Abstract
The World Health Organisation recognised human papillomavirus (HPV) as the cause of multiple cancers, including head and neck cancers. HPV is a double-stranded DNA virus, and its viral gene expression can be controlled after infection by cellular and viral promoters. In cancer cells, the HPV genome is detected as either integrated into the host genome, episomal (extrachromosomal), or a mixture of integrated and episomal. Viral integration requires the breakage of both viral and host DNA, and the integration rate correlates with the level of DNA damage. Interestingly, patients with HPV-positive head and neck cancers generally have a good prognosis except for a group of patients with fully integrated HPV who show worst clinical outcomes. Those patients present with lowered expression of viral genes and limited infiltration of cytotoxic T cells. An impediment to effective therapy applications in the clinic is the sole testing for HPV positivity without considering the HPV integration status. This review will discuss HPV integration as a potential determinant of response to therapies in head and neck cancers and highlight to the field a novel therapeutic avenue that would reduce the cancer burden and improve patient survival.
Similar content being viewed by others
Background
Head and neck squamous cell carcinomas (HNSCC) is the main type of malignancy of the head and neck region and the seventh most common cancer worldwide accounting for nearly 1 million new cases, and 470,000 deaths in 2020 [1]. The incidence of HNSCC continues to rise and is anticipated to reach an annual incidence rate of 1.37 million new cases by 2040 [2, 3].
HNSCC originates from the mucosal epithelium of the lips, oral cavity, salivary glands, larynx and pharynx. The major external risk factors associated with the incidence of HNSCC are exposure to excessive alcohol consumption, tobacco-related cancer-causing agents, or both. Along with these risk factors, intrinsic genetic, transcriptional and post-translational changes such as the expression and/or function of Fanconi Anaemia Complementation Group (FANC) [4], Grainyhead-like 3 (GRHL3) [5], Filaggrin (FLG) [6], and cellular localisation of Y-box binding protein 1 (YBX1) [7] impact HNSCC development. On the other hand, oropharyngeal squamous cell carcinoma (SCC) tumours are associated with prior infection with oncogenic strains of human papillomavirus (HPV), whereas >70% of oropharyngeal cancers are HPV-positive. Studies reveal a global trend of rising HPV-related cancers and declining HPV-unrelated subsites. Over the next two decades, the majority of HNSCC cases are expected to be HPV-positive, with some countries like the UK experiencing a higher incidence of oropharyngeal cancer than oral cavity cancer [8, 9].
In this review, we summarise the role of HPV infection in head and neck cancer development, with a focus on chromosomal abnormalities and the viral integration process. We also highlight how the integration status affects the patients’ therapeutic outcomes.
HPV: a virus causing cancer in humans
HPV is the most prevalent sexually transmitted infection worldwide that has significant social implications. Both sexually active women and men are susceptible to HPV infection [10], although not all individuals will develop associated health issues. HPV infection remains widespread worldwide as one of the leading causes of cancer [11], especially among women, making it a critical concern for public health.
HPV has different strains, each identified with a number. The strains are classified into two categories: low-risk HPVs responsible for anogenital and cutaneous warts, and high-risk HPVs responsible for oropharyngeal and anogenital cancers [12,13,14,15,16]. HPV16 is the highest-risk strain accounting for more than 90% of HPV-associated HNSCC, while other HPV strains such as HPV-18, 31, 33 and 52 have been detected in a small proportion of patients [17,18,19,20].
The currently approved HPV vaccines, including bivalent, tetravalent, and 9-valent vaccines, have demonstrated effectiveness in reducing both HPV infection and the incidence of HPV-related diseases across various geographical regions worldwide. The vaccines target and stimulate immunity against both low- and high-risk HPVs, which are responsible for most genital and cutaneous warts (70%) as well as HPV-positive cancers (90%). Despite the proven efficacy of HPV vaccines, the burden of HPV-associated cancers and diseases remains substantial [21, 22]. Therefore, epidemiological surveillance of HPV infection and related diseases is crucial in this space. This surveillance would play a vital role in monitoring and evaluating the effectiveness of available antiviral prophylactic vaccines and assessing their acceptance globally.
Molecular mechanisms involved in the cycle of HPV replication
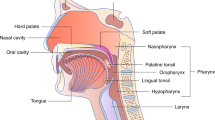
HPV is a small (50–60 nm in diameter), non-enveloped, double-stranded DNA virus belonging to the Papillomaviridae family. Unidirectional transcription of one strand of the 7–8 kb circular genomic DNA encodes for eight functional early (E1–E8), two structural late (L1 and L2) proteins [23], and a long non-coding control region also known as upstream regulatory region (URR) (Fig. 1a). Brief characteristics of expressed HPV genes are reviewed by Van Doorslaer et al. [24] and summarised in Table 1.
a Structural late L1 and L2 genes, and executive early E1–E8 genes are expressed from a nearly 8 Kbp HPV16 genome. b The HPV infection starts in the basal epithelial cells upon injury/trauma/permeability and ends with the assembly and virion release at the very top terminally differentiated epithelial cells. Early genes of E1, E2, E6, E7, E4, E5 and late genes of L1 and L2 are expressed in basal to superficial layers during the infection initiation, progression and termination. Ori origin of replication, URR upstream regulatory region.
The infection mechanism of HPV, despite the genetic variations among different strains and their diverse infection profiles, follows a similar pattern. HPV specifically targets basal epithelial cells, homed in the undifferentiated deeper layers of squamous epithelia and/or mucous membranes and possess a high mitotic capacity. To reach these target cells, the virus presence on the outer apical surface of these tissues relies on microlesions that occur during traumatic events. Upon reaching the basal epithelial cells, the virus is internalised by endocytosis. It is then transported within small vesicles through the endoplasmic reticulum (ER) and the Golgi apparatus. During this journey, a series of interactions and structural changes in the vesicles occur, leading to the removal of the viral capsid and the release of the viral genome near the nuclear membrane [25].
The viral genome enters the nucleus via nuclear pores and remains in the form of an episome (i.e., extrachromosomal) while the process of viral replication occurs via the expression and function of E1, E2, E5, E6 and E7 genes [26, 27]. Among the genes transcribed early in the viral life cycle, E2, E5, E6 and E7 play crucial roles in hijacking the cellular environment that is conducive to viral DNA replication [28, 29]. The chronology and level of HPV gene expression progress from the basal to suprabasal epithelial cells with an ultimate virus assembly in the terminally differentiated cells as presented in Fig. 1b. Of note, E6 and E7 interfere with cellular function by inhibiting two important tumour suppressor genes, i.e., p53 and Rb, to promote the proliferation of infected cells. E6 causes p53 protein degradation via mediating its interaction with E6-associated protein (E6-AP) E3-ubiquitin ligase [30]. Similarly, Rb1 undergoes sequestration and proteolysis due to E7 protein [31].
The transcriptional regulation of all functional and structural HPV genes is modulated via the upstream regulatory region (URR). The URR induces the transcription of downstream genes to start the viral genome replication upon binding of viral E1 and E2 proteins. This process is also activated by host cell transcription factors, such as nuclear factor 1 (NFI), activator protein 1 (AP-1), organic cationic transporter 1 (Oct-1), Translation elongation factor 1 (TEF-1) (also known as TF1), and specificity protein 1 (SP1) [32]. In addition to the transcriptional role of URR, the origin of replication (Ori) site lies within this region. Ori is pivotal for papillomavirus replication and is highly homologous to the mammalian autonomous replicating conserved sequences (ACS) [33]. Each papillomavirus genome contains only one Ori required for viral DNA replication [34].
To summarise, insights into the molecular characteristics and evolution of HPV infection is fundamental for understanding the distribution of HPV and its impact on HPV-related diseases. This knowledge can inform the development of new therapeutic strategies and next-generation antiviral vaccines that address the limitations of the current prophylactic regimens, such as high costs, limited spectrum of antiviral protection, and immunisation management.
