Abstract
Nonalcoholic fatty liver disease (NAFLD) is a chronic inflammatory disease in which nucleotide-binding domain of leucine-rich repeat protein 3 (NLRP3) inflammasome plays an important role. The present research was aimed to explore the protective function of ADAR1, an RNA editing enzyme, against inflammatory damages in high-fat diet (HFD)-induced NAFLD through inhibiting NLRP3 inflammasome and subsequent inflammation. A total of 30 patients with NAFLD were investigated, and ADAR1 mRNA expression in peripheral blood monocytes surveyed. The in vivo study used lentivirus to explore the function of ADAR1 overexpression in the HFD-induced mouse model of NAFLD. The in vitro study used lentivirus and siRNA to explore the function of ADAR1 on the NLRP3 inflammasome activation in THP-1 cells. Results shown that the ADAR1 expression was upregulated in NAFLD patients in comparison to healthy controls. In vivo, the upregulation of ADAR1 impaired NLRP3 inflammasome activation and alleviated liver disease in HFD mice in comparison to the control group. Moreover, ADAR1 overexpression attenuated NLRP3 inflammasome in lipopolysaccharide (LPS)+ palmitic acid (PA)-induced THP-1 cells, while ADAR1 knockdown increased the NLRP3 inflammasome activation. Furthermore, we speculated that c-Jun may participate in ADAR1’s inhibition of NLRP3 inflammasome. Our results suggested that ADAR1 is a potential treatment target for NAFLD via regulating the activation of NLRP3 inflammasome.
Similar content being viewed by others
Introduction
Non-alcoholic fatty liver (NAFLD) is one of the most frequent liver diseases worldwide and is clinical characterized by abnormal lipid accumulation in liver1,2. The clinical spectrum of NAFLD represents from simple steatosis and non-alcoholic steatohepatitis (NASH) to cirrhosis and hepatocellular carcinoma3. Despite some progress has been achieved, no evidence-based pharmacological strategy is available for the clinical treatment of NAFLD4. Thus, the identification of new target for the molecular therapy of NAFLD is an urgent requirement.
Recent research has confirmed that sterile inflammation plays a pivotal driver in triggering the transition from NAFL to NASH5 and amplifies liver damage by activation of inflammasomes, following with the secretion of the pro-inflammatory cytokines. The pro-inflammatory cytokines, including interleukin (IL)-18 and IL-1β, play an important part in the liver damage6. The nucleotide-binding domain of leucine-rich repeat protein 3 (NLRP3) inflammasome is an intracellular multiprotein complex participated in the production of mature IL-1β and IL-18 and then induces metabolic inflammation. Reportedly, NLRP3 inflammasome is essential for the development of sterile liver inflammation in NAFLD7,8. In addition, compelling evidence suggests that the abnormal activation of NLRP3 inflammasome causes the deterioration of NAFLD. The expression levels of NLRP3 inflammasome components were rose markedly in both murine models and humans with NASH9. Gain-of-function variants of NLRP3 inflammasome leads to more severe steatohepatitis in HFD-fed mice10. In addition, mice carried the NLRP3 inflammasome loss-of-function variants is mild in diet-induced NAFLD9. These findings, therefore, suggest that NLRP3 inflammasome can be regarded as a new target for therapy of NAFLD.
Adenosine deaminase acting on RNA-1 (ADAR1) is a double-stranded (ds) RNA-binding and editing protein that catalyzes the C-6 deamination of adenosine (A) to generate inosine (I) in dsRNA substrates11. ADAR1 plays a key role in both embryonic and adult livers12. Despite the research of ADAR1 focused on its RNA editing activities, editing-independent activities of the protein have also been reported13,14. Recent advances confirm the role of ADAR1 in preventing self RNAs from triggering immune responses15. Additionally, the level of ADAR1 is increased in response to various stresses then regulates inflammatory responses16. In endotoxin-stimulated mice, the up-regulation of ADAR1 was determined in peripheral lymphocytes as well as thymus and spleen. In vitro, inflammatory mediators could directly induce the expression increase of ADAR1 in macrophages17. Furthermore, ADAR1 suppresses type I interferons (IFN) production in hepatocytes and protects liver from inflammatory damage12. Thus, these studies proposed a critical role of ADAR1 in balancing immune activation. However, the mechanism underlying ADAR1-mediated hepatic homeostasis by modulating inflammation remains an enigma18.
In this study, we presented preliminary evidence on the expression pattern of ADAR1 in NAFLD patients and reported that the upregulated level of ADAR1 in HFD mouse models ameliorates the NAFLD/NASH phenotype. Focusing on the effect of ADAR1 on inflammation improvement, especially the IL-1β and IL-18, we hypothesized that ADAR1 may modulate inflammation by inhibiting the activation of NLRP3 inflammasome. This hypothesis was tested through genetic manipulation of ADAR1 gene expression in lipopolysaccharide (LPS)+ palmitic acid (PA)-induced THP-1 cells and further explored its possible mechanism.
Materials and methods
Materials
LPS and PA were obtained from Sigma (Sigma, Darmstadt, Germany). RPMI 1640 medium, FBS, and antibiotics (penicillin/streptomycin) were acquired from Gibco-BRL (Rockville, MD, USA). Anti-NLRP3 (D4D8T) and anti-β-actin (8H10D10) antibodies were acquired from Cell Signaling Technology (Beverly, MA, USA). Anti-ASC (NBP1–78977) was obtained from NovusBio (Littleton, USA). Anti-caspase-1 (EPR16883) antibodies was obtained from Abcam (Cambridge, UK). Anti-ADAR1 (sc-73408) antibody was acquired from Santa Cruz Biotechnology Inc. (CA, USA). Anti-c-Jun (24909–1-AP) antibody was acquired from Proteintech (Chicago, IL, USA).
Blood sampling
There are altogether 30 NAFLD patients (15 men and 15 women) within 5 years of diagnosis were recruited from the Diabetes Clinic in the Third Xiangya Hospital of Central South University. The NAFLD was diagnosed by sonography according to the standard criteria in previous studies19,20. A total of 20 healthy subjects (10 men and 10 women) were age- and sex-matched with the NAFLD group. None of the subjects presented evidence of infection, allergic, or other autoimmune diseases recently, and did not use immunomodulatory drugs in the 6 months before sampling. The research was approved by the Ethics Committee of the Third Xiangya Hospital of Central South University. Written informed consent was obtained from all study participants.
Venous blood was collected from fasting subjects between 6.00 and 8.00 a.m. to minimize the possible circadian variations. Peripheral blood monocytes (PBMCs) were isolated using lymphocyte-H cell separation media (VH Bio, Gateshead UK), followed by isolation of monocytes by Percoll (Sigma, Darmstadt, Germany) density gradient centrifugation21. The serum was separated by centrifugation and stored at -80 °C until further analysis.
