Abstract
Syndromes caused by copy number variations are described as reciprocal when they result from deletions or duplications of the same chromosomal region. When comparing the phenotypes of these syndromes, various clinical features could be described as reversed, probably due to the opposite effect of these imbalances on the expression of genes located at this locus. The NFIX gene codes for a transcription factor implicated in neurogenesis and chondrocyte differentiation. Microdeletions and loss of function variants of NFIX are responsible for Sotos syndrome-2 (also described as Malan syndrome), a syndromic form of intellectual disability associated with overgrowth and macrocephaly. Here, we report a cohort of nine patients harboring microduplications encompassing NFIX. These patients exhibit variable intellectual disability, short stature and small head circumference, which can be described as a reversed Sotos syndrome-2 phenotype. Strikingly, such a reversed phenotype has already been described in patients harboring microduplications encompassing NSD1, the gene whose deletions and loss-of-function variants are responsible for classical Sotos syndrome. Even though the type/contre-type concept has been criticized, this model seems to give a plausible explanation for the pathogenicity of 19p13 microduplications, and the common phenotype observed in our cohort.
Similar content being viewed by others
Introduction
Thanks to the use of chromosomal microarray as a diagnostic tool in medical genetics, copy number variations (CNVs) have been implicated in many syndromic forms of intellectual disability [1,2,3]. The pathogenic effect of these chromosomal abnormalities may arise not only from an abnormal gene dosage caused by haploinsufficiency of the deleted genes, in the case of deletions, or, conversely, by the overexpression of the duplicated genes, in the case of duplications, but also from the alteration in the expression of genes neighboring the CNV due to a breakage of topological associating domains [4].
In line with this view, several syndromes have been described as “reciprocal” at the molecular level. This phenomenon is mostly explained by non-allelic homologous recombination, the main mechanism leading to the occurrence of CNVs with recurrent breakpoints [5]. As an example, deletions and duplications at the 16p11.2 locus have been reported as having a reciprocal impact, predisposing to obesity or underweight, respectively [6]. Nevertheless, non-recurrent chromosomal rearrangements have also been identified in genetic syndromes by phenotypic comparison of patients presenting with similar clinical features and carrying overlapping CNVs. Candidate causative genes are often included in the critical minimal overlapping regions. Thus, one of these microdeletion syndromes, Sotos syndrome-2 (also referred as to Malan syndrome) (Online Mendelian Inheritance in Man (OMIM) #614753), a form of intellectual disability associated with excessive growth [7], is caused by haploinsufficiency of NFIX, a gene coding for a transcription factor involved in brain development [8]. This haploinsufficiency originates from 19p13 microdeletions encompassing the NFIX gene or from NFIX loss-of-function variants processed by nonsense-mediated mRNA decay. Until now, only one patient harboring a microduplication at this locus has been described. This patient presents moderate intellectual disability, associated with short stature and small head circumference. Here, we delineate, for the first time, the phenotype associated with 19p13 microduplications encompassing the entire NFIX gene in a series of nine additional patients.
Our findings suggest that the phenotype associated with this chromosomal imbalance could be considered in some features as the reciprocal phenotype of Sotos syndrome-2.
Materials and methods
Recruitment of the cohort
Following the identification of a 19p13 duplication in patient 9 (DECIPHER 338712) in the Medical Genetics Department of the University Hospital of Bordeaux, a national and international call for collaboration through the Association of French Language Cytogeneticists network, “Achropuces” (French acronym standing for “Réseau d’Analyse Chromosomique sur Puces à ADN) and via the DECIPHER database [9] was conducted, allowing the recruitment of eight additional patients (patients 1 to 8). The clinical and genetic data were collected and patients’ phenotypes, including the one already described in the literature by Dolan et al [10] (patient 5), were compared.
Methods for microduplication detection and confirmation
Array comparative genomic hybridization or single-nucleotide polymorphism array chips (with resolutions ranging from 44k to 180k depending on the laboratory where the analysis was performed) were used according to the manufacturers’ instructions. When possible, segregation was studied within each family, using quantitative PCR and fluorescent in situ hybridizatio (FISH) analyses. All coordinates were converted into the hg19 build of the genome via UCSC (University of California, Santa Cruz) Genome Browser Liftover tool [11]. The coordinates of the duplications correspond to the minimal intervals.
