Abstract
Thermal stabilization of proteins after ligand binding provides an efficient means to assess the binding of small molecules to proteins. We show here that in combination with quantitative mass spectrometry, the approach allows for the systematic survey of protein engagement by cellular metabolites and drugs. We profiled the targets of the drugs methotrexate and (S)-crizotinib and the metabolite 2′3′-cGAMP in intact cells and identified the 2′3′-cGAMP cognate transmembrane receptor STING, involved in immune signaling.
Similar content being viewed by others
Main
The systematic matching of small chemical compounds such as metabolites, xenobiotics, natural products and synthetic drugs to the repertoire of cellular proteins with which they have functional relationships has been a major technical hurdle and paramount objective in chemical biology.
Many successful target-deconvolution approaches use chemical modifications1 to confer properties to compounds that allow for efficient affinity purification or target labeling2,3. An ideal method for monitoring engagement with targets by small molecules would not require modification of the chemical agent, would be unbiased and thus would not rely on previous assumptions, and would operate with intact cells and proteins in their native environment.
Ligand binding can induce a conformational change in the target protein, which in turn can alter, for instance, the accessibility of protease cleavage sites4,5. Protein thermal stability in vitro has been used for many years in crystallography and drug discovery programs as a robust method to identify ligands for various protein classes6,7. The approach is based on the frequent observation that binding of a ligand to a protein yields a complex with increased thermodynamic stability compared with the free protein. This difference in stability is reflected in a change in the apparent melting temperature (Tm) of the protein and can be measured by, for example, differential scanning fluorimetry (Fig. 1a). A recent breakthrough study demonstrated that this concept can be expanded to monitor drug-target engagement in live cells (cellular thermal-shift assay)8. However, in that work, the approach was used only to investigate binding of compounds to a given cognate target.
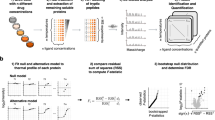
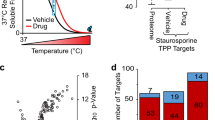
(a) Binding of a ligand to its target protein (top) increases the enthalpy required for unfolding. As a result, the melting temperature (Tm) is shifted (bottom). This can be exploited with, for example, differential scanning fluorimetry ('Tm shift assay'). A.U., arbitrary units. (b) Representative workflow. Cells are treated with either small molecule or vehicle, washed and harvested. Equal amounts of cells are then heated to increasing temperatures and lysed, and the supernatant obtained after centrifugation is digested and labeled for quantitative mass spectrometry (MS) analysis. Detection of changes in protein stability in treated versus control samples is enabled by bioinformatic processing and data plotting in the form of differential protein-abundance graphs. (c) The bioinformatics scoring parameters used for data analysis. Continuous lines depict raw data, and dashed lines are curve-fitting results.
We hypothesized that the methodology could be developed further to identify any cellular targets, including unknown and unsuspected targets, that are stabilized upon binding of a small molecule. Combined with modern mass spectrometry, the approach should provide a comprehensive view of ligand-protein interactions through the highly multiplexed determination of individual Tm shifts (Fig. 1b,c). A recent study reported the use of thermal profiling coupled with mass spectrometry9. The method was used efficiently to identify the targets of kinase inhibitors, highlighting the feasibility of such a methodology for drug discovery. However, despite its pioneering nature, that study also revealed some limitations, as the BCR-ABL inhibitor dasatinib surprisingly did not alter the cognate target BCR-ABL in K562 cells. The report also did not address the possibility of revealing target proteins other than kinases or whether the approach could be used to identify membrane protein targets of small-molecule metabolites as well as drugs.
We applied our procedure to (S)-crizotinib, an inhibitor of the Nudix hydrolase MTH1. We and others have recently shown that disruption of nucleotide-pool homeostasis by MTH1 inhibition may provide an effective new means to treat fast-replicating tumors and that (S)-crizotinib is a highly specific inhibitor of MTH1 (refs. 3,10). In our previous study, we profiled (S)-crizotinib by means of both classical chemoproteomics and extensive biochemical screening. These results therefore provide an excellent data set for evaluating the efficacy and validity of our new approach.