Animals
C57BL/6 male mice, aged 6–8 weeks (bodyweight = 23–25 g) (Slaccas Inc., Shanghai, China) were housed in a specific pathogen-free (SPF) environment with access to food and water ad libitum at the Animal Experimental Centre of Central South University. All animal protocols were approved by the Animal Care and Use Committee of the Third Xiangya Hospital of Central South University.
The mice were randomly divided into four groups: mice in the control group (n = 5) were fed normal chow for 12 weeks; 15 mice were fed HFD (60 kcal% fat, TROPHIC Animal Feed High-Tech Co. Ltd, China) for 12 weeks to induce NAFLD and then assigned into three groups: HFD group (n = 5), HFD + vector group (n = 5) injected with the LV-GFP, and HFD + ADAR1 group (n = 5) injected with LV-ADAR1-GFP. The recombinant lentivirus for LV-ADAR1-GFP and LV-GFP were acquired from GeneChem (Shanghai, China). C57BL/6 mice were infected with lentiviruses (1 × 108 infection units per mouse) by injection through the tail vein. All mice were weighed weekly. After infection, mice were weighed twice weekly. Mouse food intake was measured on a weekly basis. The total weekly food intake of a mouse was calculated by measuring the food added subtracted by the food left in the cage divided by the number of mice in the cage. Mouse daily food intake was calculated by total weekly food intake divided by 7. Two weeks after the injection, the animals were anesthetized and sacrificed. Subsequently, the blood samples isolated for further study and the liver was collected; a part of liver was fixed with formalin, and the remaining was frozen in liquid nitrogen and stored at −80 °C.
Glucose Tolerance Test (GTT) and Insulin Tolerance Test (ITT)
The GTT and ITT were performed as previous published work from our laboratory2. Glucose levels were measured using an automatic glucometer (Roche Diagnostics, Rotkreuz, Switzerland).
Cell culture
Monocytic THP-1 cells, obtained from ATCC, were cultured in RPMI 1640 medium, supplemented with 10% heat-inactivated FBS, 1% penicillin and 100 μg/mL streptomycin. Then, the cells were stimulated with 100 ng/mL LPS for 3 h, followed by stimulated with PA (0.5 mM) for another 8 h as described as previous published work from our laboratory22.
Transfection
An equivalent of 1 × 105 THP-1 cells were respectively infected with 2 × 106 LV-GFP or LV-ADAR1-GFP lentivirus, following the protocol from GeneChem (Shanghai, China). As shown in Supplementary Fig. 1, the infection efficiency of lentivirus was >90%. The efficiency of lentivirus-mediated ADAR1 up-regulation was assessed by Western blot analysis (Supplementary Fig. 2A, B).
We designed several small interfering RNA fragments that might bind to the mRNA coding sequence of ADAR1 and c-Jun and selected the sequence that inhibits most effectively. The following sense small interfering RNA (siRNA) sequences were used in the study: si-ADAR1 (5′-CCUUCUACAGUCAUGGCUUTT-3′); si-c-Jun (5′-GGCACAGCTTAAACAGAAA-3′) (GenePharma, Shanghai, China). Briefly, 3 × 105 cells/well were seeded and incubated in FBS-free medium. Subsequently, the cells were transfected with siRNA using Lipofectamine 2000 Transfection Reagent (Invitrogen, Shanghai, China), following the protocol from GenePharma (Shanghai, China). The inhibitory efficiency of siRNA probes was assessed by Western blot and qPCR analysis (Supplementary Fig. 2C, D and Supplementary Fig 6A, B).
Biochemical analyses
The plasma levels of alanine transaminase (ALT) and aspartate transaminase (AST) from mice were measured by automatic biochemical analyzer (Cobas 6000, Roche). The plasma levels of triglyceride (TG), cholesterol (CHO), high-density lipoprotein (HDL) and low-density lipoprotein (LDL) were detected using enzymatic colorimetric kits and were carried out following the instructions from manufacturer (Beyotime Biotechnology Corp.; Nanjing Jiancheng Bioengineering Institute, China). Liver tissue samples were homogenized and then centrifuged to obtain the supernatants. The level of hepatic TG was measured using enzymatic colorimetric kits (Beyotime Biotechnology Corp.; Nanjing Jiancheng Bioengineering Institute, China).
Histology
Liver slices from mice were stained with hematoxylin and eosin (H&E) or oil red O and analyzed by microscopy (Servicebio, Wuhan, China). The morphological changes were examined histologically in a blinded manner. The liver sections were scored on the basis of the criteria of the NAFLD activity score (NAS)23,24.
Immunofluorescence staining
Immunofluorescence staining was carried out following a standard protocol. Immunofluorescence microscopy of tissue was performed with a Leica TCS SP2 confocal scanning microscope adapted to an inverted Leica DMI 6000CS microscope to the acquisition of multiple tissular staining. Figures were assembled from TIF files using National Institute of Mental Health’s open software ImageJ (NIH).
Enzyme-linked immunosorbent assay (ELISA)
The protein levels of IL-1β and IL-18 in the cell culture supernatants were detected using ELISA kits (R&D Systems Inc., Minneapolis, USA).
Western blot analysis
The samples of mice liver and THP-1 cells were lysed in RIPA lysis buffer (Beyotime Biotechnology, Shanghai, China). Total protein was separated by SDS-PAGE, followed by electrophoretic transfer to PVDF membranes, and blocked in 5% non-fat milk at room temperature for 1 h. The membranes were probed with primary antibodies against NLRP3, caspase-1, ASC, ADAR1, c-Jun, and β-actin (1:1000) at 4 °C overnight. Subsequently, the blots were incubated with specific HRP-conjugated second antibody (Cell Signaling Technology) for 1 h at room temperature. Finally, the immunoreactive bands were visualized using Electrochemiluminescence (ECL) Kit (Tanon, Shanghai, China) and X-ray film.
Quantitative real-time PCR
Total RNA was extracted using TRIzol (Invitrogen, Carlsbad, CA, USA) and reverse-transcribed into cDNA using the RevertAid First Strand cDNA Synthesis Kit (Thermo Fisher Scientific, MA, USA). Quantitative PCR was implement using the SYBR GREEN PCR Kit (Toyobo, Osaka, Japan). The samples were PCR amplified at following cycles: 72 °C for 3 min, 95 °C for 30 s, then 30 cycles of 95 °C for 10 s and 58 °C for 30 s. The forward and reverse PCR primer sequences are listed in Table S1. The relative mRNA levels of the target genes were normalized to that of GAPDH and analyzed by the 2−ΔΔCt method.
NLRP3 transcription factor screening
The NLRP3 promoter sequence was retrieved using the University of California, Santa Cruz (UCSC) genomic database (http://genome.ucsc.edu)25,26. Then, the potential transcription factors bound with the NLRP3 promoter region were predicted by PROMO (http://alggen.lsi.upc.es/)27,28.
