Abstract
Single-nucleotide polymorphisms (SNPs) in the promoter region of long intergenic non-coding RNAs (lincRNAs) could play a regulatory role in its expression level and then get involved in colorectal cancer (CRC). Thus, we conducted a two-stage case–control study to investigate the associations of Tag SNPs within the promoter region of selected lincRNAs from microarray data with risk of CRC. A total of 320 cases and 319 controls were recruited in the test set to explore the associations between 16 SNPs with no deviations from Hardy–Weinberg equilibrium (HWE) and risk of CRC. Furthermore, 501 cases and 538 controls were included as the validation set to confirm the significant associations. RP11-3N2.1 rs13230517 polymorphism was found to be negatively associated with CRC in both test set (AA vs. GG, OR = 0.68, 95% CI = 0.48–0.96) and validation set (AA vs. GG, OR = 0.76, 95% CI = 0.59–0.98). Pooled analysis showed that individuals with GA/AA genotypes had a significantly decreased risk of CRC when compared with those carrying GG genotype (OR = 0.74, 95% CI = 0.60–0.90) in the combined set. The crossover analysis revealed that rs13230517 GA/AA carriers had a decreased risk of CRC than GG carriers among non-drinkers in both test and combined set. However, no gene-environment multiplicative interactions were found on risk of CRC. Our findings suggest that rs13230517 polymorphism might participate in the pathogenesis of CRC and have the potential to be a biomarker for predicting the risk of CRC.
Similar content being viewed by others
Introduction
Long non-coding RNAs (lncRNAs) represent a subgroup of non-coding RNAs with more than 200 nucleotides in length [1]. LncRNAs were previously believed to be transcriptional noise, but accumulating evidence has suggested that they play a regulatory role in a wide range of biological processes [2]. Therefore, the dysfunctions of lncRNAs are associated with various human diseases, including cancers [3,4,5].
Long intergenic non-coding RNAs (lincRNAs) that located in the interval between protein-coding genes are emerging as key regulators of diverse cellular processes [6]. Increasing studies have demonstrated that lincRNAs are regulatory elements in both oncogenic and tumor suppressive pathways [7,8,9,10]. The expression of lincRNAs can be affected by inherent genetic factors such as single-nucleotide polymorphisms (SNPs) in the promoter region, which are likely to disrupt transcription factor binding sites (TFBS). For example, a study reported that a functional SNP, rs944289, in the promoter region of lincRNA named papillary thyroid carcinoma susceptibility candidate 3 (PTCSC3) predisposeed to papillary thyroid carcinoma (PTC) through dysregulating the expression of PTCSC3 by suppressing the binding activity of both C/EBPα and C/EBPβ [11].
To date, genome-wide association studies (GWAS) have successfully identified a large number of SNPs that are associated with CRC [12,13,14,15], whereas a number of SNPs located in non-coding regions still have not been well explained. Thus, further characterization of lncRNA related SNPs may open a new avenue for functional analysis of the susceptibility of CRC. Moreover, the database developed by National Human Genome Research Institute (NHGRI) in USA (http://www.ebi.ac.uk/gwas/) indicated that disease-associated SNPs are largely located in the intergenic regions. We hypothesize that SNPs located in the promoter region of potentially related lincRNAs may contribute to the occurrence of CRC. To test this hypothesis, we focused on 10 lincRNAs (CTD-2147F2.1, RP11-384P7.7, RP11-3N2.1, RP3-525N10.2, RP11-89K21.1, CCAT1, AK055145, RP11-58A12.3, UCA1 and AY343891) with top fold-change value based on lncRNA microarray data from our previous study [16], to assess whether promoter SNPs in the lincRNAs were associated with risk of CRC.
In this study, we conducted a two-stage case–control study to identify possible biomarkers for predicting the risk of CRC. Furthermore, we evaluated gene by environment (G × E) interactions between the significant SNPs and smoking (or drinking) in relation to risk of CRC.
Materials and methods
Subjects
The source population was previously enrolled from Jiashan County, which had been approved by the Medical Ethical Committee of Zhejiang University School of Medicine. The recruitment details have been described previously [17]. Briefly, CRC cases were identified from local Cancer Surveillance and Registry System, mentally competent to complete an interview and with no previous history of familial adenomatous polyposis, ulcerative colitis, or Crohn’s disease. Healthy controls were recruited in parallel from the same population and were matched to cases by age (±5 years), gender and residential area. All participants were unrelated ethnic Han Chinese. Ultimately, a total of 821 cases and 857 controls were included in the two-stage case–control study (test set: 320 cases and 319 controls; validation set: 501 cases and 538 controls).