Molecular differences between HPV-negative and positive HNSCC
HPV-positive HNSCC displays notable variations compared to HPV-negative HNSCC regarding immune characteristics, gene expression and importantly, mutational patterns, highlighting the distinctive entity of HPV-positive HNSCC.
Immune system and microenvironment
The composition and abundance of immune cells in tumours differ significantly between HPV-positive and HPV-negative cases [35, 36]. HPV-positive tumours generally exhibit higher levels of tumour-infiltrating lymphocytes (TILs) compared to HPV-negative tumours. Notably, patients with HPV-positive tumours containing a high number of TILs (CD8+ cytotoxic T cells) tend to have favourable outcomes, while those with HPV-positive tumours and low TIL levels experience similar poor survival rates to patients with HPV-negative HNSCC [37].
HNSCC tumours manifest various mechanisms to evade immune surveillance. The tumour microenvironment (TME) in HNSCC is enriched with immunosuppressive growth factors and cytokines that promote the recruitment or activity of myeloid-derived suppressor cells (MDSCs), regulatory T cells (Treg cells), and M2 macrophages. Simultaneously, these factors hinder the anti-tumour effects of effector T cells (Teff cells) and natural killer (NK) cells. Notably, cytokines such as IL-6, IL-10, VEGF and TGF-β play a crucial role in this immunosuppressive environment [38]. Elements within the HNSCC TME, including IL-10 and TGFβ, contribute to the polarisation of macrophages into an immunosuppressive M2 phenotype [39]. Genetic and epigenetic alterations further reduce the levels of human leucocyte antigen (HLA) on tumour cells and impair antigen processing [40, 41]. Consequently, tumour cells evade recognition and elimination by the immune system. Moreover, HNSCC tumours, particularly in advanced stages, exhibit increased expression of programmed death-ligand 1 (PDL1), which suppresses the cytolytic activity of T cells [42, 43]. Likewise, MDSCs and Treg cells recruited to the HNSCC TME express PDL1 and cytotoxic T lymphocyte antigen 4 (CTLA4), respectively, both of which are immunosuppressive molecules. In the context of HPV-positive HNSCC, limited knowledge on the patient immunological status compared to HPV-negative HNSCC is available. Nevertheless, it is known that in HPV-positive HNSCC, the viral proteins E5, E6 and E7 promote immune evasion by altering the expression profile of tumour cell proteins [44, 45]. In addition, the frequent loss of TRAF3, a gene involved in antiviral immunity in HPV-positive HNSCC likely contributes to immune evasion [46, 47]. More comprehensive studies are needed in this space to understand the immune cell regulation associated with HPV infection which would shed new light for the identification of treatment strategies against HPV-positive HNSCC.
Genetic alterations
Cumulative evidence has reported a large difference between the mutational signature of HPV-negative and positive tumours, due to E6 and E7 inhibitory functions of key tumour suppressor genes TP53 and RB1. The key distinctions between HPV-negative and HPV-positive HNSCC are as follows: HPV-negative HNSCCs exhibit a high prevalence of mutations in CDKN2A, TP53, FAT1, NOTCH1, CASP8, HRAS and loss of chromosome region 3p. On the other hand, HPV-positive patients demonstrate high mutation rates in PIK3CA, ZNF750, EP300, CYLD, TRAF3, FGFR3, PTEN, B2M and RB1, along with gain of chromosome regions 3q and 19q, and loss of chromosome regions 11q, 16q and 18q. Although the top mutated genes in HPV-negative tumours are CDKN2A and TP53, loss of TRAF3 and amplification of E2F1 are the most frequently mutated genes in HPV-positive tumours [48]. Moreover, the occurrence of WGD and WGT events is significantly higher in HPV-negative cases compared to HPV-positive HNSCC. These events tend to happen earlier in the development of HPV-positive HNSCC [49]. This stark contrast suggests that the initiation and progression of HNSCC are mechanistically different depending on the presence or absence of HPV infection.
In contrast to the HPV-positive tumours, the chronological order of genetic events in HPV-negative HNSCC has been well-documented owing to available premalignant and malignant lesions throughout the steps of cancer progression from hyperplasia to invasive carcinoma [50, 51] (Fig. 2). Loss of 9p21 locus that harbours CDKN2A and ARF tumour suppressor genes occurs in the very early stages of cancer initiation from the normal mucosa to hyperplasia. These genes encode for the CDK4 and CDK6 inhibitor p16INK4A and a stabiliser of p53 protein p14. The transition to the dysplastic stage is coincident with the loss of TP53-containing locus 17p13 and 3p21. Loss of 13q21, 11q13 and 14q32 loci further facilitate the transition to carcinoma in situ, and frequent losses in chromosomes 4q27, 8, 6p and 10q23 are prevalent in invasive primary tumours [20, 48].
Major differences in the frequency of genetic lesions in HPV-negative and positive HNSCC are shown. HPV human papillomavirus, CDKN2A cyclin-dependent kinase inhibitor 2A, TP53 tumour protein 53, PTEN phosphatase and tensin homologue, PIK3CA phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit alpha.
Despite well-defined genetic changes occurring during the progression of HPV-negative HNSCC, chronological genetic alterations in HPV-positive cancers are still less understood [52]. This knowledge gap can be attributed to the challenges associated with identifying HPV-positive premalignant lesions [52, 53]. To address this issue, Leshchiner et al. employed the PhylogicNDT LeagueModel bioinformatic tool to investigate potential genetic abnormalities in 101 HPV-positive HNSCCs using whole exome sequencing (WES) data. By comparing the bioinformatic findings to alterations observed in HPV-negative cases, the authors validated the algorithm’s effectiveness in predicting pre-malignancy mutations in 421 HPV-negative HNSCCs. Of note, PhylogicNDT identified loss of 9p (CDKN2A inactivation) and 3p and mutations in TP53 as early events for HPV-negative tumour development. In addition to the sequential events detected by PhylogicNDT, whole genome duplication and triplication (WGD and WGT) were reported to occur late during the progression to HNSCC [49].
This approach was then utilised to investigate the genetic events occurring in HPV-positive HNSCC, aiming to identify the genetic alterations in early and late-stage tumour development. In HPV-positive HNSCC, specific changes such as amplification of chromosome regions 3q and 8q, coupled with the loss of chromosome regions 11q and 13q, are frequently observed in the early stages. In addition, the loss of chromosome regions 6p, 17p and 16q, as well as the amplification of chromosome regions 5p and 9q, along with mutations in the PIK3CA gene, are highly prevalent in the advanced stages of HPV-positive HNSCC progression (Fig. 2). However, this approach has not accurately predicted the specific genetic alterations that occur during the transition from hyperplasia to dysplasia, dysplasia to carcinoma in situ and invasive HNSCC.
In a more recent comprehensive study, Cheng et al. performed integrated analyses of genomic and transcriptomic profiles of 15 human HPV-negative and 11 HPV-positive HNSCC cell lines [54]. They compared their results with publicly available RNA-seq and genomics data obtained from 279 HNSCC tumours from The Cancer Genome Atlas (TCGA) [48]. In this study, the regions subjected to recurrent alterations were meaningful as they are confirmed in multiple samples, contrary to the individual patient-specific alterations discussed before. Chromosomal gains in 3q, 5p, 7p, 8q and 11q and losses in 3p, 5q, 8p, 9p, 10p, 18q and 21q were consistently detected in genomics analyses of both HNSCC TCGA and HNSCC cell lines [54].