Data source
The normalized expression data and annotation data were acquired from the Gene Expression Omnibus (GEO) database (http://www.ncbi.nlm.nih.gov/geo/). The sample set of GSE72756 included 5 NAFLD liver tissues and 5 normal liver tissues29.
Bioinformatics analyses
To eliminate discrete and abnormal samples, data were analyzed by principal component analysis (PCA)30,31. GSM1860540, GSM1860541, GSM1860543, GSM1860545 and GSM1860548 represent normal liver tissue samples. GSM1860542, GSM1860544, GSM1860546, GSM1860547, and GSM1860549 represent non-alcoholic fatty liver disease liver tissue samples (Supplementary Fig 3). Differential expression analysis and protein-protein interaction (PPI) network analysis were conducted, as described previously29,32.
Statistical analysis
Data are presented as mean ± standard deviation (SD). Two groups were compared by the unpaired Student’s t-test. The comparisons among multiple groups were made by one-way analysis of variance (ANOVA), followed by least significant difference (LSD) test as post hoc comparisons. A p-values <0.05 was considered statistically significant. Each experiment was repeated more than three times.
Results
ADAR1 gene expression was upregulated in NAFLD patients
The clinical characteristics of a total of 30 patients are shown in Table 1. As shown in Fig. 1, the level of ADAR1 expression was remarkably increased in NAFLD patients than that in healthy individuals (p < 0.05). These findings suggested that ADAR1 may play a role in NAFLD.
ADAR1 gene expression was upregulated in the liver of HFD mice
To explore the function of ADAR1 in NALFD, we detected its expression in mice with NAFLD model. Upon feeding HFD for 16 weeks, mice revealed overt NAFLD aspects. Specifically, HFD mice displayed a significant gain in bodyweight compared to the normal controls (Fig. 2A). Consistently, HFD mice developed hepatic steatosis according to H&E and oil red O staining of the liver sections (Fig. 2B, C). Moreover, the mRNA levels of pro-inflammatory factors, including IL-1β, IL-18, IL-6, and TNF-α in the liver of HFD mice were remarkably higher than that in the controls (Fig. 2D). On this basis, immunofluorescence analysis indicated that the level of CD68 expression, which is a marker of macrophage, was markedly increased in the liver of HFD groups when compared with normal groups. Interestingly, the level of ADAR1 expression in the liver of HFD mice was markedly higher than normal mice (Fig. 2E, F). Quantification analysis shown that the co-localization number of ADAR1 and CD68 in the liver of HFD mice was greater than normal mice (Fig. 2G). Similarly, Western blotting identified higher ADAR1 protein level in the liver of HFD mice in comparison with the normal mice (Fig. 2H).
A Bodyweight of the mice. B Representative microphotograph of H&E staining of hepatic lipid accumulation. C Representative microphotograph of Oil Red O staining of hepatic lipid accumulation. D mRNA abundance of inflammatory genes in liver tissue. E Confocal images of intracellular localization of ADAR1 by immunofluorescence in transversal sections of livers of control and KO mice. Slides were stained for ADAR1 (red), CD68 (green), and DAPI (blue). Original magnification ×100. F Quantification analysis of the ADAR1 expression. G Quantification analysis of the co-localization number of ADAR1 and CD68. H Abundance of ADAR1 in mouse liver measured by Western blot. The results were mean ± SD (n = 5/group). Two groups were compared by the Student’s t-test. *p < 0.05 vs. the normal group.
ADAR1 overexpression ameliorates HFD-induced NAFLD
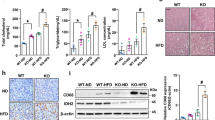
We overexpressed the molecule in mice by injecting lentiviruses through the tail vein to obtain systemic overexpression of ADAR1. The efficiency of lentivirus-mediated ADAR1 up-regulation in the liver tissue was confirmed by qPCR, and Western blotting showed an increased level of ADAR1 p150 in lentivirus-injected mice (Supplementary Fig 4). ADAR1-overexpressed had no significant effect on food intake in mice fed with HFD (Supplementary Fig 5). However, compared to the HFD group, the ADAR1-overexpressed mice gained less weight over a period (Fig. 3A). Moreover, the mean relative liver weights/bodyweight (LW/BW ratio) showed a decline in the ADAR1-overexpressed group in comparison with the HFD group. (Fig. 3B). In the GTT, ADAR1-overexpressed group’s glucose tolerance curve was apparently below HFD group’s. ITT identified an improvement in insulin tolerance in ADAR1-overexpressed mice than the HFD mice (Fig. 3C). ALT and AST are generally detected to assess hepatocellular injury. The levels of plasma ALT and AST in ADAR1-overexpressed group was lower than HFD group (Fig. 3D). Assessment of hepatic TG confirmed significantly elevated levels in HFD-fed mice. In addition, ADAR1-overexpressed group showed a marked decrease of hepatic TG level when compared with HFD group (Fig. 3E). The levels of serum TG, TC, HDL, and LDL were detected in each group. As displayed in Fig. 3F, the level of TG in ADAR1-overexpressed group was apparently lower than that in HFD group. No significant differences were found in CHO, HDL, and LDL levels between the HFD group and the ADAR1-overexpressed group (Fig. 3F). In the H&E analysis, mice on a HFD had raised levels of hepatic lipid accumulation and inflammation. Interestingly, ADAR1 lentivirus injection for 4 weeks significantly alleviates histological parameters, including steatosis, ballooning, and inflammation, on the basis of the criteria of the NAS, in comparison with mice in HFD group (Fig. 3G–I). These observations suggested that ADAR1 is a potential therapeutic target for NAFLD.
A Effect of ADAR1-overexpression on bodyweight gain of the mice (n = 5). B Effect of ADAR1-overexpression on the liver to bodyweight ratio of the mice (n = 5). C Effect of ADAR1-overexpression on the glucose tolerance test (n = 5) and insulin tolerance test (n = 5). D Serum levels of ALT and AST in mice. E Levels of hepatic TG in mice. F Serum levels of TG, CHO, HDL and LDL in mice. G Representative macroscopic liver images of the mice. H, I Representative microphotograph of H&E staining of the hepatic lipid accumulation. The results were mean ± SD (n = 5/group). Two groups were compared by the Student’s t-test. The comparisons among multiple groups were made by ANOVA, followed by LSD test as post hoc comparisons. *p < 0.05 vs. the normal group. #p < 0.05 vs. the HFD group. HFD high-fat diet.
