Abstract
Mammalian SWI/SNF complex is a key chromatin remodeler that reshapes nucleosomes and regulates DNA accessibility. Mutations in SWI/SNF subunits are found in a broad spectrum of human cancers; however, the mechanisms of how these aberrations of SWI/SNF complex would impact tumorigenesis and cancer therapeutics remain to be elucidated. Studies have demonstrated that immune checkpoint blockade (ICB) therapy is promising in cancer treatment. Nevertheless, suitable biomarkers that reliably predict the clinical response to ICB are still lacking. Emerging evidence has suggested that SWI/SNF components play novel roles in the regulation of anti-tumor immunity, and SWI/SNF deficiency can be therapeutically targeted by ICB. These findings manifest the prominence of the SWI/SNF complex as a stratification biomarker that predicts treatment (therapeutic) response to ICB. In this review, we summarize the recent advances in ICB therapy by harnessing the cancer-specific vulnerability elicited by SWI/SNF deficiency. We provide novel insights into a comprehensive understanding of the underlying mechanisms by which SWI/SNF functions as a modulator of anti-tumor immunity.
Similar content being viewed by others
Introduction
The mammalian switch/sucrose non-fermentable (SWI/SNF) family is a multi-subunit chromatin remodeling complex that utilizes the energy of ATP hydrolysis to remodel nucleosomes and regulates DNA accessibility in fundamental cellular processes, including transcription, DNA repair, and replication1. SWI/SNF complex mutations are frequently observed in a broad spectrum of human cancers2,3,4,5. There is increasing evidence regarding the critical biological functions of the SWI/SNF complex in cancer; however, the underlying mechanisms by which SWI/SNF components contribute to tumorigenesis or drug sensitivity warrant further investigation. Importantly, it remains unknown whether and how SWI/SNF mutations or defects could be exploited for therapeutic purposes6.
Cancer immunotherapy especially immune checkpoint blockade (ICB) has recently become one of the most prominent therapeutics for human cancers7. Briefly, the immune checkpoint constitutes a negative regulatory mechanism that maintains immune tolerance and prevents the overactivation of immune responses8. This mechanism can be hijacked by tumor cells to avoid elimination by immune cells, thus ICB therapy (e.g., anti-CTLA-4, anti-PD-1/PD-L1) was developed to counteract the immune suppressive microenvironment, thereby to strengthen the tumor-killing effects9. ICB therapy has been approved to treat multiple cancer types10,11,12,13,14,15,16,17,18. But the clinical efficacy of current ICB therapy is low; only a minority of patients may respond (<30% overall response rate)19. Furthermore, a considerable proportion of patients (25–30%) may develop immune-related adverse events or fatal outcomes20. In addition, up to two-thirds of ICB therapy recipients may exhibit either primary or acquired resistance21,22,23. These observations highlight the need for elucidation of resistance mechanisms, the proposal of more effective therapeutic strategies, and more importantly identification of effective biomarkers for stratifying cancer patients24.
While more effective than monotherapy, the combination ICB approaches are likely to exacerbate the incidence of immune-related adverse events that could greatly impact the clinical feasibility of combination therapy25. In this regard, the identification of biomarkers that predict patients who are more likely to respond to ICB therapy is of considerable importance. Recently, we and others have reported that SWI/SNF deficiency is related to sensitivity to immune checkpoint blockade (ICB) therapy, indicating the potential for use of the SWI/SNF complex as a target for cancer immunotherapy6,26,27,28,29. These findings suggest a novel role for SWI/SNF in modulating anti-tumor immunity and imply that aberrations of SWI/SNF components may serve as biomarkers to predict patient response to clinical ICB therapy. In addition, these results further support the notion that aberrations of SWI/SNF members can be therapeutically targeted30,31,32,33. The synthetic lethal effects and mechanisms of SWI/SNF subunits have been extensively reviewed elsewhere34,35,36. Here, we summarize the current understanding of mechanisms of molecular vulnerability mediated by SWI/SNF core members and the therapeutic applications in ICB.
SWI/SNF complex: a highly mutated chromatin remodeler in human cancers
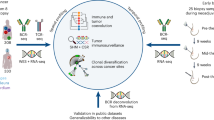
SWI/SNF complex consists of 15 subunits encoded by up to 29 genes and possessed ATP-dependent nucleosome remodeling activity (Fig. 1A)37,38. Based on the subunit composition, three major complexes in mammals: BRG1-associated factor (BAF; also known as SWI/SNF-A) complexes; polybromo BRG1-associated factor (PBAF; also known as SWI/SNF-B) complexes; and noncanonical BAF (ncBAF/GBAF) complexes. They exist in various compositions and proteins encoded by paralogous genes may alternately occupy several positions (Fig. 1B) (Table 1).
