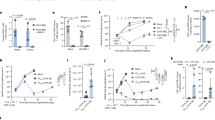
Abstract
Cell-mediated immunity critically depends on the localization of lymphocytes at sites of infection. While some memory T cells recirculate, a distinct lineage (resident memory T cells (TRM cells)) are embedded in nonlymphoid tissues (NLTs) and mediate potent protective immunity. However, the defining transcriptional basis for the establishment of TRM cells is unknown. We found that CD8+ TRM cells lacked expression of the transcription factor KLF2 and its target gene S1pr1 (which encodes S1P1, a receptor for sphingosine 1-phosphate). Forced expression of S1P1 prevented the establishment of TRM cells. Cytokines that induced a TRM cell phenotype (including transforming growth factor-β (TGF-β), interleukin 33 (IL-33) and tumor-necrosis factor) elicited downregulation of KLF2 expression in a pathway dependent on phosphatidylinositol-3-OH kinase (PI(3)K) and the kinase Akt, which suggested environmental regulation. Hence, regulation of KLF2 and S1P1 provides a switch that dictates whether CD8+ T cells commit to recirculating or tissue-resident memory populations.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Mackay, C.R., Marston, W.L. & Dudler, L. Naive and memory T cells show distinct pathways of lymphocyte recirculation. J. Exp. Med. 171, 801–817 (1990).
Klonowski, K.D. et al. Dynamics of blood-borne CD8 memory T cell migration in vivo. Immunity 20, 551–562 (2004).
Masopust, D. et al. Dynamic T cell migration program provides resident memory within intestinal epithelium. J. Exp. Med. 207, 553–564 (2010).
Wakim, L.M., Woodward-Davis, A. & Bevan, M.J. Memory T cells persisting within the brain after local infection show functional adaptations to their tissue of residence. Proc. Natl. Acad. Sci. USA 107, 17872–17879 (2010).
Hofmann, M. & Pircher, H. E-cadherin promotes accumulation of a unique memory CD8 T-cell population in murine salivary glands. Proc. Natl. Acad. Sci. USA 108, 16741–16746 (2011).
Gebhardt, T. et al. Memory T cells in nonlymphoid tissue that provide enhanced local immunity during infection with herpes simplex virus. Nat. Immunol. 10, 524–530 (2009).
Jiang, X. et al. Skin infection generates non-migratory memory CD8+ TRM cells providing global skin immunity. Nature 483, 227–231 (2012).
Mackay, L.K. et al. Long-lived epithelial immunity by tissue-resident memory T (TRM) cells in the absence of persisting local antigen presentation. Proc. Natl. Acad. Sci. USA 109, 7037–7042 (2012).
Schenkel, J.M., Fraser, K.A., Vezys, V. & Masopust, D. Sensing and alarm function of resident memory CD8+ T cells. Nat. Immunol. 14, 509–513 (2013).
Liu, L. et al. Epidermal injury and infection during poxvirus immunization is crucial for the generation of highly protective T cell–mediated immunity. Nat. Med. 16, 224–227 (2010).
Bevan, M.J. Memory T cells as an occupying force. Eur. J. Immunol. 41, 1192–1195 (2011).
Sheridan, B.S. & Lefrancois, L. Regional and mucosal memory T cells. Nat. Immunol. 12, 485–491 (2011).
Ariotti, S., Haanen, J.B. & Schumacher, T.N. Behavior and function of tissue-resident memory T cells. Adv. Immunol. 114, 203–216 (2012).
Masopust, D. & Picker, L.J. Hidden memories: frontline memory T cells and early pathogen interception. J. Immunol. 188, 5811–5817 (2012).
Shin, H. & Iwasaki, A. A vaccine strategy that protects against genital herpes by establishing local memory T cells. Nature 491, 463–467 (2012).
Casey, K.A. et al. Antigen-independent differentiation and maintenance of effector-like resident memory T cells in tissues. J. Immunol. 188, 4866–4875 (2012).
Carlson, C.M. et al. Kruppel-like factor 2 regulates thymocyte and T-cell migration. Nature 442, 299–302 (2006).