ADAR1 overexpression ameliorates HFD-induced lipid accumulation and inflammatory responses in liver
We then evaluated the effect of ADAR1 on fatty acid metabolism-related genes and crucial pro-inflammatory genes involved in NAFLD. As expected, ADAR1-overexpression significantly suppressed the expression of genes participated in fatty acid synthesis (PPARγ, SCD1, FASN, and SREBP1-c) and up-regulated the expression of genes participated in fatty acid oxidation (PPARα, ACOX1, ACADL, and ACADM) (Fig. 4A). In addition, the transcriptional analysis revealed that ADAR1-overexpression remarkably suppressed the level of mRNA from pro-inflammatory factors, including IL-1β, IL-18, IL-6, and TNF-α in comparison with HFD group (Fig. 4B). Furthermore, ADAR1-overexpression also suppressed the level of LCN2 gene expression, which is a marker for inflammation in rodents33,34, in the liver of mice when compared with HFD group (Fig. 4B).
A, B mRNA abundance of lipid metabolism and inflammatory genes in the liver tissue. C Representative Western blot bands of ADAR1, NLRP3, ASC, and caspase-1 expression in the liver of mice. D Abundance of ADAR1 and NLRP3 in mouse liver was detected by qPCR analysis, respectively. The results were mean ± SD (n = 5/group). The comparisons among multiple groups were made by ANOVA, followed by LSD test as post hoc comparisons. *p < 0.0 vs. the normal group, #p < 0.05 vs. the HFD group. HFD high-fat diet.
As hepatic inflammation is a compelling feature in NAFLD development9, the inhibition of ADAR1 on pro-inflammatory factors (i.e., IL-1β and IL-18) prompted us to explore the possible molecular mechanisms further. Recent studies suggest a pivotal role of NLRP3 inflammasome in the progression of NAFLD and regulating the secretion of IL-1β and IL-182. As expected, Western blotting showing that the levels of NLRP3 inflammasome-associated proteins, including NLRP3 and cleaved caspase-1, in the liver tissue of HFD mice were remarkably higher than normal mice (Fig. 4C). Especially, in ADAR1-overexpression mice, HFD elevated the levels of NLRP3 protein, and caspase-1 maturation were significantly abolished compared to that of vector mice (Fig. 4C). Western blots did not show any significant difference in the ASC protein expression among groups (Fig. 4C). Also, the change of mRNA level of NLRP3 was in accordance with the change of protein level (Fig. 4D). These findings suggested that ADAR1 represses lipid accumulation and inflammatory responses during HFD-induced NAFLD development.
ADAR1 suppresses NLRP3 inflammasome activation in LPS + PA-induced THP-1 cells
Previous studies revealed that LPS + PA-induced THP-1 monocytes as a putative alternative for investigating NLRP3 inflammasome relative to fatty acid metabolism disorder22,35,36. As described previously, THP-1 cells were stimulated with LPS plus PA to activate NLRP3 inflammasome22. To investigate the effect of ADAR1 in vitro, we first overexpressed ADAR1 in THP-1 cells, followed by stimulation with LPS + PA. The efficiency of lentivirus-mediated ADAR1 upregulation was verified by Western blotting, which showed an increased level of ADAR1 p150 in lentivirus-infected cells (Fig. 5A). Consequently, LPS + PA-treated THP-1 cells expressed a significant rise of NLRP3 protein level and remarkably elevated caspase-1 maturation (Fig. 5A). These effects were significantly abolished in LV-ADAR1-GFP THP-1 cells compared to LV-GFP THP-1 cells. Also, Western blots did not show any significant difference in the ASC protein expression among groups (Fig. 5A). The mRNA level of NLRP3 was similar to the protein level (Fig. 5B). The levels of IL-1β and IL-18 in the suspensions were examined. THP-1 cells expressed a remarkable rise in the secretion of IL-1β and IL-18 after LPS + PA treatment, which was significantly reduced in LV-ADAR1-GFP THP-1 cells than in LV-GFP THP-1 cells (Fig. 5C).
A Representative immunoblots for ADAR1, NLRP3, caspase-1, and ASC; B mRNA abundance of NLRP3; C ELISA of IL-1β and IL-18 level in THP-1 cells infected with LV-GFP or LV-ADAR1-GFP. D Representative immunoblots for ADAR1, NLRP3, caspase-1, and ASC; E mRNA abundance of NLRP3; F ELISA of IL-1β and IL-18 level in THP-1 cells transfected with scrambled or ADAR1 siRNA. All data represent the mean ± SD of more than three independent experiments. The comparisons among multiple groups were made by ANOVA, followed by LSD test as post hoc comparisons. *p < 0.05 vs. the untransfected LPS + PA group. LV-GFP GFP lentiviral, LV-ADAR1-GFP ADAR1-GFP lentiviral, si-NC scramble siRNA, si-ADAR1 ADAR1 siRNA.
Since ADAR1 was upregulated in the liver of HFD mice, we transfected THP-1 cells with siRNA-ADAR1 and evaluated the si-NC- and si-ADAR1-transfected THP-1 cells with or without LPS + PA stimulation, respectively, to explore the effect of ADAR1 knockdown on NLRP3 inflammasome activation. The efficiency of siRNA-mediated ADAR1 downregulation was further verified by Western blot (Fig. 5D). The basal levels of NLRP3 protein and caspase-1 maturation were slightly increased in si-ADAR1-transfected THP-1 cells in comparison with that in si-NC-transfected THP-1 cells. The protein expression was also markedly increased in LPS + PA-treated si-NC-transfected THP-1 cells and significantly enhanced in si-ADAR1-transfected THP-1 stimulated with LPS + PA as compared to that in unstimulated si-NC-transfected THP-1 cells. Consistently, Western blots did not show any significant difference in the ASC protein expression among groups (Fig. 5D). The mRNA level of NLRP3 was similar to the protein level (Fig. 5E). As shown in Fig. 5F, si-ADAR1-translated THP-1 cells were susceptible to LPS + PA-induced IL-1β and IL-18 release in comparison with the si-NC-translated THP-1 cells. Taken together, siRNA knockdown of ADAR1 significantly enhanced LPS + PA-induced NLRP3 inflammasome activation.
C-Jun may participate in the inhibitory mechanism of ADAR1 on NLRP3 inflammasome activation
Herein, we further explored the possible mediator that are implicated in the inhibiting effects of ADAR1 on NLRP3 inflammasome activation. Previous studies demonstrated that the level of NLRP3 protein is one of rate-limiting factors for the activation of the inflammasome37. The transcriptional regulation of NLRP3 expression is one of the major regulatory mechanisms for attenuating the NLRP3 inflammasome activation38. Based on our finding that ADAR1 regulates the NLRP3 expression at both mRNA and protein levels, we hypothesized that ADAR1 might modulate the transcription activity of NLRP3 by regulating its transcription factors.