A The subunits of the SWI/SNF complexes, including catalytic ATPase with bromodomain, additional bromodomain subunits, and accessory subunit. Different colors of subunits and characters in the complex indicate the function of various subunits. B Three major complexes in mammalian SWI/SNF complex: BRG1-associated factor (BAF; also known as SWI/SNF-A) complexes; polybromo BRG1-associated factor (PBAF; also known as SWI/SNF-B) complexes; and noncanonical BAF (ncBAF/GBAF) complexes.
SWI/SNF complex mutations are frequently observed in numerous human cancers, with an average mutation rate of 20%, which ranks immediately after p53 (26%) (Table 2)2,3,4. By far, the most commonly mutated BAF subunit in cancer is ARID1A (BAF250A), the AT-rich interacting domain-containing protein 1A, which is mutated in 50% of ovarian clear-cell carcinoma (OCCC), 11% of colorectal adenocarcinoma3,39,40,41,42,43 and 9% of endometrial carcinoma44, 17.5% of colon and rectal cancers45, certain pancreatic cancer46, 13% of transitional cell carcinoma of the bladder47, 27% of gastric cancer41,48, 16.7% of cholangiocarcinoma49, and 11% of childhood neuroblastoma50. BRG1/SMARCA4 is mutationally inactivated or epigenetically silenced in non-small-cell lung cancer (NSCLC), medulloblastoma, and Burkitt’s lymphoma51,52,53. In addition, ~90% of malignant rhabdoid tumors (MRT) harbor SNF5/SMARCB1 (integrase interactor 1/SWI/SNF‐related matrix‐associated actin‐dependent regulator of chromatin subfamily B member 1) mutations54. The PBAF complex component PBRM1 (BAF180) contains six bromine domains and is mutated or deleted in >50% of clear-cell renal cell carcinoma (ccRCC)55. Overall, ARID1A is the most frequently mutated SWI/SNF subunit in different cancer types; however, PBRM1 mutations are much more common in ccRCC than ARID1A mutations6. These mutations usually cause the effects of loss-of-function (LOF) to almost all the subunits; however, these incidences have shown a cancer-dependent fashion, indicating the function of the SWI/SNF complex is context-specific56. With mutations in the SWI/SNF subunits found in ~25% of cancers, the question of whether such mutations have therapeutic implications naturally arises.
SWI/SNF core subunits as potential biomarkers of ICB therapy
The investigations of biological functions of the SWI/SNF complex are currently developing and the molecular consequences resulted from subsequent mutations associated with core complex subunits in human cancers are not yet fully understood6. Here, we focused on the core subunits of the SWI/SNF complex that could serve as functional biomarkers for ICB therapy.
PBRM1
Studies in ccRCCs patients have shown that the loss of biallelic PBRM1, a PBAF component of the SWI/SNF complex, is positively associated with better response to anti-PD-1 or anti-PD-L1 treatment, regardless of mutation burden57,58. Mutations in PBRM1, ARID2, and other SWI/SNF components are also overrepresented in complete or partial responders, compared with non-responders6,57,59. Progressive ccRCC tumors exhibit high levels of CD8+ T-cell infiltration, compared with noninvasive tumors; notably, infiltrating tumors have reduced levels of favorable PBRM1 mutations. The most commonly mutated genes (e.g., VHL, PBRM1, SETD2, BAP1, and KDM5C) were recurrent in both earlier and advanced disease58. However, the precise role of PBRM1 in the ccRCC immune microenvironment remains unclear, because contradictory results have shown that loss of PBRM1 reduced the binding of BRG1 to the interferon (IFN)-γ receptor 2 promoters; STAT1 phosphorylation and subsequent expression of IFN-γ target genes, therefore correlated with a nonimmunogenic phenotype60.
ARID1A
We have reported that ARID1A mutations positively correlate with enhanced anti-tumor immunity in both experimental models and patient samples, and ARID1A-mutant tumors are sensitized to anti-PD-L1 treatment26. Okamura et al. recently used next-generation sequencing technology to analyze 3403 patients receiving ICB therapy; they found >5% prevalence of ARID1A alterations in 1540 patients with nine distinct types of cancer61. The median progression-free survival of patients with altered ARID1A was significantly longer than that of patients with wild-type ARID1A in terms of response to ICB therapy, regardless of microsatellite instability (MSI) and tumor mutation burden (TMB) statuses. Because inhibition of histone deacetylase 6 suppresses the growth of ARID1A-mutated tumors, the combination of histone deacetylase 6 suppression and ICB represents a potential treatment strategy for ARID1A-mutated cancers62. More recently, Goswami S et al. showed that ARID1A depletion indeed correlated with improved sensitivity to anti-PD-1 therapy in a murine models of bladder cancer and in clinical cohorts. Importantly, biomarker combining ARID1A mutation and CXCL13 expression exhibits even better prediction power of patient responses to ICB29. These results indicate that ARID1A status may predict patient responses to ICB therapy. Of note, Li J et al. have reported contradictory findings that ARID1A mutations led to impaired chromatin accessibility to IFN-responsive genes, and represented a poor anti-tumor immunity63. Therefore, ARID1A alteration merits further exploration as a novel biomarker for outcomes after ICB therapy64.
