Abstract
Myxoid liposarcoma (MLS) is the second most common subtype of liposarcoma, accounting for ~6% of all sarcomas. MLS is characterized by a pathognomonic FUS–DDIT3, or rarely EWSR1–DDIT3, gene fusion. The presence of ≥5% hypercellular round cell areas is associated with a worse prognosis for the patient and is considered high grade. The prognostic significance of areas with moderately increased cellularity (intermediate) is currently unknown. Here we have applied matrix-assisted laser desorption/ionization mass spectrometry imaging to analyze the spatial distribution of N-linked glycans on an MLS microarray in order to identify molecular markers for tumor progression. Comparison of the N-glycan profiles revealed that increased relative abundances of high-mannose type glycans were associated with tumor progression. Concomitantly, an increase of the average number of mannoses on high-mannose glycans was observed. Although overall levels of complex-type glycans decreased, an increase of tri- and tetra-antennary N-glycans was observed with morphological tumor progression and increased tumor histological grade. The high abundance of tri-antennary N-glycan species was also associated with poor disease-specific survival. These findings mirror recent observations in colorectal cancer, breast cancer, ovarian cancer, and cholangiocarcinoma, and are in line with a general role of high-mannose glycans and higher-antennary complex-type glycans in cancer progression.
Similar content being viewed by others
Liposarcomas are mesenchymal neoplasms displaying variable levels of adipogenic differentiation. Liposarcoma has a yearly incidence of ~30 cases per million, and represents 17–25% of all sarcoma cases [1, 2]. The World Health Organization (WHO) classification describes four different liposarcoma subtypes: (1) atypical lipomatous tumor/well-differentiated liposarcoma, (2) dedifferentiated, (3) pleomorphic, and (4) myxoid liposarcoma (MLS) [1]. MLS is the second most common subtype of liposarcoma, accounting for ~30% of all liposarcoma cases and 6% of all sarcoma cases [3]. MLS is characterized by uniform round or oval-shaped cells, admixed with a variable number of small lipoblasts, embedded in a prominent myxoid extracellular matrix (ECM) with a characteristic delicate arborizing capillary vasculature. In over 95% of cases the highly specific chromosomal translocation t(12;16)(q13;p11) is found, which results in the chimeric FUS–DDIT3 gene. The remaining cases harbor an EWSR1–DDIT3 fusion. MLS commonly occurs in the extremities of patients that are typically of a younger age compared with other liposarcoma subtypes [1]. A subset of cases has areas with increased cellularity, where tumor cells are round, with a high nuclear to cytoplasmic ratio, and arranged back to back in solid sheets, with the absence of intervening myxoid (MYX) stroma (Fig. 1) [1]. The presence of ≥5% of these so-called “round cell” areas in MLS is associated with a worse prognosis, and is therefore considered histologically high grade (previously designated as round cell (RC) liposarcoma) [1,2,3,4]. The presence of transitional areas, with increased cellularity but still retaining some MYX stroma, can be referred to as intermediate (INT) histological grade, although its prognostic significance is currently unknown (Fig. 1).
MYX areas are characterized by a low cellularity and high abundance of myxoid extracellular matrix (ECM). RC areas contain a high abundance of round cells and lack the ECM component. The cell density in transitional INT regions is too high to be considered MYX, but the ECM component is too abundant to be RC. The scale bars represent 25 µm.
Other factors with potential prognostic value include the overexpression of TP53 [5], the presence of necrosis [5], increased tumor size (>10 cm) [6], increased age (> 45 years), and male gender [7]. The treatment of MLS is multimodal, and the tumors are initially often sensitive to chemo- and radiotherapy; nonetheless 35–50% will develop either recurrence or metastases. High intra-tumor heterogeneity may complicate the determination of histological grade, especially when the tumor mass is poorly represented by a small biopsy [8]. While it is widely accepted that the presence of RCs is associated with a poor prognosis and higher risk of metastasis, the underlying molecular mechanisms of tumor progression are largely unknown.
In a previous immunohistochemistry (IHC)-based study by Demicco et al., the progression of MLS was shown to be accompanied by the activation of the PI3K/Akt pathway caused by the overexpression of insulin-like growth factor 1 receptor (IGF1R), a loss of phosphatase and tensin homolog (PTEN), or activating mutations in phosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform [9]. In addition, several studies have reported the involvement of TP53. One report indicated, based on IHC on a retrospective cohort of human MLS samples, that the overexpression of TP53 (with a missense mutation) was associated with increased chance of metastatic disease [5]. Another report, based on IHC analysis in genetically modified mice, stated that a double knockout of both Pten and TP53 resulted in the formation and progression of several liposarcoma subtypes, including MLS. Single knockout models with a loss of either Pten or TP53 did not result in any tumor formation. The combined loss of Pten and TP53 resulted in the abnormal expression of D-cyclins and CDK4/6, which, in turn, regulate enzymatic activity during the cell cycle, leading to uncontrolled proliferation [10]. A more recent report by Nezu et al. shows that the overexpression of microRNA miR-135b in MLS results in the downregulation of the THBS2 gene, coding for the glycoprotein thrombospondin-2 (TSP2), which, in turn, increases the expression of matrix metalloproteinase 2, responsible for enhanced degradation of the ECM and ultimately results in increased MLS aggressiveness [11].
