Abstract
Group II introns are self-splicing ribozymes that are essential in many organisms, and they have been hypothesized to share a common evolutionary ancestor with the spliceosome. Although structural similarity of RNA components supports this connection, it is of interest to determine whether associated protein factors also share an evolutionary heritage. Here we present the crystal structures of reverse transcriptase (RT) domains from two group II intron–encoded proteins (maturases) from Roseburia intestinalis and Eubacterium rectale, obtained at 1.2-Å and 2.1-Å resolution, respectively. These domains are more similar in architecture to the spliceosomal Prp8 RT-like domain than to any other RTs, and they share substantial similarity with flaviviral RNA polymerases. The RT domain itself is sufficient for binding intron RNA with high affinity and specificity, and it is contained within an active RT enzyme. These studies provide a foundation for understanding structure–function relationships within group II intron–maturase complexes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Pyle, A.M. The tertiary structure of group II introns: implications for biological function and evolution. Crit. Rev. Biochem. Mol. Biol. 45, 215–232 (2010).
Lambowitz, A.M. & Belfort, M. Mobile bacterial group II introns at the crux of eukaryotic evolution. Microbiol. Spectr. 3, MDNA3-0050-2014 (2015).
Lambowitz, A.M. & Zimmerly, S. Group II introns: mobile ribozymes that invade DNA. Cold Spring Harb. Perspect. Biol. 3, a003616 (2011).
Lambowitz, A.M. & Zimmerly, S. Mobile group II introns. Annu. Rev. Genet. 38, 1–35 (2004).
Michel, F. & Ferat, J.L. Structure and activities of group II introns. Annu. Rev. Biochem. 64, 435–461 (1995).
Pyle, A.M. & Lambowitz, A.M. Group II Introns: ribozymes that splice RNA and invade DNA. in The RNA World 3rd edn. (ed. Gesteland, R.F.) 469–505 (Cold Spring Harbor Laboratory Press, 2006).
Aizawa, Y., Xiang, Q., Lambowitz, A.M. & Pyle, A.M. The pathway for DNA recognition and RNA integration by a group II intron retrotransposon. Mol. Cell 11, 795–805 (2003).
Cousineau, B., Lawrence, S., Smith, D. & Belfort, M. Retrotransposition of a bacterial group II intron. Nature 404, 1018–1021 (2000).
Mohr, G., Perlman, P.S. & Lambowitz, A.M. Evolutionary relationships among group II intron-encoded proteins and identification of a conserved domain that may be related to maturase function. Nucleic Acids Res. 21, 4991–4997 (1993).
Blocker, F.J. et al. Domain structure and three-dimensional model of a group II intron-encoded reverse transcriptase. RNA 11, 14–28 (2005).
Zimmerly, S., Hausner, G. & Wu, Xc. Phylogenetic relationships among group II intron ORFs. Nucleic Acids Res. 29, 1238–1250 (2001).
Candales, M.A. et al. Database for bacterial group II introns. Nucleic Acids Res. 40, D187–D190 (2012).
Matsuura, M. et al. A bacterial group II intron encoding reverse transcriptase, maturase, and DNA endonuclease activities: biochemical demonstration of maturase activity and insertion of new genetic information within the intron. Genes Dev. 11, 2910–2924 (1997).
Wank, H., SanFilippo, J., Singh, R.N., Matsuura, M. & Lambowitz, A.M. A reverse transcriptase/maturase promotes splicing by binding at its own coding segment in a group II intron RNA. Mol. Cell 4, 239–250 (1999).
Rambo, R.P. & Doudna, J.A. Assembly of an active group II intron-maturase complex by protein dimerization. Biochemistry 43, 6486–6497 (2004).
Saldanha, R. et al. RNA and protein catalysis in group II intron splicing and mobility reactions using purified components. Biochemistry 38, 9069–9083 (1999).
Gu, S.Q. et al. Genetic identification of potential RNA-binding regions in a group II intron-encoded reverse transcriptase. RNA 16, 732–747 (2010).
Watanabe, K. & Lambowitz, A.M. High-affinity binding site for a group II intron-encoded reverse transcriptase/maturase within a stem-loop structure in the intron RNA. RNA 10, 1433–1443 (2004).