Data collection
A face-to-face interview was conducted using a structured questionnaire including demographic characteristics (e.g., age, gender, and body mass index), family history of cancer and life style (e.g., smoking and drinking). The definition of smokers was those who smoked at least 1 cigarette per day for more than 1 year, or more than 300 cigarettes in less than 3 months. An alcohol drinker was defined as someone who consumed at least one drink per day for more than 3 months. Subsequently, ~5 ml venous blood was collected from each participant and stored at −80 °C for the preparation of DNA isolation. This study was approved by the Medical Ethical Committee of Zhejiang University School of Medicine.
SNP selection and genotyping
SNPs located in the promoter region (2000 bp upstream region of the transcription start site) of each lincRNA and their minor allele frequency (MAF) value within the Chinese Han in Beijing (CHB) populations were extracted by utilizing the 1000 Genomes data (http://www.1000genomes.org/). Tag SNPs representing SNPs with pairwise correlation r 2 ≥ 0.8 and MAF ≥ 0.1 were selected.
Genomic DNA was isolated from peripheral blood samples using the modified salting-out procedure [18]. Genotyping for all SNPs was performed by the MassARRAY molecular weight array analysis system (BioMiao Biological Technology Co., Beijing, China).
Statistical analysis
Student’s t-test and Pearson χ 2-test were applied to evaluate the differences of continuous and categorical variables between cases and controls, respectively. Hardy–Weinberg equilibrium (HWE) for all SNPs was evaluated by goodness-of-fit χ 2-test. The associations between SNPs and CRC were estimated by calculating the adjusted odds ratios (ORs) and corresponding 95% confidence intervals (95% CIs) in multivariate logistic regression model. The Benjamini–Hochberg multiple testing correction method was applied, while after the P value correction, no or only few significant SNPs remained. Therefore, the uncorrected P values were used for the selection of significant SNPs between cases and controls. Pooled ORs in the combined set were estimated by the method of random-effect model when the value of I 2 > 50%, otherwise a fixed-effect model was used. Crossover analysis was used to explore the effect of gene factor only, environment factor only, and both gene and environment factors on CRC. Multiplicative model was conducted to evaluate the influence of gene–environment interactions on risk of CRC. Statistical analyses were performed using the SAS software, version 9.2 (SAS Inc., Cary, NC, USA) and Stata software, version 11.2 (StataCorp LP, TX, USA). P values < 0.05 were considered significant.
Results
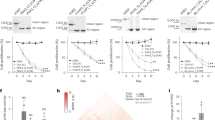
Of the 30,586 lncRNAs in the previous genome-wide expression profile, which was performed by using Arraystar Human LncRNA Microarray v3.0 in six paired CRC and adjacent normal tissues, 556 lncRNAs were found to be significantly upregulated, and 1040 lncRNAs were found to be downregulated in tumor tissues. There were 795 lincRNAs in the abnormally expressed lncRNAs, and the lincRNAs with top 10 fold-change value and corresponding –log10 P value were presented in Fig. 1.
There were no significant differences in demographic characteristics and life-style factors between cases and controls in both test and validation set. However, CRC patients were more likely to have family history of cancer than controls (P < 0.001 in the test set; P = 0.039 in the validation set) (Table 1).
Nineteen SNPs in the ten lincRNAs were selected and genotyped. The call rate for each SNP was >98%. The chromosome region of lincRNA, ID (rs number) of candidate SNPs and allele (major/minor) frequency were listed in Table 2. Moreover, RP11-384P7.7 rs146223339, RP3-525N10.2 rs2256337, and RP11-58A12.3 rs147827294 that deviated from HWE significantly among controls in the test set were excluded in further analysis.