Results
A total of nine patients were enrolled in our study through an international collaboration. Table 1 recapitulates the molecular and clinical features of the 10 subjects harboring 19p13 duplications, including patient 5, previously reported by Dolan et al. [10].
Clinical results
A detailed clinical description of the patients can be found in Supplemental Dataset 1. None of the patients were born to consanguineous parents.
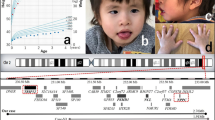
Several clinical features are common among these patients, especially short stature (7/10), defined as a height ⩽−2 SD, and a small head circumference (7/10), ⩽−2 SD. Otherwise, weight was less severely affected than height, since in 6/10 patients, the difference between these two parameters was ⩾2 SD. This reflects a potential risk of overweight or even obesity in this syndrome. Bone X-rays were performed in six patients and showed a delayed bone age in five of them. Intellectual disability was variable, ranging from severe with no language to mild disability, but special education was required in all cases. On examination, dysmorphism was rather nonspecific; however, some common features were observed, including full cheeks (6/10), brachydactyly (5/10), arched eyebrows with sparseness in their medial part (4/10) and everted, thick lower lip (4/10) (Fig. 1).
Molecular results
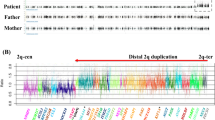
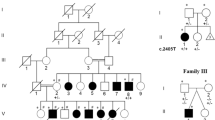
Each patient carried a heterozygous microduplication in 19p13 with non-recurrent breakpoints, but patient 9 and patient 5, from Dolan et al. [10], had similar minimal intervals (Fig. 2). In our cohort, six of the nine duplications were de novo (patients 2, 3, 4, 5, 7 and 9). Especially in two brothers (patients 2 and 3), FISH analyses with RP11-455A18 and RP11-79P23 probes performed in the parents did not find the rearrangement, thus indicating the presence of germline mosaicism in one of them. The parental study could not be performed in patient 6 and remained incomplete in patients 1 and 8 as the father’s samples were not available. Overall, microduplications ranged from 479 kb to 3.1 Mb. The 422 kb minimal region of overlap encompassed 16 coding genes. Among these, six are currently associated with a phenotype in the OMIM database: four are entirely duplicated (GCDH (OMIM: 608801), CALR (OMIM: 109091), NACC1 (OMIM: 610672) and NFIX (OMIM: 164005)) and two are partially duplicated (KLF1 (OMIM: 600599) and CACNA1A (OMIM: 601011)).
Discussion
Here, we describe a cohort of nine patients carrying various 19p13 microduplications. The minimal interval of overlap spanned 422 kb and included 16 genes. The patients share several symptoms, including intellectual disability, short stature, small head circumference and delayed bone age. Interestingly, similar manifestations were reported in patient 5 from Dolan et al. [10]. He presented mild intellectual disability, feeding difficulties with frequent vomiting, recurrent infections, nystagmus and epilepsy. Neither photographs nor morphological descriptions were provided.