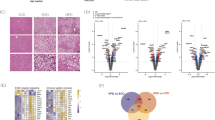
First, we obtained the promoter sequence of NLRP3 gene from UCSC genomic database. We then employed PROMO database to predict the transcriptional factor of NLRP3 genes. The target transcription factor was detected by a comparison of the number of binding sites for the promoter. As a result, we identified 25 putative transcription factors (factors predicted within a dissimilarity margin) ≤ 0% (Supporting Information Table S2). Next, the GSE72756 dataset was downloaded and analyzed to discover the critical genes involved in NAFLD progression. After normalization to exclude the bias in microarray data, the results of the heatmap and volcano plot are presented in Fig. 6A, B. The long non-coding RNAs were excluded. There are altogether 292 differentially expressed mRNAs (DE-mRNAs), including 130 upregulated and 162 downregulated mRNAs, identified. Subsequently, the genes that correspond to 25 putative transcription factors were filtered by NAFLD-related DE-mRNAs (some of these transcription factors are encoded by the same gene). Jun proto-oncogene, AP‑1 transcription factor subunit (JUN), also termed as c-Jun, was selected as the target transcriptional factor after considering the intersections. In addition, c-Jun also identified as one of the key DE-mRNAs involved in NAFLD through PPI network (Fig. 6C, D).
A Clustering heatmap for the differentially expressed genes of GSE72756. The blue-black-yellow variation indicates the alterations in the expression from low to high. B Volcano plot of differential gene expression in normal liver tissue samples vs. NAFLD liver tissue samples; A false discovery rate (FDR; adjusted p-value) <0.05 and |log2 fold-change (FC)|≥1.5 were set as thresholds. C The top 15 hub genes derived were selected using the CytoHubba plugin using the maximal clique centrality (MCC) method. Advanced ranking is represented by the intense red color. D The schematic representation of the filter strategies employed in our study. E Representative Western blot bands of c-Jun expression in the liver of mice. F mRNA abundance of c-Jun in the liver tissue. G Immunoblots for c-Jun protein and c-Jun mRNA level were measured in THP-1 cells infected with LV-GFP or LV-ADAR1-GFP. H Immunoblots for c-Jun protein and c-Jun mRNA level were detected in THP-1 cells transfected with scrambled or ADAR1 siRNA. All data represent the mean ± SD of more than three independent experiments. The comparisons among multiple groups were made by ANOVA, followed by LSD test as post hoc comparisons. *p < 0.05 vs. the untransfected LPS + PA group. LV-GFP GFP lentiviral, LV-ADAR1-GFP ADAR1-GFP lentiviral, Si-NC scrambled siRNA, Si-ADAR1 ADAR1 siRNA.
Moreover, we validated the expression of c-Jun in vivo and in vitro. Western blotting showed that the level of c-Jun in the liver tissue of HFD mice were remarkably higher than normal mice. In ADAR1-overexpression mice, HFD elevated the level of c-Jun was significantly abolished compared to that of vector mice (Fig. 6E). Similarly, the change of mRNA level of c-Jun was in accordance with the change of protein level (Fig. 6F) In vitro, after LPS + PA treatment, the mRNA and protein levels of c-Jun in different group of cells were determined, respectively. Results showed that the expression of c-Jun at both mRNA and protein level was decreased in LV-ADAR1-GFP THP-1 cells compared to that in LV-GFP THP-1 cells (Fig. 6G). Also, the expression of c-Jun at both mRNA and protein levels were markedly increased in LPS + PA-treated si-ADAR1-transfected THP-1 cells in comparison with that in si-NC-transfected THP-1 cells (Fig. 6H).
To investigate if the c-Jun was involved in the regulation of ADAR1 on NLRP3, we further induced downregulation of c-Jun in si-ADAR1-transfected THP-1 cells with LPS + PA stimulation. The efficiencies of siRNA-mediated ADAR1 downregulation and siRNA-mediated c-Jun downregulation were verified by Western blot respectively (Fig. 7A). Under the stimulation of LPS + PA, the levels of NLRP3 and c-Jun protein was significantly increased in si-ADAR1-transfected THP-1 cells in comparison with that in si-NC-transfected THP-1 cells, which consistent with the above results. Interestingly, these effects were significantly abolished in si-c-Jun- and si-ADAR1-transfected THP-1 cells compared to si-NC- and si-ADAR1-transfected THP-1 cells (Fig. 7A). The mRNA level of NLRP3 was similar to the protein level (Fig. 7B).
A Representative immunoblots for ADAR1, c-Jun, and NLRP3; B mRNA abundance of NLRP3. All data represent the mean ± SD of more than three independent experiments. The comparisons among multiple groups were made by ANOVA, followed by LSD test as post hoc comparisons. *p < 0.05 vs. the LPS + PA + siADAR1 group. si-NC scramble siRNA, si-ADAR1 ADAR1 siRNA, si-c-Jun c-Jun siRNA.
Discussion
In this research, ADAR1 expression was upregulated in both NAFLD patients and HFD mice. In subsequent experiments, we found that the overexpression of ADAR1 in mice markedly ameliorated HFD-induced liver injury. Moreover, ADAR1-overexpression significantly abolished the LPS + PA-induced NLRP3 inflammasome activation in THP-1 cells, while the inhibition of ADAR1 expression with siRNA revealed a dramatic enhancement in NLRP3 inflammasome activation. Additionally, utilizing the GEO database analysis and bioinformatics strategies, we screened out c-Jun and speculated it may participate in ADAR1’s regulation of NLRP3 inflammasome. These finding suggested that ADAR1 is functionally involved in alleviating HFD-induced liver damage through the molecular regulation of NLRP3 inflammasome.
NAFLD is one of the most common chronic liver diseases worldwide39. The presence of steatosis, oxidative stress, and inflammatory mediators are involved in the alterations of pathological condition in NAFLD40. It was an accepted fact that ADAR1 is indispensable for hepatic health and animal survival41. Moreover, ADAR1 is essential for embryonic liver development and maintaining adult liver homeostasis12. The deletion of ADAR1 gene in the mouse models resulted in severe liver damage42. Nevertheless, the mechanism by which ADAR1 prevents liver injury remains to be further explored18. In the current research, we observed that overexpression in mice ameliorated the HFD-induced NAFLD phenotype. As far as we know, this is the first research to investigate the hepatoprotective function of ADAR1 in HFD-induced NAFLD mice.