As shown in Table 3, RP11-3N2.1 rs13230517 and RP11-89K21.1 rs741813 polymorphisms were observed to be significantly associated with risk of CRC adjusted for age, gender, BMI, family history of cancer, smoking, and drinking. By additive model, RP11-89K21.1 rs741813 had an adjusted OR of 1.90 (95% CI: 1.19–3.04). By dominant model, we found that individuals with RP11-3N2.1 rs13230517 GA/AA genotypes had a reduced risk of CRC (OR = 0.70, 95% CI = 0.51–0.98) and RP11-89K21.1 rs741813 AT/TT carriers had an increased risk of CRC (OR = 1.86, 95% CI = 1.14–3.02). In the validation set, RP11-3N2.1 rs13230517 polymorphism kept to have a significantly decreased risk of CRC by dominant model (OR = 0.76, 95% CI = 0.59–0.98). However, no statistical evidence of associations between RP11-89K21.1 rs741813 and CRC were observed (Table 4). Pooled analysis revealed that individuals RP11-3N2.1 rs13230517 variants were less susceptible to CRC by additive model (OR = 0.85, 95% CI = 0.74–0.97) and dominant model (OR = 0.74, 95% CI = 0.60–0.90) in the combined set (Table 4).
Moreover, crossover analysis and gene–environment interactions were conducted for RP11-3N2.1 rs13230517. As shown in Table 5, no single effect of smoking and its combined effect with RP11-3N2.1 rs13230517 polymorphism on risk of CRC were observed. In addition, non-drinkers with GA/AA genotypes had a decreased risk of CRC when compared to those with GG genotype (OR = 0.68, 95% CI = 0.47–0.99) in the test set, and similar result was continued in the combined set (OR = 0.74, 95% CI = 0.59–0.93). However, no evidence of multiplicative interactions between smoking or drinking and RP11-3N2.1 rs13230517 polymorphism on CRC was observed.
Discussion
CRC is the third most common cancer and the fourth leading cause of cancer death worldwide [19]. The prevalence of CRC is increasing in China, but the cause of its epidemic is complicated and not entirely clear, which poses a challenge to its prevention and control. In the current study, we found that a novel SNP rs13230517, which is in the promoter region of lincRNA RP11-3N2.1 is associated with CRC.
Recently, lncRNAs were rapidly recognized as new players in tumorigenesis and tumor suppressor. They could regulate target genes by lncRNA–miRNA and lncRNA–protein interactions, or acting as miRNA precursors [20]. As a class of lncRNAs, accumulating lincRNAs were reported to participate in the complicated process of colorectal carcinogenesis. Zhai et al. [21] suggested that the expression level of lincRNA-p21 was significantly lower in CRC tumor tissues than paired normal tissues, and lincRNA-p21 level was linked to CRC stage, vascular invasion and tumor invasion. Among the selected 10 lincRNAs from microarray data, CCAT1 was proved to have promising diagnostic value in detecting CRC, because its expression was remarkably elevated in colorectal tumor tissues and adenomatous polyp tissues [22,23,24]. Besides, Han et al. [25] reported that UCA1 could affect the proliferation, apoptosis and cell cycle progression of CRC cells.
RP11-3N2.1 was found to be downregulated in CRC tumor tissues relative to paired normal tissues and its fold-change value was ranked third based on the microarray data. It is a noncoding transcript of the encoding gene zinc-finger protein 727 (ZNF727). ZNF family has been recognized as potential B-cell epitopes eliciting the production of autoantibodies in cancer [26]. The ZNF panel (ZNF346, ZNF638, ZNF700, and ZNF768) was reported to be overexpressed at the mRNA level in at least 20% of investigated tumors when compared to adjacent normal colorectal mucosa [27]. In addition, CRC patients with higher expression of ZNF545 were found to have favorable relapse-free survival than those with lower ZNF545 levels [28]. However, there are no studies on the functional similarity or association between RP11-3N2.1 and ZNF family to date. In a genome-wide analysis of lncRNA expression from the publicly available cancer microarray database Gene Expression Omnibus (GEO) under the accession number GSE95423, RP11-3N2.1 (probe name: ASHGA5P016427 in Arraystar Human LncRNA Microarray V3.0) was found to be downregulated (fold-change = 0.487, P = 0.00706) in CRC tissues with liver metastasis compared with those without metastasis, suggesting that RP11-3N2.1 might be implicated in promoting metastasis of CRC. RP11-3N2.1 is an uncharacterized RNA yet to be discovered in previous studies and limited information has been documented. RP11-3N2.1 is a type of lincRNA with a length of 748 bp located in chr7q11.21. As previous reports described that lncRNA could decrease the expression of miRNA target genes by inactivating miRNAs [29]. It was computationally predicted by DIANA-LncBase (www.microrna.gr/LncBase) that miRNA-135b may target on RP11-3N2.1. He et al. [30] indicated that miRNA-135b contributed to anti-apoptosis and chemoresistance in CRC. Besides, several lines of evidence proved that miRNA-135b was related with prognosis of CRC [31,32,33]. On the other hand, lncRNAs can also act as co-activators of transcription factors by interacting with RNA binding proteins (RBPs) and the interaction finally alters the localization and activity of the proteins [34]. The prediction tool starBase v2.0 (http://starbase.sysu.edu.cn/) presented that the RBPs might be FUS or SFRS1. However, the functional mechanisms of RP11-3N2.1 need to be further explored and confirmed at molecular level based on the above findings.
