Abstract
Programmed death-ligand 1 (PD-L1) is constitutively expressed by hypoxia-inducible factor 2α (HIF2α). It can be induced by interferon gamma (IFNγ) signaling in clear cell renal cell carcinoma (ccRCC). Clinical trials of metastatic ccRCCs have suggested that a canonical IFNγ signature is a better biomarker for therapeutic response to immune checkpoint inhibitors (ICIs) than PD-L1 expression levels in tumor cells. To understand the therapeutic response to ICIs according to PD-L1 expression levels, we analyzed transcriptional regulation of the PD-L1 promoter by HIF2α and IFNγ-inducible interferon regulatory factor-1 (IRF-1) in ccRCC cells. Here, we present two ccRCC cell models showing differential PD-L1 expression levels in response to IFNγ and hypoxia. Analysis of The Cancer Genome Atlas RNA-sequencing data revealed that PD-L1 expression correlated with JAK2 and STAT1 expression of the canonical IFNγ signature in ccRCC tissues. Upon IFNγ stimulation, PD-L1 was induced by sequential activation of JAK2/STAT1/IRF-1 signaling in both WT- and Mut- VHL ccRCC cells. IFNγ activated the IRF-1α site of the PD-L1 promoter. The IFNγ-mediated increase of PD-L1 expression in Mut-VHL cells was 4.8-fold greater than that in WT-VHL cells. Under normoxia condition, PD-L1 expression in Mut-VHL cells was significantly higher than that in WT-VHL cells due to high basal HIF2α expression. Under hypoxia condition, PD-L1 expression in WT-VHL cells was induced up to 1.8-fold through activation of hypoxia-response elements 2 and 3. In contrast, although PD-L1 in Mut-VHL cells was already highly expressed in the basal state through activation of hypoxia-response elements 2, 3, and 4, it was no longer induced by hypoxia. In conclusion, Mut-VHL ccRCC cells displayed higher PD-L1 expression due to high basal HIF2α expression and a stronger response to IFNγ stimulation than WT-VHL cells. The fact that HIF2α antagonists can potentially reduce PD-L1 expression levels should be considered in ICI combination therapy.
Similar content being viewed by others
Introduction
Programmed death ligand-1 (PD-L1) is a glycoprotein receptor encoded by the CD274 gene on chromosome 9p24.1, which is usually expressed on the cell surface of some epithelial cells under inflammatory conditions, as well as, in activated B and T lymphocytes, macrophages, and dendritic cells. PD-L1 binds to programmed cell death protein 1 (PD-1) on activated T cells, especially CD8 (+) cytotoxic T lymphocytes, leading to suppression of an effective immune response1,2,3,4. PD-L1 is upregulated in various epithelial and lymphoid tumors, including clear cell renal cell carcinomas (ccRCCs)5,6. Anti-PD-1 antibodies such as nivolumab and pembrolizumab that can disrupt the PD-L1/PD-1 axis in combination with anti-cytotoxic T lymphocyte antigen-4 or vascular endothelial growth factor (VEGF) receptor tyrosine kinase inhibitors (TKIs), have been approved by the United States Food and Drug Administration as immune checkpoint inhibitors (ICIs) for patients with metastatic ccRCC1,7. Nivolumab has also been authorized as second-line treatment alone after the failure of VEGF receptor-TKIs in patients with advanced or metastatic ccRCC8.
PD-L1 is constitutively expressed and/or induced upon extracellular interferon gamma (IFNγ) and hypoxia stimulation. Constitutive PD-L1 expression is regulated by hypoxia-inducible factor 2α (HIF2α) in von Hippel-Lindau (VHL)-deficient ccRCC cells because VHL gene mutations lead to the accumulation of HIF2α in cancer cells9,10. HIF2α binds to the hypoxia-response elements (HREs) in the PD-L1 promoter, leading to PD-L1 expression10. Moreover, PD-L1 expression can be induced by extracellular signals1. IFNγ is primarily secreted by tumor–infiltrating activated T cells such as CD8 cytotoxic T cells, CD4 T helper type 1 cells, γδ T cells, and natural killer cells1. IFNγ strikingly induces PD-L1 expression by activating the janus kinase 2 (JAK2)/signal transducer and activator of transcription 1 (STAT1)/interferon regulatory factor 1 (IRF-1) signaling pathway, particularly in Epstein Barr virus-positive gastric cancer (EBVGC) cells11 or by activating JAK2/STAT3 in triple-negative breast cancer with amplification of JAK2 and PD-L1 genes12. Taken together, it is well-known that constitutive PD-L1 expression is regulated by HIF2α in ccRCCs9,10; when ccRCC cells are exposed to hypoxia, the expression and accumulation of intracellular HIF2α increases, which can bind to HREs of the PD-L1 promoter and induce PD-L1 expression9,10. In addition, a bioinformatics study has suggested that PD-L1 expression could be induced by IFNγ in ccRCCs, very similar to that in EBVGCs11,13,14.
The level of PD-L1 expression in tumor cells is known to be the best biomarker for predicting the therapeutic response of patients receiving ICI therapy. However, clinical trials enrolling patients with metastatic ccRCC treated with ICIs and/or VEGF receptor TKIs have raised many questions regarding the use of PD-L1 expression as a readily available and good biomarker for patient selection15,16. For example, the KEYNOTE-426 trial of pembrolizumab + axitinib versus sunitinib did not show any difference in overall survival between tumor cell PD-L1-expressing and non-expressing groups, although the overall survival benefit of the combination therapy was evident in all patient subgroups15,16. In recent investigations using RNA-sequencing (RNA-seq) data, a canonical or non-canonical IFNγ mRNA signature rather than PD-L1 expression levels has emerged as a better biomarker in melanoma, non-small cell lung cancer, and urothelial carcinoma17,18,19,20.
To understand the therapeutic response of ICIs according to PD-L1 expression levels, we analyzed changes in PD-L1 expression levels according to the activation of the PD-L1 promoter by IFNγ and hypoxia in ccRCCs. First, through bioinformatics analysis of The Cancer Genome Atlas (TCGA) RNA-seq data from ccRCC tissues, we examined the association between two therapeutic biomarkers of ICIs. That is, we examined the association between PD-L1 expression and the expression of IFNγ signaling-related proteins, the so-called canonical IFNγ signature. Second, we investigated how PD-L1 expression was induced upon IFNγ stimulation and hypoxia exposure by analyzing the PD-L1 promoter containing IRF-1-binding interferon-stimulated response elements (ISREs) and HIF2α-binding HREs sites in ccRCC cells with or without VHL mutations. Finally, we proposed two ccRCC models showing differential PD-L1 expression levels in response to IFNγ and hypoxia.
Materials and methods
TCGA analysis via cBioportal
We analyzed TCGA data from 538 tissue samples of ccRCC (TCGA, Firehose Legacy ccRCC) using cBioportal, a web-based interface that enables integrative analysis of complex cancer genomics and clinical profiles21,22. OncoPrint was used to visualize genomic alterations and mRNA expression levels of queried genes in genomic and RNA-seq datasets from tissue samples to identify trends. For co-occurrence analysis of a pair of query genes, log2 odds ratios were calculated, which indicate the likelihood that the events of two genes were mutually exclusive or co-occurring in the selected case. In addition, p-values derived using Fisher’s exact test and q-values derived using Benjamini Hochberg false discovery rate correction procedure for multiple comparisons were calculated to assess statistical significance. Spearman’s correlation coefficient and Pearson’s correlation coefficient were obtained for correlation analysis between two genes. R square value was also calculated.