SMARCB1/SNF5
Inactivation of biallelic SMARCB1/SNF5 is largely restricted to the rare pediatric rhabdoid tumors. The vast majority of the rhabdoid tumors (over 95%) harbor SMARCB1 mutations, or rarely SMARCA4 mutations (<5%)65. Experimental rhabdoid tumor models have demonstrated significant regression of established tumors under ICB therapy, up to 67% of the mice with the SMARCB1-mutant rhabdomyoma readily responded to anti-PD-1 treatment66. Though the underlying mechanisms are not fully understood, SMARCB1-dependent activation of interferon signaling was observed in the modulation of immunogenicity66. The genetic background of rhabdoid tumors and a significantly higher responding rate suggest that SMARCB1-mutant tumors can be therapeutically targeted by ICB therapy. However, further studies employing clinical cohorts are required to test the prediction ability of SMARCB1 status to ICB.
SMARCA4/BRG1
SMARCA4 deficiency reportedly correlated with improved responses to ICB therapy. A 3-year follow-up study of 126 squamous cell carcinoma of the head and neck (SCCHN) patients receiving anti-PD-1/L1 therapy demonstrated that SMARCA4 mutation and/or frameshift were more frequently observed in responders than non-responders67. Consistently, a case report of thoracic sarcoma showed that SMARCA4 deficiency resulted in notable clinical response to Nivolumab (anti-PD-1) treatment68. Of note, in a small cohort study, four small-cell carcinomas of the ovary, hypercalcemic type (SCCOHT) patients showed a notable response to anti-PD-1 immunotherapy. Given that SCCOHT is a SMARCA4 mutation-driven, highly aggressive monogenic cancer type, it may reflect the enhanced immunogenicity mediated by loss of SMARCA4 function69. Emerging data indicate that alternative epigenetic enzymes, such as lysine-specific demethylase 1 (LSD1), could induce an anti-tumor immune response in the scenario of SWI/SNF inactivation. LSD1 is highly expressed in SWI/SNF-mutant SCCOHT tumors and the inhibition of LSD1 activity exhibits remarkable anti-tumor efficacy when combined with immune checkpoint blockade70. These findings suggest the potential of LSD1 as a target for the combinational immunotherapy of SWI/SNF-mutated tumors. However, these results require studies with a larger cohort of patients and longer follow-up to validate.
ICB clinical trials targeting SWI/SNF-mutant tumors
Multiple immune checkpoint inhibitors (listed in Table 3) are currently under investigation to exploit aberrations of SWI/SNF components71: nivolumab, a fully human IgG4 PD-1 antibody; pembrolizumab (i.e., MK-3475 or lambrolizumab), a high-affinity humanized IgG4 monoclonal antibody targeting PD-1; and MPDL3280A, an engineered IgG anti-PD-L1 antibody. Notably, the feasibility of the SWI/SNF complex as a biomarker is still, to a certain extent, under debate. One recent cohort study showed that loss-of-function mutations in SWI/SNF components failed to predict improvements in overall survival, time to treatment failure, and disease control rate. However, patients harboring PBRM1 mutations exhibited significantly improved overall survival and time to treatment failure72. These results further highlight the context-dependent functions of the SWI/SNF complex, which warrant large cohort and multicancer studies.
Mechanisms of SWI/SNF-mediated immune modulation
Silencing of IFN-stimulated genes
Mutations in PBRM1 are associated with the enrichment of gene expressions that stimulate immune responses (e.g., hypoxia responses and JAK-STAT signaling) in ccRCC57. In addition, inactivation of PBRM1 sensitizes tumor cells to T-cell-mediated cytotoxicity and results in a more favorable tumor microenvironment59. An important role of PBRM1 in immune modulation is the suppression of IFN-γ-responsive gene expression, thus conferring T-cell-mediated killing resistance to tumor cells. Pan et al. have found that inactivation of PBRM1 enhances the chromatin accessibility of transcription factors on the promoters of many IFN-γ-inducible genes, therefore leads to the suboptimal response to ICB therapies59. They also discovered that decreased interferon signaling can lead to lost opportunities for the upregulation of PD-L1, an established downstream target of IFN-γ59. Moreover, PBRM1 functions as a synergistic factor with EZH2, which promotes its silencing effect upon IFN-stimulated genes; this suggests a possible mechanistic explanation for the positive relationship of PBAF loss with the reduction of IFN-stimulated gene expression73. Notably, Th1-type chemokines (e.g., CCL5, CXCL9, CXCL10, and CXCL11) are top-level genes that are differentially regulated by both EZH2 and ARID1A. Furthermore, the expression level of ARID1A was positively correlated with the levels of IFN-responsive genes63. EZH2 has been previously shown to suppress Th1-type chemokine (primarily CXCL9 and CXCL10) expression and alter effector T-cell tumor trafficking74,75. Therefore, PBRM1 also downregulates the expression of innate immune-related chemokines76.