We heated aliquots of SW480 colon carcinoma cells treated with either (S)-crizotinib or dimethyl sulfoxide (DMSO) to different temperatures and digested equal volumes of supernatants after cell lysis and centrifugation with trypsin (Fig. 1b). To enable reliable and comparative quantification of protein abundance in the samples at varying temperatures, we used a labeling strategy based on isobaric tandem mass tags (TMTs)11. The pooled 10-plex samples were fractionated offline before being analyzed on a Q-Exactive mass spectrometer, which provided data for more than 3,400 proteins after stringent filtering. We normalized the data (Supplementary Fig. 1) and compared DMSO-treated samples to (S)-crizotinib–treated samples by fitting a sigmoid model to each and scoring the thermal shift (ΔTm; Fig. 1c). We identified MTH1 (also known as NUDT1) as the top cellular target of (S)-crizotinib on the basis of a pronounced stabilizing effect, which was notably absent for the related protein-family member NUDT5 (Fig. 2a). We also observed shifts for other proteins linked to DNA repair, such as POLA12 (Supplementary Fig. 2 and Supplementary Table 1).
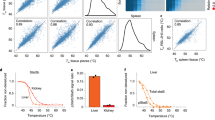
(a) Protein-abundance graph derived from mass spectrometry TMT 10-plex experiments showing data for the cognate (S)-crizotinib target MTH1 and the Nudix family member NUDT5. Data are representative of two independent experiments (n = 2). (b) Western blot confirming the stabilization of MTH1 by (S)-crizotinib in intact SW480 cells. Images are cropped for clarity; full-length blots are presented in Supplementary Figure 3. (c) Protein-abundance graphs derived from mass spectrometry TMT 10-plex experiments profiling DHFR as a target of MTX. Data are representative of two independent experiments (n = 2). (d) Immunoblot confirming the stabilization of DHFR by MTX in intact K562 cells. Images are cropped for clarity; full-length blots are presented in Supplementary Figure 6. (e) Results of mass spectrometry analysis of RAW macrophages treated with 2′3′-cGAMP to profile the stabilization of the transmembrane receptor STING. Data are representative of two independent experiments (n = 2). In a, c and e, individual data points are marked by ×; curves represent lines of best fit. (f) Western blot validating the increased thermal stability observed for STING after treatment with 2′3′-cGAMP. Images are cropped for clarity; full-length blots are presented in Supplementary Figure 7.
To validate the thermal stabilization of MTH1 by (S)-crizotinib as detected by mass spectrometry under these experimental conditions, we performed an analogous experiment using SW480 colon carcinoma cells and analyzed the supernatants by immunoblotting. Indeed, (S)-crizotinib protected MTH1 from unfolding, whereas DMSO-treated control cells showed a sharp decrease in MTH1 content (Fig. 2b and Supplementary Fig. 3).
To investigate potential differences between intact cells and lysates, we performed an analogous experiment using SW480 cell lysates (Supplementary Table 2). As reported for other compounds tested with the immunoblotting method8, we applied (S)-crizotinib at a higher final concentration (100 μM) to avoid potential dilution effects in the lysate. In contrast to the experiment with intact cells, the analysis revealed not only MTH1 but also several kinases as targets, many of which had been highlighted by the previous in vitro kinase screen (Supplementary Figs. 4 and 5 and Supplementary Table 3). This supports the notion that cellular compartmentalization may affect a compound's target profile and that thermal profiling experiments using intact cells may be more likely to reflect the actual physiological context than experiments with lysates.
Next we evaluated the cellular-target profile of the antimetabolite methotrexate (MTX) in K562 cells. Our analysis identified the cognate and well-studied target dihydrofolate reductase (DHFR) as the top candidate (Fig. 2c and Supplementary Table 4). We confirmed the stabilization by immunoblotting (Fig. 2d and Supplementary Fig. 6). We also detected a highly reproducible shift for thymidylate synthase (TYMS), an important mediator of MTX action. MTX exhibits inhibitory activity toward TYMS only at concentrations above 10 μM, whereas it has been shown that MTX polyglutamate metabolites are active at concentrations in the nanomolar range13. As our treatment conditions were compatible with the formation of MTX polyglutamate species, the data indicate that our method—in contrast to classical target-identification approaches—could enable tracking of both drug and drug metabolite effects.