Conventional omics-analyses (e.g., genomics, proteomics) have had limited success in MLS, possibly due to its high degree of intra-tumor heterogeneity [12, 13]. Nezu et al. used laser-capture microdissection to separate the contributions from the RC and MYX areas in MLS [11]. The success of this microdissection approach demonstrated that new biomolecular insights can be obtained with intra-tumoral-grade-specific analysis.
Matrix-assisted laser desorption/ionization mass spectrometry imaging (MALDI-MSI) is an analytical technique that allows the spatially correlated molecular analysis of various molecular classes directly from tissue [14, 15]. By combining the molecular analysis of modern biomolecular mass spectrometry with the imaging capabilities of microscopy MALDI-MSI enables the simultaneous registration of the spatial distributions of 100–1000 s of biomolecules without labeling and is untargeted in nature. Furthermore, MALDI-MSI does not adversely affect the tissue’s underlying histology; consequently, the tissue may be histologically stained after the MSI experiment and the histological image aligned with the MSI dataset. These multimodal data, MSI and histology, allow the tissue’s molecular content to be examined in its correct histopathological context, and thus, for instance, to compare low- and high-grade areas within a single tissue section. In a process termed virtual microdissection cell-type-specific molecular signatures can be extracted, and then used to identify molecular signatures [16,17,18,19].
MALDI-MSI can be used to analyze different molecular classes, using essentially the same technology but different tissue preparation procedures. On-tissue digestion MALDI-MSI using proteolytic enzymes is used to increase the range of proteins amenable to MALDI-MSI, aid their identification, and allow the analysis of formalin-fixed, paraffin-embedded (FFPE) [20,21,22,23]. On-tissue digestion MALDI-MSI using the endoglycosidase PNGase F enables the analysis of N-linked glycans, from both fresh frozen and FFPE tissue sections [24, 25]. N-glycans are oligosaccharides, and a class of common co-translational protein modifications. Attached during translation in the endoplasmatic reticulum, high-mannose-type N-glycans (consisting of the N-glycan-core, and up to nine additional hexoses) are checkpoints during the protein folding process, and are considered precursors for the rest of the N-glycan maturing process that takes place in the Golgi system. During N-glycan maturation, high-mannose-type glycans are truncated, and redecorated with a mixture of N-acetylhexosamines, hexoses, fucoses, and sialic acids, resulting in either complex-type or hybrid-type N-glycan structures [26]. Proteins, previously associated with MLS tumor progression, such as TSP2, IGF1R, and other receptor tyrosine kinases, involved in the activation of the PIK3/Akt pathway, are known to be glycoproteins [27, 28]. While these proteins have a known role in MLS progression, the role of the N-glycans is unknown. Therefore, we have analyzed the N-glycans by on-tissue digestion MALDI-MSI, to investigate the glycomic alterations underlying tumor progression.
Materials and methods
Chemicals and reagents
All chemicals were purchased from Sigma-Aldrich (Steinheim, Germany) except ethanol and xylene (Merck, Darmstadt, Germany). Recombinant Peptide N-Glycosidase F (PNGase F) from Flavobacterium meningosepticum was expressed and purified as previously described [25].
Tissues, tissue microarray (TMA) construction, and patient grouping
A TMA was previously constructed from FFPE MLS tissues from 32 patients diagnosed with MLS at Leiden University Medical Center [29, 30]. Patients did not receive neoadjuvant therapy. The array consists of 141 tissue cores of 1.5 mm diameter, divided in two paraffin blocks, here referred to as MLS-1 and MLS-2. Both blocks also contain six control cores, originating from human placenta, colon, spleen, neuronal, lung, and tonsil tissues. Cores were obtained from three distinct morphological regions in the tumor, MYX areas, INT areas, and RC areas. If a tissue block presented only a single morphology, three needle cores were included in the TMA; if more than one morphology was present (MYX + INT, MYX + RC, or INT + RC), two cores per morphology were included. An overview of the TMA is reported in Table 1. The use of the human samples and clinical follow-up data was approved by the Leiden University Medical Center ethical committee (B17.012).
N-glycan MALDI-MSI
Sections of 6 µm thickness were cut using a microtome and mounted on indium tin oxide coated glass slides (Bruker Daltonics GmbH, Bremen, Germany), previously coated with poly-L-lysine for improved adhesion. Importantly, TMA sections were not transferred to the glass slide using a tape transfer system due to the risk of polymer contamination. Slides were dried overnight at 37 °C and stored at 4 °C in the dark until further processing. On-tissue digestion was performed after the application of 15 layers (10 µL/min) of PNGase F (100 ng/µL in deionized water) using a SunCollect sprayer (SunChrom GmbH, Friedrichsdorf, Germany). Following overnight incubation at 37 °C, 7 mg/mL α-cyano-4-hydroxycinnamic acid in 50% acetonitrile and 0.1% trifluoroacetic acid was applied to the tissue using the SunCollect (SunChrom GmbH) system. A comprehensive description of the sample preparation has been previously reported [31]. N-glycan MALDI-MSI analysis was performed using a RapifleX MALDI-TOF/TOF instrument (Bruker Daltonics). Data were recorded in reflectron, positive ion mode covering the mass ranges between m/z 900 and 3200, using 500 laser shots per spot and a 50 × 50 µm pixel size.