Matsuura, M., Noah, J.W. & Lambowitz, A.M. Mechanism of maturase-promoted group II intron splicing. EMBO J. 20, 7259–7270 (2001).
Cech, T.R. The generality of self-splicing RNA: relationship to nuclear mRNA splicing. Cell 44, 207–210 (1986).
Zimmerly, S., Guo, H., Perlman, P.S. & Lambowitz, A.M. Group II intron mobility occurs by target DNA-primed reverse transcription. Cell 82, 545–554 (1995).
Sharp, P.A. On the origin of RNA splicing and introns. Cell 42, 397–400 (1985).
Xiong, Y. & Eickbush, T.H. Origin and evolution of retroelements based upon their reverse transcriptase sequences. EMBO J. 9, 3353–3362 (1990).
Zimmerly, S. & Semper, C. Evolution of group II introns. Mob. DNA 6, 7 (2015).
Madhani, H.D. & Guthrie, C. A novel base-pairing interaction between U2 and U6 snRNAs suggests a mechanism for the catalytic activation of the spliceosome. Cell 71, 803–817 (1992).
Fica, S.M., Mefford, M.A., Piccirilli, J.A. & Staley, J.P. Evidence for a group II intron-like catalytic triplex in the spliceosome. Nat. Struct. Mol. Biol. 21, 464–471 (2014).
Robart, A.R., Chan, R.T., Peters, J.K., Rajashankar, K.R. & Toor, N. Crystal structure of a eukaryotic group II intron lariat. Nature 514, 193–197 (2014).
Toor, N., Keating, K.S., Taylor, S.D. & Pyle, A.M. Crystal structure of a self-spliced group II intron. Science 320, 77–82 (2008).
Marcia, M. & Pyle, A.M. Visualizing group II intron catalysis through the stages of splicing. Cell 151, 497–507 (2012).
Shukla, G.C. & Padgett, R.A. A catalytically active group II intron domain 5 can function in the U12-dependent spliceosome. Mol. Cell 9, 1145–1150 (2002).
Fica, S.M. et al. RNA catalyses nuclear pre-mRNA splicing. Nature 503, 229–234 (2013).
Galej, W.P., Oubridge, C., Newman, A.J. & Nagai, K. Crystal structure of Prp8 reveals active site cavity of the spliceosome. Nature 493, 638–643 (2013).
Dlakić, M. & Mushegian, A. Prp8, the pivotal protein of the spliceosomal catalytic center, evolved from a retroelement-encoded reverse transcriptase. RNA 17, 799–808 (2011).
Nguyen, T.H. et al. The architecture of the spliceosomal U4/U6.U5 tri-snRNP. Nature 523, 47–52 (2015).
Yan, C. et al. Structure of a yeast spliceosome at 3.6-angstrom resolution. Science 349, 1182–1191 (2015).
Belancio, V.P., Hedges, D.J. & Deininger, P. Mammalian non-LTR retrotransposons: for better or worse, in sickness and in health. Genome Res. 18, 343–358 (2008).
Rodić, N. et al. Retrotransposon insertions in the clonal evolution of pancreatic ductal adenocarcinoma. Nat. Med. 21, 1060–1064 (2015).
Lambowitz, A.M. & Perlman, P.S. Involvement of aminoacyl-tRNA synthetases and other proteins in group I and group II intron splicing. Trends Biochem. Sci. 15, 440–444 (1990).
Mohr, S. et al. Thermostable group II intron reverse transcriptase fusion proteins and their use in cDNA synthesis and next-generation RNA sequencing. RNA 19, 958–970 (2013).
Gillis, A.J., Schuller, A.P. & Skordalakes, E. Structure of the Tribolium castaneum telomerase catalytic subunit TERT. Nature 455, 633–637 (2008).
Ding, J. et al. Structure and functional implications of the polymerase active site region in a complex of HIV-1 RT with a double-stranded DNA template-primer and an antibody Fab fragment at 2.8 A resolution. J. Mol. Biol. 284, 1095–1111 (1998).