Bai, A., Hu, H., Yeung, M. & Chen, J. Kruppel-like factor 2 controls T cell trafficking by activating L-selectin (CD62L) and sphingosine-1-phosphate receptor 1 transcription. J. Immunol. 178, 7632–7639 (2007).
Matloubian, M. et al. Lymphocyte egress from thymus and peripheral lymphoid organs is dependent on S1P receptor 1. Nature 427, 355–360 (2004).
Grayson, J.M., Murali-Krishna, K., Altman, J.D. & Ahmed, R. Gene expression in antigen-specific CD8+ T cells during viral infection. J. Immunol. 166, 795–799 (2001).
Schober, S.L. et al. Expression of the transcription factor lung Kruppel-like factor is regulated by cytokines and correlates with survival of memory T cells in vitro and in vivo. J. Immunol. 163, 3662–3667 (1999).
Cyster, J.G. & Schwab, S.R. Sphingosine-1-phosphate and lymphocyte egress from lymphoid organs. Annu. Rev. Immunol. 30, 69–94 (2012).
Weinreich, M.A. et al. KLF2 transcription-factor deficiency in T cells results in unrestrained cytokine production and upregulation of bystander chemokine receptors. Immunity 31, 122–130 (2009).
Odumade, O.A., Weinreich, M.A., Jameson, S.C. & Hogquist, K.A. Kruppel-like factor 2 regulates trafficking and homeostasis of gammadelta T cells. J. Immunol. 184, 6060–6066 (2010).
Anderson, K.G. et al. Cutting edge: intravascular staining redefines lung CD8 T cell responses. J. Immunol. 189, 2702–2706 (2012).
Masopust, D., Vezys, V., Wherry, E.J., Barber, D.L. & Ahmed, R. Cutting edge: gut microenvironment promotes differentiation of a unique memory CD8 T cell population. J. Immunol. 176, 2079–2083 (2006).
Kuo, C.T., Veselits, M.L. & Leiden, J.M. LKLF: a transcriptional regulator of single-positive T cell quiescence and survival. Science 277, 1986–1990 (1997).
Shiow, L.R. et al. CD69 acts downstream of interferon-α/β to inhibit S1P1 and lymphocyte egress from lymphoid organs. Nature 440, 540–544 (2006).
Bankovich, A.J., Shiow, L.R. & Cyster, J.G. CD69 suppresses sphingosine 1-phosophate receptor-1 (S1P1) function through interaction with membrane helix 4. J. Biol. Chem. 285, 22328–22337 (2010).
Masopust, D., Vezys, V., Marzo, A.L. & Lefrancois, L. Preferential localization of effector memory cells in nonlymphoid tissue. Science 291, 2413–2417 (2001).
Sallusto, F., Geginat, J. & Lanzavecchia, A. Central memory and effector memory T cell subsets: function, generation, and maintenance. Annu. Rev. Immunol. 22, 745–763 (2004).
Lo, C.G., Xu, Y., Proia, R.L. & Cyster, J.G. Cyclical modulation of sphingosine-1-phosphate receptor 1 surface expression during lymphocyte recirculation and relationship to lymphoid organ transit. J. Exp. Med. 201, 291–301 (2005).
Gräler, M.H., Huang, M.C., Watson, S. & Goetzl, E.J. Immunological effects of transgenic constitutive expression of the type 1 sphingosine 1-phosphate receptor by mouse lymphocytes. J. Immunol. 174, 1997–2003 (2005).
Ledgerwood, L.G. et al. The sphingosine 1-phosphate receptor 1 causes tissue retention by inhibiting the entry of peripheral tissue T lymphocytes into afferent lymphatics. Nat. Immunol. 9, 42–53 (2008).
Hart, G.T., Hogquist, K.A. & Jameson, S.C. Kruppel-like factors in lymphocyte biology. J. Immunol. 188, 521–526 (2012).
Moran, A.E. et al. T cell receptor signal strength in Treg and iNKT cell development demonstrated by a novel fluorescent reporter mouse. J. Exp. Med. 208, 1279–1289 (2011).
Sinclair, L.V. et al. Phosphatidylinositol-3-OH kinase and nutrient-sensing mTOR pathways control T lymphocyte trafficking. Nat. Immunol. 9, 513–521 (2008).