Currently, supporting data indicated that a number of disease-associated SNPs reside in lincRNAs [35]. As the binding of a transcription factor (TF) to promoter region can result in local structural modification, making the removal of a preexisting component or recruitment of a new component [36], SNPs located in the promoter region are regarded as functional SNPs to influence gene expression and subsequently disease involvement. In our study, we detected that a promising SNP RP11-3N2.1 rs13230517 was implicated in CRC. This SNP is about 1842 bp upstream of RP11-3N2.1 and was selected as one of the Tag SNPs by candidate SNP strategy. It is plausible that RP11-3N2.1 rs13230517 mutant may affect the process of transcription by binding with a certain TF, and the abnormal transcription inhibited the expression of RP11-3N2.1. According to HaploReg v4.1 (http://compbio.mit.edu/HaploReg), rs13230517 polymorphism has been predicted to alter seven regulatory motifs, including TFII-I, T3R, MAZR, and so on. Furthermore, the SNP is detected within a region with promoter histone markers (H3K4m3) in colonic and rectal mucosa. However, the lack of experimental data requires further investigation into the functional relevance of rs13230517 to be performed, to conclude the nature of the conferred phenotypic effects.
As for the environmental factors, we focused on the factors of CRC related life styles (smoking and drinking), which have been identified and established in epidemiological studies [37, 38]. Crossover analysis suggested that RP11-3N2.1 rs13230517 polymorphism was negatively associated with CRC among non-drinkers, which proved the protective role of rs13230517 polymorphism in CRC as well. Even though other associations did not reach statistical significance, their directions (positive/negative) were in accordance with previous studies, such as the deleterious role of smoking and drinking on CRC. Nevertheless, we found no multiplicative gene–environment interactions in related to CRC. Accordingly, the sample size still needs to be enlarged for exploring the interactions.
Although the findings from our genetic association analysis suggest a link between RP11-3N2.1 rs13230517 polymorphism and CRC, several limitations of our study should be noted. Firstly, the participants in the case–control study were ethnic Han Chinese living in a small rural county, confining its representativeness for Chinese population. Besides, because of the relatively small sample size of our study, we might not be able to detect weak genetic-disease associations and gene–environment interactions. However, it is less of concern as we combined the results from the test set and validation set to cover the shortage. Thirdly, due to the lack of staging information of CRC patients in the current study, we cannot evaluate the association between RP11-3N2.1 rs13230517 polymorphism and clinical features among CRC patients. Finally, no functional data were achieved for the novel SNP, so the mechanism at the molecular level remains largely unknown and needs to be elucidated.
In conclusion, the present study first provides the evidence of the association between RP11-3N2.1 rs13230517 polymorphism and colorectal carcinogenesis in a Han Chinese population. The findings might provide alternative biomarkers for the risk prediction, prevention and early diagnosis of CRC. Additional epidemiological studies with large sample size and further mechanistic investigations into the function of the biomarker are warranted to improve our understanding of the role in colorectal tumorigenesis more comprehensively.
References
Mercer TR, Dinger ME, Mattick JS. Long non-coding RNAs: insights into functions. Nat Rev Genet. 2009;10:155–9.
Chen G, Wang ZY, Wang DQ, Qiu CX, Liu MX, Chen X, et al. LncRNADisease: a database for long-non-coding RNA-associated diseases. Nucleic Acids Res. 2013;41:D983–D986.
Yang F, Bi JW, Xue XC, Zheng LM, Zhi KK, Hua JD, et al. Up-regulated long non-coding RNA H19 contributes to proliferation of gastric cancer cells. FEBS J. 2012;279:3159–65.
Gutschner T, Hammerle M, Eissmann M, Hsu J, Kim Y, Hung G, et al. The noncoding RNA MALAT1 is a critical regulator of the metastasis phenotype of lung cancer cells. Cancer Res. 2013;73:1180–9.
