Abstract
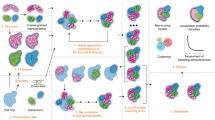
Crosslinking mass spectrometry is increasingly used for structural characterization of multisubunit protein complexes. Chemical crosslinking captures conformational heterogeneity, which typically results in conflicting crosslinks that cannot be satisfied in a single model, making detailed modeling a challenging task. Here we introduce an automated modeling method dedicated to large protein assemblies ('XL-MOD' software is available at http://aria.pasteur.fr/supplementary-data/x-links) that (i) uses a form of spatial restraints that realistically reflects the distribution of experimentally observed crosslinked distances; (ii) automatically deals with ambiguous and/or conflicting crosslinks and identifies alternative conformations within a Bayesian framework; and (iii) allows subunit structures to be flexible during conformational sampling. We demonstrate our method by testing it on known structures and available crosslinking data. We also crosslinked and modeled the 17-subunit yeast RNA polymerase III at atomic resolution; the resulting model agrees remarkably well with recently published cryoelectron microscopy structures and provides additional insights into the polymerase structure.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Change history
07 February 2018
In the version of this article initially published, an important funding source, the Agence National de Recherche (ANR-10-BINF-0003 BIP:BIP to M.N.), was omitted. The error has been corrected in the HTML and PDF versions of the article.
References
Merkley, E.D., Cort, J.R. & Adkins, J.N. Crosslinking and mass spectrometry methodologies to facilitate structural biology: finding a path through the maze. J. Struct. Funct. Genomics 14, 77–90 (2013).
Rappsilber, J. The beginning of a beautiful friendship: crosslinking/mass spectrometry and modelling of proteins and multi-protein complexes. J. Struct. Biol. 173, 530–540 (2011).
Wu, C.-C., Lin, Y.-C. & Chen, H.-T. The TFIIF-like Rpc37/53 dimer lies at the center of a protein network to connect TFIIIC, Bdp1, and the RNA polymerase III active center. Mol. Cell. Biol. 31, 2715–2728 (2011).
Wu, C.-C. et al. RNA polymerase III subunit architecture and implications for open promoter complex formation. Proc. Natl. Acad. Sci. USA 109, 19232–19237 (2012).
Forné, I., Ludwigsen, J., Imhof, A., Becker, P.B. & Mueller-Planitz, F. Probing the conformation of the ISWI ATPase domain with genetically encoded photoreactive crosslinkers and mass spectrometry. Mol. Cell. Proteomics 11, M111.012088 (2012).
Politis, A. et al. A mass spectrometry–based hybrid method for structural modeling of protein complexes. Nat. Methods 11, 403–406 (2014).
Lasker, K. et al. Molecular architecture of the 26S proteasome holocomplex determined by an integrative approach. Proc. Natl. Acad. Sci. USA 109, 1380–1387 (2012).
Erzberger, J.P. et al. Molecular architecture of the 40S·eIF1·eIF3 translation initiation complex. Cell 158, 1123–1135 (2014).
Street, T.O. et al. Elucidating the mechanism of substrate recognition by the bacterial Hsp90 molecular chaperone. J. Mol. Biol. 426, 2393–2404 (2014).
de Vries, S.J. et al. HADDOCK versus HADDOCK: new features and performance of HADDOCK2.0 on the CAPRI targets. Proteins 69, 726–733 (2007).
Kahraman, A. et al. Crosslink guided molecular modeling with ROSETTA. PLoS ONE 8, e73411 (2013).
Kalisman, N., Adams, C.M. & Levitt, M. Subunit order of eukaryotic TRiC/CCT chaperonin by crosslinking, mass spectrometry, and combinatorial homology modeling. Proc. Natl. Acad. Sci. USA 109, 2884–2889 (2012).
Chen, Z.A. et al. Architecture of the RNA polymerase II-TFIIF complex revealed by crosslinking and mass spectrometry. EMBO J. 29, 717–726 (2010).