Among the 16 genes included in the critical minimum interval, six (KLF1, GCDH, CALR, NFIX, NACC1 and CACNA1A) are described in the OMIM database as associated with a pathological phenotype. KLF1 encodes a transcriptional activator of the β-globin promoter and some heterozygous variants were found to cause the rare red blood cell phenotype in(Lu) lacking Lutherian antigen [12]. GCDH encodes an acyl-dehydrogenase involved in the metabolism of lysine, hydroxylysine and tryptophan. Biallelic variants altering the function of this gene are associated with glutaric aciduria type 1 [13]. CALR encodes a major protein implicated in calcium storage in the lumen of the endoplasmic reticulum. Somatic variants of CALR were found in primitive myelofibrosis [14]. CACNA1A encodes a transmembrane protein involved in the formation of calcium channel P/Q or Cav2.1 [15]. CACNA1 truncating variants and exon deletions were reported in episodic ataxia type 2 [16]. Moreover, a patient presenting with episodic ataxia and carrying an intragenic CACNA1A duplication was also described, while his son exhibited diplopia without ataxia [17]. Otherwise, numerous CACNA1A missense variants affecting the protein function have been associated with not only familial hemiplegic migraine [18], but also various epileptic syndromes, either as a form of idiopathic generalized epilepsy [18] or epileptic encephalopathy [19]. Finally, coding CAG repeat expansions cause spinocerebellar ataxia type 6 [20]. NACC1 encodes for a transcriptional repressor implicated in gene expression, and has been recently identified as implicated in an autosomal dominant neurodevelopmental phenotype characterized by epilepsy, cataracts and severe intellectual disability [21]. The seven reported individuals harbored the same de novo heterozygous missense variant, and the authors postulated that this missense variant exerted a specific dominant negative or a gain-of-function effect on the protein. They also pointed out that none of the patients with a copy number variant that included NACC1 showed the same syndromic phenotype.
NFIX is the sixth OMIM gene included in the minimal region of overlap and several data suggest that it is involved in various clinical features. This gene spans 74 kb, contains nine or ten coding exons according to the isoform, and encodes a transcription factor belonging to the NFI family, which comprises four paralogs in vertebrates: NFIA, NFIB, NFIC and NFIX [22]. Proteins encoded by these genes act as homodimers or heterodimers and bind to palindromic sequence TTGGCNNNNNGCCAA to regulate the expression of target genes. NFI transcription factors are crucial for brain development, especially gliogenesis [23,24,25]. Knockout murine models Nfia−/− [26, 27] and Nfib−/− [28] exhibit agenesis of the corpus callosum and an abnormal frontal cortex, with hydrocephalus in Nfia−/− mice. Several mouse models of Nfix−/− were generated and highlighted the central role of this gene in the development of several brain regions such as the neocortex, the hippocampus and the cerebellum [29,30,31]. In particular, Nfix induces the differentiation of neural stem cells. Postnatal hydrocephalus was also observed [32]. Previous functional studies in neural stem cells showed that NFIX overexpression induces quiescence of neural progenitors [33, 34]. Additionally, the expression of Nfix in the backbone suggests that it plays a role in chondrocyte differentiation, with a potential negative effect on endochondral ossification [8]. It was also demonstrated that Nfix regulates the survival of hematopoietic progenitors [35] and plays a role in muscle regeneration [36].
In human disease, genetic alterations of NFIX have been associated with two distinct phenotypes: Marshall–Smith syndrome and Sotos syndrome-2. Marshall–Smith syndrome is caused by NFIX splice site or frameshift variants, escaping the nonsense-mediated mRNA decay (NMD) system, with a presumed dominant negative effect [8, 37, 38]. This syndrome is characterized by intellectual disability, overgrowth with accelerated bone maturation and skeletal malformations such as severe scoliosis, osteopenia and multiple fractures. Significant respiratory and feeding difficulties worsen the prognosis. The facial phenotype is characterized by a high forehead, proptosis, bluish sclera, midface retrusion and retrognathia. Sotos syndrome-2 (also referred as to Malan syndrome in the literature) is also associated with NFIX variants, either 19p13 microdeletions encompassing the entire gene or truncating variants. However, unlike the previous syndrome, the mutations are processed by the NMD system, suggesting a haploinsufficiency mechanism [8]. The phenotype of these patients is characterized by intellectual disability, overgrowth, macrocephaly, prominent forehead, high anterior hairline, upslanted palpebral fissures and prominent chin [8, 10,39,40,41,42,43]. Some patients carrying similar deletions or truncating variants have been described with a marfanoid phenotype, or with some skeletal features, such as pectus excavatum and scoliosis. These patients exhibit mild intellectual disability, with feeding difficulties and hypotonia in the neonatal period. Ophthalmologic disorders have also been described, including strabismus, nystagmus or papillary pallor.