Regardless of HPV infection, gain of 3q26–q28 harbouring PIK3CA, sex-determining region Y-box 2 (SOX2), TP63, and Telomerase RNA Component (TERC) and gain of 11q are the dominant genomic alterations. Consistent with TCGA data, a gain of chromosome 11q that harbours Fas-Associated protein with Death Domain (FADD), Cyclin D1 (CCND1), Cortactin (CTTN), Fibroblast Growth Factor (FGF3/4/19), Protein Tyrosine Phosphatase, Receptor Type, F Interacting Protein Alpha 1 (PPFIA1), Myeloma Overexpressed (MYEOV), and Anoctamin 1 (ANO1), and gain of 11q22 that contains Baculoviral IAP Repeat Containing (BIRC2/3), Yes-associated protein (YAP1), and Platelet-Derived Growth Factor D (PDGFD), and loss of 9p21 that carries CDKN2A were predominately amplified in HPV-negative lines. Copy number alterations combined with transcriptomic analyses further revealed that gains in 3q, 11q and 5p increase the expression levels of HNSCC-related oncogenes. Chromosome 3q26 loci play a pivotal role as its amplification and TP53 mutation coincidence significantly correlate with the poor prognosis of HNSCC patients [54]. These findings collectively confirm that the HNSCC cell lines recapitulate genomic alterations of more aggressive HNSCC tumour subtypes, thus the cell lines might not be the ideal models to study cancer initiation.
HPV integration is a dynamic process during the tumourigenic process
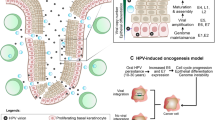
HPV integration refers to the process by which the HPV DNA becomes physically integrated into the host cell’s genome. The identification of multiple integration sites within early gains in HPV-positive HNSCC suggests that the integration process is a precursor to the genetic alterations observed during tumorigenesis. Notably, the integration events can occur as early as 25 years before the onset of primary tumours and many years before the WGD/WGT occurs. These highlight that HPV integration is the main contributor to HPV-positive cancer initiation [49]. Notwithstanding, the integration traces may or may not remain throughout the cancer development. For instance, Parfenov et al. [55] detected HPV integration in 25 of the 35 studied cases, meaning that nearly 30% of HPV-positive tumours had only HPV episomes. In addition, the detection of the viral genome or its fragments, integrated into the genome of high-grade HNSCC [55, 56] and cervical cancers [57,58,59,60] suggest that the integration process occurs more frequently in the late stages of cancer progression. In line with this, the heightened integration levels of HPV16 into the host genome of 56 TCGA HPV-positive HNSCC correlates with advanced stages of cancer and poorer survival of HNSCC patients [61]. Thus, although the integration would start many years prior to tumour formation, HPV-positive cancer cells with viral integration status may be enhanced in advanced HNSCC (Fig. 3). This is in line with the analysis of integration sites in subclonal samples by PhylogicNDT revealing that integration is a dynamic process in HPV-positive HNSCC [49]. Furthermore, Puram et al. recently studied the HPV-positive oropharyngeal SCC samples at the single-cell resolution. Analysing the HPV transcripts in subclones of malignant epithelial single cells revealed that not all HPV-positive tumour cells express HPV genes. For instance, HPV was detected in 90%, 4% and 0% of cells in three subclones. Considering all the malignant epithelial cells from 11 samples showed reads of HPV in only 66% of HPV-positive samples. Besides differences in the frequency of HPV detection, some subclones differed in the relative detection of distinct HPV genes as well. Therefore, the overall frequency and relative expression patterns of HPV genes are subject to changes during tumour evolution [62].
Following an infection with HPV, the integration process could take place in infected cells that evade the immune system and are not cleared by apoptosis. During chronic infection, infected cells can become immortalised (transformed) and by accumulating various alterations in tumour suppressors and oncogenes eventually progress to cancer. The HPV-positive pre-cancer and cancer cells harbour integrated and non-integrated HPV. The HPV-positive cells with a higher rate of viral integration are enriched at the final stages of HNSCC and correlate with a poor patient prognosis. HPV human papillomavirus, HNSCC head and neck cancer squamous cell carcinoma.
HPV integration is facilitated by DNA damage during the progression of HPV-positive HNSCC
During the process of viral DNA integration, both viral and host cell genomes undergo a molecular breakage. Akagi et al. discovered that breakpoints occurred throughout the viral genome, leading to the reduction of E2 and retainment of E6 and E7 in a mixture of HPV16-positive cervical and HNSCC tumours [63]. E2 plays a pivotal role in maintaining the life cycle of HPV by initiating DNA replication. It is also known that the E2 protein suppresses the expression and function of E6 and E7 oncoproteins [64,65,66], and induces apoptosis of infected cells [67,68,69]. Disruption of the E2 open-reading frame (ORF) is one of the main events caused by HPV integration, which eventually leads to enhanced expression of E6 and E7, thereby promoting cell growth [65, 70]. Along with this critical event, the integration process confers transforming capabilities to the host cells by stabilising the viral transcripts (produced from the integrated gene vs. episome) and also by deregulating the host genome/transcriptome depending on the integration sites [55, 71,72,73].
The rate of breakpoints in the host genome is very low (less than ten breakpoints in the genome), suggesting that a broader fragmentation in the host genome might be necessary for the full integration of the viral genome. Consistently, Parfenov et al. demonstrated that breakpoints of the host cell genome also tend to occur in regions of microhomology (1–10 bp) among the viral and host genome, and most frequently into genic or near-genic and miRNA regions [55]. Although the viral integration is linked to an increase in somatic DNA copy number of the integrated region, there is no significant association between the integration and mutational burden, indicating that HPV infection enhances the mutational burden in HNSCC regardless of integration. There is also no correlation between the occurrence of integration and the stage of cancer in the studied HPV-positive HNSCCs, which is in contrast with results showing an association between higher stages and poorer prognosis of HNSCCs and heightened levels of integration [55, 56, 61]. Consistent with other work, higher levels of E2 and E5 expression, and lower levels of E6 and E7 expression are observed in tumours without integration compared with integration-positive tumours. The overall results of these studies need further assessments as the outcomes have been generated from a small set of samples, i.e., n = 3 HPV-positive HNSCC + 2 HPV-positive cervical lines + 2 HPV-positive primary HNSCC tumours in Akagi et al.’s; and n = 35 HPV-positive HNSCCs in Parfenov et al.’s studies.
HPV genome integration is associated with genetic alterations in HNSCC
The overall mutational burden in HPV-negative and -positive HNSCCs was reported to be in the same range, while many other groups have reported a higher load in HPV-negative cases [74,75,76]. Of note, in HPV-positive HNSCCs, the elevated frequency of mutations, deletions, amplifications, and structural translocations at sites of HPV integration was reported [77]. While the integration process seems to be random, Akagi et al. found HPV integrants cluster at specific regions in different cell lines. For example, the integrants cluster at chromosome 3p11 in UM-SCC-47, chromosome Xq21 in UD-SCC-2 cells, and chromosomes 3p12, 6p21 and 9q22 in SCC090 cells. This highlights that cell-specific vulnerability might depend on the genetic and epigenetic backgrounds. These alterations eventually lead to the amplification and expression of integrated E6 and E7 viral genes and thereby the genomic instability of host cells. The key cancer-related genes affected by HPV integrants are Diaphanous-related formin 2 (DIAPH2) gene rearrangements and Tumour protein 63 (TP63) gene promoter aberration and Proviral integration site for Moloney murine leukaemia virus 1 (PIM1) and Forkhead box E1 (FOXE1) gene amplification [63].