Schluns, K.S. & Lefrancois, L. Cytokine control of memory T-cell development and survival. Nat. Rev. Immunol. 3, 269–279 (2003).
Takada, K. et al. Kruppel-like factor 2 is required for trafficking but not quiescence in postactivated T cells. J. Immunol. 186, 775–783 (2011).
Freeman, B.E., Hammarlund, E., Raue, H.P. & Slifka, M.K. Regulation of innate CD8+ T-cell activation mediated by cytokines. Proc. Natl. Acad. Sci. USA 109, 9971–9976 (2012).
Hedrick, S.M. The cunning little vixen: Foxo and the cycle of life and death. Nat. Immunol. 10, 1057–1063 (2009).
Zhang, Y.E. Non-Smad pathways in TGF-β signaling. Cell Res. 19, 128–139 (2009).
Plas, D.R. & Thompson, C.B. Akt activation promotes degradation of tuberin and FOXO3a via the proteasome. J. Biol. Chem. 278, 12361–12366 (2003).
Mackay, L.K. et al. The development pathway for CD103+CD8+ tissue-resident memory T cells of skin. Nat. Immunol. 10.1038/ni.2744 (27 October 2013).
Zhu, J. et al. Immune surveillance by CD8αα+ skin-resident T cells in human herpes virus infection. Nature 497, 494–497 (2013).
Debes, G.F. et al. Chemokine receptor CCR7 required for T lymphocyte exit from peripheral tissues. Nat. Immunol. 6, 889–894 (2005).
Bromley, S.K., Thomas, S.Y. & Luster, A.D. Chemokine receptor CCR7 guides T cell exit from peripheral tissues and entry into afferent lymphatics. Nat. Immunol. 6, 895–901 (2005).
Finlay, D. & Cantrell, D.A. Metabolism, migration and memory in cytotoxic T cells. Nat. Rev. Immunol. 11, 109–117 (2011).
Ohkura, N., Kitagawa, Y. & Sakaguchi, S. Development and maintenance of regulatory T cells. Immunity 38, 414–423 (2013).
Zhu, J. & Paul, W.E. Peripheral CD4+ T-cell differentiation regulated by networks of cytokines and transcription factors. Immunol. Rev. 238, 247–262 (2010).
Rutishauser, R.L. & Kaech, S.M. Generating diversity: transcriptional regulation of effector and memory CD8 T-cell differentiation. Immunol. Rev. 235, 219–233 (2010).
Araki, K. et al. mTOR regulates memory CD8 T-cell differentiation. Nature 460, 108–112 (2009).
Valcu, M. & Valcu, C.M. Data transformation practices in biomedical sciences. Nat. Methods 8, 104–105 (2011).
Acknowledgements
We thank J. Cyster (University of California, San Francisco) for the vector MSCV-S1PR1-hCD4; J. Chen (Massachusetts Institute of Technology) for the retroviral vector MSCV-IRES-Thy1.1 (MiT); K. Walkowiak and J. Schenkel for input on parabiosis; L. Mackay and D. Kaplan for advice on DNFB studies; and the Jamequist laboratory for intellectual support. Supported by the US National Institutes of Health (R37 AI38903 to S.C.J.; R37 AI39560 to K.A.H.; T32 AI07313 to C.N.S.; and T90 DE022732 to K.G.A.).
Author information
Authors and Affiliations
Contributions
C.N.S., J.-Y.L., D.M., K.A.H. and S.C.J. designed the experiments; C.N.S., J.-Y.L. and K.G.A. did experiments; and C.N.S., J.-Y.L. and S.C.J. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 CD69 expression and KLF2 downregulation correlate with parenchymal P14 T cells in NLTs.
Analysis of KLF2-GFP expression in adoptively transferred P14 cells 8 days (a) or 30 days post LCMV infection (b). (A) IV anti-CD8 antibody was used to distinguish P14 cells in tissue parenchyma (black) versus vascular-associated (red). Data are overlaid with wildtype P14 cells (grey filled) as control. Representative of n=9 from 3 independent experiments. (b) Gated on live congenically marked P14 cells isolated from different tissues. Showing CD8 IV versus CD69 (left) or KLF2-GFP (right). Horizontal dotted line represents the separation between red pulp and white pulp of the spleen, based on previous studies (25). Representative of 4 independent experiments with 3 mice each. Abbreviations as in Fig. 1.