ADAR1 was originally identified as an RNA-editing enzyme that modifies cellular and viral RNA sequences by adenosine deamination43. Previous studies on ADAR1 were focused on exploring the RNA editing activities and identifying the RNA target44. However, some RNA editing independent activities of ADAR1 were reported12. Recent advances indicated that ADAR1 plays a role in inhibiting adipogenesis and obesity. Yu et al. reported that overexpression of ADAR1 could inhibit the adipogenic differentiation, with the decrease of cell size and intracellular lipid droplets45. Studies have also shown that ADAR1 prevents autoimmunity and regulates innate immunity15. For example, Chen et al. demonstrated that ADAR1 is implicated in the regulation of IL-1β-induced ICAM-1 and VCAM-1 expression16. Wang et al. reported that deficiency of ADAR1 in hepatocytes induces inflammatory response. The study also demonstrated that ADAR1 prevents the liver from inflammatory injury by suppressing IFN signal pathways12. Consist with previous research, the current study demonstrated that ADAR1 critically reduces LPS + PA-induced IL-1β and IL-18 secretion in THP-1 cells. Similarly, we observed that ADAR1 inhibits the expression of pro-inflammatory genes during HFD-induced NAFLD development in vivo. Together, these findings suggested the potential role of ADAR1 as an inflammatory regulator in the NAFLD model. Although the high expression of ADAR1 in NAFLD patients and mice seemingly contradicted the conclusion of its protective effect, it is reasonable to infer that ADAR1 is up-regulated and may play a protective role when mice in abnormal conditions, such as obese state or under inflammatory conditions in our experiment. There’s a growing body of evidence that inflammatory stress also engages regulatory feedback mechanisms to resolve and tightly regulate the inflammatory response. And ADAR1 may operate in a similar fashion, induced by inflammation to limit inflammatory-mediated damage46. Thus, we guess the “natural” increased expression of ADAR1 induced by inflammatory mediators in NAFLD mice was just not sufficient to stop disease progression. But the overexpression of ADAR1 can be beneficial. The seemingly contradictory effect may also be explained by differences in ADAR1 expression in various cell types within the liver. For instance, HFD-diet increased the “natural” expression of ADAR1 while overexpression treatment increased the level of ADAR1 expression in not only hepatocytes but also Kuffper cells, which played a better beneficial function to limit inflammatory. And further studies of this remains to be investigated.
Inflammasome is pivotal in the development of obesity-associated liver disease. Whereas, the related mechanisms still remain largely unknown. NLRP3 inflammasome has been well-characterized with respect to the pathophysiological roles and clinical implications47. NLRP3 inflammasome is a molecular platform that modulates innate immune functions via the maturation of caspase-1 and the production of IL-1β and IL-1848. There is accumulating evidence indicating that the NLRP3 inflammasome drives sterile inflammation and is pivotal in NAFLD progression49. In the current research, we detected significantly increased levels of NLRP3 protein and caspase-1 maturation in LPS + PA-induced THP-1 cells. These changes were abrogated by ADAR1 overexpression while enhanced by ADAR1 knockdown. Moreover, the upregulation of ADAR1 impaired HFD-induced NLRP3 inflammasome activation. Together, these results suggested that the NLRP3 inflammasome pathway mediates the inhibiting effect of ADAR1 on inflammation. Therefore, the current study suggested that ADAR1 has an anti-NLRP3 inflammasome quality and can be an effective therapeutic approach to NLRP3-related diseases, such as NAFLD.
JUN, also known as AP-1 or c-Jun, was increased in a previous study of NAFLD50. A previous study suggested that enhancing the activity of c-Jun accelerates the progression and development of NASH51,52. As an oncogene in the liver, the expression of c-Jun is up-regulated in response to inflammatory stimuli, promoting liver tumorigenesis53. Moreover, activation of c-Jun is the critical step in the transcriptional regulation of inflammasome-related genes54,55. Herein, we employed GEO database analysis in combination with bioinformatics strategies to explore the possible function of c-Jun implicated in the inhibitory effects of ADAR1 on NLRP3 inflammasome activation. Our results showed that c-Jun not only is one of the putative transcription factors of NLRP3 protein but also the key DE-mRNA in NAFLD. Consist with the current research, Wu et al. reported that JUN was highly expressed as compared to the other nodes in the PPI network, based on the GSE72756 dataset analysis; it might also participate in the pathogenic mechanisms of NAFLD32. Recent studies have reported the role of c-Jun in regulating the activation of NLRP3 inflammasome. It was shown that the activation of c-Jun is essential for the transcriptional regulation of inflammasome-related genes, including NLRP3 and Caspase-1. And inhibition of JNK/c-Jun signaling can promote human retinal pigment epithelial cell survival by suppressing the NLRP3 inflammasome55. Also, Wang et al. reported that miR-139 exerted neuroprotection against oxygen-glucose deprivation/reoxygenation (OGD/R)-induced nerve injury by negatively regulating c-Jun/NLRP3 inflammasome signaling. In their study, miR-139 overexpression ameliorated OGD/R-induced inflammation response through reducing NLRP3 inflammasome activation. And the benefits of miR-139 mimics were abolished by overexpression of c-Jun. Moreover, silence of c-Jun suppressed the promoting effects of miR-139 inhibitor on NLRP3 inflammasome activation. Thus, their study demonstrated that c-Jun acted as a transcription factor of NLRP3 protein to regulate inflammation56. Similarly, our mechanistic research showed that ADAR1 overexpression reduced, while ADAR1 knockdown increased LPS + PA-induced c-Jun’s expression. These results were in consistent with the change of NLRP3 inflammasome activation in the same condition. Moreover, our result also showed that the effect of ADAR1 knockdown on the protein level of NLRP3 in LPS + PA-treated THP-1 cells was significantly abolished by c-Jun knockdown. Collectively, we provided preliminary evidence to support that ADAR1 regulates NLRP3 inflammasome via c-Jun. However, more detail research about the role of c-Jun in ADAR1’s regulation on NLRP3 need to be investigated in our near future.
In summary, we provided a preliminary evidence for the in vitro role of ADAR1 in regulating NLRP3 inflammasome activation. Moreover, ADAR1 overexpression ameliorates HFD-induced liver injury, suggesting that upregulation of ADAR1 could be an effective therapeutic approach for the treatment of obesity-induced liver steatosis. Taken together, the current research revealed that ADAR1 is a potential NAFLD suppressor by regulating NLRP3 inflammasome.
Data availability
The sample set of GSE72756 was acquired from the Gene Expression Omnibus (GEO) database (http://www.ncbi.nlm.nih.gov/geo/). All the data and materials supporting the conclusion of the present study have been included within the article.