Parfenov et al. reported the integrants to propose critical mechanisms by which the integration process could initiate tumorigenesis. A virus-host genome rearrangement resulting in the disruption of viral E1, E4 and E5 and the human RecA homologue 51B (RAD51B) gene could be one potential path to induce cancer development. Insertion of the HPV16 genome to the coding regions of E26 transformation-specific 2 (ETS2), and programmed death-ligand 1 (PDL1) was proposed as the second potential mechanism. Moreover, HPV integrants upstream of nuclear receptor subfamily 4, group A, member 2 (NR4A2) gene with subsequent effects on the amplification and overexpression of this oncogene were reported as key genetic changes in one of the HPV-positive HNSCC patients. Interestingly, the expression levels of E6 and E7 were very low, suggesting that such integration-mediated genetic change can override the roles of viral oncogenes. Another interesting mode of action was related to the insertion of HPV16 in nongenic regions which results in intrachromosomal rearrangements between chromosomes 3 and 13. Such remodelling affects TP63 regulated 1 (TPRG1) and TP63 genes on chromosome 3 and the Kruppel-like factor 5 (KLF5) on chromosome 13. Integration-mediated overexpression of these genes could therefore drive tumourigenesis [55]. The HPV16 integration-mediated changes on the TP63 gene in both Akagi and Parfenov studies highlight the importance of this gene in HNSCC tumorigenesis.
Interestingly, HPV integration could also affect infected cells epigenetically [55]. Hypomethylation and overexpression of BarH-like homeobox 2 (BARX2) and Iroquois homeobox protein 1 (IRX4) genes were shown in HPV-positive with integration status. These genes were shown to contribute to tumour formation [78, 79]. Conversely, hypermethylation and downregulation of Single-minded family bHLH transcription factor 2 (SIM2) and Cathepsin E (CTSE) were depicted in these cells. These genes are also shown to contribute to tumour formation [80, 81].
In a nutshell, the HPV virus integration process is a random yet cell-specific process leading to increased genetic alterations at the integration sites. Depending on the integration site, affected tumour suppressor genes could be downregulated/inactivated or affected oncogenes upregulated/activated. The integration process could also impact the epigenome and confer oncogenic properties. Integration-induced genetic and epigenetic changes are important determinants to what extent the pre-cancer cells require the viral oncoproteins E6 and E7 to drive the tumourigenesis process. Thus, the function of these oncogenes might not be required in all HNSCC patients.
Consistent with this postulation and, as mentioned earlier, the characterisation of oropharyngeal SCC tumours at single-cell resolution has unveiled the presence of a distinct subset of HPV-positive cells in which the expression of HPV is either lost or diminished. These cells are referred to as HPVoff, in contrast to the HPV-positive cells that continue to express E6 and E7 oncoproteins, referred to as HPVon. The similar genomic copy number of HPV genes between HPVon and HPVoff cells indicates that epigenetic regulation controls HPV gene expression in on and off subclones. This was concluded based on in vitro results following the inhibition of enhancer of zeste 2 polycomb repressive complex 2 subunit (EZH2) and DNA methyltransferase (DNMT) that result in the reduction of HPV gene expression [62]. While the findings were described in cancer cell lines rather than a progressive HNSCC model, the possibility of losing the HPV genomic content during cancer progression is not ruled out.
Intriguingly, functional enrichment analysis revealed a significant association of G1/S with HPVon cells compared to both HPVoff and HPV-negative cells. Conversely, HPVoff cells exhibited an enrichment of the epithelial senescence-associated (EpiSen) meta-programme compared to HPVon cells [62]. Consistent outcomes were concluded by analysing the bulk TCGA and HPV-positive cells (93VU147T and SCC47), supporting the concept that HPVon cells have higher proliferation and less senescence compared to HPVoff and HPV-negative cells. Importantly, HPVoff tumours also tend to have reduced recurrence-free survival. Further in vitro analysis of the HPVon and HPVoff subclones of 93VU147T and SCC47 revealed higher sensitivity of HPVon cells to therapeutic agents such as Cisplatin and radiation. HPVoff cells, however, showed treatment resistance and tend to migrate more significantly [62].
These findings suggest that HPVon may drive tumourigenesis by destabilising the genome and inhibiting key tumour suppressors such as p53 and Rb1. Consequently, a subset of HPV-positive malignant cells undergoes a loss of active transcription state, resulting in the suppression of hyperproliferative characteristics. This phenomenon ultimately confers resistance to conventional treatments for HPV-positive HNSCC, increases the risk of metastasis and worsens the prognosis for patients.
Prognosis of HPV-positive HNSCC patients
Survival rates for HNSCC patients have seen modest improvement in the last 30 years. An analysis spanning numerous HNSCC subgroups revealed better survival rates across different age groups, except for patients over the age of 75, while survival rates have improved for most anatomical sites. The overall increase in survival can be partially attributed to the emergence of HPV-associated HNSCC, a subgroup that benefits from a better prognosis [20, 82].
With the HPV-positive HNSCC, a subset exhibits extremely unfavourable treatment outcomes similar to HPV-negative patients. Only a limited number of studies have provided insights into comprehending the molecular basis of this phenomenon. The presence of lymphocyte infiltration in HPV-positive tumours has demonstrated superior outcomes, while the absence of local infiltration by these cytotoxic cells mirrors the unfavourable survival observed in HPV-negative cases [37].
By single-cell analysis of immune and non-immune cell interactions in head and neck tumours, Kurten et al. showed no clear association between HPV status and inflammation, where among 6 HPV-positive samples, one was in the low inflammation, two were in the medium, and three were in the high group, confirming the heterogeneity of HPV-positive tumours and their microenvironment [83]. Nevertheless, it remains uncertain whether these patients have an insufficient adaptive immune response or, more likely, alterations within HNSCC with HPV integration (such as TRAF3 and PIK3CA mutations) facilitate the evasion of tumour cells from the immune system. This is speculated to impart resistance against therapy and decreased survival rates for these patients.
The impact of HPV integration on reduced survival rates could be bolstered by reports indicating the prevalence of HPV genome integration within the genomes of high-grade HNSCCs [55, 56] and cervical cancers [57,58,59,60]. More importantly, elevated integration levels of HPV are reported to correlate with poorer survival of HNSCC patients [61]. These findings suggest that a higher frequency of integration, along with its downstream effects on the transcriptome and proteome of the host cell, could potentially alter the tumour microenvironment and promote immune evasion (Fig. 4).
Furthermore, there appears to be a correlation between HPV integration and the HPVon/HPVoff status presented by Puram et al. [62]. In fact, the link established between HPVoff status and reduced survival outcomes could potentially be attributed to increased integration events. Consistently, Zhang et al. demonstrated that an HPV subgroup with elevated levels of keratinocyte differentiation and oxidation-reduction process (HPV-KRT) has higher levels of viral integration and lower expression of viral genes E2/E4/E5, compared to the other HPV-positive samples with strong immune response and mesenchymal differentiation (HPV-IMU) [84]. This highlights the link between the integration status, expression states of the viral genes, and immune cell infiltration. To test this proposed hypothesis shown in Fig. 4, it would be pivotal to conduct an analysis on a large cohort of HPV-positive HNSCC samples with varying prognoses. The investigation would involve tracking the HPV genomics and transcriptomics content to determine the extent of integration in association with clinical outcomes. This can be complemented by tracing the TILs in samples with good or bad prognoses.
Signalling pathways associated with HPV integration
Over the last few decades, there has been a notable surge in the incidence of HPV-positive HNSCCs, particularly oropharyngeal cancers. Besides, the prevalence of HPV-positive HNSCC is higher among younger individuals, imposing a greater economic, medical, and psychological burden on societies. Given that the integration of the HPV genome is recognised as a primary event in HPV-positive tumour development, it is crucial to identify and target the molecular pathways that control the integration process. Moreover, such an approach can enhance the survival of HPV-positive HNSCC patients by preventing additional integrations and maintaining the tumour cells in an episomal state, likely resembling an HPNon status. This strategy would help preserve the sensitivity of HPV-positive HNSCC to standard therapeutic interventions.