Robinson, P.J. et al. Molecular architecture of the yeast Mediator complex. Elife 4, e08719 (2015).
Rieping, W., Habeck, M. & Nilges, M. Inferential structure determination. Science 309, 303–306 (2005).
Nilges, M. et al. Accurate NMR structures through minimization of an extended hybrid energy. Structure 16, 1305–1312 (2008).
Habeck, M., Rieping, W. & Nilges, M. Weighting of experimental evidence in macromolecular structure determination. Proc. Natl. Acad. Sci. USA 103, 1756–1761 (2006).
Bouvier, G., Desdouits, N., Ferber, M., Blondel, A. & Nilges, M. An automatic tool to analyze and cluster macromolecular conformations based on self-organizing maps. Bioinformatics 31, 1490–1492 (2015).
Armache, K.-J., Mitterweger, S., Meinhart, A. & Cramer, P. Structures of complete RNA polymerase II and its subcomplex, Rpb4/7. J. Biol. Chem. 280, 7131–7134 (2005).
Méndez, R., Leplae, R., De Maria, L. & Wodak, S.J. Assessment of blind predictions of protein-protein interactions: current status of docking methods. Proteins 52, 51–67 (2003).
Hoffmann, N.A. et al. Molecular structures of unbound and transcribing RNA polymerase III. Nature 528, 231–236 (2015).
Fernández-Tornero, C. et al. Conformational flexibility of RNA polymerase III during transcriptional elongation. EMBO J. 29, 3762–3772 (2010).
Vannini, A. et al. Molecular basis of RNA polymerase III transcription repression by Maf1. Cell 143, 59–70 (2010).
Fernández-Tornero, C. et al. Crystal structure of the 14-subunit RNA polymerase I. Nature 502, 644–649 (2013).
Engel, C., Sainsbury, S., Cheung, A.C., Kostrewa, D. & Cramer, P. RNA polymerase I structure and transcription regulation. Nature 502, 650–655 (2013).
Lefèvre, S. et al. Structure-function analysis of hRPC62 provides insights into RNA polymerase III transcription initiation. Nat. Struct. Mol. Biol. 18, 352–358 (2011).
Zhang, Y. I-TASSER: fully automated protein structure prediction in CASP8. Proteins 77 (suppl. 9), 100–113 (2009).
Merkley, E.D. et al. Distance restraints from crosslinking mass spectrometry: mining a molecular dynamics simulation database to evaluate lysine-lysine distances. Protein Sci. 23, 747–759 (2014).
Ferri, M.L. et al. A novel subunit of yeast RNA polymerase III interacts with the TFIIB-related domain of TFIIIB70. Mol. Cell. Biol. 20, 488–495 (2000).
He, Y., Fang, J., Taatjes, D.J. & Nogales, E. Structural visualization of key steps in human transcription initiation. Nature 495, 481–486 (2013).
Thuillier, V., Stettler, S., Sentenac, A., Thuriaux, P. & Werner, M. A mutation in the C31 subunit of Saccharomyces cerevisiae RNA polymerase III affects transcription initiation. EMBO J. 14, 351–359 (1995).
Kosinski, J. et al. Xlink Analyzer: software for analysis and visualization of crosslinking data in the context of three-dimensional structures. J. Struct. Biol. 189, 177–183 (2015).
Moreno-Morcillo, M. et al. Solving the RNA polymerase I structural puzzle. Acta Crystallogr. D Biol. Crystallogr. 70, 2570–2582 (2014).
Leitner, A. et al. Expanding the chemical crosslinking toolbox by the use of multiple proteases and enrichment by size exclusion chromatography. Mol. Cell Proteomics 11, M111.014126 (2012).
Kettenberger, H., Armache, K.-J. & Cramer, P. Complete RNA polymerase II elongation complex structure and its interactions with NTP and TFIIS. Mol. Cell 16, 955–965 (2004).
Xu, H. & Freitas, M.A. MassMatrix: a database search program for rapid characterization of proteins and peptides from tandem mass spectrometry data. Proteomics 9, 1548–1555 (2009).