Supplementary Figure 2 Kinetics of KLF2 downregulation in lymphoid and nonlymphoid tissues.
Wildtype (grey filled) and KLF2-GFP P14 cells (black or red) were adoptively co-transferred and host mice infected with LCMV. (a) KLF2-GFP expression on live, non-vascular-associated P14 cells at the indicated days following LCMV infection, in lymphoid tissues (black) and NLT (red) (representative of n=9 from 3 independent experiments). Cells in blood exhibit slightly lower KLF2-GFP levels relative to spleen/LN, especially at memory timepoints, for unclear reasons. (b) GFP MFI from KLF2-GFP P14 cells isolated from various tissues 30 days post LCMV infection. GFP MFI from Wildtype P14 cells was subtracted from KLF2-GFP MFI to eliminate background variability from tissue to tissue (this graph was compiled from 2 independent, representative experiments; n=6).
Supplementary Figure 3 Schematic for parabiosis experiments.
Congenically distinct KLF2-GFP P14 cells were transferred into normal C57BL/6 mice, which were infected with LCMV the following day. A parallel set of mice were infected but did not receive P14 adoptive transfer. At 30-65 days post-infection, paris of transferred and non-transferred mice underwent parabiotic surgery, and 13-17 days after surgery, paried animals were sacrificed and tissues harvested. The mice originally receiving the KLF2-GFP P14 cell population is termed the “Donor” while the other animal in a parabiotic pair is termed the “Parabiont”.
Supplementary Figure 4 Retroviral transduction system used for forced expression of S1PR1 and KLF2.
(a) Retroviral constructs used for transduction contained S1PR1 cDNA in the first expression casette, or lacked a gene at this site (Empty vector, “EV”). A similar construct was used for forced KLF2 expression. In all cases, Thy-1.1 was used as a transduction marker. (b-d) Characterization of transduction system using S1PR1 vector. (b) Mock, EV or S1PR1-transduced P14 cells incubated in vitro for 2 days with 10 ng/ml hIL-2 and 250 nM gp33 peptide. Cells were stained for Thy-1.1 (the transduction marker) and for CD69 or the Flag-epitope (which was cloned into the N-terminus of the retroviral S1PR1). Expression of the retroviral S1PR1 is indicated by cell surface Flag-epitope staining, and loss of staining for CD69 (which competes with S1PR1 for surface expression). Gated on live CD8+ cells. (c) Mock (grey), S1PR1 (red), and EV (black) transduced P14 cells were cultured in vitro for additional days with 20ng/ml hIL-2. Data show the fold-change in live cell numbers over 2 day increments. Data were compiled from at least 3 independent experiments (n=5-8). (d) Transduction efficiency of live S1PR1 (top) and EV (bottom) transduced P14 cells cultured in vitro as in (b). Each line is from an independent experiment (n=7). Note that proliferation is not impaired in the S1PR1 transduced population (relative to mock or EV transduced) (c), and that there is no selective disadvantage of S1PR1 transduced cells for expansion (d).
Supplementary Figure 5 Gating strategy and number of bulk and transduced P14 cells for transduction model in vivo.
(a) Gating strategy for calculating percent transduction per tissue and equation for calculating normalized transduction relative to the spleen. Adoptively co-transferred P14 cells were identified using congenic markers, non-vascular-associated cells were detected using CD8b IV administration, and percent transduction was calculated using the Thy1.1 marker. (b) Number of total non-vascular-associated P14 T cells from spleen and LN that underwent S1PR1 (red) or EV (black) transduction prior to adoptive transfer. The date ranges are the times of sacrifice following in vivo LCMV infection. (c) Number of transduced, non-vascular-associated P14 T cells for EV (black) or S1PR1 (red) vectors in indicated tissues within the time ranges following LCMV infection in vivo. (b-c) Data are compiled from 4 independent experiments (n=9-15).
Supplementary Figure 6 Compared to empty vector–transduced P14 cells, there is no skewing toward CD103 expression on the few P14 cells -tranduced to express S1P1 that remain in nonlymphoid tissues.