References
Hernandez-Alvarez MI, Sebastian D, Vives S, Ivanova S, Bartoccioni P, Kakimoto P, et al. Deficient endoplasmic reticulum-mitochondrial phosphatidylserine transfer causes liver disease. Cell 177, 881–895 (2019)
Wang F, Liu Y, Yuan J, Yang W, Mo Z, Compound C protects mice from HFD-induced obesity and nonalcoholic fatty liver disease. Int. J. Endocrinol. 2019, 3206587 (2019)
Chalasani N, Younossi Z, Lavine JE, Diehl AM, Brunt EM, Cusi K, et al. The diagnosis and management of non-alcoholic fatty liver disease: practice guideline by the American Gastroenterological Association, American Association for the Study of Liver Diseases, and American College of Gastroenterology. Gastroenterology 142, 1592–1609 (2012)
Lu Y, Jiang Z, Dai H, Miao R, Shu J, Gu H, et al. Hepatic leukocyte immunoglobulin-like receptor B4 (LILRB4) attenuates nonalcoholic fatty liver disease via SHP1-TRAF6 pathway. Hepatology 67, 1303–1319 (2018)
Kubes P, Mehal WZ, Sterile inflammation in the liver. Gastroenterology 143, 1158–1172 (2012)
Lamkanfi M, Dixit VM, Mechanisms and functions of inflammasomes. Cell 157, 1013–1022 (2014)
Mridha AR, Wree A, Robertson A, Yeh MM, Johnson CD, Van Rooyen DM, et al. NLRP3 inflammasome blockade reduces liver inflammation and fibrosis in experimental NASH in mice. J Hepatol 66, 1037–1046 (2017)
Szabo G, Csak T, Inflammasomes in liver diseases. J Hepatol 57, 642–654 (2012)
Wan X, Xu C, Yu C, Li Y. Role of NLRP3 Inflammasome in the Progression of NAFLD to NASH. Can J Gastroenterol Hepatol 2016, 6489012 (2016)
Wree A, McGeough MD, Pena CA, Schlattjan M, Li H, Inzaugarat ME, et al. NLRP3 inflammasome activation is required for fibrosis development in NAFLD. J Mol Med 92, 1069–1082 (2014)
Li Z, Wolff KC, Samuel CE. RNA adenosine deaminase ADAR1 deficiency leads to increased activation of protein kinase PKR and reduced vesicular stomatitis virus growth following interferon treatment. Virology 396, 316–322 (2010)
Wang G, Wang H, Singh S, Zhou P, Yang S, Wang Y, et al. ADAR1 Prevents Liver Injury from Inflammation and Suppresses Interferon Production in Hepatocytes. Am J Pathol 185, 3224–3237 (2015)
Ota H, Sakurai M, Gupta R, Valente L, Wulff BE, Ariyoshi K, et al. ADAR1 forms a complex with Dicer to promote microRNA processing and RNA-induced gene silencing. Cell 153, 575–589 (2013)
Yang W, Wang Q, Howell KL, Lee JT, Cho DS, Murray JM, et al. ADAR1 RNA deaminase limits short interfering RNA efficacy in mammalian cells. J Biol Chem 280, 3946–3953 (2005)
Lamers MM, van den Hoogen BG, Haagmans BL. ADAR1: “Editor-in-Chief” of Cytoplasmic Innate Immunity. Front Immunol 10, 1763 (2019)
Chen Y, Peng H, Zhou S, Zhuang Y. ADAR1 is targeted by miR-143 to regulate IL-1beta-induced endothelial activation through the NFkappaB pathway. Int J Biochem Cell Biol 89, 25–33 (2017)
Yang JH, Luo X, Nie Y, Su Y, Zhao Q, Kabir K, et al. Widespread inosine-containing mRNA in lymphocytes regulated by ADAR1 in response to inflammation. Immunology 109, 15–23 (2003)
Wang Q. RNA editing catalyzed by ADAR1 and its function in mammalian cells. Biochemistry (Mosc) 76, 900–911 (2011)
Ahmed HS, Pedersen N, Jayanna MB, Ten EP, Sanchez A, Murali AR. Predictive Factors and Time to Development of Hepatic Decompensation in Patients with Non-alcoholic Fatty Liver Disease. J Gen Intern Med 35, 1523–1529 (2020)
Bae YS, Ko YS, Yun JM, Eo AY, Kim H. Association and Prediction of Subclinical Atherosclerosis by Nonalcoholic Fatty Liver Disease in Asymptomatic Patients. Can J Gastroenterol Hepatol 2020, 8820445 (2020)
Pertoft H, Johnsson A, Warmegard B, Seljelid R. Separation of human monocytes on density gradients of Percoll. J Immunol Methods 33, 221–229 (1980)
Liu Y, He H, Fan L, Yuan J, Huang H, Yang W, et al. Compound C attenuates NLRP3 inflammasome despite AMPK knockdown in LPS plus palmitate-induced THP-1 cells. Naunyn Schmiedebergs Arch Pharmacol 393, 67–76 (2020)
Kleiner DE, Brunt EM, Van Natta M, Behling C, Contos MJ, Cummings OW, et al. Design and validation of a histological scoring system for nonalcoholic fatty liver disease. Hepatology 41, 1313–1321 (2005)
Brunt EM, Neuschwander-Tetri BA, Oliver D, Wehmeier KR, Bacon BR. Nonalcoholic steatohepatitis: histologic features and clinical correlations with 30 blinded biopsy specimens. Hum Pathol 35, 1070–1082 (2004)
Lee CM, Barber GP, Casper J, Clawson H, Diekhans M, Gonzalez JN, et al. UCSC Genome Browser enters 20th year. Nucleic Acids Res. 48, D756-D761 (2020)
Tang T, Tao X, Bao X, Chen J, Dai J, Ye J, et al. Mitofusin-2 (Mfn-2) Might Have Anti-Cancer Effect through Interaction with Transcriptional Factor SP1 and Consequent Regulation on Phosphatidylinositol Transfer Protein 3 (PITPNM3) Expression. Med Sci Monit 26, e918599 (2020)
Messeguer X, Escudero R, Farre D, Nunez O, Martinez J, Alba MM. PROMO: detection of known transcription regulatory elements using species-tailored searches. Bioinformatics 18, 333–334 (2002)
Zhao H, Chen M, Wang J, Cao G, Chen W, Xu J. PCNA-associated factor KIAA0101 transcriptionally induced by ELK1 controls cell proliferation and apoptosis in nasopharyngeal carcinoma: an integrated bioinformatics and experimental study. Aging 12, 5992–6017 (2020)
Sun C, Liu X, Yi Z, Xiao X, Yang M, Hu G, et al. Genome-wide analysis of long noncoding RNA expression profiles in patients with non-alcoholic fatty liver disease. Iubmb Life 67, 847–852 (2015)
Reich D, Price AL, Patterson N. Principal component analysis of genetic data. Nat Genet 40, 491–492 (2008)
Zheng X, Levine D, Shen J, Gogarten SM, Laurie C, Weir BS. A high-performance computing toolset for relatedness and principal component analysis of SNP data. Bioinformatics 28, 3326–3328 (2012)
Wu H, Song X, Ling Y, Zhou J, Tao Z, Shen Y. Comprehensive bioinformatics analysis of critical lncRNAs, mRNAs and miRNAs in nonalcoholic fatty liver disease. Mol Med Rep 19, 2649–2659 (2019)
