supported data acquisition. R.S. and B.J.M. analyzed and interpreted the cryo-EM data. V.S., S.A.F., and N.K. performed the biochemical experiments and analyzed them with R.L. U.L. performed the mass spectrometric analyses including the iodoTMT experiments which were evaluated with V.S. J.O. and M.H.H. acquired the Mössbauer data which were evaluated together with Vo.S. V.S., R.S., and J.O. prepared the figures and tables. Writing of the original draft was by V.S., R.S., Vo.S., B.J.M., and R.L. Review and editing of the manuscript was by all authors. Visualization was by V.S., R.S., J.O., S.A.F., and R.L. Supervision and funding acquisition was by Vo.S., B.J.M., and R.L.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Schulz, V., Steinhilper, R., Oltmanns, J. et al. Mechanism and structural dynamics of sulfur transfer during de novo [2Fe-2S] cluster assembly on ISCU2. Nat Commun 15, 3269 (2024). https://doi.org/10.1038/s41467-024-47310-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-024-47310-8
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.