Cell culture
ACHN, Caki-1, Caki-2, A498, and SNU719 cell lines were purchased from the Korean Cell Line Bank (Seoul, Korea). Caki-1, Caki-2, and SNU719 cells were maintained in RPMI medium (Welgene, Seoul, Korea), while ACHN and A498 cells were maintained in Dulbecco’s modified Eagle’s medium (Welgene). All growth media were supplemented with 10% fetal bovine serum (Welgene) and 1% antibiotic–antimycotic solution (Thermo Fisher Scientific, Waltham, MA, USA).
Reverse transcription-quantitative polymerase chain reaction (RT-qPCR)
Total mRNA was isolated from cancer cells using the RNeasy® Mini Kit (Qiagen, Valencia, CA, USA). First-strand cDNA was synthesized from 2 μg total RNA using a LeGene Premium Express cDNA synthesis system (LeGene Biosciences, San Diego, CA, USA) in a 20 μL reaction containing random hexamer primers. RT-qPCRs were conducted in a 20 μL reaction with 1 μL cDNA, 1 μL of each 10 μM primer, and 10 μL SYBR® Green Master Mix (Applied Biosystems, Foster City, CA, USA) using a StepOnePlus® Real-Time PCR system (Thermo Fisher Scientific). The PCR program was as follows: denaturation at 95 °C for 2 min, followed by 40 cycles of 95 °C for 10 s, 60 °C for 15 s, and 72 °C for 30 s. Each sample was run in triplicate. The expression of a specific gene was normalized to that of β-actin. Relative mRNA expression was determined using the comparative cycle threshold (2−ΔΔCT) method. Primers used in this study were the same as those described previously11.
Western blot analysis
Cells were lysed in radioimmunoprecipitation assay buffer (50 mM Tris-Cl pH 7.4, 0.1% NaN3, 1% Nonidet P-40, 0.25% sodium deoxycholate, 1 mM EDTA, 1 mM Na3VO4, 1 mM NaF, and protease inhibitor mixture) supplemented with 0.4 M NaCl. Cell lysates were centrifuged and resulting supernatants were subjected to western blot analysis. A total of 30 μg of cell lysate was resolved by SDS-PAGE and then transferred to polyvinylidene difluoride membranes. After membranes were blocked with 5% non-fat dry milk, they were incubated with anti-JAK2, STAT1, p-STAT1 (Y701), PD-L1 (Cell Signaling Technology, Beverly, MA, USA), and IRF-1 (Santa Cruz Biotechnology, Dallas, TX, USA) primary antibodies. Blots were probed with anti-β-actin (Sigma-Aldrich, Darmstadt, Germany) as a loading control. These membranes were incubated with anti-mouse or anti-rabbit HRP-conjugated secondary antibodies (Cell Signaling Technology) and visualized using an ECL detection kit (Thermo Fisher Scientific).
PD-L1 promoter luciferase reporter assay
Different fragments of the PD-L1 promoter were generated using overlap extension PCR and cloned into the pGL4.15-basic vector at KpnI and XhoI sites. Obtained clones were verified using DNA sequencing. Primer sequences are summarized in Supplementary Table S1. Cells were seeded into six-well plates and transfected with a 1 μg/well luciferase reporter plasmid. To normalize the transfection efficiency, cells were co-transfected with 100 ng of pRL-CMV (Renilla luciferase). At 4 h after transfection, the culture medium was replaced with complete medium with or without 10 ng/mL IFNγ (R&D Systems, Minneapolis, MN, USA) or 100 μM cobalt chloride (CoCl2), a hypoxia-mimetic chemical (Sigma-Aldrich)23. Twenty-four hours after IFNγ stimulation or hypoxia exposure, luciferase activity was measured using a Dual-Luciferase® Reporter Assay System kit (Promega, Madison, WI, USA) according to the manufacturer’s instructions.
Chromatin immunoprecipitation analysis
Using a chromatin immunoprecipitation (ChIP) assay kit (Millipore, Bedford, MA, USA), cells were cross-linked with 1% formaldehyde for 10 min at 37 °C, incubated in ChIP lysis buffer, and sonicated. Fragmented chromatin was immunoprecipitated with either anti-IRF-1 (Santa Cruz Biotechnology), IgG (Cell Signaling Technology) as a negative control, or anti-histone 3 antibody (Abcam, Cambridge, UK) as a positive control. After antibody-bound complexes were pulled down with protein A-agarose/salmon sperm DNA (Millipore) and cross-links were reversed, eluted DNA was quantified using RT-PCR. Primers used in RT-PCR were as the same as those described previously11.
Electrophoretic mobility shift assay (EMSA)
Nuclear extracts were obtained using a nuclear extract kit (Active Motif, Carlsbad, CA, USA) and incubated with a biotin-labeled probe for 20 min in binding buffer (Active Motif). For supershift analysis, extracts were incubated with an anti-IRF-1 antibody (Santa Cruz Biotechnology) and a control antibody (Cell Signaling Technology). Reaction mixtures were separated on a 6% acrylamide gel. Wild-type and mutant probe sequences were used as previously described11.
Blockade of IFNγ signaling with JAK2 and STAT1 inhibitors
Cells were seeded into six-well plates at a density of 1 × 106 cells and treated with 5, 10, and 20 μM of AZD1480 or fludarabine (Selleckchem, Houston, TX, USA) dissolved in dimethyl sulfoxide (DMSO) for 2 h, followed by treatment with or without 10 ng/mL IFNγ for 24 h. Cells were then harvested to analyze mRNA/protein expression. EMSA was conducted using nuclear extracts. DMSO was added to the culture medium alone as a vehicle control.
HIF2α chemical inhibition and siRNA silencing
Cells were seeded into six-well plates at a density of 1 × 106 cells and treated with 5 or 10 μM PT2385 (Selleckchem) dissolved in 100% DMSO for 24 h or transfected with HIF2α siRNA mixtures using Lipofectamine® RNAi MAX (Thermo Fisher Scientific) for 4 h. These cells were then harvested to analyze mRNA/protein expression.
For the PD-L1 promoter luciferase reporter assay, cells were transfected with the WT pPD-L1-luc vector, as previously mentioned11. At 4 h after transfection, cells were treated with 5 or 10 μM PT2385 (Selleckchem) for 24 h or transfected with HIF2α siRNAs. Luciferase activity was measured. HIF2α siRNAs used in this study are described in Supplementary Table S2.
Statistical analysis
All experiments were repeated at least thrice. Mean values were compared using two-tailed Student’s t-tests and one-way analysis of variance (with Tukey’s post-hoc test). Differences were considered statistically significant at p < 0.05. All statistical analyses were performed using SPSS software for Windows 10.0 (IBM, Chicago, IL, USA).