Alwahsh SM, Xu M, Seyhan HA, Ahmad S, Mihm S, Ramadori G, et al. Diet high in fructose leads to an overexpression of lipocalin-2 in rat fatty liver. World J Gastroenterol 20, 1807–21 (2014)
Sultan S, Pascucci M, Ahmad S, Malik IA, Bianchi A, Ramadori P, et al. LIPOCALIN-2 is a major acute-phase protein in a rat and mouse model of sterile abscess. Shock 37, 191–6 (2012)
Wen H, Gris D, Lei Y, Jha S, Zhang L, Huang MT, et al. Fatty acid-induced NLRP3-ASC inflammasome activation interferes with insulin signaling. Nat Immunol 12, 408–415 (2011)
Lendermon EA, Coon TA, Bednash JS, Weathington NM, McDyer JF, Mallampalli RK. Azithromycin decreases NALP3 mRNA stability in monocytes to limit inflammasome-dependent inflammation. Respir Res 18, 131 (2017)
Haneklaus M, O’Neill LA, Coll RC. Modulatory mechanisms controlling the NLRP3 inflammasome in inflammation: recent developments. Curr Opin Immunol 25, 40–45 (2013)
Huai W, Zhao R, Song H, Zhao J, Zhang L, Zhang L, et al. Aryl hydrocarbon receptor negatively regulates NLRP3 inflammasome activity by inhibiting NLRP3 transcription. Nat Commun 5, 4738 (2014)
Byrne CD, Targher G. NAFLD: a multisystem disease. J Hepatol 62, S47–S64 (2015)
Cobbina E, Akhlaghi F. Non-alcoholic fatty liver disease (NAFLD) - pathogenesis, classification, and effect on drug metabolizing enzymes and transporters. Drug Metab Rev 49, 197–211 (2017)
Hartner JC, Schmittwolf C, Kispert A, Muller AM, Higuchi M, Seeburg PH. Liver disintegration in the mouse embryo caused by deficiency in the RNA-editing enzyme ADAR1. J Biol Chem 279, 4894–4902 (2004)
Wang Q, Miyakoda M, Yang W, Khillan J, Stachura DL, Weiss MJ, et al. Stress-induced apoptosis associated with null mutation of ADAR1 RNA editing deaminase gene. J Biol Chem 279, 4952–4961 (2004)
Nie Y, Hammond GL, Yang JH. Double-stranded RNA deaminase ADAR1 increases host susceptibility to virus infection. J Virol 81, 917–923 (2007)
Gatsiou A, Vlachogiannis N, Lunella FF, Sachse M, Stellos K. Adenosine-to-Inosine RNA Editing in Health and Disease. Antioxid Redox Signal 29, 846–863 (2018)
Yu Z, Luo R, Li Y, Li X, Yang Z, Peng J, et al. ADAR1 inhibits adipogenesis and obesity by interacting with Dicer to promote the maturation of miR-155-5P. J Cell Sci 135, jcs259333 (2022)
Meltzer M, Long K, Nie Y, Gupta M, Yang J, Montano M. The RNA editor gene ADAR1 is induced in myoblasts by inflammatory ligands and buffers stress response. Clin Transl Sci 3, 73–80 (2010)
Broz P, Dixit VM. Inflammasomes: mechanism of assembly, regulation and signalling. Nat Rev Immunol 16, 407–420 (2016)
Rathinam VA, Fitzgerald KA. Inflammasome Complexes: Emerging Mechanisms and Effector Functions. Cell 165, 792–800 (2016)
Neumann K, Schiller B, Tiegs G. NLRP3 Inflammasome and IL-33: Novel Players in Sterile Liver Inflammation. Int J Mol Sci 19, 2732 (2018)
Dorn C, Engelmann JC, Saugspier M, Koch A, Hartmann A, Muller M, et al. Increased expression of c-Jun in nonalcoholic fatty liver disease. Lab Invest 94, 394–408 (2014)
Singh R, Wang Y, Xiang Y, Tanaka KE, Gaarde WA, Czaja MJ. Differential effects of JNK1 and JNK2 inhibition on murine steatohepatitis and insulin resistance. Hepatology 49, 87–96 (2009)
Karin M, Gallagher E. From JNK to pay dirt: jun kinases, their biochemistry, physiology and clinical importance. Iubmb Life 57, 283–295 (2005)
Min L, Ji Y, Bakiri L, Qiu Z, Cen J, Chen X, et al. Liver cancer initiation is controlled by AP-1 through SIRT6-dependent inhibition of survivin. Nat Cell Biol 14, 1203–1211 (2012)
Lin Z, Altaf N, Li C, Chen M, Pan L, Wang D, et al. Hydrogen sulfide attenuates oxidative stress-induced NLRP3 inflammasome activation via S-sulfhydrating c-Jun at Cys269 in macrophages. Biochim Biophys Acta Mol Basis Dis 1864, 2890–2900 (2018)
Jin X, Wang C, Wu W, Liu T, Ji B, Zhou F. Cyanidin-3-glucoside Alleviates 4-Hydroxyhexenal-Induced NLRP3 Inflammasome Activation via JNK-c-Jun/AP-1 Pathway in Human Retinal Pigment Epithelial Cells. J Immunol Res 2018, 5604610 (2018)
Wang QS, Luo XY, Fu H, Luo Q, Wang MQ, Zou DY. MiR-139 protects against oxygen-glucose deprivation/reoxygenation (OGD/R)-induced nerve injury through targeting c-Jun to inhibit NLRP3 inflammasome activation. J Stroke Cerebrovasc Dis 29, 105037 (2020)
Funding
This study was funded by the National Natural Science Foundation of China (81970403, 81400831, 82000427), National Natural Science Foundation of Hunan province (2020JJ5785, 2021JJ40948), Changsha Municipal Natural Science Foundation (kq2014256), Scientific research project of Hunan Provincial Health Commission (202203065271), the Fundamental Research Funds for the Central Universities of Central South University (2021zzts0079) and Hunan Provincial Innovation Foundation for Postgraduate (CX20210177).
Author information
Authors and Affiliations
Contributions
FW and RX conceived and designed this study. YXL conducted experiments. LLF contributed new reagents or analytical tools. BYJ analyzed the data. FW and YXL wrote the manuscript. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
This study was approved by the Ethics Committee of the Third Xiangya Hospital of the Central South University. Written informed consent was obtained from all study participants. All animal protocols were approved by the Institutional Animal Care and Use Committee of Central South University.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Xiang, R., Liu, Y., Fan, L. et al. RNA adenosine deaminase (ADAR1) alleviates high-fat diet-induced nonalcoholic fatty liver disease by inhibiting NLRP3 inflammasome. Lab Invest 102, 1088–1100 (2022). https://doi.org/10.1038/s41374-022-00805-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41374-022-00805-8
This article is cited by
-
Deficiency of ADAR2 ameliorates metabolic-associated fatty liver disease via AMPK signaling pathways in obese mice
Communications Biology (2024)