Our results prompted us to further investigate the possibility of using the methodology to elucidate the molecular interaction partners of cellular metabolites, which would make it an option for applications beyond drug-target deconvolution. We examined whether the approach could recover the cellular targets of the recently reported cGAMP synthase metabolite 2′3′-cGAMP14. Binding of 2′3′-cGAMP to its receptor STING (stimulator of interferon genes; also known as TMEM173) elicits the transcription of proinflammatory cytokines. To ensure sufficient engagement of STING by its endogenous ligand, we transfected RAW macrophages with 2′3′-cGAMP and analyzed the samples by mass spectrometry. Bioinformatic filtering showed a reproducible shift for STING as well as for well-known downstream mediators of innate immune signaling such as NF-κB and STATs (signal transducers and activators of transcription)15 (Fig. 2e and Supplementary Table 5). We also confirmed a stabilizing effect on STING by immunoblotting (Fig. 2f and Supplementary Fig. 7), demonstrating not only that the approach can recapitulate the activation of the inflammatory response by the cGAMP synthase metabolite, but also that the procedure is compatible with transmembrane proteins.
Our study demonstrates that this type of approach is applicable to enzymes other than kinases (as was shown by Savitski et al.9), such as hydrolases and oxidoreductases, as well as to the detection of transmembrane protein targets. Moreover, our procedure differed from that of Savitski et al.9 in sample processing, mass spectrometric analysis, and computational and statistical interpretation of the data (Supplementary Note 1). Our approach may be further tailored to specific target classes, subcellular compartments or tissues.
The choice of temperature steps may be critical for capturing differences in individual stabilities in particular groups of proteins. Therefore, several experiments with different temperature ranges may be required to identify the unknown targets of less well-studied compounds. Future systematic work will undoubtedly show correlations between ligand pocket architecture and many other parameters. Moreover, thorough charting of the general thermal stability of the proteins in a cell should uncover many interesting features regarding the thermal stability of cellular proteins, as well as their subcellular localization, abundance, multiprotein-complex formation and post-translational modifications. Finally, there is an increasing appreciation of the fundamental role of cellular metabolism in human disease, and an exact mapping of the functional relationships between metabolites and their protein counterparts could not only increase our mechanistic understanding but also reveal new potential drug targets. Importantly, the approach should allow for target engagement in human biopsies. Combined with targeted quantitative proteomics approaches, the method could focus on specific biomarkers and thus act in synergy with other efforts in this area16.
In summary, we have presented a methodology that takes advantage of alterations in protein thermal stability after ligand binding, providing a straightforward approach for identifying the cellular proteins that are engaged by a metabolite or a drug in intact, living cells.
Methods
Cell culture.
K562 and RAW 264.7 cells were obtained from DSMZ and ATCC, respectively. SW480 cells were kindly provided by W. Berger (Institute of Cancer Research). All cells were cultured in the recommended media containing 10% FBS and 10 U ml−1 penicillin-streptomycin (Gibco) and checked for mycoplasma by PCR or ELISA before experimental use.
Immunoblotting.
The following antibodies were used according to the manufacturer's instructions: mouse anti-DHFR (A-9, sc-377091, Santa Cruz Biotechnology)8, rabbit anti-MTH1 (NB100-109, Novus Biologicals)3 and rabbit anti-STING (3337, Cell Signaling)17.
Sample preparation.
SW480 cells were cultured in 12-well cell culture plates to 80% confluency and treated with media containing either DMSO or 5 μM (S)-crizotinib for 3 h. K562 cells were cultured in T75 cell culture flasks and, upon reaching about 80% confluency, aliquoted into 12-well cell culture plates and treated with media containing either DMSO or 10 μM MTX for 3 h. After treatment, cells were detached with trypsin (SW480 only), collected by centrifugation and resuspended in PBS. Cell suspensions were transferred into 0.2-ml PCR tubes and heated for 3 min. After a subsequent 3-min incubation at room temperature, cells were lysed by the addition of 30 μl of lysis buffer and three repeated freeze-thaw cycles using liquid nitrogen. Precipitated proteins were separated from the soluble fraction by centrifugation at 17,000g for 20 min at 4 °C. For metabolite experiments, RAW cells were cultured in 100-mm culture plates to 80% confluency and transfected with either water or 2′3′-cGAMP (10 μM, InvivoGen) using Lipofectamine 2000. After 4 h cells were harvested and treated as described above. For cell lysate experiments, cells were lysed with lysis buffer (50 mM Tris-HCl, 100 mM NaCl, 0.2% NP-40, 5% glycerol, 1.5 mM MgCl2, 25 mM NaF, 1 mM Na3VO4, 1 mM phenylmethylsulfonyl fluoride, 1 mM dithiothreitol (DTT), 10 μg/mL TLCK, 1 μg/mL leupeptin, 1 μg/mL aprotinin, and 10 μg/mL soybean trypsin inhibitor (Sigma), pH 7.5) and, after centrifugation, incubated with DMSO or (S)-crizotinib (100 μM) for 30 min (6 mg total protein per condition). Six aliquots were prepared for each sample and heated for 3 min in a PCR machine. Precipitated proteins were separated from the soluble fraction by centrifugation as described above.