ARID1A aberrations also have been shown to restrict the accessibility of chromatin to IFN-responsive genes, resulting in attenuated IFN gene expression and poor Th1-type chemokine expression (Fig. 2B)63. Through a synergistic effect with phosphatidylinositol 3-kinase signaling, ARID1A inhibits inflammation-driven tumorigenesis by limiting the production of interleukin-677. In addition, ARID1A interacts with EZH2, an enzymatic subunit of the Polycomb complex, through its carboxyl-terminal; this interaction antagonizes EZH2-mediated IFN reactivity63. Importantly, the loss of ARID1A may synergize with phosphatidylinositol 3-kinase/AKT signaling activation, further enhancing oncogenic signaling36.
A ARID1A modulates DNA mismatch repair and the expression of IFN-responsive genes. ARID1A restricts the chromatin accessibility of EZH2 and PRC2 complex to IFN-responsive genes. The aberration of ARID1A attenuates IFN gene expression and poor Th1-type chemokine expression (e.g., CXCL9 and CXCL10). ARID1A interacts with MMR protein MSH2 and functionally regulates MSH2 positioning at DNA mismatch sites, tumor mutation load, and the subsequent immunogenicity. B SNF5 and BRG1 modulate the expression of IFN-responsive genes through MYC and MAX. SWI/SNF subunits SNF5 and BRG1 interact with MYC and MAX, respectively, therefore alleviate the inhibitory function of MYC and MAX upon IFN-responsive genes. SNF5 interacts directly with MYC, through MYC HLH-LZ and SNF5 Rpt motifs. BRG1 interacts and regulates MAX, a functional partner of MYC.
Interactions with MYC in immune regulation
SWI/SNF complexes also interact with MYC, a well-characterized transcription factor, and master regulator of anti-tumor immune responses78,79,80. The link between MYC and epigenetic regulation is the interaction of MYC and SNF5 (SMARCB1/INI/BAF47), a core member of the SWI/SNF chromatin remodeling complex81.
Liu et al. described the negative regulation of the MYC-mediated BRD7 subunit, which is consistent with our findings regarding the co-expression correlation analysis of BRD782. The oncogene MYC is regarded as the central hub, which may regulate the expression of SWI/SNF subunits: (1) MYC protein interacts with the SWI/SNF subunit BAF47/hSNF583; (2) MYC localizes the SWI/SNF complex to the target gene locus that will undergo transcriptional regulation84; (3) the SWI/SNF complex can regulate MYC through transcription. MYC and SNF5 interact directly, through important functional domains in both proteins, including MYC HLH-LZ and SNF5 Rpt motifs85. The interaction of SNF5 with MYC impedes the DNA-binding activity of MYC at certain target genes83. In addition, BRG1 is reported to regulate MAX, a functional partner of MYC, therefore alleviate the inhibitory function of MAX and/or MYC at the chromatin region of IFN-responsive genes86.
Regulation of DNA mismatch repair
DNA mismatch repair (MMR) maintains replication fidelity by correcting mismatched nucleotides bound by DNA polymerases87,88. The loss of DNA mismatch repair activity leads to MSI, a hypermutable phenotype89. ARID1A has been reported to interact with MMR protein MSH2 and functionally regulate MSH2 positioning at DNA mismatch sites without affecting MSH2 expression. ARID1A deficiency impairs MMR efficiency and causes a mutator phenotype in both cancer cell lines and in vivo tumor samples (Fig. 2A). ARID1A deficiency is associated with the genomic features of a C > T mutation pattern and markedly elevated TMB, which are commonly observed in MMR-deficient samples. Importantly, ARID1A-deficient tumors are sensitive to anti-PD-L1 treatment in syngeneic mouse models26.
Clinical studies have shown that MSI and/or MMR deficiency sensitizes tumors to ICB therapy; therefore, MSI and MMR deficiency have received expedited the United States Food and Drug Administration approval for use as a patient stratification biomarker in the treatment of solid tumors, regardless of cancer types17,18. However, current methods for MSI determination are based on the detection of abnormalities at genomic loci or the loss of MMR proteins (e.g., MLH1); thus, functional defects in MMR may not be readily identified. For instance, trimethylation of the histone H3 lysine 36 recruits Mutsα to the replicating chromatin and facilitates MMR; defects in histone H3 lysine 36 activity result in functional impairment of MMR, rather than clinically validated MSI87. These results manifest the importance of functional biomarker of MMR (e.g., ARID1A status) to predict the responses to ICB therapy.