Results
PD-L1 mRNA expression correlates with the canonical IFNγ mRNA signature required for the IFNγ response in ccRCC tissues
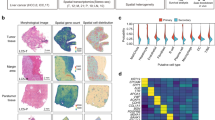
OncoPrint analysis using TCGA RNA-seq data from 538 ccRCC tissue samples showed genomic alterations and mRNA expression of PD-L1, VHL, and canonical IFNγ signaling genes including IFNγ, interferon gamma receptor 1 (IFNGR1), IFNGR2, JAK2, STAT1, and IRF-1. In OncoPrint, patients with high PD-L1 (CD274) mRNA expression were subdivided into mutant (Mut) and wild-type (WT) VHL subgroups according to the presence or absence of VHL mutations. The black box represents the Mut-VHL subgroup of patients with high PD-L1 mRNA expression. High expression levels of JAK2 and STAT1 mRNAs belonging to the canonical IFNγ signature were more frequent in the Mut-VHL subgroup than in the WT-VHL subgroup (Fig. 1A). Co-occurrence analysis showed that PD-L1 mRNA expression significantly co-occurred with JAK2 or STAT1 mRNA expression (P < 0.001, q < 0.001) (Fig. 1B). In addition, PD-L1 mRNA expression was significantly associated with JAK2 (Spearman’s correlation coefficient r = 0.58, Pearson’s correlation coefficient r = 0.60, R2 = 0.35, P < 0.001) or STAT1 (Spearman’s correlation coefficient r = 0.46, Pearson’s correlation coefficient r = 0.46, R2 = 0.21, P < 0.001) mRNA expression.
A OncoPrint represents genomic alterations and mRNA expression of PD-L1 (CD274), VHL, and canonical IFNγ signaling genes, including IFNγ, IFNGR1, IFNGR2, JAK2, STAT1, and IRF-1 in 538 tissue samples of ccRCC (TCGA, Firehose Legacy). The black box shows that canonical IFNγ mRNA signature is more frequent in the Mut-VHL subgroup of patients with high PD-L1 mRNA expression than in the WT-VHL subgroup. B Co-occurrence analysis revealed that PD-L1 (CD274) mRNA expression was significantly associated with that of JAK2 or STAT1. C Relative protein expression levels of IFNGR1, IFNGR2, JAK2, STAT1, and IRF-1 in Caki-1 and A498 ccRCC cells and ACHN and Caki-2 pRCC cells were determined using western blot. D Relative protein expression levels of HIF2α and PD-L1 in Caki-1 and A498 ccRCC cells and ACHN and Caki-2 pRCC cells were determined using western blot. All experiments were performed in triplicate. β-actin protein was used as a loading control.
For cancer cells to respond to IFNγ stimulation, IFNGR1/IFNGR2/JAK2/STAT1/IRF-1 signaling proteins need to be expressed in the basal state. Thus, we examined expression levels of canonical IFNγ signaling proteins in ccRCC and papillary renal cell carcinoma (pRCC) cell lines24,25,26,27. Caki-1 and A498 ccRCC cell lines and ACHN and Caki-2 pRCC cell lines could be reclassified as WT-VHL ACHN and Caki-1 and Mut-VHL Caki-2 and A498 cells24,25,26,27. Canonical IFNγ signaling proteins including IFNGR1, IFNGR2, JAK2, STAT1, and IRF-1 were constitutively expressed at varying levels in ACHN, Caki-1, Caki-2, and A498 cells. In particular, Caki-2 pRCC cell line did not express STAT1, indicating that this cell line would not respond to IFNγ stimulation, whereas A498 ccRCC cells showed higher levels of IFNGR2, JAK2, STAT1, and IRF-1 expression in the basal state than other cell lines (Fig. 1C).
HIF2α is known to be a driver of carcinogenesis and is a well-known major transcription factor of PD-L1 expression in ccRCC. Under normal oxygen conditions, we investigated whether HIF2α and PD-L1 were constitutively expressed in ccRCC and pRCC cell lines. PD-L1 protein in the basal state was constitutively overexpressed in ACHN and Caki-2 pRCC cells as well as Mut-VHL A498 ccRCC cells. However, PD-L1 was least expressed in WT-VHL Caki-1 ccRCC cells. As expected, constitutive HIF2α and PD-L1 expression levels in Mut-VHL A498 cells were higher than those in WT-VHL Caki-1 cells (Fig. 1D).
IFNγ induces significantly higher PD-L1 expression through JAK2/STAT1/IRF-1 signaling in Mut-VHL than in WT-VHL ccRCC cells
WT-VHL Caki-1 and Mut-VHL A498 ccRCC cell lines were selected for the experiment of PD-L1 induction upon IFNγ stimulation. IFNγ-non-reactive Caki-2 pRCC cells were used as a negative control, while the IFNγ-sensitive EBVGC SNU719 cell line was used as a positive control11. We have previously determined the appropriate concentration and duration of IFNγ stimulation using EBVGC SNU719 cells11. Effects of IFNγ stimulation on PD-L1 expression were dose-dependent. The optimal concentration of IFNγ used for stimulation was 10 ng/mL. PD-L1 mRNA level was found to reach a peak at 24 h after stimulation. We examined expression of JAK2, STAT1, IRF-1, and PD-L1 post-IFNγ treatment in ccRCC and control cell lines. Similar to positive control SNU719 cells, IFNγ stimulation dose-dependently induced mRNA and protein levels of JAK2, STAT1, IRF-1, and PD-L1 in Caki-1 and A498 cells (Fig. 2A). The signaling response of A498 cells to IFNγ was much more sensitive and stronger than that of Caki-1 cells. When 10 ng/mL IFNγ was administered to ccRCC cells for 24 h, the level of PD-L1 mRNA in A498 cells was increased ~30-fold as compared to that in unstimulated A498 cells, whereas that in Caki-1 cells was increased sixfold (Fig. 2A) (Supplementary Table S3). In Caki-2 pRCC cells, when IFNγ concentration was increased from 5 to 10 ng/mL, mRNA and protein levels of JAK2 were also increased, but not those of STAT1 and IRF-1. Since STAT1 was not expressed in the basal state of Caki-2 cells (Fig. 1C), it was expected that the signal would not be transduced to STAT1 and IRF-1 downstream of JAK2 upon IFNγ stimulation (Fig. 2A).
A After treatment with increasing doses of IFNγ for 24 h, relative mRNA and protein expression levels of JAK2, STAT1, IRF-1, and PD-L1 in Caki-1 ccRCC, A498 ccRCC, Caki-2 pRCC, and SNU719 cells were determined using RT-qPCR and western blot, respectively. Upon detection using western blot, PD-L1 showed a heterogeneous pattern ranging from a fully glycosylated ~50 kDa to a non-glycosylated ~33 kDa form due to the varying degree of N-linked glycosylation among cancer cell lines. IFNγ non-responsive Caki-2 and highly responsive SNU719 cells were used as negative and positive controls, respectively. β-actin was used as a loading control for mRNA and protein. All experiments were performed in triplicate. The mRNA data are presented as mean ± SD (n = 3). B Schematic representation of deletion reporter constructs and nucleotide sequence of the PD-L1 promoter. The transcription start site is indicated using +1. Underlined sequences represent two putative IRF-1 binding ISREs designated as IRF-1α (−171 to −159) and IRF-1β (−109 to −97), which were deleted one by one or both using PCR-driven overlap extension, to create reporter constructs ΔIRF-1α pPD-L1-luc, ΔIRF-1β pPD-L1-luc, and ΔIRF-1αβ pPD-L1-luc. C WT-VHL Caki-1, Mut-VHL A498, negative control Caki-2, and positive control SNU719 cells were transfected with the wild-type PD-L1 promoter luciferase vector (WT pPD-L1-luc). At 4 h after transfection, these cells were stimulated with increasing doses of IFNγ for 24 h. D Caki-1, A498, Caki-2, and SNU719 cells were transfected with the WT and ISRE-deleted pPD-L1-luc vectors. At 4 h after transfection, these cells were stimulated with 10 ng/mL IFNγ for an additional 24 h. Relative luciferase activities determined using the dual firefly/renilla luciferase assay were compared to that of the pGL4.15 basic reporter. Data are presented as mean ± SD (n = 3). *P < 0.05; **P < 0.01; and ***P < 0.001 compared to WT pPD-L1-luc reporter (Student’s t test and ANOVA).