Lesburg, C.A. et al. Crystal structure of the RNA-dependent RNA polymerase from hepatitis C virus reveals a fully encircled active site. Nat. Struct. Biol. 6, 937–943 (1999).
Holm, L. & Rosenström, P. Dali server: conservation mapping in 3D. Nucleic Acids Res. 38, W545–W549 (2010).
Zhang, Y. & Skolnick, J. TM-align: a protein structure alignment algorithm based on the TM-score. Nucleic Acids Res. 33, 2302–2309 (2005).
Jamburuthugoda, V.K. & Eickbush, T.H. Identification of RNA binding motifs in the R2 retrotransposon-encoded reverse transcriptase. Nucleic Acids Res. 42, 8405–8415 (2014).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Brookes, E., Demeler, B., Rosano, C. & Rocco, M. The implementation of SOMO (SOlution MOdeller) in the UltraScan analytical ultracentrifugation data analysis suite: enhanced capabilities allow the reliable hydrodynamic modeling of virtually any kind of biomacromolecule. Eur. Biophys. J. 39, 423–435 (2010).
Dai, L. et al. A three-dimensional model of a group II intron RNA and its interaction with the intron-encoded reverse transcriptase. Mol. Cell 30, 472–485 (2008).
Mitchell, M., Gillis, A., Futahashi, M., Fujiwara, H. & Skordalakes, E. Structural basis for telomerase catalytic subunit TERT binding to RNA template and telomeric DNA. Nat. Struct. Mol. Biol. 17, 513–518 (2010).
Doublié, S. & Zahn, K.E. Structural insights into eukaryotic DNA replication. Front. Microbiol. 5, 444 (2014).
Doublié, S., Sawaya, M.R. & Ellenberger, T. An open and closed case for all polymerases. Structure 7, R31–R35 (1999).
Keating, K.S., Toor, N., Perlman, P.S. & Pyle, A.M. A structural analysis of the group II intron active site and implications for the spliceosome. RNA 16, 1–9 (2010).
Yean, S.L., Wuenschell, G., Termini, J. & Lin, R.J. Metal-ion coordination by U6 small nuclear RNA contributes to catalysis in the spliceosome. Nature 408, 881–884 (2000).
Sigel, R.K. et al. Solution structure of domain 5 of a group II intron ribozyme reveals a new RNA motif. Nat. Struct. Mol. Biol. 11, 187–192 (2004).
Toor, N., Hausner, G. & Zimmerly, S. Coevolution of group II intron RNA structures with their intron-encoded reverse transcriptases. RNA 7, 1142–1152 (2001).
Beck, C.R., Garcia-Perez, J.L., Badge, R.M. & Moran, J.V. LINE-1 elements in structural variation and disease. Annu. Rev. Genomics Hum. Genet. 12, 187–215 (2011).
Lohmann, V., Körner, F., Herian, U. & Bartenschlager, R. Biochemical properties of hepatitis C virus NS5B RNA-dependent RNA polymerase and identification of amino acid sequence motifs essential for enzymatic activity. J. Virol. 71, 8416–8428 (1997).
Kew, Y., Olsen, L.R., Japour, A.J. & Prasad, V.R. Insertions into the beta3-beta4 hairpin loop of HIV-1 reverse transcriptase reveal a role for fingers subdomain in processive polymerization. J. Biol. Chem. 273, 7529–7537 (1998).
Kabsch, W. Xds. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
Sheldrick, G.M. Experimental phasing with SHELXC/D/E: combining chain tracing with density modification. Acta Crystallogr. D Biol. Crystallogr. 66, 479–485 (2010).
Terwilliger, T.C. et al. Decision-making in structure solution using Bayesian estimates of map quality: the PHENIX AutoSol wizard. Acta Crystallogr. D Biol. Crystallogr. 65, 582–601 (2009).
McCoy, A.J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Afonine, P.V. et al. Towards automated crystallographic structure refinement with phenix.refine. Acta Crystallogr. D Biol. Crystallogr. 68, 352–367 (2012).
Chen, V.B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
Chillon, I. et al. Native purification and analysis of long RNAs. Methods Enzymol. 558, 3–37 (2015).