The HPV genome integration into the host’s genome is still poorly understood. The current knowledge strongly suggests that DNA damage contributes to HPV integration. Eukaryotic cells sense the DNA damage and repair it mainly by Ataxia telangiectasia mutated (ATM), Ataxia telangiectasia and Rad3-related (ATR), and DNA protein kinases (Fig. 5). While ATM is activated in response to double-stranded breaks (DSBs), ATR responds to single-stranded lesions. DSBs can occur in chromosomal DNA due to replication fork collapse, programmed rearrangements, physical stress, or exposure to damaging agents like reactive oxygen and nitrogen species (ROS and RNS). These lesions pose a threat to cell viability and can lead to chromosomal translocations and genomic instability. DSBs can be repaired through two main pathways being the non-homologous end-joining (NHEJ) or homologous recombination (HR), depending on the availability of repair templates and the cell cycle stage [85, 86]. HR relies on the functions of the Rad51 family of proteins, while NHEJ is initiated by the association of Ku70/80 proteins with DNA ends and the recruitment of the DNA-dependent protein kinases [87]. Auto-phosphorylation of these kinases allows their release from DNA, enabling the processing of DNA ends for repair by DNA ligase IV and X-ray repair cross-complementing group (XRCC4).
HPV infection could manipulate the normal DNA repair systems in the cells and thus enhance the level of chromosomal instability and DNA breakage. This phenomenon ultimately increases the rate of HPV integration into the host genome. HPV human papillomavirus, RAD DNA repair protein, MRE11 meiotic recombination 11 homologue A, NBS1 Nijmegen breakage syndrome 1, ATM ataxia telangiectasia mutated, Chk checkpoint kinase, 53BP1 p53-binding protein 1, MDC1 mediator of DNA damage checkpoint protein 1, RPA replication protein A, HUS haemolytic-uraemic syndrome, ATRIP ATR-interacting protein, ATR ataxia telangiectasia and Rad3-related protein, TOPBP1 DNA topoisomerase 2-binding protein 1.
The Mre11-Rad50-Nbs (MRN) complex is considered a sensor for DSBs [88]. It is crucial for effective activation of ATM, which undergoes auto-phosphorylation and then signals to numerous downstream targets involved in checkpoints and repair mechanisms [89]. The DNA damage cascade phosphorylates various proteins, including mediators (53BP1 and Mdc1) and effectors (Chk1 and Chk2) of checkpoint responses. One of the initial proteins to be phosphorylated upon DNA damage is the histone variant H2AX, which acts as a signal for recruiting DNA damage proteins to DSBs [90]. ATR is specifically activated during the S phase to regulate the firing of origins and repair damaged replication forks [91]. ATR is recruited to sites of single-stranded DNA (ssDNA) damage through its association with ATR-interacting protein (ATRIP), which recognises the RPA complex coating ssDNA. The kinase activity of ATR-ATRIP is stimulated by TopBP1 [92]. Upon recruitment, ATR phosphorylates substrates, including Chk1 for checkpoint activation, and RPA32, Smc1, and Rad9 to facilitate repair processes [93]. Viral interactions with components of these networks are likely to unveil regulatory processes and determine the pathways exploited or inactivated. For instance, silencing a key regulator of NHEJ, Ku70, resulted in the generation of double-strand DNA breaks in episomal W12 cells and was shown to induce new HPV16 viral integration events [94]. Consistently, Someya et al. reported a significantly lower activity of DNA-dependent protein kinases Ku70 and Ku86 in cervical cancer patients compared to normal volunteers [95]. These findings suggest that enhancing DSBs is a primary mechanism HPV employs to integrate into the host cell genome.
The most deregulated pathway in HNSCC, i.e., PI3K-AKT-mTOR, could be another bona fide target to prevent viral integration. In HNSCC tumours, genetic alterations in components of this pathway are frequently observed [20, 48, 96]. PIK3CA, the catalytic domain of PI3K kinase undergoes mutations and gene amplifications at high frequency in HNP-positive HNSCC. Furthermore, loss of function in PTEN, which negatively regulates the PI3K-AKT signalling, is identified in numerous HNSCC tumours, resulting from both genetic and epigenetic changes [20, 97]. A systematic review of 95 HNSCC patients presented that 39% of cases with PIK3CA mutations are associated with nearly 5 times increase in the risk of recurrence or death in HPV-positive patients [98]. Furthermore, it is reported that the HPV16 E7 protein can promote the phosphorylation of Akt, even in the absence of PTEN downregulation. E7 is known to physically bind and sequester the PP2A protein, which in turn maintains the activation of the PI3K/Akt signalling pathway [99]. On the other hand, E6 protein can activate mTOR complex 1 leading to oncogenic activation of the PI3K-AKT-mTOR pathway [100]. Although there are no reports elaborating on the exact mechanism by which the PI3K-AKT-mTOR pathway causes HPV integration, it appears that constitutive activation of this oncogenic pathway maintains cellular transformation and increases the rate of DNA breakages over multiple uncontrolled cell divisions.
Future directions and perspectives
HPV-positive HNSCC cases are expected to continue to rise. HPV-positive patients generally exhibit improved survival rates compared to HPV-negative patients, but a subset of HPV-positive HNSCC patients with high HPV integration levels experience poorer prognostic outcomes. The HPV integration is a dynamic process occurring years before tumour formation which leads to genetic alterations, potentially activating oncogenes and inactivating tumour suppressors. The integration-induced genomic instability and altered host gene expression play a key role in the initiation and sustained promotion of cancer growth.
Furthermore, dynamic HPV integration plays a pivotal role in shaping the diverse expression profiles of HPV genes within the host cell. Typically, HPV integration events have been associated with the disruption of the viral E2 gene and the subsequent overexpression of oncogenic E6 and E7 genes at the initial stage of HPV-positive tumour development. As the tumour evolves over time, increasing genetic changes are driven by heightened HPV integration levels which further promote viral gene expression. This phenomenon can result in the loss of sensitivity to standard therapies, allowing the tumour to persist and recur.
Despite HPV-positive HNSCC being characterised as immune-hot, the process of HPV integration can significantly impact the tumour microenvironment. While oncogenic viral genes E6 and E7 are highly expressed, viral genes responsible for viral replication and transcription, such as E1^4 and E2, are either absent or expressed at low levels in HNSCC with HPV integration. More importantly, viral antigen E2 is known to strongly trigger the activation of antiviral antibody-secreting cells and T cells, responsible for viral clearance. This suggests that loss of E2 during HPV integration contributes to the immune evasion of HPV-positive tumour cells.
It is now clear that HPV integration is responsible for de-novo genetic alterations, diverse HPV gene expression patterns, and a tumour-permissive microenvironment, ultimately contributing to the heterogeneity of HPV-positive HNSCC. Given the significant role of HPV integration in the development and progression of HNSCC, targeting this process holds significant therapeutic implications. Inhibiting HPV integration could maintain the episomal state of the viral genome, reducing the oncogenic effects of integrated E6 and E7 oncogenes, and preserving the sensitivity of tumour cells to standard therapies.
One potential therapeutic strategy to inhibit HPV integration into the host genome involves targeting the DNA repair mechanism. Genes essential for the repair of DNA DSB are frequently lost or highly mutated in HNSCC with integrated HPV, proposing vulnerabilities of these tumours to therapies that exploit DNA repair defects, such as Poly(ADP-ribose) polymerase (PARP) inhibitors. Indeed, clinical trials are currently investigating the use of PARP inhibitor Olaparib as radiosensitisers to enhance local control in patients with advanced HPV-positive HNSCC, highlighting the significance of understanding the intricate interactions between HPV integration and DNA repair pathways towards developing novel treatment strategies for HPV-positive cancers.
Overall, fundamental research and preclinical studies aiming to better understand the molecular mechanisms of HPV integration and its impact on HNSCC progression are urgently needed. Ultimately, therapeutic strategies that prevent HPV integration may offer promising opportunities for heterogeneous HPV-positive HNSCC, consequently improving the management and outcomes for HPV-positive HNSCC patients.
Data availability
Not applicable.
References
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: Cancer J Clin. 2021;71:209–49.