Histology staining and annotation
Following MALDI-MSI analysis, excess MALDI matrix was removed with two ethanol washes (70%, 2 × 1 min) and the TMAs stained with haematoxylin and eosin (H&E). Slides were then scanned using a digital slide scanner (IntelliSite Pathology Ultra-Fast Scanner, Philips, Eindhoven, The Netherlands). The scanned histology images were co-registered to the MALDI-MSI data in flexImaging 5.0 (Bruker Daltonics). The individual TMA cores were annotated by an expert pathologist (JVMGB). The annotations demarcate the MYX, “intermediate” (INT), and RC areas within each core (Fig. 1). INT areas were defined as transitional areas with modestly increased cellularity without elevated nuclear grade and mitotic activity, thus too cellular to be annotated as MYX and containing too much ECM to be considered an RC area. Following annotation, the average mass spectra were extracted from 138 regions of interest (ROIs), originating from 32 patients. Differentiation between patients with low- and high-grade disease (<5% and ≥5% RC morphology, respectively) was based on the clinical diagnostic evaluation.
Data preprocessing and analysis
The total ion current normalized average spectrum for each of the 138 annotated ROIs was exported in CSV file format using flexImaging 5.0. An overall average spectrum representing the N-glycan content of the full TMA was created in mMass [32]. Spectral processing of the overall average mass spectrum was performed in mMass using the following parameters: Savitsky–Golay smoothing—window size: 0.1 m/z, cycles: 2; baseline subtraction—precision 45, relative offset 15; peak picking—signal-to-noise ratio (S/N) threshold: 3; deisotoping—maximum charge: 1, isotope mass tolerance: 0.15 m/z, isotope intensity tolerance: 50%. A total of 130 peaks (S/N ≥ 3) were detected and which were then assigned to glycan composition using GlycoWorkBench (v1.2.4105) [33], based on the observed mass and previously performed MALDI-MS/MS experiments [31, 34]. Assigned N-glycans were added to the “composition list” required for data extraction by MassyTools (version 0.1.8.1) [35]. In MassyTools, a recalibration of all ROI spectra was performed on the basis of a predefined list of calibrants (Table S1, Supplementary information). Spectra with at least four calibrants (S/N ≥ 9) were recalibrated and considered for further analysis. Feature extraction of the assigned N-glycan compositions [31, 34] was performed through integration of the areas of at least 70% of the theoretical isotopic envelope. A feature-specific baseline correction was performed by subtracting the baseline detected within a window of 20 m/z. Following an assessment of the quality of the extracted mass spectra led to the exclusion of three ROI spectra because the “fraction of the analytes above S/N 9” was below 3 standard deviations of the mean (mean and SD: 89 ± 10%). Analytes were considered for further analysis if they had a mass error below 20 ppm and S/N ≥ 9 in the majority (≥50%) of the ROI spectra. A total of 43 N-glycans met the curation criteria and their relative intensities were rescaled to 100% and submitted to statistical analysis. In addition, derived traits were calculated based on the compositional features of the N-glycan species (Table S2, Supplementary information) [36].
Two approaches were taken to assess molecular differences associated with disease progression: (1) morphology-based analysis: the different annotated tumor areas (MYX, INT, RC) within the different tumors were compared with each other. Thus, of one tumor or patient, different morphological areas (if present) were included in the analysis as it was purely based on morphology; (2) patient-based analysis: the different tumors/patients were compared with each other: of every tumor, the area most representative (i.e., most cellular) of the histological grade as determined on routine diagnostic evaluation of the entire tumor specimen was used for the analysis. Statistical comparisons were performed in MatLab (R2017a, Mathworks, Natick, MA, USA) to find derived traits and individual N-glycans correlated with MLS progression.
The Kruskal–Wallis test, a nonparametric one-way ANOVA-type analysis, was used for feature selection. N-glycans and derived traits with a p value < 0.05 were considered for pair-wise comparisons between the three morphology groups (MYX vs. INT, MYX vs. RC, INT vs. RC) and patient groups (low grade vs. high grade), using a nonparametric alternative to the t-test (the Mann–Whitney U test). Survival analyses (Kaplan–Meier) were performed for the significant hits from both the morphology-based and the patient-based analysis using SPSS (version 25, IBM corporation). The survival curves among low- and high-intensity groups (based on third quartile dichotomization) were compared using the low-rank test. All reported p values were corrected for multiple testing using the Benjamini–Hochberg method.
Results
In this study, MLS morphological disease progression was studied in two different ways: we compared the N-glycan content between (1) annotated areas displaying MYX, INT, and RC morphology, and (2) between tumors diagnosed as low grade vs. high grade based on overall morphology (<5% vs. ≥5% RC component, respectively).