IFNγ activates the IRF-1α ISRE of the PD-L1 promoter in ccRCC cells
To determine whether putative ISREs might be functional, we compared PD-L1 promoter activity upon stimulation of PD-L1 promoter luciferase (pPD-L1-luc) constructs-transfected ccRCC cells with IFNγ (Fig. 2B–D). The PD-L1 promoter sequence from −456 to +151 contains two IRF-1 binding ISRE sites designated as IRF-1α and IRF-1β sites. The 607-bp promoter sequence was amplified using PCR and cloned into the pGL4.15 basic luciferase vector. IRF-1α site, IRF-1β site, or both was deleted using PCR-driven overlap extension (Fig. 2B). After WT-VHL Caki-1, Mut-VHL A498, Caki-2, and SNU719 cells were transfected with WT pPD-L1-luc, these cells were stimulated with increasing doses of IFNγ. In Caki-1 and A498 cells, PD-L1 promoter activity increased proportionally with increasing concentrations of IFNγ. Promoter activity increased sharply in A498 cells as compared to that in Caki-1 cells. In Caki-2 pRCC cells refractory to IFNγ stimulation, there was no change in the promoter activity as expected. However, the activity level of the PD-L1 promoter was increased most steeply in SNU719 cells. When 10 ng/mL IFNγ was used to treat WT pPD-L1-luc-transfected cells for 24 h, the PD-L1 promoter activity increased approximately 1.7-fold in Caki-1 cells as compared to that in the unstimulated state. On the other hand, it increased approximately 3.1-times in A498 cells. In addition, the PD-L1 promoter activity showed no change in Caki-2 cells, while it displayed an increase from threefold to a maximum of fivefold in SNU719 cells (Figs. 2C, and 5B) (Supplementary Table S3).
In the basal state, luciferase activities of ISRE-deleted PD-L1 promoter constructs in Caki-1, Caki-2, and SNU719 cells showed no significant changes compared to the WT PD-L1 promoter activity, whereas luciferase activities of ΔIRF-1α and ΔIRF-1αβ pPD-L1-luc in A498 cells decreased by 50% and 52%, respectively, compared to WT pPD-L1-luc activity (Fig. 2D). These results suggest that even in the absence of IFNγ stimulation in A498 cells, some constitutive IRF-1 might have bound to the IRF-1α site of the PD-L1 promoter. However, this possibility needs to be verified by combining results of chromatin immunoprecipitation and EMSA experiments.
When IFNγ was administered to ccRCC cells and control cells transfected with pPD-L1-luc constructs, luciferase activities of WT and ΔIRF-1β pPD-L1-luc reporters increased ~1.7-fold and 3.1-fold as compared to those in basal states of Caki-1 and A498 cells, respectively. Luciferase activities of ΔIRF-1α and ΔIRF-1αβ pPD-L1-luc reporters were significantly reduced to those in unstimulated WT pPD-L1-luc-transfected Caki-1 and A498 cells as compared to that of the WT pPD-L1-luc reporter (Fig. 2D) (Supplementary Table S3). These results indicate that IFNγ-induced IRF-1 can bind to the IRF-1α element of the PD-L1 promoter rather than the IRF-1β element in both Caki-1 and A498 cells.
IFNγ-induced IRF-1 binds to the IRF-1α sequence of PD-L1 promoter in ccRCC cells in vivo and in vitro
Chromatin immunoprecipitation assays were performed using IFNγ-stimulated or -unstimulated Caki-1, A498, Caki-2, and SNU719 cells, to validate whether IFNγ-induced IRF-1 could bind to IRF-1α in the PD-L1 promoter in vivo. IRF-1 did not specifically bind to IRF-1α or IRF-1β ISRE in the basal state of Caki-1 or A498 cells. In contrast, upon IFNγ stimulation in Caki-1 and A498 cells, large amounts of IRF-1 bound only to the IRF-1α site of the PD-L1 promoter. Caki-2 and SNU719 nuclear extracts were used as negative and positive controls, respectively. IgG and anti-H3 antibodies were used as experimental controls. Data were normalized to DNA input results (Fig. 3A).
A Chromatin immunoprecipitation assays were performed using IFNγ-stimulated or -unstimulated Caki-1, A498, Caki-2, and SNU719 cells. The latter two were used as negative and positive control cells, respectively. Anti-IRF-1 antibody and sequence-specific primers (IRF-1α and IRF-1β) targeting the IRF-1 binding ISREs of the PD-L1 promoter were used. After the ccRCC and control cells were stimulated with 10 ng/mL IFNγ for 24 h, immunoprecipitated DNA–protein complexes were analyzed using RT-PCR and normalized against input DNA. IgG and anti-H3 antibody were used as negative and positive controls, respectively. DW, distilled water. B Competitive electrophoretic mobility shift assays (EMSAs) for evaluating the specificity of the IRF-1α binding site were performed with IFNγ-stimulated or -unstimulated Caki-1 and SNU719 nuclear extracts and biotin-labeled WT (Hot), unlabeled WT (Cold), unlabeled mutant (Cold mut), and biotin-labeled mutant (Hot mut) IRF-1α probes. C Supershift assays for evaluating the binding of IRF-1 at the IRF-1α ISRE were performed using IFNγ-stimulated or -unstimulated Caki-1 and SNU719 nuclear extracts, WT IRF-1α probe, and anti-IRF-1 antibody. IgG was used as a negative control. Arrowheads and asterisks indicate DNA-protein complexes.
To investigate whether IRF-1 could specifically bind to the PD-L1 promoter region upon IFNγ stimulation in vitro, we conducted EMSA, competitive EMSA, and supershift assays. When stimulated with IFNγ, IRF-1 in nuclear proteins of Caki-1 and A498 cells could bind to the IRF-1α DNA probe significantly more than it did to the IRF-1β DNA probe to form a protein–DNA complex. Nonspecific protein–DNA complexes of various sizes were observed in the negative control Caki-2 nuclear extracts regardless of IFNγ stimulation. A single band of protein-DNA complexes was clearly seen in the nuclear extracts of the positive control SNU719 cells due to IFNγ-induced binding of IRF-1 to IRF-1α DNA probes (Supplementary Fig. 1).
We then performed competitive EMSA and supershift assays using Caki-1 and SNU-719 nuclear extracts to confirm whether IFNγ-induced IRF-1 could specifically bind to the IRF-1α site. Protein-DNA complexes formed by the binding of IFNγ-stimulated Caki-1 and SNU719 nuclear extracts with biotin-labeled hot wild type IRF-1α probes were competitively inhibited with over 50-fold excess of unlabeled cold wild type IRF-1α probes. Complex formation was restored with unlabeled cold mutant IRF-1α probes. No protein–DNA complex was formed when only biotin-labeled hot mutant IRF-1α probes were added to IFNγ-stimulated nuclear extracts (Fig. 3B). In addition, supershift EMSA analysis showed a decrease in the level of IRF-1–DNA complexes in the presence of anti-IRF-1 antibodies as compared to the presence of IgG (Fig. 3C). These results indicate that IFNγ-induced IRF-1 can specifically bind to the IRF-1α site of the PD-L1 promoter in ccRCC cells.