WHO. Internation Agency for Research on Cancer. Lyon: WHO; 2023.
Ferlay J, Colombet M, Soerjomataram I, Mathers C, Parkin DM, Piñeros M, et al. Estimating the global cancer incidence and mortality in 2018: GLOBOCAN sources and methods. Int J Cancer. 2019;144:1941–53.
Velleuer E, Dietrich R. Fanconi anemia: young patients at high risk for squamous cell carcinoma. Mol Cell Pediatr. 2014;1:1–6.
Georgy SR, Cangkrama M, Srivastava S, Partridge D, Auden A, Dworkin S, et al. Identification of a novel proto-oncogenic network in head and neck squamous cell carcinoma. J Natl Cancer Inst. 2015;107:djv152.
Bai Y, Zhao Z, Boath J, van Denderen BJ, Darido C. The functional GRHL3-filaggrin axis maintains a tumor differentiation potential and influences drug sensitivity. Mol Ther. 2021;29:2571–82.
Bai Y, Gotz C, Chincarini G, Zhao Z, Slaney C, Boath J, et al. YBX1 integration of oncogenic PI3K/mTOR signalling regulates the fitness of malignant epithelial cells. Nat Commun. 2023;14:1591.
Gormley M, Creaney G, Schache A, Ingarfield K, Conway DI. Reviewing the epidemiology of head and neck cancer: definitions, trends and risk factors. Br Dent J. 2022;233:780–6.
Louie KS, Mehanna H, Sasieni P. Trends in head and neck cancers in England from 1995 to 2011 and projections up to 2025. Oral Oncol. 2015;51:341–8.
Chesson HW, Dunne EF, Hariri S, Markowitz LE. The estimated lifetime probability of acquiring human papillomavirus in the United States. Sex Transm Dis. 2014;41:660.
de Martel C, Georges D, Bray F, Ferlay J, Clifford GM. Global burden of cancer attributable to infections in 2018: a worldwide incidence analysis. Lancet Glob Health. 2020;8:e180–e90.
De Martel C, Ferlay J, Franceschi S, Vignat J, Bray F, Forman D, et al. Global burden of cancers attributable to infections in 2008: a review and synthetic analysis. Lancet Oncol. 2012;13:607–15.
Forman D, de Martel C, Lacey CJ, Soerjomataram I, Lortet-Tieulent J, Bruni L, et al. Global burden of human papillomavirus and related diseases. Vaccine. 2012;30:F12–F23.
Buchanan TR, Graybill WS, Pierce JY. Morbidity and mortality of vulvar and vaginal cancers: impact of 2-, 4-, and 9-valent HPV vaccines. Hum Vaccines Immunother. 2016;12:1352–6.
Asiaf A, Ahmad ST, Mohammad SO, Zargar MA. Review of the current knowledge on the epidemiology, pathogenesis, and prevention of human papillomavirus infection. Eur J Cancer Prev. 2014;23:206–24.
De Martel C, Plummer M, Vignat J, Franceschi S. Worldwide burden of cancer attributable to HPV by site, country and HPV type. Int J cancer. 2017;141:664–70.
Stein AP, Saha S, Kraninger JL, Swick AD, Yu M, Lambertg PF, et al. Prevalence of human papillomavirus in oropharyngeal cancer: a systematic review. Cancer J. 2015;21:138.
Isayeva T, Li Y, Maswahu D, Brandwein-Gensler M. Human papillomavirus in non-oropharyngeal head and neck cancers: a systematic literature review. Head Neck Pathol. 2012;6:104–20.
Michaud DS, Langevin SM, Eliot M, Nelson HH, Pawlita M, McClean MD, et al. High‐risk HPV types and head and neck cancer. Int J Cancer. 2014;135:1653–61.
Johnson DE, Burtness B, Leemans CR, Lui VWY, Bauman JE, Grandis JR. Head and neck squamous cell carcinoma. Nat Rev Dis Prim. 2020;6:92.
Yousefi Z, Aria H, Ghaedrahmati F, Bakhtiari T, Azizi M, Bastan R, et al. An update on human papilloma virus vaccines: history, types, protection, and efficacy. Front Immunol. 2022;12:6036.
Rosenblum HG, Lewis RM, Gargano JW, Querec TD, Unger ER, Markowitz LE. Human papillomavirus vaccine impact and effectiveness through 12 years after vaccine introduction in the United States, 2003 to 2018. Ann Intern Med. 2022;175:918–26.
Zur Hausen H. Papillomaviruses and cancer: from basic studies to clinical application. Nat Rev Cancer. 2002;2:342–50.
Van Doorslaer K, Chen Z, Bernard H-U, Chan PK, DeSalle R, Dillner J, et al. ICTV virus taxonomy profile: papillomaviridae. J Gen Virol. 2018;99:989–90.
Urbanelli L, Buratta S, Tancini B, Sagini K, Delo F, Porcellati S, et al. The role of extracellular vesicles in viral infection and transmission. Vaccines. 2019;7:102.
Aydin I, Weber S, Snijder B, Samperio Ventayol P, Kühbacher A, Becker M, et al. Large scale RNAi reveals the requirement of nuclear envelope breakdown for nuclear import of human papillomaviruses. PLoS Pathog. 2014;10:e1004162.
Zhang W, Kazakov T, Popa A, DiMaio D. Vesicular trafficking of incoming human papillomavirus 16 to the Golgi apparatus and endoplasmic reticulum requires γ-secretase activity. MBio. 2014;5:e01777–14.
Pol SBV, Klingelhutz AJ. Papillomavirus E6 oncoproteins. Virology. 2013;445:115–37.
Roman A, Munger K. The papillomavirus E7 proteins. Virology. 2013;445:138–68.
Bharadwaj M, Hussain S, Tripathi R, Singh N, Mehrotra R. Human papillomavirus (HPV): diagnosis and treatment. In: Verma AS, Singh A, editors. Animal biotechnology. Amsterdam: Elsevier; 2014. p. 95–120.
Oh K-J, Kalinina A, Bagchi S. Destabilization of Rb by human papillomavirus E7 is cell cycle dependent: E2-25K is involved in the proteolysis. Virology. 2010;396:118–24.
O’Connor M, Chan S-Y, Bernard H-U. Transcription factor binding sites in the long control region of genital HPVs. Hum Papillomaviruses. 1995;21–40.
Li N, Lam WH, Zhai Y, Cheng J, Cheng E, Zhao Y, et al. Structure of the origin recognition complex bound to DNA replication origin. Nature. 2018;559:217–22.
Yu L, Majerciak V, Zheng Z-M. HPV16 and HPV18 genome structure, expression, and post-transcriptional regulation. Int J Mol Sci. 2022;23:4943.
Partlová S, Bouček J, Kloudová K, Lukešová E, Zábrodský M, Grega M, et al. Distinct patterns of intratumoral immune cell infiltrates in patients with HPV-associated compared to non-virally induced head and neck squamous cell carcinoma. Oncoimmunology. 2015;4:e965570.
Mandal R, Şenbabaoğlu Y, Desrichard A, Havel JJ, Dalin MG, Riaz N, et al. The head and neck cancer immune landscape and its immunotherapeutic implications. JCI Insight. 2016;1:e89829.
Ward M, Thirdborough S, Mellows T, Riley C, Harris S, Suchak K, et al. Tumour-infiltrating lymphocytes predict for outcome in HPV-positive oropharyngeal cancer. Br J Cancer. 2014;110:489–500.
Canning M, Guo G, Yu M, Myint C, Groves MW, Byrd JK, et al. Heterogeneity of the head and neck squamous cell carcinoma immune landscape and its impact on immunotherapy. Front Cell Dev Biol. 2019;7:52.
Costa NL, Valadares MC, Souza PPC, Mendonça EF, Oliveira JC, Silva TA, et al. Tumor-associated macrophages and the profile of inflammatory cytokines in oral squamous cell carcinoma. Oral Oncol. 2013;49:216–23.