N-glycan changes with increased histological grade
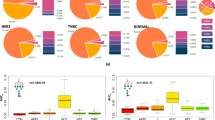
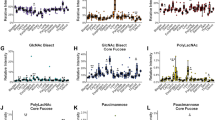
A total of 43 N-glycans were detected and assigned with a glycan composition (= direct traits; Fig. 2 and Table S3, Supplementary information). N-glycan species were grouped on the basis of shared structural features that reflect the enzymatic steps of N-glycan biosynthesis (= derived traits) [36]. These N-glycan traits were compared between the three different morphologies of MLS, as well as between the patients with low- and high-grade disease. The most prominent difference was observed in the abundance of high-mannose type N-glycans between MYX and RC morphologies (FCRC/MYX = 1.53, p = 0.006), as well as between INT and RC morphologies (FCRC/INT = 1.43, p = 0.015) (Fig. 3a). Moreover, a 1.4-fold higher level of high-mannose type N-glycans was observed in patients with high-grade tumors, compared with low-grade disease (FChigh/low = 1.43, p = 0.002) (Fig. S1a, Supplementary information), indicating an overall increase of high-mannose type N-glycans with morphological progression. In addition, a slight, but significant increase in the average number of mannoses attached to the N-glycan core was observed between MYX and RC areas (MYX: average of 6.99 mannoses, RC: average 7.16 mannoses, p = 0.027). Also, between patients with high- and low-grade disease a significant increase in the number of mannose residues on high-mannose species was observed (low grade: 7.01 mannoses, high grade: 7.16 mannoses, p = 0.006) (Fig. S1b, Supplementary information).
Boxplots showing the relative area distributions in MYX, INT, and RC areas for the (a–d) derived N-glycan traits, d–g single high-mannose type N-glycans, and h–k single complex-type N-glycans that were significantly altered in the morphology-based statistical analysis. Asterisks represent various significance levels (*p < 0.05; **p ≤ 0.01; ***p ≤ 0.005).
The direct trait contributing most to the high-mannose type derived trait was H9N2 (where H = hexose, and N = N-acetylhexosamine), found at m/z 1905.63 [M + Na]+, which showed the highest fold change between the MYX and RC areas (FCRC/MYX = 2.26, p = 0.003), and INT and RC areas (FCRC/INT = 1.95, p = 0.015). Moreover, the same N-glycan exhibited an almost twofold increase (FChigh/low = 1.91, p = 0.001) in patients with high-grade MLS, compared with patients with low-grade MLS (Fig. 4). In addition, three other high-mannose type N-glycans (H6N2, m/z 1419.48 [M + Na]+; H7N2, m/z 1581.53 [M + Na]+; H8N2, m/z 1743.58 [M + Na]+) were also significantly associated with changes in morphology and tumor histological grade (Figs. 3e–h and S1e–h, Supplementary information).
Examples of the molecular distribution of high-mannose glycan H9N2 (m/z 1905.63 [M + Na]+) in a myxoid (MYX) morphology of patient L413, b “intermediate” (INT) morphology of patient L3242, and c round cell (RC) morphology of patient L3403. d–f Corresponding H&E images to panels (a)–(c). g Zoom view on m/z 1905.63 in the average spectra of L413-MYX, L3242-INT, and L3403-RC.
The increase in high-mannose type N-glycans with morphological disease progression, together with the equality of hybrid-type N-glycans, led to a concomitant decrease in the relative abundance of complex-type N-glycans as an effect of total area normalization/rescaling to 100% (Figs. 3b and S1c, Supplementary information). Interestingly, within the group of complex-type N-glycans, the composition/types/abundances of N-glycans also changed with progression. A significant increase in tri- and tetra-antennary complex-type N-glycans was observed in the transition from MYX to RC areas (Fig. 3c, d). Accordingly, in the total tumor histological grade-based analysis the tri-antennary complex-type N-glycans were more abundant in the high-grade tumors, compared with the low-grade tumors (FChigh/low = 1.18, p = 0.012) (Fig. S1d, Supplementary information).
Four complex-type N-glycans (H6N5F1, m/z 2174.77 [M + Na]+; H6N5F2, m/z 2320.83 [M + Na]+; H6N5F1S1, m/z 2487.85 [M + Na]+; H7N6F1, m/z 2539.90 [M + Na]+, where H = Hexose, N = N-acetylhexosamine, F = fucose, and S = N-acetylneuraminic acid) were found with significant variation between MLS morphologies (Fig. 3i–l). Three of these species, H6N5F1, H6N5F2, and H6N5F1S1, were fucosylated tri-antennary complex-type glycans, the fourth, H7N6F1, was a mono-fucosylated tetra-antennary complex-type glycan, suggesting this glycan to drive the derived trait.
When comparing patients with high- and low-grade MLS, the molecular picture was very similar; two of the tri-antennary complex-type glycans that correlated with disease morphology, H6N5F2 (FChigh/low = 1.26, p = 0.012) and H6N5F1S1 (FChigh/low = 1.41, p = 0.002), had a similar correlation with clinical grade. In addition, H5N4F1S1 (FChigh/low = 1.23, p = 0.047) and H5N5F1S1 (FChigh/low = 1.20, p = 0.041) correlated positively with progression in grade (Fig. S1i–l, Supplementary information).