Nie Y, Liu X, Qu SH, Song E, Zou H, Gong C. Long non-coding RNA HOTAIR is an independent prognostic marker for nasopharyngeal carcinoma progression and survival. Cancer Sci. 2013;104:458–64.
St-Laurent G, Wahlestedt C, Kapranov P. The landscape of long noncoding RNA classification. Trends Genet. 2015;31:239–51.
Tsai MC, Spitale RC, Chang HY. Long intergenic noncoding RNAs: new links in cancer progression. Cancer Res. 2011;71:3–7.
Gibb EA, Brown CJ, Lam WL. The functional role of long non-coding RNA in human carcinomas. Mol Cancer. 2011;10:38.
Huarte M, Guttman M, Feldser D, Garber M, Koziol MJ, Kenzelmann-Broz D, et al. A large intergenic noncoding RNA induced by p53 mediates global gene repression in the p53 response. Cell. 2010;142:409–19.
Szafron LM, Balcerak A, Grzybowska EA, Pienkowska-Grela B, Podgorska A, Zub R, et al. The putative oncogene, CRNDE, is a negative prognostic factor in ovarian cancer patients. Oncotarget. 2015;6:43897–910.
Jendrzejewski J, He HL, Radomska HS, Li W, Tomsic J, Liyanarachchi S, et al The polymorphism rs944289 predisposes to papillary thyroid carcinoma through a large intergenic noncoding RNA gene of tumor suppressor type. Proc Natl Acad Sci USA. 2012;109:8646–51.
Zanke BW, Greenwood CMT, Rangrej J, Kustra R, Tenesa A, Farrington SM, et al. Genome-wide association scan identifies a colorectal cancer susceptibility locus on chromosome 8q24. Nat Genet. 2007;39:989–94.
Nan HM, Hutter CM, Lin Y, Jacobs EJ, Ulrich CM, White E, et al. Association of aspirin and NSAID use with risk of colorectal cancer according to genetic variants. JAMA. 2015;313:1133–42.
Houlston RS, Cheadle J, Dobbins SE, Tenesa A, Jones AM, Howarth K, et al. Meta-analysis of three genome-wide association studies identifies susceptibility loci for colorectal cancer at 1q41, 3q26.2, 12q13.13 and 20q13.33. Nat Genet. 2010;42:973–89.
Cui R, Okada Y, Jang SG, Ku JL, Park JG, Kamatani Y, et al Common variant in 6q26-q27 is associated with distal colon cancer in an Asian population. Gut. 2011;60:799–805.
Jing FY, Jin HC, Mao YY, Li YJ, Ding Y, Fan CH, et al. Genome-wide analysis of long non-coding RNA expression and function in colorectal cancer. Tumor Biol. 2017;39:1010428317703650.
Zhang YJ, Liu B, Jin MJ, Ni Q, Liang X, Ma XY, et al. Genetic polymorphisms of transforming growth factor-beta 1 and its receptors and colorectal cancer susceptibility: a population-based case-control study in China. Cancer Lett. 2009;275:102–8.
Nasiri H, Forouzandeh M, Rasaee MJ, Rahbarizadeh F. Modified salting-out method: High-yield, high-quality genomic DNA extraction from whole blood using laundry detergent. J Clin Lab Anal. 2005;19:229–32.
Ferlay J, Soerjomataram I, Dikshit R, Eser S, Mathers C, Rebelo M, et al. Cancer incidence and mortality worldwide: sources, methods and major patterns in GLOBOCAN 2012. Int J Cancer. 2015;136:E359–E386.
Guttman M, Rinn JL. Modular regulatory principles of large non-coding RNAs. Nature. 2012;482:339–46.
Zhai HY, Fesler A, Schee K, Fodstad O, Flatmark K, Ju JF. Clinical significance of long intergenic noncoding RNA-p21 in colorectal cancer. Clin Colorectal Cancer. 2013;12:261–6.
Nissan A, Stojadinovic A, Mitrani-Rosenbaum S, Halle D, Grinbaum R, Roistacher M, et al. Colon cancer associated transcript-1: a novel RNA expressed in malignant and pre-malignant human tissues. Int J Cancer. 2012;130:1598–606.