Conclusions
The discovery that the SWI/SNF complex plays an essential role in determining the therapeutic efficacy of cancer immunotherapy highlights several important future goals. First, we note that controversial results were reported regarding the function of SWI/SNF aberration in predicting the clinical responses to ICB therapy (e.g., ARID1A and PBRM1 alterations from different research groups), therefore further studies require longer follow-up durations and larger cohorts of patients to further determine the clinical feasibility. Second, most of the current findings are based on phenotypic observations that lack mechanistic insights. Thus, the underlying mechanisms by which SWI/SNF complex members could modulate anti-tumor immunity and/or responses to ICB therapy warrant further examination. It is crucial to determine how SWI/SNF complex-mediated chromatin remodeling could modify immune cell function in the tumor microenvironment, to elucidate whether these regulatory mechanisms of SWI/SNF could be exploited to refine the immune checkpoint networks and to identify the context-dependent binding partners of SWI/SNF that could be targeted to achieve durable ICB therapeutic effects. In addition, a single-target biomarker may lack enough prediction power. In this regard, a combinational biomarker that based on SWI/SNF complex and other markers (e.g., PD-L1, CXCL13) could be further investigated in the future.
References
Wilson, B. G. & Roberts, C. W. SWI/SNF nucleosome remodellers and cancer. Nat. Rev. Cancer 11, 481–492 (2011).
Hodges, C., Kirkland, J. G. & Crabtree, G. R. The many roles of BAF (mSWI/SNF) and PBAF complexes in cancer. Cold Spring Harb. Perspect. Med. 6, a026930 (2016).
Kadoch, C. et al. Proteomic and bioinformatic analysis of mammalian SWI/SNF complexes identifies extensive roles in human malignancy. Nat. Genet. 45, 592–601 (2013).
Workman, J. L. & Kingston, R. E. Alteration of nucleosome structure as a mechanism of transcriptional regulation. Annu. Rev. Biochem. 67, 545–579 (1998).
Shain, A. H. & Pollack, J. R. The spectrum of SWI/SNF mutations, ubiquitous in human cancers. PLoS ONE 8, e55119 (2013).
Mittal, P. & Roberts, C. W. M. The SWI/SNF complex in cancer-biology, biomarkers and therapy. Nat. Rev. Clin. Oncol. 17, 1–14 (2020).
He, X. & Xu, C. Immune checkpoint signaling and cancer immunotherapy. Cell Res. 30, 660–669 (2020).
Topalian, SuzanneL., CGDaDMP. Immune checkpoint blockade: a common denominator approach to cancer therapy. Cancer Cell 27, 450–461 (2015).
Pardoll, D. M. The blockade of immune checkpoints in cancer immunotherapy. Nat. Rev. Cancer 12, 252–264 (2012).
Robert, C. T. L. et al. Ipilimumab plus dacarbazine for previously untreated metastatic melanoma. N. Engl. J. Med. 364, 2517–2526 (2011).
Reck, M. et al. Pembrolizumab versus chemotherapy for PD-L1-positive non-small-cell lung cancer. N. Engl. J. Med. 375, 1823–1833 (2016).
Motzer, R. J. et al. Nivolumab versus everolimus in advanced renal-cell carcinoma. N. Engl. J. Med. 373, 1803–1813 (2015).
Armand, P. et al. Programmed death-1 blockade with pembrolizumab in patients with classical Hodgkin lymphoma after brentuximab vedotin failure. J. Clin. Oncol. 34, 3733–3739 (2016).
Powles, T. et al. Efficacy and safety of durvalumab in locally advanced or metastatic urothelial carcinoma: updated results from a phase 1/2 open-label study. JAMA Oncol. 3, e172411 (2017).
Ferris, R. L. et al. Nivolumab for recurrent squamous-cell carcinoma of the head and neck. N. Engl. J. Med. 375, 1856–1867 (2016).
Nghiem, P. T. et al. PD-1 blockade with pembrolizumab in advanced merkel-cell carcinoma. N. Engl. J. Med. 374, 2542–2552 (2016).
Le, D. T. et al. PD-1 blockade in tumors with mismatch-repair deficiency. N. Engl. J. Med. 372, 2509–2520 (2015).
Le, D. T. et al. Mismatch repair deficiency predicts response of solid tumors to PD-1 blockade. Science 357, 409–413 (2017).
Martins, F. et al. Adverse effects of immune-checkpoint inhibitors: epidemiology, management and surveillance. Nat. Rev. Clin. Oncol. 16, 563–580 (2019).
Suzanne, L. et al. Safety, activity, and immune correlates of Anti-PD-1 antibody in cancer. N. Engl. J. Med. 366, 2443–2454 (2012).
Julie, R. et al. Safety and activity of anti-PD-L1 antibody in patients with advanced cancer. N. Engl. J. Med. 366, 2455–2465 (2012).
Wang, Q. & Wu, X. Primary and acquired resistance to PD-1/PD-L1 blockade in cancer treatment. Int. Immunopharmacol. 46, 210–219 (2017).