N-glycan changes with tumor histological grade within areas with “intermediate” morphology
To find out whether the “intermediate” areas, the transitional morphology between typical MYX and RC MLS, harbor any clinical relevance the N-glycan content of the INT areas was compared between patients with low- and high-grade MLS. It was found that the high-mannose type N-glycan H9N2 (FChigh/low = 1.48, p = 0.046) and complex-type N-glycan H5N4F1S1 (FChigh/low = 1.25, p = 0.033) were associated with patients with high-grade disease. However, these results were not maintained following multiple testing correction, which may be due to the small number of patients analyzed here. In addition, a similar approach was taken comparing MYX areas of patients with low- and high-grade MLS. No significant differences were observed between MYXlow-grade and MYXhigh-grade.
Survival analysis for altered N-glycan species and traits
To assess the prognostic potential of the significantly altered N-glycan species and derived traits, Kaplan–Meier analyses for disease-specific survival were performed. It was found that the high abundance of the CA3 trait (tri-antennary complex-type N-glycans) resulted in a significantly poorer prognosis than compared with the low-abundance group (Fig. 5). The Kaplan–Meier analyses for the high-mannose trait, as well as the individual H9N2 and H8N2 species, resulted in significant log rank test results. Notably, these results were not maintained following Benjamini–Hochberg multiple testing correction, which may be due to the small patient series used here.
Discussion
An increased relative abundance of high-mannose type N-glycans was found to be associated with morphological progression of MLS. Significantly higher levels of high-mannose type species (H6N2, H7N2, H8N2, and H9N2) were found between MYX and RC areas. As expected, the same molecular species differentiated between patients with low- and high-grade tumors. Interestingly, the association between high-mannose type glycans and tumor progression was previously described for breast cancer, ovarian cancer, colorectal cancers, and cholangiocarcinoma [37,38,39,40,41]. It should be noted that for MLS, the term “morphological progression” refers to the increased presence of morphological features associated with poor prognosis (increased cellularity, presence of RC component), rather than a clinical progression of the disease (e.g., the progression of a carcinoma from in situ, to invasive, to metastatic). In normal cells, high-mannose type glycans play a crucial role in both protein folding and the protection of proteins against degradation during intracellular re-localization. However, the exact role of high-mannose glycans in cancer is still unknown.
Several complex-type N-glycans, characterized by increased antennarity, were detected at higher levels in high-grade tumors. Increases in tri-antennary complex-type N-glycans, similar to those observed in the current study, have been previously reported in cholangiocarcinoma [41]. The increase in branching of N-glycans is likely caused by the higher expression of the MGAT4 and MGAT5 genes, which code for N-acetylglucosaminyltransferase IV (addition of β1,4-GlcNAc) and N-acetylglucosaminyltransferase V (GnT-V; addition of β1,6- GlcNAc), which previously have been associated with invasion and metastasis in both breast and colorectal cancer [42,43,44,45]. Overexpression of GnT-V and the resulting increase in N-glycan branching resulted in increased cell motility, a loss of contact inhibition, and caused morphological transformation of normal human epithelial cells [46, 47]. In addition, mice with a MGAT5 deficiency showed a suppression of mammary tumor cell growth and metastasis [46,47,48].
It is known that both PI3K and Ras/Mapk signaling stimulate MGAT5 and MGAT4 gene expression in human carcinomas [45, 46]. In MLS, increased PI3K/Akt signaling, through either a loss of PTEN homolog activity or overexpression of the IGF1R, is involved in tumor progression, which could indicate similar molecular mechanisms being active in the different tumor types [9].
All complex N-glycan structures that correlated with morphological progression carried at least one fucose residue. N-glycans can carry a fucose residue on both the N-glycan core and the antennae. However, the present MALDI-MSI approach could not differentiate between these structural isomers. Aberrant core- and antenna fucosylation have been implicated in tumor progression, metastasis, and resistance to chemotherapy in colorectal cancer, pancreatic cancer, and hepatocellular carcinoma [45, 49, 50]. The role of fucosylation in MLS progression remains to be established, as well as whether fucosylation is another general adaptation that accompanies all tumors.
Although the presence of INT areas in MLS is now well recognized, the prognostic significance of these transitional areas with more-limited hypercellularity is less certain. Here we have studied the potential prognostic value of the INT morphology by comparing the INT areas of patients with low-grade MLS with the INT areas of patients with high-grade MLS. It was found that a selection of direct N-glycan traits differentiated between the INT areas of patients with low- and high-grade MLS. These N-glycans were the same as those significant between patients with low- and high-grade MLS in the patient-based analysis discussed previously. When looking at the morphology-based analysis, the N-glycan species differentiating between MYX and RC areas were always less abundant in the INT areas compared with RC areas, but more abundant compared with MYX areas. This finding indicates that the INT areas are not just reflecting a morphological transition, but also a molecular transition, and that the INT areas likely harbor prognostic potential. Previous work by Haniball et al. has shown that patients with “intermediate grade” MLS (up to 5% RC component) have a higher risk of metastatic disease, compared with patients without RC component [51]. The definition for “intermediate grade” MLS, however, is based on the assessment of the full tumor histological grade, and does not take into account areas with increased cellularity compared with low-grade MLS. This shows that, in order to fully exploit the INT areas for prognostic evaluation, a careful definition of the INT morphology and INT grade in the WHO description of MLS would be required.