Ferris RL, Hunt JL, Ferrone S. Human leukocyte antigen (HLA) class I defects in head and neck cancer: molecular mechanisms and clinical significance. Immunol Res. 2005;33:113–33.
Ferris RL, Whiteside TL, Ferrone S. Immune escape associated with functional defects in antigen-processing machinery in head and neck cancer. Clin Cancer Res. 2006;12:3890–5.
Ferris RL, Blumenschein G Jr, Fayette J, Guigay J, Colevas AD, Licitra L, et al. Nivolumab for recurrent squamous-cell carcinoma of the head and neck. New Engl J Med. 2016;375:1856–67.
Seiwert TY, Burtness B, Mehra R, Weiss J, Berger R, Eder JP, et al. Safety and clinical activity of pembrolizumab for treatment of recurrent or metastatic squamous cell carcinoma of the head and neck (KEYNOTE-012): an open-label, multicentre, phase 1b trial. Lancet Oncol. 2016;17:956–65.
DiMaio D, Petti LM. The E5 proteins. Virology. 2013;445:99–114.
Grabowska AK, Riemer AB. Suppl 2: the invisible enemy–how human papillomaviruses avoid recognition and clearance by the host immune system. Open Virol J. 2012;6:249.
Karim R, Tummers B, Meyers C, Biryukov JL, Alam S, Backendorf C, et al. Human papillomavirus (HPV) upregulates the cellular deubiquitinase UCHL1 to suppress the keratinocyte’s innate immune response. PLoS Pathog. 2013;9:e1003384.
Gu Z, Shi W, Zhang L, Hu Z, Xu C. USP19 suppresses cellular type I interferon signaling by targeting TRAF3 for deubiquitination. Future Microbiol. 2017;12:767–79.
Network CGA. Comprehensive genomic characterization of head and neck squamous cell carcinomas. Nature. 2015;517:576.
Leshchiner I, Mroz EA, Cha J, Rosebrock D, Spiro O, Bonilla-Velez J, et al. Inferring early genetic progression in cancers with unobtainable premalignant disease. Nat Cancer. 2023;4:550–63.
Pai SI, Westra WH. Molecular pathology of head and neck cancer: implications for diagnosis, prognosis, and treatment. Annu Rev Pathol: Mechan Dis. 2009;4:49–70.
Califano J, Van Der Riet P, Westra W, Nawroz H, Clayman G, Piantadosi S, et al. Genetic progression model for head and neck cancer: implications for field cancerization. Cancer Res. 1996;56:2488–92.
Gillison ML, Akagi K, Xiao W, Jiang B, Pickard RK, Li J, et al. Human papillomavirus and the landscape of secondary genetic alterations in oral cancers. Genome Res. 2019;29:1–17.
Chae J, Park WS, Kim MJ, Jang SS, Hong D, Ryu J, et al. Genomic characterization of clonal evolution during oropharyngeal carcinogenesis driven by human papillomavirus 16. BMB Rep. 2018;51:584.
Cheng H, Yang X, Si H, Saleh AD, Xiao W, Coupar J, et al. Genomic and transcriptomic characterization links cell lines with aggressive head and neck cancers. Cell Rep. 2018;25:1332–45. e5.
Parfenov M, Pedamallu CS, Gehlenborg N, Freeman SS, Danilova L, Bristow CA, et al. Characterization of HPV and host genome interactions in primary head and neck cancers. Proc Natl Acad Sci USA. 2014;111:15544–9.
Begum S, Cao D, Gillison M, Zahurak M, Westra WH. Tissue distribution of human papillomavirus 16 DNA integration in patients with tonsillar carcinoma. Clin Cancer Res. 2005;11:5694–9.
Cullen AP, Reid R, Campion M, Lörincz A. Analysis of the physical state of different human papillomavirus DNAs in intraepithelial and invasive cervical neoplasm. J Virol. 1991;65:606–12.
Daniel B, Mukherjee G, Seshadri L, Vallikad E, Krishna S. Changes in the physical state and expression of human papillomavirus type 16 in the progression of cervical intraepithelial neoplasia lesions analysed by PCR. J Gen Virol. 1995;76:2589–93.
Klaes R, Woerner SM, Ridder R, Wentzensen N, Duerst M, Schneider A, et al. Detection of high-risk cervical intraepithelial neoplasia and cervical cancer by amplification of transcripts derived from integrated papillomavirus oncogenes. Cancer Res. 1999;59:6132–6.
Pirami L, Giache V, Becciolini A. Analysis of HPV16, 18, 31, and 35 DNA in pre-invasive and invasive lesions of the uterine cervix. J Clin Pathol. 1997;50:600–4.
Nulton TJ, Kim NK, DiNardo LJ, Morgan IM, Windle B. Patients with integrated HPV16 in head and neck cancer show poor survival. Oral Oncol. 2018;80:52–5.
Puram SV, Mints M, Pal A, Qi Z, Reeb A, Gelev K, et al. Cellular states are coupled to genomic and viral heterogeneity in HPV-related oropharyngeal carcinoma. Nat Genet. 2023;55:640–50.
Akagi K, Li J, Broutian TR, Padilla-Nash H, Xiao W, Jiang B, et al. Genome-wide analysis of HPV integration in human cancers reveals recurrent, focal genomic instability. Genome Res. 2014;24:185–99.
Fields BN, Knipe DM, Howley PM, Everiss KD, Kung H-J. Fundamental virology. PA: Lippincott-Raven Philadelphia; 1996.
Romanczuk H, Howley PM. Disruption of either the E1 or the E2 regulatory gene of human papillomavirus type 16 increases viral immortalization capacity. Proc Natl Acad Sci USA. 1992;89:3159–63.
Pal A, Kundu R. Human papillomavirus E6 and E7: the cervical cancer hallmarks and targets for therapy. Front Microbiol. 2020;10:3116.
Jamal DF, Rozaimee QA, Osman NH, Mohd Sukor A, Elias MH, Shamaan NA, et al. Human papillomavirus 16 E2 as an apoptosis-inducing protein for cancer treatment: a systematic review. Int J Mol Sci. 2022;23:12554.
Parish JL, Kowalczyk A, Chen H-T, Roeder GE, Sessions R, Buckle M, et al. E2 proteins from high-and low-risk human papillomavirus types differ in their ability to bind p53 and induce apoptotic cell death. J Virol. 2006;80:4580–90.
Burns JE, Walker HF, Schmitz C, Maitland NJ. Phenotypic effects of HPV-16 E2 protein expression in human keratinocytes. Virology. 2010;401:314–21.
Thierry F, Yaniv M. The BPV1‐E2 trans‐acting protein can be either an activator or a repressor of the HPV18 regulatory region. EMBO J. 1987;6:3391–7.
Jeon S, Lambert PF. Integration of human papillomavirus type 16 DNA into the human genome leads to increased stability of E6 and E7 mRNAs: implications for cervical carcinogenesis. Proc Natl Acad Sci USA. 1995;92:1654–8.
Lehoux M, D’Abramo CM, Archambault J. Molecular mechanisms of human papillomavirus-induced carcinogenesis. Public Health Genomics. 2009;12:268–80.
Groves IJ, Drane EL, Michalski M, Monahan JM, Scarpini CG, Smith SP, et al. Short-and long-range cis interactions between integrated HPV genomes and cellular chromatin dysregulate host gene expression in early cervical carcinogenesis. PLoS Pathog. 2021;17:e1009875.
Agrawal N, Frederick MJ, Pickering CR, Bettegowda C, Chang K, Li RJ, et al. Exome sequencing of head and neck squamous cell carcinoma reveals inactivating mutations in NOTCH1. Science. 2011;333:1154–7.