Comparison of the phenotype observed in all patients carrying 19p13 microduplications encompassing NFIX suggests a type/contre-type effect of NFIX CNVs, especially regarding growth parameters. Indeed, 8/10 patients exhibited growth retardation, with bone age delay confirmed in 5/6, while 7/10 presented a small head circumference. Unfortunately, in most of our cases, growth parameters and head circumferences (occipital frontal circumference) were not available in patients’ parents for the assessment of the familial background.
Mirror phenotypes associated with 19p13 microdeletions and microduplications could be explained by the opposite effects of NFIX haploinsufficiency and overexpression. Evidence from the literature suggests that NFIX could be a repressor of endochondral ossification. It has been hypothesized that the tall stature of patients with Marshall–Smith syndrome and Sotos syndrome-2 could be related to a decrease in this repression. Conversely, NFIX overexpression in patients with duplications could lead to increased repression, resulting in growth deficiency. Likewise, several studies described patients carrying 5q35 duplications encompassing the NSD1 gene and presenting with intellectual disability, growth delay, microcephaly and delayed bone age [44,45,46]. Strikingly, this contrasts with the phenotype associated with NSD1 deletions and NSD1 loss-of-function variants, which are responsible for classical Sotos syndrome-1. The resulting phenotype associated with NSD1 duplication has been described as “reversed Sotos syndrome-1 phenotype”. This hypothesis was also supported by a recent study in which three different Xq25q26 microduplications were described in male and female individuals who exhibited various degrees of growth retardation, microcephaly and intellectual disability. In this study, growth impairment was attributed, at least in part, to an increased dosage of GPC3. Loss-of-function variants of this gene are involved in another well-known multiple congenital anomaly syndrome characterized by overgrowth, namely Simpson–Golabi–Behmel syndrome. The study of transgenic mouse models overexpressing GPC3 revealed that they were smaller and had a 6 to 19% lower body weight compared with wild-type littermates [47]. Some criticisms have been raised about this concept of “reversed phenotype” [48], and the same criticisms can be made about this potential “reversed Sotos syndrome-2 phenotype”. Indeed, only growth parameters seem to be relevant, whereas facial features and cognitive abilities do not exhibit the mirror phenotype. None of the other genes in the minimal interval of overlap seem to play a role in growth or bone maturation, although overexpression of several genes located within the minimal interval of overlap have not been studied yet.
Regarding intellectual disability, CACNA1A, which is expressed in the brain, could also be involved in the phenotype. Until now, CACNA1A duplications restricted to the entire gene have never been reported. Moreover, none of our patients, although still young, have epilepsy or ataxia, and the phenotype severity in our cohort does not appear to be affected by the fact that the duplications fully encompass CACNA1A or not; only the patient from Dolan et al. [10] suffered from seizures, though the description regarding the evolution was limited. We also cannot exclude the possibility that NACC1 is overexpressed and contributes to the neurodevelopmental phenotype of our patients. However, the pathophysiological mechanism implicated in the NACC1-associated phenotype known to date is different.
The two most severely affected patients were patients 2 and 3. These siblings from the same family harbored the largest duplication among this cohort. This finding is probably due to the fact that the duplication also included the genes AKAP8 and AKAP8L, for which gene dosage anomalies have been correlated with head size and associated with autism [49]. In particular, microduplications encompassing AKAP8 and AKAP8L were found to be associated with macrocephaly, but patient 2, who had been included in the DECIPHER 257523 study, was the only one who presented a small head circumference. This finding suggests a stronger effect of the NFIX gene on head size and the additive effects of the deregulated expression of at least these three genes on cognitive alterations. It is noteworthy that this family illustrates once again the risk of germline mosaicism in such de novo chromosomal disorders and this has to be kept in mind for genetic counseling; such mosaicism at this locus was also reported in a family in which two children carried a de novo 399 kb deletion disrupting NFIX and CACNA1A [50].
Although we cannot definitely prove that NFIX is the major gene accounting for the phenotype reported in this cohort, there is strong evidence that it does play a role.
Conclusion
We report here a cohort of nine patients carrying various de novo microduplications at the 19p13 locus, all of them encompassing the NFIX gene. The most striking features include intellectual disability of variable severity, short stature associated with delayed bone age, small head circumference and several minor nonspecific morphological features.