Restifo, N. P., Smyth, M. J. & Snyder, A. Acquired resistance to immunotherapy and future challenges. Nat. Rev. Cancer 16, 121–126 (2016).
Yan, Y. et al. Combining immune checkpoint inhibitors with conventional cancer therapy. Front. Immunol. 9, 1739 (2018).
Larkin, J. et al. Combined nivolumab and ipilimumab or monotherapy in untreated melanoma. N. Engl. J. Med. 373, 23–34 (2015).
Shen, J. et al. ARID1A deficiency promotes mutability and potentiates therapeutic antitumor immunity unleashed by immune checkpoint blockade. Nat. Med. 24, 556–562 (2018).
Braun, D. A. et al. Clinical validation of PBRM1 alterations as a marker of immune checkpoint inhibitor response in renal cell carcinoma. JAMA Oncol. 5, 1631–1633 (2019).
Li, J. et al. Epigenetic driver mutations in ARID1A shape cancer immune phenotype and immunotherapy. J. Clin. Investig. 130, 2712–2726 (2020).
Goswami, S. et al. ARID1A mutation plus CXCL13 expression act as combinatorial biomarkers to predict responses to immune checkpoint therapy in mUCC. Sci. Transl. Med. 12, eabc4220 (2020).
Shen, J. et al. ARID1A deficiency impairs the DNA damage checkpoint and sensitizes cells to PARP inhibitors. Cancer Discov. 5, 752–767 (2015).
Bitler, B. G. et al. Synthetic lethality by targeting EZH2 methyltransferase activity in ARID1A-mutated cancers. Nat. Med. 21, 231–238 (2015).
Bitler, B. G. et al. ARID1A-mutated ovarian cancers depend on HDAC6 activity. Nat. Cell Biol. 19, 962–973 (2017).
Williamson, C. T. et al. ATR inhibitors as a synthetic lethal therapy for tumours deficient in ARID1A. Nat. Commun. 7, 13837 (2016).
Sasaki, M. & Ogiwara, H. Synthetic lethal therapy based on targeting the vulnerability of SWI/SNF chromatin remodeling complex-deficient cancers. Cancer Sci. 111, 774–782 (2020).
Chabanon, R. M., Morel, D. & Postel-Vinay, S. Exploiting epigenetic vulnerabilities in solid tumors: novel therapeutic opportunities in the treatment of SWI/SNF-defective cancers. Semin. Cancer Biol. 61, 180–198 (2020).
Fukumoto, T., Magno, E. & Zhang, R. SWI/SNF complexes in ovarian cancer: mechanistic insights and therapeutic implications. Mol. Cancer Res. 16, 1819–1825 (2018).
Kadoch, C. C. G. et al. Mammalian SWI/SNF chromatin remodeling complexes and cancer: mechanistic insights gained from human genomics. Sci. Adv 1, e1500447 (2015).
Wu, J. I., Lessard, J. & Crabtree, G. R. Understanding the words of chromatin regulation. Cell 136, 200–206 (2009).
Wiegand, K. C. ARID1A mutations in endometriosis-associated ovarian carcinomas. N. Engl. J. Med. 363, 1532–1543 (2010).
Jones, S. et al. Frequent mutations of chromatin remodeling gene ARID1A in ovarian clear cell carcinoma. Science 330, 228–231 (2010).
Wang, K. et al. Exome sequencing identifies frequent mutation of ARID1A in molecular subtypes of gastric cancer. Nat. Genet. 43, 1219–1223 (2011).
Giannakis, M. et al. Genomic correlates of immune-cell infiltrates in colorectal carcinoma. Cell Rep. 15, 857–865 (2016).
Cajuso, T. et al. Exome sequencing reveals frequent inactivating mutations in ARID1A, ARID1B, ARID2 and ARID4A in microsatellite unstable colorectal cancer. Int. J. Cancer 135, 611–623 (2014).
Cancer Genome Atlas Research Network. et al. Integrated genomic characterization of endometrial carcinoma. Nature 497, 67–73 (2013).
Network. CGA. Comprehensive molecular characterization of human colon and rectal cancer. Nature 487, 330–337 (2012).
Biankin, A. V. et al. Pancreatic cancer genomes reveal aberrations in axon guidance pathway genes. Nature 491, 399–405 (2012).
Gui, Y. et al. Frequent mutations of chromatin remodeling genes in transitional cell carcinoma of the bladder. Nat. Genet. 43, 875–878 (2011).
Wang, K. et al. Whole-genome sequencing and comprehensive molecular profiling identify new driver mutations in gastric cancer. Nat. Genet. 46, 573–582 (2014).
Chan-On, W. et al. Exome sequencing identifies distinct mutational patterns in liver fluke-related and non-infection-related bile duct cancers. Nat. Genet. 45, 1474–1478 (2013).
Sausen, M. et al. Integrated genomic analyses identify ARID1A and ARID1B alterations in the childhood cancer neuroblastoma. Nat. Genet. 45, 12–17 (2013).