While little is still known about glycosylation changes in MLS, the current findings would support a possible role for N-glycans in tumor progression in MLS. The biosynthesis of complex-type N-glycans is dependent on the activity of a large number of different glycosyltransferases, and changes in the expression of these enzymes result in changed glycosylation profiles. The increase in high-mannose type N-glycans and decreased abundance of complex-type glycans indicate that the maturation of glycans is disrupted. Nevertheless, the processing of N-glycans in the endoplasmic reticulum and Golgi system does not appear to be uniformly reduced, as an increase in N-glycan branching was observed in addition to the accumulation of high-mannose type glycans. Expression analysis of the genes coding for the various glycosyltransferases, or a quantitative (micro)proteomics approach, could help identify which enzymes cause the altered glycosylation profiles in the different MLS grades. While the results clearly display a positive correlation between the abundance of several N-glycans and the morphological progression of MLS, a potential complicating factor is that MLS tumor progression is associated with an increase in cellularity [1]. A normalization using the number of cells per pixel would solve this issue. However, the majority of glycoproteins are either secreted or membrane bound. The N-glycans analyzed by MALDI-MSI must first be released from their carrier proteins (achieved here with on-tissue PNGase F digestion) and were analyzed using a 50 × 50 µm2 pixel size. These parameters make it impossible to establish whether a glycan was bound to a membrane protein or a secreted, extracellular protein. Therefore, a cell density normalization would not result in a more accurate representation of the N-glycan distributions in the tissues.
In summary, the results indicate that the INT MLS morphology is not just transitional state between MYX and RC MLS on a morphological level, but also on the level of N-glycan expression. More importantly, we have found an association between the increase of high-mannose-type N-glycans, an increase in their mannose content, a decrease of complex-type glycans, and an increase of their β1,4- and β1,6-linked GlcNAc branching with morphological MLS progression; a molecular pattern that mirrors recent findings in other tumor types, and would indicate a more general role for N-glycans in tumor biology.
References
Fletcher CDM, Bridge JA, Hogendoorn PCW, Mertens F. Pathology and genetics of tumours of soft tissue and bone. In: World Health Organization classification of tumours. 4th ed. Lyon: IARC Press; 2013.
Lemeur M, Mattei J-C, Souteyrand P, Chagnaud C, Curvale G, Rochwerger A. Prognostic factors for the recurrence of myxoid liposarcoma: 20 cases with up to 8 years follow-up. Orthop Traumatol Surg Res. 2015;101:103–7.
Nishida Y, Tsukushi S, Nakashima H, Ishiguro N. Clinicopathologic prognostic factors of pure myxoid liposarcoma of the extremities and trunk wall. Clin Orthop Relat Res. 2010;468:3041–6.
De Graaff MA, Yu JS, Beird HC, Ingram DR, Nguyen T, Juehui Liu J, et al. Establishment and characterization of a new human myxoid liposarcoma cell line (DL-221) with the FUS-DDIT3 translocation. Lab Invest. 2016;96:885–94.
Antonescu CR, Tschernyavsky SJ, Decuseara R, Leung DH, Woodruff JM, Brennan MF, et al. Prognostic impact of P53 status, TLS-CHOP fusion transcript structure, and histological grade in myxoid liposarcoma: a molecular and clinicopathologic study of 82 cases. Clin Cancer Res. 2001;7:3977–87.
Moreau LC, Turcotte R, Ferguson P, Wunder J, Clarkson P, Masri B, et al. Myxoid\round cell liposarcoma (MRCLS) revisited: an analysis of 418 primarily managed cases. Ann Surg Oncol. 2012;19:1081–8.
Hoffman A, Ghadimi MPH, Demicco EG, Creighton CJ, Torres K, Colombo C, et al. Localized and metastatic myxoid/round cell liposarcoma. Cancer. 2013;119:1868–77.
Fritchie KJ, Goldblum JR, Tubbs RR, Sun Y, Carver P, Billings SD, et al. The expanded histologic spectrum of myxoid liposarcoma with an emphasis on newly described patterns: implications for diagnosis on small biopsy specimens. Am J Clin Pathol. 2012;137:229–39.
Demicco E, Torres K, Ghadimi M, Colombo C, Bolshakov S, Hoffman A, et al. Involvement of the PI3K/Akt pathway in myxoid/round cell liposarcoma. Mod Pathol. 2011;25:212–21.
Puzio-Kuter AM, Laddha SV, Castillo-Martin M, Sun Y, Cordon-Cardo C, Chan CS, et al. Involvement of tumor suppressors PTEN and p53 in the formation of multiple subtypes of liposarcoma. Cell Death Differ. 2015;22:1785–91.
Nezu Y, Hagiwara K, Yamamoto Y, Fujiwara T, Matsuo K, Yoshida A, et al. miR-135b, a key regulator of malignancy, is linked to poor prognosis in human myxoid liposarcoma. Oncogene. 2016;35:6177–88.
Suehara Y, Kondo T, Juji K, Hasegawa T, Kawai A, Seki K, et al. Proteomic signatures corresponding to histological classification and grading of soft-tissue sarcomas. Proteomics. 2006;6:4402–9.
Kanojia D, Nagata Y, Garg M, Lee DH, Sato A, Yoshida K, et al. Genomic landscape of liposarcoma. Oncotarget. 2015;6:42429–44.