IFNγ-induced PD-L1 signaling is reduced in the presence of JAK2 and STAT1 inhibitors
To verify whether IFNγ-induced PD-L1 is upregulated through the sequential activation of JAK2/STAT1/IRF-1 signaling in ccRCC, we investigated changes in WT PD-L1 promoter activity and JAK2/STAT1/IRF-1/PD-L1 signaling mRNA and protein expression in IFNγ-stimulated Caki-1 cells after pretreatment with a JAK2 inhibitor AZD1480, a STAT1 inhibitor fludarabine, or both.
When Caki-1 cells were pretreated with AZD1480, fludarabine, or both and then stimulated with IFNγ, the luciferase activity of WT pPD-L1-luc was significantly decreased in a dose-dependent manner as compared to that seen in Caki-1 cells stimulated with IFNγ without pretreatment with AZD1480 and/or fludarabine (Fig. 4A). Upon IFNγ stimulation, AZD1480 and/or fludarabine significantly prevented the overexpression of JAK2, STAT1, IRF-1, and PD-L1 mRNAs and proteins as well as the phosphorylation of STAT1 (Fig. 4B, C). Given that AZD1480 competes with ATP for binding to JAK2 and fludarabine depletes STAT1 mRNA and protein, it was noteworthy that the expression of JAK2 mRNA or protein itself decreased when AZD1480 or fludarabine was used. These results suggest the possible existence of a positive feedback loop through JAK2 in IFNγ/JAK2/STAT1/IRF-1/PD-L1 signaling. In addition, in EMSA experiments, AZD1480 and/or fludarabine reduced the formation of IRF-1 protein–IRF-1α DNA complexes, although these complexes were increased upon IFNγ stimulation (Fig. 4D). These results demonstrate that IFNγ-induced PD-L1 expression is mediated by the sequential expression of JAK2/STAT1/IRF1 signaling in ccRCC.
A Caki-1 cells transfected with WT pPD-L1-luc vector were treated with increasing concentrations (5, 10, and 20 μM) of AZD1480 (an ATP-competitive JAK2 inhibitor) or fludarabine (a STAT1 inhibitor that depletes STAT1 mRNA and protein) for 2 h, followed by stimulation with 10 ng/mL IFNγ for 24 h and measurement of luciferase activities. B, C After treatments with 10 μM of AZD1480 and/or fludarabine for 24 h, mRNA and protein levels of JAK2, STAT1, IRF-1, and PD-L1 in Caki-1 cells were determined using RT-qPCR and immunoblotting, respectively. D EMSAs were performed using nuclear extracts from IFNγ-stimulated or -unstimulated Caki-1 cells and biotin-labeled double-strand oligonucleotide IRF-1α probes, after pretreatment with 10 μM of AZD1480 and/or fludarabine for 24 h. Data are presented as mean ± SD (n = 3). *P < 0.05; **P < 0.01; and ***P < 0.001 compared to IFNγ-stimulated Caki-1 cells (Student’s t test and ANOVA). Arrowhead indicates DNA–protein complexes.
Hypoxia further induces PD-L1 overexpression by activating HREs of the PD-L1 promoter in WT-VHL cells, but not in Mut-VHL ccRCC cells
We investigated activation patterns of the PD-L1 promoter for hypoxia and identified HRE sites that were activated in WT-VHL and Mut-VHL ccRCC cells under basal and hypoxic conditions. Three ΔHRE pPD-L1-luc reporters were created by deleting both IRF-1α (−171 to −159) and IRF-1β (−109 to −97) ISREs to eliminate effects of IFNγ stimulation, followed by removal of three HRE sites, resulting in reporters designated as ΔHRE 4-, ΔHRE 3, 4-, and ΔHRE 2, 3, 4-pPD-L1-luc reporters (Fig. 5A).
A Schematic representation of pPD-L1-luc reporter created by first deleting both IRF-1α (−171 to −159) and IRF-1β (−109 to −97) ISREs and then sequentially removing four HREs. The PD-L1 nucleotide sequence containing 456-bp upstream and 151-bp downstream of the transcription start site (+1) is shown. Underlined sequences represent two IRF-1 binding ISREs (IRF-1α and -1β) and four HREs (HRE 1, 2, 3, and 4). B Caki-1, A498, Caki-2, and SNU719 cells were transfected with WT pPD-L1-luc reporter. At 4 h after transfection, cells were stimulated with 10 ng/mL IFNγ or 100 μM CoCl2 for an additional 24 h. C Under the same conditions as B, cells were transfected with HREs-deleted pPD-L1-luc reporters. Relative luciferase activities compared to pGL4.15 basic vector are shown. Data are expressed as mean ± SD (n = 3). *P < 0.05; **P < 0.01; and ***P < 0.001 compared to WT pPD-L1-luc reporter (Student’s t test and ANOVA).
When hypoxia was induced in Caki-1 cells, the PD-L1 promoter activity increased by ~1.4-times than that in normoxic cells. Upon stimulation with IFNγ, the PD-L1 promoter activity increased by 1.7-times than that in unstimulated cells. When hypoxia was induced in A498 cells, no significant change was observed in PD-L1 promoter activity. However, when A498 cells were stimulated with IFNγ, the PD-L1 promoter activity increased threefold as compared to that in unstimulated cells. When hypoxia was induced in Caki-2 and SNU719 cells, no significant change in PD-L1 promoter activity was observed. Upon IFNγ stimulation, the PD-L1 promoter activity in Caki-2 cells was not changed, but increased threefold in SNU719 cells as compared to that in unstimulated SNU719 cells. Taken together, these results indicate that PD-L1 promoter in Caki-1 cells could be activated by IFNγ and hypoxic stimulation. The PD-L1 promoter in A498 and SNU719 cells was only activated by IFNγ, but refractory to hypoxia. PD-L1 promoter in Caki-2 cells was refractory to both IFNγ and hypoxia stimulations (Fig. 5B) (Supplementary Table S3).
As mentioned earlier, hypoxia activated the WT PD-L1-luc vector in Caki-1 cells up to 1.4-times as compared to the basal state of Caki-1 cells. Even under basal and hypoxic conditions, the ΔHRE 4-PD-L1-luc reporter showed little activity change compared to the WT PD-L1-luc vector, while the luciferase activity of ΔHRE 3, 4- and ΔHRE 2, 3, 4-PD- L1-luc decreased in a step-wise manner. In particular, the activity of the ΔHRE 2, 3, 4- PD-L1-luc reporter was significantly reduced. This indicates that HRE 2 of the PD-L1 promoter is the most important functional site in WT-VHL ccRCC under both basal and hypoxic conditions. In contrast to Caki-1 cells, activities of the WT PD-L1-luc vector in A498, Caki-2, and SNU719 cells did not differ between normoxia and hypoxia. Activities of ΔHRE 4-, ΔHRE 3,4-, and ΔHRE 2, 3, 4-PD-L1-luc constructs sequentially decreased to approximately the same extent, starting with the activity of WT PD-L1-luc in A498 cells under normoxic and hypoxic conditions (Fig. 5C).
PD-L1 expression is downregulated in the presence of HIF2α antagonist and siRNAs in Mut-VHL ccRCC cells
To confirm the feasibility of ICI combination therapy with HIF2α antagonists in ccRCCs, we investigated whether PD-L1 expression was downregulated in A498 cells using HIF2α- dimerization antagonist PT2385 and HIF2α siRNAs. HIF2α, the only HIF expressed in Mut-VHL A498 cells, is not only the major transcription factor of PD-L1 gene, but also the driving force for ccRCC carcinogenesis. HIF2α target genes including PD-L1, PAI-1, VEGFα, CCND1, and SLC2A1 were used as positive controls for assessing effects of PT2385, while HIF1α target genes including LDHA and PGK1 were used as negative controls.