Regarding growth parameters, this phenotype contrasts with that of patients suffering from Sotos syndrome-2, and shares similar clinical manifestations with patients carrying NSD1 and GPC3 duplications. These “mirror” features could be explained by the opposite effect of haploinsufficiency and overexpression of NFIX, especially on endochondral ossification. Intellectual disability is hypothesized to be caused by the role of NFIX in the multiplication and the differentiation of neuronal progenitors. Further functional studies and descriptions of additional patients carrying 19p13 microduplications are needed to potentially refine the associated phenotype and to reduce the critical minimal interval.
References
Miller DT, Adam MP, Aradhya S, et al. Consensus statement: chromosomal microarray is a first-tier clinical diagnostic test for individuals with developmental disabilities or congenital anomalies. Am J Hum Genet 2010;86:749–64.
Redon R, Ishikawa S, Fitch KR, et al. Global variation in copy number in the human genome. Nature 2006;444:444–54.
Zarrei M, MacDonald JR, Merico D, Scherer SW. A copy number variation map of the human genome. Nat Rev Genet 2015;16:172–83.
Franke M, Ibrahim DM, Andrey G, et al. Formation of new chromatin domains determines pathogenicity of genomic duplications. Nature 2016;538:265–9.
Liu P, Carvalho CM, Hastings P, Lupski JR. Mechanisms for recurrent and complex human genomic rearrangements. Curr Opin Genet Dev 2012;22:211–20.
Jacquemont S, Reymond A, Zufferey F, et al. Mirror extreme BMI phenotypes associated with gene dosage at the chromosome 16p11.2 locus. Nature 2011;478:97–102.
Sotos JF. Sotos syndrome 1 and 2. Pediatr Endocrinol Rev 2014;12:2–16.
Malan V, Rajan D, Thomas S, et al. Distinct effects of allelic NFIX mutations on nonsense-mediated mRNA decay engender either a Sotos-like or a Marshall-Smith syndrome. Am J Hum Genet 2010;87:189–98.
Firth HV, Richards SM, Bevan AP, et al. DECIPHER: Database of Chromosomal Imbalance and Phenotype in Humans Using Ensembl Resources. Am J Hum Genet 2009;84:524–33.
Dolan M, Mendelsohn NJ, Pierpont ME, Schimmenti LA, Berry SA, Hirsch B. A novel microdeletion/microduplication syndrome of 19p13.13. Genet Med 2010;12:503–11.
Kent WJ, Sugnet CW, Furey TS, et al. The Human Genome Browser at UCSC. Genome Res 2002;12:996–1006.
Singleton BK, Burton NM, Green C, Brady RL, Anstee DJ. Mutations in EKLF/KLF1 form the molecular basis of the rare blood group In(Lu) phenotype. Blood 2008;112:2081–88.
Goodman SI, Stein DE, Schlesinger S, et al. Glutaryl-CoA dehydrogenase mutations in glutaric acidemia (type I): review and report of thirty novel mutations. Hum Mutat 1998;12:141–4.
Nangalia J, Massie CE, Baxter EJ, et al. Somatic CALR mutations in myeloproliferative neoplasms with nonmutated JAK2. N Engl J Med 2013;369:2391–405.
Kordasiewicz HB, Thompson RM, Clark HB, Gomez CM. C-termini of P/Q-type Ca2+ channel alpha1A subunits translocate to nuclei and promote polyglutamine-mediated toxicity. Hum Mol Genet 2006;15:1587–99.
Ophoff RA, Terwindt GM, Vergouwe MN, et al. Familial hemiplegic migraine and episodic ataxia type-2 are caused by mutations in the Ca2+ channel gene CACNL1A4. Cell 1996;87:543–52.
Labrum RW, Rajakulendran S, Graves TD, et al. Large scale calcium channel gene rearrangements in episodic ataxia and hemiplegic migraine: implications for diagnostic testing. J Med Genet 2009;46:786–91.
Chioza B, Wilkie H, Nashef L, et al. Association between the alpha(1a) calcium channel gene CACNA1A and idiopathic generalized epilepsy. Neurology 2001;56:1245–6.