Fitzgerald, M.E., Vela, A. & Pyle, A.M. Dicer-related helicase 3 forms an obligate dimer for recognizing 22G-RNA. Nucleic Acids Res. 42, 3919–3930 (2014).
Schuck, P. Size-distribution analysis of macromolecules by sedimentation velocity ultracentrifugation and lamm equation modeling. Biophys. J. 78, 1606–1619 (2000).
Daugherty, M.A. & Fried, M.G. Protein-DNA interactions studied at sedimentation equilibrium. in Analytical Ultracentrifugation: Techniques and Methods (eds. Scott, D., Harding, S.E. & Rowe, A.) 195–209 (The Royal Society of Chemistry, 2005).
Folta-Stogniew, E. & Williams, K.R. Determination of molecular masses of proteins in solution: implementation of an HPLC size exclusion chromatography and laser light scattering service in a core laboratory. J. Biomol. Tech. 10, 51–63 (1999).
Acknowledgements
The authors acknowledge E. Folta-Stogniew at the Biophysics Resource of the Keck Facility at Yale University for performing the SEC-MALS experiments. Additionally, we thank F. Liu for kindly providing the plasmid, radiolabeled primer and ladders for the primer extension assay; J.A. Liberman and S. Somarowthu for helpful discussions; and T.H. Dickey for reading the manuscript. The SEC-LS/UV/RI instrumentation is supported by NIH award no. 1S10RR023748-01. This work was supported by National Institutes of Health (NIH) grant RO1GM50313 (A.M.P.) and the Howard Hughes Medical Institute (A.M.P.). C.Z. is supported by a Yale University Fellowship and a Gruber Science Fellowship.
Author information
Authors and Affiliations
Contributions
C.Z. and A.M.P. designed the project. C.Z. performed the experiments and solved the structure. C.Z. and A.M.P. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
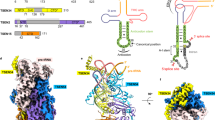
Supplementary Figure 1 Domain construction, sequence, overall structure and active site of R.i. and E.r. maturases.
a. Organization of the maturase RT construct. All maturases contain an N-terminal RT domain (blue) and a C-terminal thumb/X domain (orange). b. Sequence alignment of the full-length maturase, comparing E.r. (Eu.re.I2) and R.i. (Ro.in.I1). The black boxes indicate identical residues, the grey boxes indicate similar residues, and the white box indicates residues with substantial differences. The red bar indicates the boundary for the RT domain used for crystallization. The blue arrow indicates the N-terminal RT domain and the orange arrow indicates the C-terminal X or thumb domain. c. R.i. and E.r. RT domain structures in cartoon representation. The R.i. RT is shown in green and the E.r. RT is shown in gray. d. Comparison of the E.r. RT active site and primer grip with an active telomerase (PDBID: 3KYL)49. The E.r. RT is shown in grey and telomerase is shown in cyan. For the residues shown as sticks, red color indicates oxygen and blue color indicates nitrogen atoms. The potassium in the R.i. RT active site is shown in purple, and the Mg2+ in the telomerase active site is shown in yellow. Metal ion coordination is indicated by orange dashed lines.
Supplementary Figure 2 Maturase RT forms a dimer.
a. Each panel shows the content of an ASU in different crystals, and the organism names and the space groups are indicated at the bottom of each panel. In the dimer, one monomer is colored in grey and the other is colored in green. The ASU of R.i. RT crystallized in P21 space group from condition A (see online Methods) contains 2 molecules. The ASU of R.i. RT crystallized in P212121 space group from condition B (see online methods) contains 4 molecules. The ASU of R.i. RT crystallized in P1 space group from condition C (see online methods) contains 4 molecules. The ASU of E.r. RT crystallized in P21 space group from condition D (see online methods) contains 8 molecules. b. The active site within the R.i. RT domain is partially obstructed by the dimerization interface. On the left panel, monomer A is represented as a cartoon diagram in grey, and monomer B is represented as a space-filling model in green. For monomer A, the active site loop is colored in red, the primer grip is colored in magenta and the insertion loop containing a short α-helix a10 is colored in yellow. The right panel is a close-up view of the interactions that stabilize the insertion loop shown in sticks. Oxygen atoms are colored in red, nitrogen atoms are colored in blue and hydrogen bonds are shown as orange dashed lines.