PT2385 and HIF2α siRNA treatments reduced expression levels of PD-L1, HIF2α itself, and HIF2α target genes, including PAI-1, VEGFα, CCND1, and SLC2A1 (Fig. 6A, B, D, E). These treatments also reduced the activity of the PD-L1 promoter (Fig. 6C, F). PD-L1 expression was further decreased when PT2385 was used for treatment at 10 μM instead of 5 μM (Fig. 6A, B, C). However, PT2385 and HIF2α siRNA treatments did not lead to any significant changes in mRNA expression levels of HIF1α or its target genes, including LDHA and PGK1 used as negative controls, in A498 cells (Fig. 6A, D).
A Mut-VHL A498 cells were treated with 5 μM or 10 μM of PT2385 for 24 h. The mRNA levels of HIF2α and its target genes PD-L1, PAI-1, VEGFα, CCND1, and SLC2A1 were measured using RT-qPCR. The mRNA levels of HIF1α and its target genes LDHA and PGK1 were used as negative controls. Data are presented as mean ± SD (n = 3). *P < 0.01 compared to DMSO-treated cells (Student’s t test). B Under the same conditions as A, protein levels of HIF2α and its target genes PD-L1 and PAI-1 were determined using western blot. C A498 cells were incubated with 5 μM or 10 μM of PT2385 for 2 h, and then transfected with WT pPD-L1-luc reporter. Relative luciferase activities compared to untreated WT pPD-L1-luc reporter are shown. Data are expressed as mean ± SD (n = 3). *P < 0.01, as compared to DMSO-treated cells (Student’s t test). D For gene silencing of HIF2α, A498 cells were transfected with HIF2α siRNAs #1 and #2 for 24 h. The mRNA levels of HIF2α and its target genes PD-L1, PAI-1, VEGFα, CCND1, and SLC2A1 were measured using RT-qPCR. The mRNA levels of HIF1α and its target genes LDHA and PGK1 were used as negative controls. Data are presented as mean ± SD (n = 3). *P < 0.01, as compared to scrambled siRNA-transfected cells (Student’s t test). E Under the same conditions as D, protein levels of HIF2α, PD-L1, and PAI-1 were determined using western blot. F A498 cells were transfected with HIF2α siRNAs #1 and #2 for 24 h, and then transfected with WT pPD-L1-luc for 24 h. Relative luciferase activities compared to untreated WT pPD-L1-luc reporter are shown. Data are expressed as mean ± SD (n = 3). *P < 0.05; **P < 0.01; and ***P < 0.001 compared to scrambled siRNA transfected cells (Student’s t test and ANOVA).
Since the mechanism of action of PT2385 and its analog PT2399 is to block the binding of HIF2α to HIF1β, they should not inhibit HIF2α expression itself, but only decrease the expression of HIF2α target genes such as PD-L1 and PAI-1. Interestingly, consistent with results of other studies28,29, our results confirmed that HIF2α mRNA and protein expression levels decreased upon PT2385 treatment (Fig. 6A, B). These results suggest that a positive feedback loop may exist between HIF2α and its target proteins in ccRCC. This positive feedback mechanism is currently under active investigation in our laboratory.
Two PD-L1 expression models are presented through PD-L1 promoter analysis
Our results suggested two cellular models with differential levels of PD-L1 expression, a well-known biomarker for predicting the therapeutic response of ICIs. Moreover, these cellular models revealed differences in PD-L1 expression mechanisms, providing clues to consider novel targets for ICI combination therapy. Dissection of the PD-L1 promoter showed that constitutive and inducible PD-L1 expression levels were differentially regulated by IFNγ and hypoxia signaling pathways in WT-VHL and Mut-VHL ccRCC cells (Fig. 7). Upon IFNγ stimulation, WT-VHL Caki-1 cells induced IRF-1, which bound to IRF-1α ISRE, leading to a 1.7-fold increase in PD-L1 promoter activity. In response to hypoxia, HIF2α activated HRE 2 predominantly and HRE 3 partly, resulting in a 1.4-fold increase in PD-L1 promoter activity. Thus, PD-L1 expression could be induced by both IFNγ and hypoxia in WT-VHL Caki-1 cells (Fig. 7, left panel) (Supplementary Table S3).
Left panel PD-L1 expression in WT-VHL Caki-1 cells is induced by both IFNγ and hypoxia. Right panel In contrast, PD-L1 expression in Mut-VHL A498 cells is induced only by IFNγ. Mut-VHL A498 cells show higher PD-L1 expression due to high basal HIF2α expression and a stronger response to IFNγ stimulation than WT-VHL Caki-1 cells.
On the other hand, when Mut-VHL A498 cells with high basal expression of IFNγ signaling proteins were stimulated using IFNγ, IRF-1 bound to IRF-1α ISRE, resulting in a 3.1-fold increase in PD-L1 promoter activity. However, in response to hypoxia, Mut-VHL A498 cells showed little change in PD-L1 promoter activity. A498 cells were already in a pseudohypoxic state because of the high basal HIF2α expression caused by VHL mutations. Thus, in the basal state, the PD-L1 promoter in Mut-VHL A498 cells was already saturated with basal HIF2α and activated HREs 2, 3, and 4 to a similar extent, resulting in a 2.7-fold higher basal PD-L1 promoter activity than that in WT-VHL Caki-1 cells. Therefore, PD-L1 expression in Mut-VHL A498 cells was induced only by IFNγ stimulation. It was no longer induced by hypoxia (Fig. 7, right panel) (Supplementary Table S3).
Discussion
Using TCGA RNA-seq data, we found that two therapeutic biomarkers of ICIs were closely related to each other. Specifically, PD-L1 expression correlated with the canonical IFNγ signature, JAK2, or STAT1 expression in ccRCC tissues. Under normoxia, PD-L1 expression was much higher in Mut-VHL ccRCC cells than in WT-VHL ccRCC cells because of the high basal expression of HIF2α, a well-known transcription factor of the PD-L1 promoter. Interestingly, basal expression levels of JAK2/STAT1/IRF-1 signaling proteins as components required for IFNγ-induced PD-L1 expression were higher in Mut-VHL cells than in WT-VHL cells. Although PD-L1 expression was induced by IFNγ and hypoxia stimulation in WT-VHL ccRCC cells, it was only induced by IFNγ in Mut-VHL ccRCC cells. The increase of PD-L1 expression upon IFNγ stimulation was significantly higher in Mut-VHL ccRCC cells than in WT-VHL ccRCC cells. To the best of our knowledge, this is the first study to demonstrate that IFNγ can induce PD-L1 overexpression by sequentially activating canonical JAK2/STAT1/IRF-1 signaling in both WT- and Mut-VHL ccRCC cells. The induced IRF-1 binds to the IRF-1α ISRE of the PD-L1 promoter in ccRCC cells. In contrast, constitutive and inducible HIF2α activates multiple HREs interspersed with the PD-L1 promoter.