Rinner, O. et al. Identification of crosslinked peptides from large sequence databases. Nat. Methods 5, 315–318 (2008).
Walzthoeni, T. et al. False discovery rate estimation for crosslinked peptides identified by mass spectrometry. Nat. Methods 9, 901–903 (2012).
Kosinski, J., Barbato, A. & Tramontano, A. MODexplorer: an integrated tool for exploring protein sequence, structure and function relationships. Bioinformatics 29, 953–954 (2013).
Söding, J. Protein homology detection by HMM-HMM comparison. Bioinformatics 21, 951–960 (2005).
Eswar, N. et al. Comparative protein structure modeling using MODELLER. Curr. Protoc. Protein Sci. 51, 2.9.1–2.9.31 (2007).
Nilges, M., Malliavin, T. & Bardiaux, B. in Solid-State NMR Studies of Biopolymers (eds. McDermott, A.E. & Polenova, T.) Ch. 22 (John Wiley & Sons, Ltd., 2010).
Nilmeier, J.P., Crooks, G.E., Minh, D.D.L. & Chodera, J.D. Nonequilibrium candidate Monte Carlo is an efficient tool for equilibrium simulation. Proc. Natl. Acad. Sci. USA 108, E1009–E1018 (2011).
Brünger, A.T. et al. Crystallography & NMR system: A new software suite for macromolecular structure determination. Acta Crystallogr. D Biol. Crystallogr. 54, 905–921 (1998).
Joosten, R.P. et al. A series of PDB related databases for everyday needs. Nucleic Acids Res. 39, D411–D419 (2011).
Marks, D.S. et al. Protein 3D structure computed from evolutionary sequence variation. PLoS ONE 6, e28766 (2011).
Acknowledgements
We thank B. Bardiaux for his help and expertise with the CNS software and N. Hoffmann for discussions. We acknowledge support from the EMBL Proteomics Core Facility. This work was supported by the EMBL Interdisciplinary Postdoc Programme under Marie Curie COFUND Actions (J.K., grant number 291772), postdoctoral fellowships from the Alexander von Humboldt foundation and Marie Curie Actions (A.O.), the Agence National de Recherche (ANR-10-BINF-0003 BIP:BIP to M.N.), and the European Union (FP7-IDEAS-ERC 294809 to M.N. and ERC-2013-AdG 340964-POL1PIC to C.W.M.). M.M.-M. and U.J.R. acknowledge support by EMBO Long-Term fellowships and by the Marie-Curie fellowship (FP7-PEOPLE-2011-IEF 301002 to M.M.-M.).
Author information
Authors and Affiliations
Contributions
M.F., J.K., P.R.B. and M.N. designed and performed modeling, analyzed data and wrote the manuscript; A.O., M.M.-M. and U.J.R. performed experiments and analyzed data; A.O. and U.J.R. performed crosslinking; B.S. analyzed data; G.B. analyzed structure distributions; C.W.M., M.B. and M.N. designed experiments, oversaw the project and wrote the manuscript. All authors contributed to editing the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Distribution of distances between cross-linked lysine pairs resembles a log-normal distribution
(a) Distribution of distances between cross-linked lysine pairs (blue) in the Pol II cross-link dataset, while the distribution of all lysine pairs is normal (green). Cross-links are mapped on the Pol II crystal structure. The cross-links are colored blue if the linked positions are satisfied (Cα-Cα less than 30 Å apart) or red if violated (Cα-Cα further than 30 Å apart). (b) Distribution of distances between cross-linked lysine pairs in the Pol III core structural model and mapped cross-links. Coloring as in (a). Both in (a) and (b) structures are depicted in cross-eye stereo view.
Supplementary Figure 2 Cross-link satisfaction by the representative model of Pol II
Cross-links of Pol II mapped on (a) the X-ray structure (pdb: 1WCM) and (b) the selected model. The cross-links are colored blue if the linked positions are satisfied (Cα-Cα less than 30 Å apart) or red if violated (Cα-Cα further than 30 Å apart).