In-solution tryptic digestion, TMT derivatization and 2D reversed-phase liquid chromatography.
The protein concentration of cell lysates was determined using the Bradford method (Bio-Rad). Thirty (SW480), fifty (K562) and eighty (RAW) micrograms of total protein were denatured with 8 M urea and reduced with DTT (10 mM), and cysteine residues were alkylated with iodoacetamide (55 mM) and digested with modified porcine trypsin (1:100, Worthington Biochemical) overnight at 37 °C after the dilution of urea to less than 2 M. After quenching with trifluoroacetic acid (TFA), peptides were purified by solid-phase extraction using C18 SPE columns (The Nest Group). Eluted peptides were lyophilized in a vacuum centrifuge and resuspended in TEAB buffer to a final concentration of 100 mM before being labeled with the TMT reagents (Thermo Fisher Scientific). Labeling was performed according to the instructions provided by the manufacturer. After quenching, all samples were pooled, purified by solid-phase extraction and separated at pH 10 by reversed-phase liquid chromatography as described18 using an Agilent 1200 series HPLC system. Separation was performed at a flow rate of 100 μl min−1 on a Phenomenex column (150 × 2.0 mm Gemini-NX, 3 μm C18, 110 Å) with a 50-min linear gradient of 5–70% (vol/vol) acetonitrile containing 20 mM ammonium formate. Twenty time-based fractions were automatically collected, and peptides were lyophilized in a vacuum centrifuge and reconstituted in 5% (vol/vol) formic acid for on-line liquid chromatography–mass spectrometry (LC-MS). The peptide abundance per fraction was estimated on the basis of the UV trace, and samples were diluted accordingly to avoid overloading the nano–LC-MS system. Fractions were injected onto a Dionex Ultimate 3000 system (Thermo Fisher Scientific) coupled to a Q-Exactive mass spectrometer (Thermo Fisher Scientific). Peptides were loaded onto a trap column (Zorbax 300SB-C18 5 μm, 5 × 0.3 mm, Agilent Biotechnologies) at 45 μl min−1 and separated on a customized 16 cm × 50 μm inner diameter reversed-phase column packed with 3-μm C18 particles (ReproSil-Pur 120 C18-AQ, Dr. Maisch). Peptides were separated at a constant flow rate of 100 nl min−1 with an 85-min gradient of 3–35% (vol/vol) solvent B that was increased to 70% (vol/vol) solvent B within 12 min and, finally, to 100% (vol/vol) solvent B for 10 min before re-equilibration to 3% solvent B. Solvent A consisted of 0.4% (vol/vol) formic acid and 70% (vol/vol) methanol, and solvent B consisted of 20% (vol/vol) isopropanol and 0.4% (vol/vol) formic acid. Analysis on the Q-Exactive was performed in data-dependent mode with the top ten most intense ions selected for fragmentation. MS and higher-energy collision-induced dissociation (HCD) MS spectra2 were acquired at 70,000 and 35,000 resolution (at m/z 200), respectively. Automatic gain control was used to prevent overfilling of the ion trap and was set to 3 × 106 and 2 × 105 ions for MS and HCD, respectively. Maximum ion accumulation times were set to 250 and 120 ms for MS and HCD, respectively. The dynamic exclusion for selected ions was 60 s. The singly charged siloxane mass at m/z 445.120024 was used for lock mass correction19. The threshold for switching from MS to tandem mass spectrometry (MS/MS) was 1% of the target ion value. The normalized collision energy was 29%, with a fixed first mass of 100 m/z. Software versions used for the operation of the Q-Exactive were Tune 2.3 SP1 and Xcalibur 3.0.63.
Bioinformatic analysis.
All MS/MS spectra were searched against UniProtKB/Swiss-Prot human with Mascot20 and Phenyx21. Results were merged and false discovery rate imposed a maximum of 1% according to a previously described procedure22. Only spectra providing a signal above the estimated background noise23 in all ten TMT channels were considered, and we further limited the analysis to proteins identified by at least two such spectra.
Data normalization.