Our study revealed that the level of constitutive PD-L1 expression was significantly higher in Mut-VHL cells than in WT-VHL cells. WT-VHL Caki-1 cells express HIF1α and HIF2α even under normal oxygen conditions. Expression levels of HIF1α and HIF2α are increased under hypoxia condition. Mut-VHL A498 cells fail to express the full-length HIF1α protein due to deletion of exons 2-6 of HIF1α under normoxia. In addition, HIF2α accumulates in these cells because the mutant VHL protein fails to degrade HIF2α30. Thus, A498 cells overexpress only HIF2α under normal oxygen conditions. Unlike Caki-1 cells, A498 cells exhibit a pseudohypoxic state without increasing HIF2α expression even under hypoxic exposure. Although HIF1α and HIF2α are transcription factors for PD-L1 expression in many cancers, only HIF2α, a key player in oncogenesis of ccRCC, is known to increase PD-L1 expression in ccRCCs9,10.
Recently, RCC cell lines are more accurately classified into ccRCC and pRCC by re-evaluating them with genomic data obtained from next-generation sequencing24,25,26,27. Therefore, in this experiment, WT-VHL Caki-1 and Mut-VHL A498 cells were used as ccRCC cells24,25,26,27, while WT-VHL ACHN and Mut-VHL Caki-2 cells were used as pRCC cell lines25,27. Caki-2 cell line exhibits high levels of MET and leucine-rich repeat kinase 2 expression as well as chromosome 8 aberrations and MYC activation31,32. Moreover, orthotopic and subcutaneous mouse xenografts using Caki-2 cells display typical features of cystic pRCC33. However, the lineage and genetic status of Caki-2 cells are still open to discussion.
Basal expression levels of JAK2/STAT1/IRF-1 signaling proteins required for IFNγ-induced PD-L1 expression were higher in Mut-VHL cells than in WT-VHL cells. For cancer cells to express PD-L1 upon IFNγ stimulation, IFNGR1, IFNGR2, JAK2, STAT1, and IRF-1 should be expressed to some extent even in the basal state of cells34,35. Therefore, it was expected that Mut-VHL A498 cells would respond better to IFNγ stimulation because they strongly expressed signaling components necessary for IFNγ response in the basal state than WT-VHL Caki-1 cells.
IFNγ induced PD-L1 overexpression via sequential activation of JAK2/STAT1/IRF-1 signaling in a dose-dependent manner in both WT-VHL and Mut-VHL ccRCC cells. A498 cells responded very sensitively to IFNγ stimulation and sequentially activated JAK2, STAT1, and IRF-1 to induce PD-L1 overexpression. Compared to WT-VHL Caki-1 cells, Mut-VHL A498 cells showed dramatic increases of JAK2, STAT1, IRF-1 and PD-L1 protein expression levels with increasing IFNγ concentration. The IFNγ response of Mut-VHL A498 cells was stronger than that of WT-VHL Caki-1 cells, similar to that of SNU719 cells, the positive control11. Caki-2 pRCC cells as the negative control with loss of STAT1 and polybromo 1 (PBRM1) expression (Supplementary Fig. 2) were refractory to IFNγ stimulation, consistent with results of other studies13,36,37. PBRM1 deficiency during IFNγ stimulation can inhibit STAT1 phosphorylation and reduce the binding of brahma-associated gene 1 (BRG1) to the IFNGR2 promoter, which in turn can decrease IFNGR2 expression levels. However, it does not affect the total protein level of JAK2, although it can reduce JAK2 phosphorylation probably due to decreased IFNGR2 expression37. In Caki-2 cells with loss of STAT1 and PBRM1 expression, our results showed that IFNγ stimulation increased JAK2 expression in a dose-dependent manner, although PD-L1 expression was not induced by IFNγ signaling activation. This is thought to be because STAT1 is an essential protein just downstream of JAK2 for IFNγ signaling, and PBRM1 is an enhancer of IFNGR2 expression that plays an important role in JAK2 and STAT1 phosphorylation.
IFNγ-induced IRF-1 binds to the functional IRF-1α sequence of the PD-L1 promoter in ccRCC cells both in vivo and in vitro. In contrast, most of the induced HIF2α in WT-VHL ccRCC cells specifically bind to HRE 2 and the rest to HRE 3, while constitutive HIF2α in Mut-VHL ccRCC cells bind to HRE 2, HRE 3, and HRE 4 to a similar extent. In fact, our results could not be compared with those of other studies as HRE 3 and HRE 4 regions of the PD-L1 promoter in ccRCC cells were not reported in other previous investigations10,38. Consistent with results of a previous study10, in which the HRE 2 site was named as HRE 4, using Mut-VHL 786-O cells expressing only HIF2α, HIF2α increased constitutive PD-L1 expression by binding directly to the same site we termed HRE 2 in the PD-L1 promoter. Therefore, HRE 2 is a functionally important site for constitutive and hypoxia-induced PD-L1 expression in ccRCC cells regardless of VHL mutations. Two previous studies have shown that PD-L1 expression is regulated by HIF2α in Mut-VHL A498 and 786-O cells expressing only HIF2α9,10. Moreover, results of these two studies were consistent with each other in that PD-L1 is regulated by HIF2α even in Mut-VHL RCC4, which expresses both HIF1α and HIF2α. However, these two studies showed contradictory results as to whether PD-L1 is regulated by HIF1α9,10. It is currently controversial as to whether RCC4 cells are true cellular models of ccRCC24,27.
This study has some limitations. First, cellular models using only IFNγ and HIF2α known to be major transcription factors for PD-L1 expression in ccRCCs were presented. However, other minor transcription factors may exist. Second, to make simpler and more intuitive cellular models, we did not perform experiments with more ccRCC cell lines. Third, effects of the non-canonical IFNγ signaling pathway on PD-L1 expression were excluded. Finally, although we analyzed TCGA RNA-seq data from ccRCC tissues to investigate the association between PD-L1 expression and canonical IFNγ signature, there is a need for subsequent large-scale validation of therapeutic effects of ICIs according to the expression of biomarker proteins.
In summary, our results revealed that two therapeutic biomarkers of ICIs were closely related to each other. PD-L1 expression was closely associated with the canonical IFNγ signature in ccRCC tissues. For an increase in PD-L1 expression upon IFNγ stimulation, IFNγ signaling component proteins should be expressed in the basal state of ccRCC cells. PD-L1 expression was much more induced upon IFNγ stimulation in Mut-VHL ccRCC cells than in WT-VHL ccRCC cells. Thus, Mut-VHL ccRCC cells displayed higher levels of PD-L1 expression due to high basal HIF2α expression and a stronger induction of IFNγ stimulation than WT-VHL ccRCC cells. Mut-VHL cells are expected to respond well to ICI therapy. The fact that HIF2α antagonists can potentially decrease PD-L1 expression levels should be considered in combination therapy with ICIs.
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author upon reasonable request. TCGA datasets are freely available at https://www.cbioportal.org/
References
Wu, Y., Chen, W., Xu, Z. P. & Gu, W. PD-L1 distribution and perspective for cancer immunotherapy-blockade, knockdown, or inhibition. Front. Immunol. 10, 2022 (2019).
Sun, C., Mezzadra, R. & Schumacher, T. N. Regulation and function of the PD-L1 checkpoint. Immunity 48, 434–452 (2018).
Yi, M., Niu, M., Xu, L., Luo, S. & Wu, K. Regulation of PD-L1 expression in the tumor microenvironment. J. Hematol. Oncol. 14, 10 (2021).
Chen, J., Jiang, C. C., Jin, L. & Zhang, X. D. Regulation of PD-L1: a novel role of pro-survival signalling in cancer. Ann. Oncol. 27, 409–416 (2016).
Stone, L. Kidney cancer: programmed death ligand 1 regulation in ccRCC. Nat. Rev. Urol. 13, 65 (2016).