Mountzios G, Rampias T, Psyrri A. The mutational spectrum of squamous-cell carcinoma of the head and neck: targetable genetic events and clinical impact. Ann Oncol. 2014;25:1889–900.
Kim HAJ, Zeng PY, Shaikh MH, Mundi N, Ghasemi F, Di Gravio E, et al. All HPV-negative head and neck cancers are not the same: analysis of the TCGA dataset reveals that anatomical sites have distinct mutation, transcriptome, hypoxia, and tumor microenvironment profiles. Oral Oncol. 2021;116:105260.
Labarge B, Hennessy M, Zhang L, Goldrich D, Chartrand S, Purnell C, et al. Human papillomavirus integration strictly correlates with global genome instability in head and neck cancer. Mol Cancer Res. 2022;20:1420–8.
Sellar GC, Li L, Watt KP, Nelkin BD, Rabiasz GJ, Stronach EA, et al. BARX2 induces cadherin 6 expression and is a functional suppressor of ovarian cancer progression. Cancer Res. 2001;61:6977–81.
Ha Nguyen H, Takata R, Akamatsu S, Shigemizu D, Tsunoda T, Furihata M, et al. IRX4 at 5p15 suppresses prostate cancer growth through the interaction with vitamin D receptor, conferring prostate cancer susceptibility. Hum Mol Genet. 2012;21:2076–85.
Lu B, Asara JM, Sanda MG, Arredouani MS. The role of the transcription factor SIM2 in prostate cancer. PLoS ONE. 2011;6:e28837.
Konno-Shimizu M, Yamamichi N, Inada K-i, Kageyama-Yahara N, Shiogama K, Takahashi Y, et al. Cathepsin E is a marker of gastric differentiation and signet-ring cell carcinoma of stomach: a novel suggestion on gastric tumorigenesis. PLoS ONE. 2013;8:e56766.
Chaturvedi AK, Engels EA, Pfeiffer RM, Hernandez BY, Xiao W, Kim E, et al. Human papillomavirus and rising oropharyngeal cancer incidence in the United States. J Clin Oncol. 2011;29:4294.
Kürten CH, Kulkarni A, Cillo AR, Santos PM, Roble AK, Onkar S, et al. Investigating immune and non-immune cell interactions in head and neck tumors by single-cell RNA sequencing. Nat Commun. 2021;12:7338.
Zhang Y, Koneva LA, Virani S, Arthur AE, Virani A, Hall PB, et al. Subtypes of HPV-Positive head and neck cancers are associated with HPV characteristics, copy number alterations, PIK3CA mutation, and pathway signatures. Clin Cancer Res. 2016;22:4735–45.
Shrivastav M, De Haro LP, Nickoloff JA. Regulation of DNA double-strand break repair pathway choice. Cell Res. 2008;18:134–47.
Branzei D, Foiani M. Regulation of DNA repair throughout the cell cycle. Nat Rev Mol Cell Biol. 2008;9:297–308.
Sancar A, Lindsey-Boltz LA, Ünsal-Kaçmaz K, Linn S. Molecular mechanisms of mammalian DNA repair and the DNA damage checkpoints. Annu Rev Biochem. 2004;73:39–85.
Petrini JH, Stracker TH. The cellular response to DNA double-strand breaks: defining the sensors and mediators. Trends Cell Biol. 2003;13:458–62.
Lavin M. ATM and the Mre11 complex combine to recognize and signal DNA double-strand breaks. Oncogene. 2007;26:7749–58.
Huen MS, Chen J. The DNA damage response pathways: at the crossroad of protein modifications. Cell Res. 2008;18:8–16.
Cimprich KA, Cortez D. ATR: an essential regulator of genome integrity. Nat Rev Mol Cell Biol. 2008;9:616–27.
Mordes DA, Glick GG, Zhao R, Cortez D. TopBP1 activates ATR through ATRIP and a PIKK regulatory domain. Genes Dev. 2008;22:1478–89.
Chaurushiya MS, Weitzman MD. Viral manipulation of DNA repair and cell cycle checkpoints. DNA Repair. 2009;8:1166–76.
Winder D, Pett M, Foster N, Shivji M, Herdman M, Stanley M, et al. An increase in DNA double‐strand breaks, induced by Ku70 depletion, is associated with human papillomavirus 16 episome loss and de novo viral integration events. J Pathol. 2007;213:27–34.
Someya M, Sakata K-i, Matsumoto Y, Yamamoto H, Monobe M, Ikeda H, et al. The association of DNA-dependent protein kinase activity with chromosomal instability and risk of cancer. Carcinogenesis. 2005;27:117–22.
Tan FH, Bai Y, Saintigny P, Darido C. mTOR signalling in head and neck cancer: heads up. Cells. 2019;8:333.
Krump NA, You J. Molecular mechanisms of viral oncogenesis in humans. Nat Rev Microbiol. 2018;16:684–98.
Cochicho D, Esteves S, Rito M, Silva F, Martins L, Montalvão P, et al. PIK3CA Gene mutations in HNSCC: systematic review and correlations with HPV status and patient survival. Cancers. 2022;14:1286.
Pim D, Massimi P, Dilworth SM, Banks L. Activation of the protein kinase B pathway by the HPV-16 E7 oncoprotein occurs through a mechanism involving interaction with PP2A. Oncogene. 2005;24:7830–8.
Surviladze Z, Sterk RT, DeHaro SA, Ozbun MA. Cellular entry of human papillomavirus type 16 involves activation of the phosphatidylinositol 3-kinase/Akt/mTOR pathway and inhibition of autophagy. J Virol. 2013;87:2508–17.
Bergvall M, Melendy T, Archambault J. The E1 proteins. Virology. 2013;445:35–56.
McBride AA. The papillomavirus E2 proteins. Virology. 2013;445:57–79.
McBride AA, Oliveira JG, McPhillips MG. Partitioning viral genomes in mitosis: same idea, different targets. Cell Cycle. 2006;5:1499–502.
Doorbar J. The E4 protein; structure, function and patterns of expression. Virology. 2013;445:80–98.
Doorbar J, Ely S, Sterling J, McLean C, Crawford L. Specific interaction between HPV-16 E1–E4 and cytokeratins results in collapse of the epithelial cell intermediate filament network. Nature. 1991;352:824–7.
Wang Q, Kennedy A, Das P, McIntosh PB, Howell SA, Isaacson ER, et al. Phosphorylation of the human papillomavirus type 16 E1^ E4 protein at T57 by ERK triggers a structural change that enhances keratin binding and protein stability. J Virol. 2009;83:3668–83.
McIntosh PB, Laskey P, Sullivan K, Davy C, Wang Q, Jackson DJ, et al. E1^ E4-mediated keratin phosphorylation and ubiquitylation: a mechanism for keratin depletion in HPV16-infected epithelium. J Cell Sci. 2010;123:2810–22.
Dreer M, van de Poel S, Stubenrauch F. Control of viral replication and transcription by the papillomavirus E8^ E2 protein. Virus Res. 2017;231:96–102.
Buck CB, Day PM, Trus BL. The papillomavirus major capsid protein L1. Virology. 2013;445:169–74.
Wang JW, Roden RB. L2, the minor capsid protein of papillomavirus. Virology. 2013;445:175–86.
Acknowledgements
The authors were supported by a grant from the Australian Medical Research Future Fund, Genomics Health Futures Mission (MRFF, APP2008888). The figures have been created by BioRender.
Funding
Open Access funding enabled and organized by CAUL and its Member Institutions.
Author information
Authors and Affiliations
Contributions
CD participated in the conception and edition of the manuscript. HT, YB, RH and AC participated in the perception and writing of the manuscript. HT also participated in the preparation of figures. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tabatabaeian, H., Bai, Y., Huang, R. et al. Navigating therapeutic strategies: HPV classification in head and neck cancer. Br J Cancer (2024). https://doi.org/10.1038/s41416-024-02655-1
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41416-024-02655-1