Supplementary Figure 3 The performance of our method depends on the number of cross-links and their localization in the structures.
(a) Modeling test on the designed complex of colicin E7 DNase and the Im7 Immunity protein55 using previously published cross-links11. The convergence location of the mobile subunit (shown in red) during the simulation is shown as a light red volume density. Despite the apparent compatibility between the cross-links and the crystal structure, the native conformation could not be explored because only two lysine residues are coss-linked on E7. (b) Modeling test on ovotransferrin56 using previously published cross-links11. The convergence location of the mobile subunit (shown in red) during the simulation is shown as a light red volume density. Despite the availability of 6 cross-links, two of them are identified as false positives on the crystal structure. In turn, the mobile subunit converges in a location that is very different from the native location. This illustrates the limits of comparison between cross-linking data and X-ray data. Both in (a) and (b) cross-links are shown in blue if satisfied, and in red if violated.
55. Kortemme, T. et al. Computational redesign of protein-protein interaction specificity. Nat. Struct. Mol. Biol. 11, 371–379 (2004).
56. Mizutani, K., Mikami, B., Aibara, S. & Hirose, M. Structure of aluminium-bound ovotransferrin at 2.15 Angstroms resolution. Acta Crystallogr. D Biol. Crystallogr. 61, 1636–1642 (2005).
Supplementary Figure 4 Starting conformations of the mobile subunits of Pol III
(a) Model of C31. (b) Model of C82. (c) Model of C34. The C82 insertions and the C34 linkers between WH domains (i.e. regions missing in the homology models that were added as flexible loops) are highlighted in purple.
Supplementary Figure 5 Lysine-lysine XL-MS cross-links of Pol III obtained in this work
Pol III subunits are shown as rectangular bars except C160 and C128, which are shown as ovals for the sake of clarity. Inter-links are shown as lines connecting the protein bars, while intra-links are shown as curves. Inter-links to C31 are colored yellow, to C34 - gold, to C37 – violet, to C53 - cyan. The remaining inter-links are colored gray. Domains of C82 and C34 discussed in this work are indicated. Regions missing in crystal structures or homology models are colored black. The figure was created with xiNET57.
57. Combe, C. W., Fischer, L. & Rappsilber, J. xiNET: cross-link network maps with residue resolution. Mol. Cell Proteomics 14, mcp.O114.042259–1147 (2015).
Supplementary Figure 6 Analysis of the results of the Pol III simulation
(a) Cross-linked Nζ-Nζ distance distribution and photo-cross-linked d-20-summed distance (see Online Methods) distribution on the entire Pol III modeling trajectory. The peak around 15 Å was consistent with data from Pol II and from the core complex of Pol III (Supplementary Fig. 1). A second peak was observed at around 45 Å, and corresponded to the restraints that were down-weighted during the conformational search and thus followed a distribution resembling that of not cross-linked residues2 (Supplementary Fig. 1a). (b) MS-cross-links satisfaction (Cα-Cα distance <30 Å) projected onto the SOM space and scatter plot of the relationship between the U-matrix score (local similarity) and the cross-links satisfaction. (c) Average iRMSD to reference structure projected onto the SOM space and scatter plot of the relationship between the U-matrix score and the iRMSD.
Supplementary Figure 7 Venn diagram summary of satisfied XL-MS cross-links between clusters in the Pol III simulation.
See Fig. 3 for visualization of the clusters.
Supplementary Figure 8 Agreement of the Pol III model with photo-cross-links
Photo-cross-linking residues3,4 are depicted as spheres. The residues cross-linking to C82 are colored orange, to C34 - gold, to C160 – cyan, to C31 – yellow. Only the photo-cross-links involving the C31/C82/C34 trimer are shown.