Tatli Dogan, H. et al. Prognostic significance of the programmed death ligand 1 expression in clear cell renal cell carcinoma and correlation with the tumor microenvironment and hypoxia-inducible factor expression. Diagn. Pathol. 13, 60 (2018).
Pan, D. et al. A major chromatin regulator determines resistance of tumor cells to T cell-mediated killing. Science 359, 770–775 (2018).
Xu, J. X. et al. FDA approval summary: nivolumab in advanced renal cell carcinoma after anti-angiogenic therapy and exploratory predictive biomarker analysis. Oncologist 22, 311–317 (2017).
Ruf, M., Moch, H. & Schraml, P. PD-L1 expression is regulated by hypoxia inducible factor in clear cell renal cell carcinoma. Int. J. Cancer 139, 396–403 (2016).
Messai, Y. et al. Renal cell carcinoma programmed death-ligand 1, a new direct target of hypoxia-inducible factor-2 alpha, is regulated by von Hippel-Lindau gene mutation status. Eur. Urol. 70, 623–632 (2016).
Moon, J. W. et al. IFN gamma induces PD-L1 overexpression by JAK2/STAT1/IRF-1 signaling in EBV-positive gastric carcinoma. Sci. Rep. 7, 17810 (2017).
Chen, M. et al. JAK2 and PD-L1 amplification enhance the dynamic expression of PD-L1 in triple-negative breast cancer. Clin. Breast Cancer 18, e1205–e1215 (2018).
Hanze, J., Wegner, M., Noessner, E., Hofmann, R. & Hegele, A. Co-regulation of immune checkpoint PD-L1 with interferon-gamma signaling is associated with a survival benefit in renal cell cancer. Target Oncol. 15, 377–390 (2020).
Chen, S. et al. Mechanisms regulating PD-L1 expression on tumor and immune cells. J. Immunother. Cancer 7, 305 (2019).
Mori, K. et al. The predictive value of programmed death ligand 1 in patients with metastatic renal cell carcinoma treated with immune-checkpoint inhibitors: a systematic review and meta-analysis. Eur. Urol. 79, 783–792 (2021).
Albiges, L., Flippot, R. & Escudier, B. Immune checkpoint inhibitors in metastatic clear-cell renal cell carcinoma: is PD-L1 expression useful? Eur. Urol. 79, 793–795 (2021).
Grasso, C. S. et al. Conserved interferon-gamma signaling drives clinical response to immune checkpoint blockade therapy in melanoma. Cancer Cell 39, 122 (2021).
Higgs, B. W. et al. Interferon gamma messenger RNA signature in tumor biopsies predicts outcomes in patients with non-small cell lung carcinoma or urothelial cancer treated with durvalumab. Clin Cancer Res. 24, 3857–3866 (2018).
Ayers, M. et al. IFN-gamma-related mRNA profile predicts clinical response to PD-1 blockade. J. Clin. Invest. 127, 2930–2940 (2017).
Tu, M. M. et al. Molecular biomarkers of response to PD-1/PD-L1 immune checkpoint blockade in advanced bladder cancer. Bladder Cancer 5, 131–145 (2019).
Cerami, E. et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2, 401–404 (2012).
Gao, J. et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci. Signal. 6, pl1 (2013).
Wu, D. & Yotnda, P. Induction and testing of hypoxia in cell culture. J. Vis. Exp. 54, e2899 (2011).
Wolf, M. M., Kimryn Rathmell, W. & Beckermann, K. E. Modeling clear cell renal cell carcinoma and therapeutic implications. Oncogene 39, 3413–3426 (2020).
Brodziak, A. et al. Drug resistance in papillary RCC: from putative mechanisms to clinical practicalities. Nat. Rev. Urol. 16, 655–673 (2019).
Sinha, R. et al. Analysis of renal cancer cell lines from two major resources enables genomics-guided cell line selection. Nat. Commun. 8, 15165 (2017).
Brodaczewska, K. K., Szczylik, C., Fiedorowicz, M., Porta, C. & Czarnecka, A. M. Choosing the right cell line for renal cell cancer research. Mol. Cancer 15, 83 (2016).
Cho, H. et al. On-target efficacy of a HIF-2alpha antagonist in preclinical kidney cancer models. Nature 539, 107–111 (2016).
Wallace, E. M. et al. A small-molecule antagonist of HIF2alpha is efficacious in preclinical models of renal cell carcinoma. Cancer Res. 76, 5491–5500 (2016).
Shen, C. et al. Genetic and functional studies implicate HIF1alpha as a 14q kidney cancer suppressor gene. Cancer Discov. 1, 222–235 (2011).
Looyenga, B. D. et al. Chromosomal amplification of leucine-rich repeat kinase-2 (LRRK2) is required for oncogenic MET signaling in papillary renal and thyroid carcinomas. Proc Natl Acad Sci USA 108, 1439–1444 (2011).
Furge, K. A. et al. Detection of DNA copy number changes and oncogenic signaling abnormalities from gene expression data reveals MYC activation in high-grade papillary renal cell carcinoma. Cancer Res. 67, 3171–3176 (2007).
Pulkkanen, K. J. et al. HSV-tk gene therapy for human renal cell carcinoma in nude mice. Cancer Gene Ther. 8, 529–536 (2001).
Castro, F., Cardoso, A. P., Goncalves, R. M., Serre, K. & Oliveira, M. J. Interferon-gamma at the crossroads of tumor immune surveillance or evasion. Front. Immunol. 9, 847 (2018).
de Weerd, N. A. & Nguyen, T. The interferons and their receptors–distribution and regulation. Immunol. Cell Biol. 90, 483–491 (2012).
Dovhey, S. E., Ghosh, N. S. & Wright, K. L. Loss of interferon-gamma inducibility of TAP1 and LMP2 in a renal cell carcinoma cell line. Cancer Res. 60, 5789–5796 (2000).
Liu, X. D. et al. PBRM1 loss defines a nonimmunogenic tumor phenotype associated with checkpoint inhibitor resistance in renal carcinoma. Nat. Commun. 11, 2135 (2020).
Noman, M. Z. et al. PD-L1 is a novel direct target of HIF-1alpha, and its blockade under hypoxia enhanced MDSC-mediated T cell activation. J. Exp. Med. 211, 781–790 (2014).
Funding
This work was supported by the Korea Medical Device Development Fund grant funded by the Korea government (the Ministry of Science and ICT, the Ministry of Trade, Industry and Energy, the Ministry of Health & Welfare, and the Ministry of Food and Drug Safety) (9991007312, KMDF_PR_20200901_0133), Bridging Research Program through the Ministry of Education of the Republic of Korea and the National Research Foundation of Korea (NRF-2019R1H1A2101464), and the Korea University Ansan Hospital Grant (K1916631).
Author information
Authors and Affiliations
Contributions
S.K.K. and Y.S.K. conceptualized and designed the study; S.K.K., B.S.K., H.L., and H.J.K. performed experiments; S.K.K. and Y.S.K. wrote the manuscript; Y.S.K. supervised the study; All authors reviewed and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval and consent to participate
This study did not require ethical approval.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Kong, SK., Kim, B.S., Lim, H. et al. Dissection of PD-L1 promoter reveals differential transcriptional regulation of PD-L1 in VHL mutant clear cell renal cell carcinoma. Lab Invest 102, 352–362 (2022). https://doi.org/10.1038/s41374-021-00703-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41374-021-00703-5
This article is cited by
-
CD30 stimulation induces multinucleation and chromosomal instability in HTLV-1-infected cell lines
International Journal of Hematology (2023)