Caprioli RM, Farmer TB, Gile J. Molecular imaging of biological samples: localization of peptides and proteins using MALDI-TOF MS. Anal Chem. 1997;69:4751–60.
McDonnell LA, Heeren RM. Imaging mass spectrometry. Mass Spectrom Rev. 2007;26:606–43.
Elsner M, Rauser S, Maier S, Schöne C, Balluff B, Meding S, et al. MALDI imaging mass spectrometry reveals COX7A2, TAGLN2 and S100-A10 as novel prognostic markers in Barrett’s adenocarcinoma. J Proteomics. 2012;75:4693–704.
Meding S, Nitsche U, Balluff B, Elsner M, Rauser S, Schöne C, et al. Tumor classification of six common cancer types based on proteomic profiling by MALDI imaging. J Proteome Res. 2012;11:1996–2003.
Balluff B, Frese CK, Maier SK, Schöne C, Kuster B, Schmitt M, et al. De novo discovery of phenotypic intratumour heterogeneity using imaging mass spectrometry. J Pathol. 2015;235:3–13.
Lou S, Balluff B, de Graaff MA, Cleven AH, Briaire-de Bruijn I, Bovée JVMG, et al. High-grade sarcoma diagnosis and prognosis: biomarker discovery by mass spectrometry imaging. Proteomics. 2016;16:1802–13.
Schober Y, Guenther S, Spengler B, Römpp A. High-resolution matrix-assisted laser desorption/ionization imaging of tryptic peptides from tissue. Rapid Commun Mass Spectrom. 2012;26:1141–6.
Gustafsson OJR, Eddes JS, Meding S, McColl SR, Oehler MK, Hoffmann P. Matrix-assisted laser desorption/ionization imaging protocol for in situ characterization of tryptic peptide identity and distribution in formalin-fixed tissue. Rapid Commun Mass Spectrom. 2013;27:655–70.
Heijs B, Carreira RJ, Tolner EA, de Ru AH, Maagdenberg AM, van Veelen PA, et al. Comprehensive analysis of the mouse brain proteome sampled in mass spectrometry imaging. Anal Chem. 2015;87:1867–75.
Kriegsmann M, Casadonte R, Kriegsmann J, Dienemann H, Schirmacher P, Kobarg JH, et al. Reliable entity subtyping in non-small cell lung cancer by matrix-assisted laser desorption/ionization imaging mass spectrometry on formalin-fixed paraffin-embedded tissue specimens. Mol Cell Proteomics. 2016;15:3081–9.
Powers TW, Jones EE, Betesh LR, Romano PR, Gao P, Copland JA, et al. Matrix assisted laser desorption ionization imaging mass spectrometry workflow for spatial profiling analysis of N-linked glycan expression in tissues. Anal Chem. 2013;85:9799–806.
Powers TW, Neely BA, Shao Y, Tang H, Troyer DA, Mehta AS, et al. MALDI imaging mass spectrometry profiling of N-glycans in formalin-fixed paraffin embedded clinical tissue blocks and tissue microarrays. PLoS ONE. 2014;9:e106255.
Freeze HH. Genetic defects in the human glycome. Nat Rev Genet. 2006;7:537–51.
Carlson B, Bernstein D, Annis D, Misenheimer T, Hannah B, Mosher D, et al. Structure of the calcium-rich signature domain of human thrombospondin-2. Nat Struct Mol Biol. 2005;12:910–4.
Negri T, Virdis E, Brich S, Bozzi F, Tamborini E, Tarantino E, et al. Functional mapping of receptor tyrosine kinases in myxoid liposarcoma. Clin Cancer Res. 2010;16:3581–93.
Endo M, Graaff M, Ingram D, Lim S, Lev D, Bruijn I, et al. NY-ESO-1 (CTAG1B) expression in mesenchymal tumors. Mod Pathol. 2015;28:587–95.
de Graaff MA, Malu S, Guardiola I, Kruisselbrink A, de Jong Y, Corver W, et al. High-throughput screening of myxoid liposarcoma cell lines reveals survivin as a potential novel druggable target. Leiden University; Leiden, the Netherlands; 2017.
Heijs B, Holst S, Briaire-de Bruijn IH, Pelt G, de Ru AH, van Veelen PA, et al. Multimodal mass spectrometry imaging of N-glycans and proteins from the same tissue section. Anal Chem. 2016;88:7745–53.
Strohalm M, Hassman M, Kosata B, Kodícek M. mMass data miner: an open source alternative for mass spectrometric data analysis. Rapid Commun Mass Spectrom. 2008;22:905–8.
Ceroni A, Maass K, Geyer H, Geyer R, Dell A, Haslam SM. GlycoWorkbench: a tool for the computer-assisted annotation of mass spectra of glycans. J Proteome Res. 2008;74:1650–9.
Holst S, Heijs B, de Haan N, van Zeijl RJM, Briaire-de Bruijn IH, van Pelt GW, et al. Linkage-specific in situ sialic acid derivatization for N-glycan mass spectrometry imaging of formalin-fixed paraffin-embedded tissues. Anal Chem. 2016;88:5904–13.
Jansen BC, Reiding KR, Bondt A, Ederveen AL, Palmblad M, Falck D, et al. MassyTools: a high-throughput targeted data processing tool for relative quantitation and quality control developed for glycomic and glycoproteomic MALDI-MS. J Proteome Res. 2015;14:5088–98.