Classical normalization approaches could not be applied because the total amount of proteins in samples at increasing temperatures was reduced on purpose. Nonetheless, it could be appreciated that un-normalized data, especially those obtained with DMSO treatment, showed a departure from the expected sigmoid curve at the first three temperatures above the lowest temperature (Supplementary Fig. 1). This was observed in the original western blotting procedure8, and we hypothesize that this phenomenon represents an artifact due to protein-crowding effects at the lower temperatures. We thus reasoned that data could be normalized by their shape instead of their values. To this end we summed 50,000 randomly selected spectra in each sample separately, modeling the average temperature-dependent abundance decrease, which we fitted by a sigmoid of the form

where t is the temperature and a and b are the curve parameters (determined by nonlinear least-square optimization). At each temperature, the summed spectrum was compared with the sigmoid and a relative shift was determined that was subsequently applied to all the proteins of the corresponding sample as a means of normalization (Supplementary Fig. 1).
Target scoring.
Two scores were computed to capture different features of the thermal shift that should be observed in targets: a temperature shift at 50% reduction of protein abundance in the sample, and a signed area between the DMSO- or vehicle-treated sample curve and the compound-treated curve (Fig. 1c and Supplementary Fig. 8). The sigmoid curves required to estimate the thermal shift at 50% concentration reduction were of the form above. The two scores were necessary because more than 3,000 proteins had to be scored in each sample comparison, and odd curves were able to yield good values according to one of the two scores by chance. In addition, it was required that a potential drug target feature at least a 25% protein abundance reduction at the highest temperature under compound treatment and at least a 75% reduction under vehicle for MTX; these values were, respectively, 25% and 50% for (S)-crizotinib and 10% and 10% for 2′3′-cGAMP. These nonstringent conditions limited the analysis to proteins for which the assay generated meaningful, interpretable data. This procedure also eliminated some oddly shaped melting profiles that generated tentatively decent scores by coincidence and would be discarded immediately upon expert inspection.
Target selection.
Exploiting the two replicates available in each cell line for each compound and vehicle control, we created two pairs of vehicle versus compound-treated samples. Proteins identified in a pair were scored as described above. In each pair we retained proteins in the top p% of both scores and intersected such selections across the two biological replicates. We used P = 10 for MTX, P = 15 for (S)-crizotinib and P = 20 for 2′3′-cGAMP. We calculated the coefficient of variation (CV) values reported in Supplementary Tables 1, 4 and 5 by applying the classical definition (s.d. divided by the mean) to the two replicates of a given score (temperature shift or signed area). Namely, if s1 and s2 are the replicates of one of the two scores, CV = (√2)|s1 – s2|/(s1 + s2).
Accession codes.
The raw mass spectrometric data used in this study are available via PeptideAtlas with the identifier PASS00693.
References
Lee, J. & Bogyo, M. Curr. Opin. Chem. Biol. 17, 118–126 (2013).
Kambe, T., Correia, B.E., Niphakis, M.J. & Cravatt, B.F. J. Am. Chem. Soc. 136, 10777–10782 (2014).
Huber, K.V.M. et al. Nature 508, 222–227 (2014).
Feng, Y. et al. Nat. Biotechnol. 32, 1036–1044 (2014).
Lomenick, B. et al. Proc. Natl. Acad. Sci. USA 106, 21984–21989 (2009).
Niesen, F.H., Berglund, H. & Vedadi, M. Nat. Protoc. 2, 2212–2221 (2007).
Fedorov, O. et al. Proc. Natl. Acad. Sci. USA 104, 20523–20528 (2007).
Martinez Molina, D. et al. Science 341, 84–87 (2013).
Savitski, M.M. et al. Science 346, 1255784 (2014).
Gad, H. et al. Nature 508, 215–221 (2014).
Dayon, L. et al. Anal. Chem. 80, 2921–2931 (2008).
Parlanti, E., Locatelli, G., Maga, G. & Dogliotti, E. Nucleic Acids Res. 35, 1569–1577 (2007).
Allegra, C.J. et al. J. Biol. Chem. 260, 9720–9726 (1985).
Ablasser, A. et al. Nature 498, 380–384 (2013).
Cai, X., Chiu, Y.-H. & Chen, Z.J. Mol. Cell 54, 289–296 (2014).
Lambert, J.-P. et al. Nat. Methods 10, 1239–1245 (2013).
Yoh, S.M. et al. Cell 161, 1293–1305 (2015).
Gilar, M., Olivova, P., Daly, A.E. & Gebler, J.C. J. Sep. Sci. 28, 1694–1703 (2005).