Alaiyan B, Ilyayev N, Stojadinovic A, Izadjoo M, Roistacher M, Pavlov V, et al. Differential expression of colon cancer associated transcript1 (CCAT1) along the colonic adenoma-carcinoma sequence. BMC Cancer. 2013;13:196.
Ye ZY, Zhou M, Tian B, Wu B, Li JC. Expression of IncRNA-CCAT1, E-cadherin and N-cadherin in colorectal cancer and its clinical significance. Int J Clin Exp Med. 2015;8:3707–15.
Han Y, Yang YN, Yuan HH, Zhang TT, Sui H, Wei XL, et al. UCA1, a long non-coding RNA up-regulated in colorectal cancer influences cell proliferation, apoptosis and cell cycle distribution. Pathology. 2014;46:396–401.
Backes C, Ludwig N, Leidinger P, Harz C, Hoffmann J, Keller A, et al. Immunogenicity of autoantigens. BMC Genomics. 2011;12:340.
Kijanka G, Hector S, Kay EW, Murray F, Cummins R, Murphy D, et al. Human IgG antibody profiles differentiate between symptomatic patients with and without colorectal cancer. Gut. 2010;59:69–78.
Xiang SL, Xiang TX, Xiao Q, Li YH, Shao BF, Luo T. Zinc-finger protein 545 is inactivated due to promoter methylation and functions as a tumor suppressor through the Wnt/β-catenin, PI3K/AKT and MAPK/ERK signaling pathways in colorectal cancer. Int J Oncol. 2017;51:801–11.
Jalali S, Bhartiya D, Lalwani MK, Sivasubbu S, Scaria V. Systematic transcriptome wide analysis of lncRNA-miRNA interactions. PLoS ONE. 2013;8:e53823.
He YQ, Wang JX, Wang J, Yung VY, Hsu E, Li A, et al. MicroRNA-135b regulates apoptosis and chemoresistance in colorectal cancer by targeting large tumor suppressor kinase 2. Am J Cancer Res. 2015;5:1382–95.
Valeri N, Braconi C, Gasparini P, Murgia C, Lampis A, Paulus-Hock V, et al. MicroRNA-135b promotes cancer progression by acting as a downstream effector of oncogenic pathways in colon cancer. Cancer Cell. 2014;25:469–83.
Wu WY, Wang Z, Yang PC, Yang JF, Liang J, Chen YH, et al. MicroRNA-135b regulates metastasis suppressor 1 expression and promotes migration and invasion in colorectal cancer. Mol Cell Biochem. 2014;388:249–59.
Li JL, Liang HW, Bai M, Ning T, Wang C, Fan Q, et al. miR-135b promotes cancer progression by targeting transforming growth factor beta receptor II (TGFBR2) in colorectal cancer. PLoS ONE. 2015;10:e0145589.
Wang X, Arai S, Song X, Reichart D, Du K, Pascual G, et al. Induced ncRNAs allosterically modify RNA-binding proteins in cis to inhibit transcription. Nature. 2008;454:126–30.
Ning SW, Zhao ZXL, Ye JR, Wang P, Zhi H, Li RH, et al. LincSNP: a database of linking disease-associated SNPs to human large intergenic non-coding RNAs. BMC Bioinformatics. 2014;15:152.
Ko E, Seo HW, Jung ES, Kim BH, Jung G. The TERT promoter SNP rs2853669 decreases E2F1 transcription factor binding and increases mortality and recurrence risks in liver cancer. Oncotarget. 2016;7:684–99.
Liang PS, Chen TY, Giovannucci E. Cigarette smoking and colorectal cancer incidence and mortality: systematic review and meta-analysis. Int J Cancer. 2009;124:2406–15.
Fedirko V, Tramacere I, Bagnardi V, Rota M, Scotti L, Islami F, et al. Alcohol drinking and colorectal cancer risk: an overall and dose-response meta-analysis of published studies. Ann Oncol. 2011;22:1958–72.
Acknowledgements
This work was supported by the National Natural Science Foundation of China (no. 81373087, 81602917), the National Key Basic Research Program (973 program no. 2015CB554003) and the Natural Science Foundation of Zhejiang Province, China (no. LQ15H260001).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Ye, D., Hu, Y., Jing, F. et al. A novel SNP in promoter region of RP11-3N2.1 is associated with reduced risk of colorectal cancer. J Hum Genet 63, 47–54 (2018). https://doi.org/10.1038/s10038-017-0361-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s10038-017-0361-3