Supplementary Figure 3 A possible model for association of the maturase dimer with a group II intron RNA.
A cartoon diagram of oceanobacillus iheyensis group II intron (PDBID: 4FAQ)29 is shown on the left, where domain 1 (D1) is colored yellow and domain 4 (D4) is colored in cyan. All other domains are grey. The electrostatic surface of the E.r. RT dimer is shown at right, where blue indicates positive charge and red indicates negative charge. The sizes of the group II intron model and the maturase RT dimer model are not to scale. Based on the electrostatic surface potentials of the RT dimer, and previous results showing that maturase interacts with the intron as a dimer15-16 through a high-affinity binding site in intron D414 and additional auxiliary sites in D148, the potential surfaces involved in RNA-protein interactions are marked by arcs, and the potential interactions are indicated by dashed lines.
Supplementary Figure 4 E.r. RT binds RNA with high affinity and specificity.
a. Comparison of the secondary structures for D4 in the L1.LtrB14 and E.r. introns. The residues colored in red belong to the D4A region. The start codon is highlighted in green. The rest of the open reading frame (ORF) encoding the maturase is indicated by a circle. b. Electrophoretic mobility shift assay (EMSA) for E.r. RT binding to D4A or D2 (Fig. 3b and Fig. 3c). The protein concentrations (nM) for each lane are indicated at the top. c. Phosphate binding site on the R.i. RT domain. The R.i. RT is shown as a surface rendering in grey. The phosphorus atom is orange and the oxygens are red. The simulated annealing omit map for the phosphate ion is shown in blue at the 3σ level. The interactions stabilizing the phosphate are shown on the right panel. Carbon atoms are grey, oxygens are red and nitrogen are blue. Hydrogen bonds are shown as orange dashed lines. d. Molecular weight analysis of E.r. D4A RNA using SEC-MALS. The experiments for E.r. D4A RNA (shades of grey) were performed over two concentrations (in mg/mL) as indicated, and the MW at each concentration was plotted as triangles and circles respectively (upper right legend) on the right y-axis. For each concentration, the UV trace (curve) was plotted on the left y-axis with the elution volume indicated on the x-axis. The MW at elution peak is 23 kDa at 0.03 mg/mL and 24 kDa at 0.07 mg/mL.
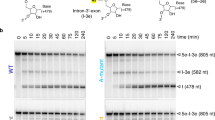
Supplementary Figure 5 Primer extension assay for the maturase.
Lanes 1-3 are DNA sequencing ladders (corresponding to nucleotides A, G and T respectively) showing the sequence and the resolution of the primer extension products on the gel. Lane 4 is the control lane, which only contains the primer. Lane 5 is the primer extension product from the E.r. RT construct, and lane 6 is the primer extension product from the E.r. full-length maturase. In order to resolve DNA products of vastly different length on the same gel (enabling us to compare nonprocessive and processive activity by the RT domain and full-length maturase constructs, respectively), we used a long sequencing gel. The gel is slightly twisted, so each corresponding lane is also labeled at the bottom of the gel to avoid confusion.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5 and Supplementary Table 1 (PDF 1766 kb)
Rights and permissions
About this article
Cite this article
Zhao, C., Pyle, A. Crystal structures of a group II intron maturase reveal a missing link in spliceosome evolution. Nat Struct Mol Biol 23, 558–565 (2016). https://doi.org/10.1038/nsmb.3224
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3224
This article is cited by
-
Engineered CRISPR prime editors with compact, untethered reverse transcriptases
Nature Biotechnology (2023)
-
Structural Analysis of Monomeric RNA-Dependent Polymerases Revisited
Journal of Molecular Evolution (2022)
-
Discovery and evolution of RNA and XNA reverse transcriptase function and fidelity
Nature Chemistry (2020)
-
Using bioinformatic and phylogenetic approaches to classify transposable elements and understand their complex evolutionary histories
Mobile DNA (2017)
-
Globular domain structure and function of restriction-like-endonuclease LINEs: similarities to eukaryotic splicing factor Prp8
Mobile DNA (2017)