Imielinski, M. et al. Mapping the hallmarks of lung adenocarcinoma with massively parallel sequencing. Cell 150, 1107–1120 (2012).
Robinson, G. et al. Novel mutations target distinct subgroups of medulloblastoma. Nature 488, 43–48 (2012).
Love, C. et al. The genetic landscape of mutations in Burkitt lymphoma. Nat. Genet. 44, 1321–1325 (2012).
Versteege, I. et al. Truncating mutations of hSNF5/INI1 in aggressive paediatric cancer. Nature 394, 203–206 (1998).
Varela, I. et al. Exome sequencing identifies frequent mutation of the SWI/SNF complex gene PBRM1 in renal carcinoma. Nature 469, 539–542 (2011).
Schick, S. et al. Systematic characterization of BAF mutations provides insights into intracomplex synthetic lethalities in human cancers. Nat. Genet. 51, 1399–1410 (2019).
Miao, D. et al. Genomic correlates of response to immune checkpoint therapies in clear cell renal cell carcinoma. Science 359, 801–806 (2018).
Braun, D. A. et al. Interplay of somatic alterations and immune infiltration modulates response to PD-1 blockade in advanced clear cell renal cell carcinoma. Nat. Med. 26, 909–918 (2020).
Deng Pan, A. K. et al. A major chromatin regulator determines resistance of tumor cells to T cell–mediated killing. Science 359, 770–775 (2018).
Liu, X. D. et al. PBRM1 loss defines a nonimmunogenic tumor phenotype associated with checkpoint inhibitor resistance in renal carcinoma. Nat. Commun. 11, 2135 (2020).
Okamura, R. et al. ARID1A alterations function as a biomarker for longer progression-free survival after anti-PD-1/PD-L1 immunotherapy. J. Immunother. Cancer 8, e000438 (2020).
Fukumoto, T. et al. HDAC6 inhibition synergizes with anti-PD-L1 therapy in ARID1A-inactivated ovarian cancer. Cancer Res. 79, 5482–5489 (2019).
Li, J. et al. Epigenetic driver mutations in ARID1A shape cancer immune phenotype and immunotherapy. J. Clin. Investig. 130, 2712–2726 (2020).
Goodman, A. M. et al. Tumor mutational burden as an independent predictor of response to immunotherapy in diverse cancers. Mol. Cancer Ther. 16, 2598–2608 (2017).
Fruhwald, M. C., Biegel, J. A., Bourdeaut, F., Roberts, C. W. & Chi, S. N. Atypical teratoid/rhabdoid tumors-current concepts, advances in biology, and potential future therapies. Neuro-Oncol. 18, 764–778 (2016).
Leruste, A. et al. Clonally expanded T cells reveal immunogenicity of rhabdoid tumors. Cancer Cell 36, 597–612 e598 (2019).
Hanna, G. J. et al. Frameshift events predict anti-PD-1/L1 response in head and neck cancer. JCI Insight 3, e98811 (2018).
Fiala, O. et al. Immune-related adverse effects and outcome of patients with cancer treated with immune checkpoint inhibitors. Anticancer Res. 40, 1219–1227 (2020).
Jin, K. T. et al. Immune-mediated adverse effects of immune-checkpoint inhibitors and their management in cancer. Immunol. Lett. 221, 61–71 (2020).
Soldi, R. et al. The novel reversible LSD1 inhibitor SP-2577 promotes anti-tumor immunity in SWItch/Sucrose-NonFermentable (SWI/SNF) complex mutated ovarian cancer. PLoS ONE 15, e0235705 (2020).
Carbognin, L. et al. Differential activity of nivolumab, pembrolizumab and MPDL3280A according to the tumor expression of programmed death-ligand-1 (PD-L1): sensitivity analysis of trials in melanoma, lung and genitourinary cancers. PLoS ONE 10, e0130142 (2015).
Abou Alaiwi, S. et al. Mammalian SWI/SNF complex genomic alterations and immune checkpoint blockade in solid tumors. Cancer. Immunol. Res. 8, 1075–1084 (2020).
Shu, X. S. et al. The epigenetic modifier PBRM1 restricts the basal activity of the innate immune system by repressing retinoic acid-inducible gene-I-like receptor signalling and is a potential prognostic biomarker for colon cancer. J. Pathol. 244, 36–48 (2018).
Nagarsheth, N. et al. PRC2 epigenetically silences Th1-type chemokines to suppress effector T-cell trafficking in colon cancer. Cancer Res. 76, 275–282 (2016).
Peng, D. et al. Epigenetic silencing of TH1-type chemokines shapes tumour immunity and immunotherapy. Nature 527, 249–253 (2015).
Neely, K. E. et al. Activation domain-mediated targeting of the SWI/SNF complex to promoters stimulates transcription from nucleosome arrays. Mol. Cell 4, 649–655 (1999).