(a) Shows percent transduction (relative to spleen) of EV (black) and S1PR1 (red) vector-transduced P14 cells in the parenchyma of IEL 28-60 days post LCMV. The frequency of S1PR1 transduced cells was more variable in the IEL versus other non-lymphoid tissue (see figure 4b,4d). N=15-18 from at least 5 independent experiments. (b) The percentage of CD103+ P14 cells transduced by EV (black) or S1PR1 (red) vectors in the indicated tissue parenchyma isolated at 5, 8 and 30 days post LCMV infection (n=9 from 3 independent experiments). All analyses used gating on live, non-vascular-associated CD8+ P14 T cells.
Supplementary Figure 7 TGF-β and IL-33 induce loss of KLF2 expression in a PI(3)K-Akt-dependent pathway.
(a-c, e) Wildtype and KLF2-GFP P14 cells were co-transferred and recipients infected with LCMV for 4.5 days before sacrifice and splenocyte preparation. (a) Splenocytes were cultured with TGF-β and IL-33 (grey) or no additional cytokines (black) (set as one) for 10-40 hours (n=11-13 from at least 4 independent experiments). (b) Splenocytes were cultured with the indicated cytokines for 40 hrs ex vivo (as in Fig. 7)(n=8 from 3 independent experiments). The graph shows mean (+/- SEM) of KLF2-GFP expression in P14 cells, normalized to P14 cells cultured with no added cytokines. (c) Splenocytes were exposed to TGF-β and IL-33 (left) or no additional cytokines (right) in the presence of indicated LY294002 concentrations for 40 hrs ex vivo). Graph shows KLF2-GFP mean fluorescent intensity of P14 cells (representative of 4 independent experiments (n=8)). (d) Wildtype and KLF2-GFP P14 CD8+ T cells were activated in vitro for 48 hours and then cultured with cytokines and inhibitors as in Fig. 7d. Data compiled from 4 independent experiments (e) Percentage of CD103+ P14 cells from splenocytes cultured in indicated cytokines and inhibitors (similar to Fig. 7c). Data are compiled from 4 experiments (n=12).
Supplementary Figure 8 In vivo administration of the PI(3)K inhibitor LY294002 leads to upregulation of KLF2 in the salivary gland and LPL; Foxo1 expression is reduced in memory P14 CD8+ T cells from NLTs versus lymphoid sites.
(a) Wildtype and KLF2-GFP P14 CD8+ T-cells were co-transferred into C57BL/6 animals which were then infected with LCMV. Four days post infection, animals were treated with LY294002 or vehicle only, as in Fig. 7f. N=12 from 5 independent experiments. (a) KLF2-GFP gMFI (minus wildtype gMFI) was calculated and analyzed (with vehicle only group set as 1). (b) P14 cells were isolated from spleen and salivary gland 30-60 days post LCMV and stained for intracellular Foxo1 protein (n=9 from 3 independent experiments). Data are normalized on isotype control and gated on live P14 CD8+ T cells. Statistical analysis for (a) used ANOVA, while (b) used Student's t-test.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 and Supplementary Tables 1–2 (PDF 1382 kb)
Rights and permissions
About this article
Cite this article
Skon, C., Lee, JY., Anderson, K. et al. Transcriptional downregulation of S1pr1 is required for the establishment of resident memory CD8+ T cells. Nat Immunol 14, 1285–1293 (2013). https://doi.org/10.1038/ni.2745
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.2745
This article is cited by
-
Getting under the skin: resident memory CD8+ T cells have a second residence in the draining lymph node
Genes & Immunity (2024)
-
Prognostic values of tissue-resident CD8+T cells in human hepatocellular carcinoma and intrahepatic cholangiocarcinoma
World Journal of Surgical Oncology (2023)
-
Prmt5 deficiency inhibits CD4+ T-cell Klf2/S1pr1 expression and ameliorates EAE disease
Journal of Neuroinflammation (2023)
-
T cell egress via lymphatic vessels is tuned by antigen encounter and limits tumor control
Nature Immunology (2023)
-
Assessing the generation of tissue resident memory T cells by vaccines
Nature Reviews Immunology (2023)