Reiding KR, Hipgrave Ederveen AL, Rombouts Y, Wuhrer M. Murine plasma N-glycosylation traits associated with sex and strain. J Proteome Res. 2016;15:3489–99.
de Leoz ML, Young LJ, An HJ, Kronewitter SR, Kim J, Miyamoto S, et al. High-mannose glycans are elevated during breast cancer progression. Mol Cell Proteomics. 2011;10:M110.
Liu X, Nie H, Zhang Y, Yao Y, Maitikabili A, Qu Y, et al. Cell surface-specific N-glycan profiling in breast cancer. PloS ONE. 2013;8:e72704.
Anugraham M, Jacob F, Nixdorf S, Everest-Dass AV, Heinzelmann-Schwarz V, Packer NH. Specific glycosylation of membrane proteins in epithelial ovarian cancer cell lines: glycan structures reflect gene expression and DNA methylation status. Mol Cell Proteomics. 2014;13:2213–32.
Kaprio T, Satomaa T, Heiskanen A, Hokke CH, Deelder AM, Mustonen H, et al. N-glycomic profiling as a tool to separate rectal adenomas from carcinomas. Mol Cell Proteomics. 2015;14:277–88.
Talabnin K, Talabnin C, Ishihara M, Azadi P. Increased expression of the high-mannose M6N2 and NeuAc3H3N3M3N3F tri-antennary N-glycans in cholangiocarcinoma. Oncol Lett. 2018;15:1030–6.
Dennis JW, Laferte S, Waghorne C, Breitman ML, Kerbel RS. Beta 1-6 branching of Asn-linked oligosaccharides is directly associated with metastasis. Science. 1987;236:582–5.
Fernandes B, Sagman U, Auger M, Demetriou M, Dennis JW. β1–6 branched oligosaccharides as a marker of tumor progression in human breast and colon neoplasia. Cancer Res. 1991;51:718–23.
Dennis JW, Granovsky M, Warren CE. Glycoprotein glycosylation and cancer progression. Biochim Biophys Acta. 1999;1473:21–34.
D’Arrigo A, Belluco C, Ambrosi A, Digito M, Esposito G, Bertola A, et al. Metastatic transcriptional pattern revealed by gene expression profiling in primary colorectal carcinoma. Int J Cancer. 2005;115:256–62.
Holst S, Wuhrer M, Rombouts Y. Glycosylation characteristics of colorectal cancer. Adv Cancer Res. 2015;126:203–56.
Demetriou M, Nabi I, Coppolino M, Dedhar S, Dennis J. Reduced contact-inhibition and substratum adhesion in epithelial cells expressing GlcNAc-transferase V. J Cell Biol. 1995;130:383–92.
Granovsky M, Fata J, Pawling J, Muller WJ, Khokha R, Dennis JW. Suppression of tumor growth and metastasis in Mgat5-deficient mice. Nat Med. 2000;6:306–12.
Pendu J, Marionneau S, Cailleau-Thomas A, Rocher J, Moullac-Vaidye B, Clément M. ABH and Lewis histo-blood group antigens in cancer. APMIS. 2001;109:9–26.
Blanas A, Sahasrabudhe NM, Rodríguez E, van Kooyk Y, van Vliet SJ. Fucosylated antigens in cancer: an alliance toward tumor progression, metastasis, and resistance to chemotherapy. Front Oncol. 2018;8:39.
Haniball J, Sumathi VP, Kindblom LG, Abudu A, Carter SR, Tillman RM, et al. Prognostic factors and metastatic patterns in primary myxoid/round-cell liposarcoma. Sarcoma. 2011; 2011:538085.
Acknowledgements
This work was supported by the ZonMW Zenith project “Imaging Mass Spectrometry-Based Molecular Histology: Differentiation and Characterization of Clinically Challenging Soft Tissue Sarcomas” (No. 93512002; BH) and the Liddy Shriver Sarcoma Initiative. The authors would like to thank Drs Sarantos Kostidis, Pierre Åman, Alexander Lazar, Neeta Somaiah, Hannah Beird, Chia Chin Wu, Torsten Nielsen, and Jamie Yu for their valuable input and helpful discussions and Prof. Dr Richard Drake and Dr Anand Mehta for kindly producing and providing the PNGase F required for this study.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Heijs, B., Holst-Bernal, S., de Graaff, M.A. et al. Molecular signatures of tumor progression in myxoid liposarcoma identified by N-glycan mass spectrometry imaging. Lab Invest 100, 1252–1261 (2020). https://doi.org/10.1038/s41374-020-0435-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41374-020-0435-2
This article is cited by
-
An N-glycome tissue atlas of 15 human normal and cancer tissue types determined by MALDI-imaging mass spectrometry
Scientific Reports (2024)
-
Matrix-assisted laser desorption/ionization mass spectrometry imaging (MALDI-MSI) reveals potential lipid markers between infrapatellar fat pad biopsies of osteoarthritis and cartilage defect patients
Analytical and Bioanalytical Chemistry (2023)
-
Proteomic and Metabolomic Profiling in Soft Tissue Sarcomas
Current Treatment Options in Oncology (2022)