Chandler, R. L. et al. Coexistent ARID1A-PIK3CA mutations promote ovarian clear-cell tumorigenesis through pro-tumorigenic inflammatory cytokine signalling. Nat. Commun. 6, 6118 (2015).
Stojanova, A. et al. MYC interaction with the tumor suppressive SWI/SNF complex member INI1 regulates transcription and cellular transformation. Cell Cycle 15, 1693–1705 (2016).
Casey, S. C. et al. MYC regulates the antitumor immune response through CD47 and PD-L1. Science 352, 227–231 (2016).
Casey, S. C., Baylot, V. & Felsher, D. W. The MYC oncogene is a global regulator of the immune response. Blood 131, 2007–2015 (2018).
Cheng, S. W. et al. c-MYC interacts with INI1/hSNF5 and requires the SWI/SNF complex for transactivation function. Nat. Genet. 22, 102–105 (1999).
Lu, Y. et al. MYC targeted long noncoding RNA DANCR promotes cancer in part by reducing p21 levels. Cancer Res. 78, 64–74 (2018).
Weissmiller, A. M. et al. Inhibition of MYC by the SMARCB1 tumor suppressor. Nat. Commun. 10, 2014 (2019).
Pal, S. et al. mSin3A/histone deacetylase 2- and PRMT5-containing Brg1 complex is involved in transcriptional repression of the Myc target gene cad. Mol. Cell Biol. Nov. 23, 7475–87 (2003).
Sammak, S., Allen, M. D., Hamdani, N., Bycroft, M. & Zinzalla, G. The structure of INI1/hSNF5 RPT1 and its interactions with the c-MYC:MAX heterodimer provide insights into the interplay between MYC and the SWI/SNF chromatin remodeling complex. FEBS J. 285, 4165–4180 (2018).
Srikanth, S., Ramachandran, S. & Mohan, S. S. Construction of the gene regulatory network identifies MYC as a transcriptional regulator of SWI/SNF complex. Sci. Rep. 10, 158 (2020).
Huang, Y. & Li, G. M. DNA mismatch repair in the context of chromatin. Cell Biosci. 10, 10 (2020).
Goellner, E. M. Chromatin remodeling and mismatch repair: access and excision. DNA Repair 85, 102733 (2020).
Dietmaier, W., Buttner, R. & Ruschoff, J. Microsatellite instability: review of methods and applications. Der Pathol. 40, 313–327 (2019).
Boileve, A. et al. Immune checkpoint inhibitors in MITF family translocation renal cell carcinomas and genetic correlates of exceptional responders. J. Immunother. Cancer 6, 159 (2018).
Lheureux, S. et al. A clinical and molecular phase II trial of oral ENMD-2076 in ovarian clear cell carcinoma (OCCC): a study of the Princess Margaret Phase II Consortium. Clin. Cancer Res. 24, 6168–6174. (2018).
Kuroda, T. et al. Therapeutic preferability of gemcitabine for ARID1A-deficient ovarian clear cell carcinoma. Gynecol. Oncol. 155, 489–498 (2019).
Lu, M. et al. Efficacy, safety and biomarkers of toripalimab in patients with recurrent or metastatic neuroendocrine neoplasms: a multiple-center phase Ib trial. Clin. Cancer Res. 26, 2337–2345 (2020).
Thomas, A. R. C. & Sciuto, L. et al. Phase I study of ATR inhibitor M6620 in combination with topotecan in patients with advanced solid tumors. J. Clin. Oncol. 36, 1594–602 (2018).
Acknowledgements
This work was supported by The Program for Professor of Special Appointment (Eastern Scholar) at Shanghai Institutions of Higher Learning (No. TP2018046 to J.S.), Shanghai Municipal Education Commission-Two Hundred Talent (No. 20191817 to J.S.), and General Program of National Natural Science Foundation of China (No. 81972667 to J.S.)
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhou, M., Yuan, J., Deng, Y. et al. Emerging role of SWI/SNF complex deficiency as a target of immune checkpoint blockade in human cancers. Oncogenesis 10, 3 (2021). https://doi.org/10.1038/s41389-020-00296-6
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41389-020-00296-6
This article is cited by
-
Primary Oropharyngeal SMARCA4-Deficient Carcinoma: Expanding the Diagnostic Spectrum in Head and Neck Cancer
Head and Neck Pathology (2024)
-
Top 10 Basaloid Neoplasms of the Sinonasal Tract
Head and Neck Pathology (2023)
-
TMBur: a distributable tumor mutation burden approach for whole genome sequencing
BMC Medical Genomics (2022)
-
Discovering biomarkers for hormone-dependent tumors: in silico study on signaling pathways implicated in cell cycle and cytoskeleton regulation
Molecular Genetics and Genomics (2022)
-
Dedifferentiation-like tubular and solid carcinoma of the stomach shows phenotypic divergence and association with deficient SWI/SNF complex
Virchows Archiv (2022)