Supplementary Figure 9 The representative Pol III model of cluster 1 better explains available experimental data than the previously published models
(a) BPA photo-cross-links from C82 to C160 used in modeling mapped on the model. The positions making the photo-cross-links are marked as cyan spheres. (b) BPA photo-cross-links from C82 to C31 not used in modeling. The positions making the photo-cross-links are marked as yellow spheres. (c) The positioning of the WH2 domain of the C34 subunit agrees with the lysine-lysine cross-links not used in modeling and the photo-cross-links from position 187 of C373. From the lysine-lysine cross-links, only the inter-cross-links involving C34 subunit are shown. The cross-links are colored blue if the linked positions are satisfied (Cα-Cα less than 30 Å apart) or red if violated (Cα-Cα further than 30 Å apart). The photo-cross-linking position is labeled.
Supplementary Figure 10 Control modeling simulations of Pol III without including missing regions.
To demonstrate the benefit of including missing regions in the Pol III conformational search, we performed a control simulation without including these regions. In particular, the insertions of C82 (Supplementary Figure 2) and the whole C31 subunit were removed. (a) After clustering, the U-matrix revealed one main convergence basin. (b) Average iRMSD to the crystal structure projected onto the SOM space. The simulation does not converge towards the best conformations anymore. (c) The resulting structure shows that the C82/C34 subcomplex is disconnected from the stalk due to the absence of C31 that mediates the contacts. The C34 WH3 has undefined positioning due to the omission of the cross-links with the C82 insertions.
Supplementary Figure 11 Localization densities of the Pol III subcomplex computed in three distinct simulations with three distinct C31 models.
Only the most constant domains remain, while the highly fluctuating parts of the subcomplex, such as C34 WH3, do not appear. Pol III core complex is shown in gray. C31, C82 and C34 densities are represented in metallic yellow, orange and yellow, respectively. C31 model 1 (a) model 2 (b) and model 3 (c) confer the same overall conformation to the subcomplex.
Supplementary Figure 12 Per-residue Root Mean Square Fluctuation on three selected subunits during Pol III conformational sampling.
(a) C160 fluctuations are around 0.5Å, except for the region located at the interface with the mobile subunits. (b) C82, as a mobile subunit, is more flexible by design. Unrestrained insertions are located with red bars. (c) Internal restraints of C31 were down-weighted individually prior to the simulation, by a factor that depended on the per-residue reliability of the starting model (given by I-TASSER webserver, which was used to model the initiation C31 structure). The most drastic effect was a reduction of some weights by up to 99%. It resulted in higher conformational fluctuations.
Supplementary Figure 13 Projection of the 20 sub-trajectories of Pol III onto the SOM space.
Most trajectories explore a wide part of the conformational space and are not stuck in local minima. Running parallel trajectories is therefore equivalent to running a single longer trajectory.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–13, Supplementary Tables 4 and 5, and Supplementary Note 1 (PDF 1740 kb)
Supplementary Table 1
Table of experimental Pol II cross-links (obtained from Chen et al.13) (XLSX 46 kb)
Supplementary Table 2
Table of experimental Pol III cross-links in xQuest format38 (XLS 66 kb)
Supplementary Table 3
Table of experimental Pol III cross-links that involve the C34/C82/C31 subcomplex (XLSX 11 kb)
Supplementary Software
XL-MOD Software (ZIP 4666 kb)
Rights and permissions
About this article
Cite this article
Ferber, M., Kosinski, J., Ori, A. et al. Automated structure modeling of large protein assemblies using crosslinks as distance restraints. Nat Methods 13, 515–520 (2016). https://doi.org/10.1038/nmeth.3838
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.3838
This article is cited by
-
Development of R7BP inhibitors through cross-linking coupled mass spectrometry and integrated modeling
Communications Biology (2019)
-
Efficient and robust proteome-wide approaches for cross-linking mass spectrometry
Nature Protocols (2018)
-
Erratum: Corrigendum: Automated structure modeling of large protein assemblies using crosslinks as distance restraints
Nature Methods (2018)
-
M3: an integrative framework for structure determination of molecular machines
Nature Methods (2017)
-
Facilitating identification of minimal protein binding domains by cross-linking mass spectrometry
Scientific Reports (2017)