Olsen, J.V. et al. Mol. Cell. Proteomics 4, 2010–2021 (2005).
Perkins, D.N., Pappin, D.J.C., Creasy, D.M. & Cottrell, J.S. Electrophoresis 20, 3551–3567 (1999).
Colinge, J., Masselot, A., Giron, M., Dessingy, T. & Magnin, J. Proteomics 3, 1454–1463 (2003).
Burkard, T.R. et al. BMC Syst. Biol. 5, 17 (2011).
Breitwieser, F.P. et al. J. Prot. Res. 10, 2758–2766 (2011).
Acknowledgements
We are grateful to W. Berger (Institute of Cancer Research, Vienna, Austria) for providing SW480 cells and P. Majek (CeMM, Vienna, Austria) for assistance with data processing. This work was supported by the Austrian Academy of Sciences, the European Union (FP7 259348, ASSET) and the Austrian Science Fund (FWF F4711, MPN).
Author information
Authors and Affiliations
Contributions
K.V.M.H. designed experiments and jointly performed them with K.M.O.; A.C.M. and K.L.B. performed mass spectrometry; C.S.H.T. and J.C. performed bioinformatics analysis; and K.V.M.H. and G.S.-F. conceived the study and wrote the manuscript. All authors contributed to the discussion of results and participated in preparation of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Data normalization.
Average thermal profiles are plotted for DMSO- and (S)-crizotinib-treated cells. Note that the two profiles (broken straight lines) are similar (no thermal shift) but imperfect. The two fitted sigmoid models are very close (dashed smooth lines), thus further indicating the absence of influence of (S)-crizotinib on the average thermal stability. Vertical arrows (only displayed for the DMSO sample for simplicity), expressed as shift in normalized coordinates, can be applied to spectra specific to a given protein for subsequent scoring of the whole dataset and can be regarded as a structural normalization considering the different protein content at each temperature.
Supplementary Figure 2 Known interactions between proteins found to be stabilized in experiments with intact SW480 cells.
Edge thickness represents interaction confidence. Data and graphical representation taken from STRING1.
Supplementary Figure 3 Original full-length anti-MTH1 western blots used for Figure 2b.
Data from SW480 intact cell experiments. (a) DMSO; (b) (S)-crizotinib. Asterisk indicates unspecific band.
Supplementary Figure 4 Melting-temperature profiles obtained for MTH1 (NUDT1) from SW480 cell lysate experiments.
Similar to intact cell experiments, treatment of SW480 cell extracts with (S)‑crizotinib leads to a stabilization of MTH1 (NUDT1). Data indicate one technical replicate.
Supplementary Figure 5 Kinases with differential melting profiles observed in cell lysates treated with 100 µM (S)-crizotinib.
Treatment of SW480 cell lysates with 100 µM (S)‑crizotinib leads to a stabilization of (a) PI3K-beta (PIK3CB) and (b) CDK2. Data indicate one technical replicate.
Supplementary Figure 6 Original full-length anti-DHFR western blots used for Figure 2d.
Data from K562 intact cell experiments. (a) DMSO; (b) methotrexate.
Supplementary Figure 7 Original full-length anti-STING western blots used for Figure 2f.
Data from RAW intact cell experiments. (a) Water; (b) 2ʹ3ʹ-cGAMP.
Supplementary Figure 8 Definition of the area score.
The area score considers a signed area between the vehicle-treated sample curve and the compound-treated curve.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8, Supplementary Tables 1–5 and Supplementary Note 1 (PDF 2462 kb)
Source data
Rights and permissions
About this article
Cite this article
Huber, K., Olek, K., Müller, A. et al. Proteome-wide drug and metabolite interaction mapping by thermal-stability profiling. Nat Methods 12, 1055–1057 (2015). https://doi.org/10.1038/nmeth.3590
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.3590
This article is cited by
-
Learning the metabolic language of cancer
Nature (2023)
-
The emerging role of mass spectrometry-based proteomics in drug discovery
Nature Reviews Drug Discovery (2022)
-
Host and microbiota metabolic signals in aging and longevity
Nature Chemical Biology (2021)
-
Callyspongiolide kills cells by inducing mitochondrial dysfunction via cellular iron depletion
Communications Biology (2021)
-
The analog of cGAMP, c-di-AMP, activates STING mediated cell death pathway in estrogen-receptor negative breast cancer cells
Apoptosis (2021)