Myers C, McMahon J, Schneider A, et al. De novo mutations in SLC1A2 and CACNA1A are important causes of epileptic encephalopathies. Am J Hum Genet 2016;99:287–98.
Zhuchenko O, Bailey J, Bonnen P, et al. Autosomal dominant cerebellar ataxia (SCA6) associated with small polyglutamine expansions in the α1A-voltage-dependent calcium channel. Nat Genet 1997;15:62–9.
Schoch K, Meng L, Szelinger S, et al. A recurrent de novo variant in NACC1 causes a syndrome characterized by infantile epilepsy, cataracts, and profound developmental delay. Am J Hum Genet 2017;100:343–51.
Roulet E, Bucher P, Schneider R, et al. Experimental analysis and computer prediction of CTF/NFI transcription factor DNA binding sites. J Mol Biol 2000;297:833–48.
Deneen B, Ho R, Lukaszewicz A, Hochstim CJ, Gronostajski RM, Anderson DJ. The transcription factor NFIA controls the onset of gliogenesis in the developing spinal cord. Neuron 2006;52:953–68.
Kang P, Lee HK, Glasgow SM, et al. Sox9 and NFIA coordinate a transcriptional regulatory cascade during the initiation of gliogenesis. Neuron 2012;74:79–94.
Wilczynska KM, Singh SK, Adams B, et al. Nuclear factor I isoforms regulate gene expression during the differentiation of human neural progenitors to astrocytes. Stem Cells 2009;27:1173–81.
das Neves L, Duchala CS, Tolentino-Silva F, et al. Disruption of the murine nuclear factor I-A gene (Nfia) results in perinatal lethality, hydrocephalus, and agenesis of the corpus callosum. Proc Natl Acad Sci USA 1999;96:11946–51.
Shu T, Butz KG, Plachez C, Gronostajski RM, Richards LJ. Abnormal development of forebrain midline glia and commissural projections in Nfia knock-out mice. J Neurosci 2003;23:203–12.
Piper M, Moldrich RX, Lindwall C, et al. Multiple non-cell-autonomous defects underlie neocortical callosal dysgenesis in Nfib-deficient mice. Neural Dev 2009;4:43.
Driller K, Pagenstecher A, Uhl M, et al. Nuclear factor I X deficiency causes brain malformation and severe skeletal defects. Mol Cell Biol 2007;27:3855–67.
Campbell CE, Piper M, Plachez C, et al. The transcription factor Nfix is essential for normal brain development. BMC Dev Biol 2008;8:52.
Heng YHE, Zhou B, Harris L, et al. NFIX regulates proliferation and migration within the murine SVZ neurogenic niche. Cereb Cortex 2015;25:3758–78.
Vidovic D, Harris L, Harvey TJ, et al. Expansion of the lateral ventricles and ependymal deficits underlie the hydrocephalus evident in mice lacking the transcription factor NFIX. Brain Res 2015;1616:71–87.
Martynoga B, Mateo JL, Zhou B, et al. Epigenomic enhancer annotation reveals a key role for NFIX in neural stem cell quiescence. Genes Dev 2013;27:1769–86.
Zhou B, Osinski JM, Mateo JL, et al. Loss of NFIX transcription factor biases postnatal neural stem/progenitor cells toward oligodendrogenesis. Stem Cells Dev 2015;24:2114–26.
Holmfeldt P, Pardieck J, Saulsberry AC, et al. Nfix is a novel regulator of murine hematopoietic stem and progenitor cell survival. Blood 2013;122:2987–96.
Rossi G, Antonini S, Bonfanti C, et al. Nfix regulates temporal progression of muscle regeneration through modulation of myostatin expression. Cell Rep 2016;14:2238–49.
Schanze D, Neubauer D, Cormier-Daire V, et al. Deletions in the 3’ part of the NFIX gene including a recurrent Alu-mediated deletion of exon 6 and 7 account for previously unexplained cases of Marshall-Smith syndrome. Hum Mutat 2014;35:1092–100.
Martinez F, Marín-Reina P, Sanchis-Calvo A, et al. Novel mutations of NFIX gene causing Marshall-Smith syndrome or Sotos-like syndrome: one gene, two phenotypes. Pediatr Res 2015;78:533–39.
Priolo M, Grosso E, Mammì C, et al. A peculiar mutation in the DNA-binding/dimerization domain of NFIX causes Sotos-like overgrowth syndrome: a new case. Gene 2012;511:103–5.
Yoneda Y, Saitsu H, Touyama M,et al. Missense mutations in the DNA-binding/dimerization domain of NFIX cause Sotos-like features. J Hum Genet 2012;57:207–11.
Shimojima K, Okamoto N, Tamasaki A, Sangu N, Shimada S, Yamamoto T. An association of 19p13.2 microdeletions with Malan syndrome and Chiari malformation. Am J Med Genet A 2015;167A:724–30.
Klaassens M, Morrogh D, Rosser EM, et al. Malan syndrome: Sotos-like overgrowth with de novo NFIX sequence variants and deletions in six new patients and a review of the literature. Eur J Hum Genet 2015;23:610–15.
Gurrieri F, Cavaliere ML, Wischmeijer A, et al. NFIX mutations affecting the DNA-binding domain cause a peculiar overgrowth syndrome (Malan syndrome): a new patients series. Eur J Med Genet 2015;58:488–91.
Zhang H, Lu X, Beasley J,et al. Reversed clinical phenotype due to a microduplication of Sotos syndrome region detected by array CGH: Microcephaly, developmental delay and delayed bone age. Am J Med Genet A 2011;155:1374–78.
Rosenfeld JA, Kim KH, Angle B, et al. Further evidence of contrasting phenotypes caused by reciprocal deletions and duplications: duplication of NSD1 causes growth retardation and microcephaly. Mol Syndromol 2013;3:247–254
Novara F, Stanzial F, Rossi E, et al. Defining the phenotype associated with microduplication reciprocal to Sotos syndrome microdeletion. Am J Med Genet A 2014;164A:2084–90.
Møller RS, Jensen LR, Maas SM, et al. X-linked congenital ptosis and associated intellectual disability, short stature, microcephaly, cleft palate, digital and genital abnormalities define novel Xq25q26 duplication syndrome. Hum Genet 2014;133:625–38.
Dikow N, Maas B, Gaspar H, et al. The phenotypic spectrum of duplication 5q35.2-q35.3 encompassing NSD1: Is it really a reversed sotos syndrome? Am J Med Genet A 2013;; 161:2158–66.
Nebel RA, Kirschen J, Cai J, Woo YJ, Cherian K, Abrahams BS. Reciprocal relationship between head size, an autism endophenotype, and gene dosage at 19p13.12 points to AKAP8 and AKAP8L. PLOS ONE 2015;10:e0129270.
Nimmakayalu M, Horton VK, Darbro B, et al. Apparent germline mosaicism for a novel 19p13.13 deletion disrupting NFIX and CACNA1A. Am J Med Genet A 2013;161:1105–9.
Acknowledgements
We thank the patients and their families for participating in this study. We thank Wallid Deb from Universitary Hospital of Nantes for additional data supply, Dr Lucile Pinson and Dr Anouck Schneider from Universitary Hospital of Montpelier and Hilde van Esch from KU Leuven for the constructive discussions about their patients. This study makes use of data generated by the DECIPHER community. A full list of centers who contributed to the generation of the data is available from http://decipher.sanger.ac.uk and via email from decipher@sanger.ac.uk. Funding for this project was provided by the Wellcome Trust. We thank Arthur Sorlin, Dr Julia Lauer Zillhardt and Pr Damien Sanlaville for their help during the use of this database.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interests
The authors declare that they have no competing financial interests.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Trimouille, A., Houcinat, N., Vuillaume, ML. et al. 19p13 microduplications encompassing NFIX are responsible for intellectual disability, short stature and small head circumference. Eur J Hum Genet 26, 85–93 (2018). https://doi.org/10.1038/s41431-017-0037-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41431-017-0037-7