Abstract
In vertebrates, the first haematopoietic stem cells (HSCs) with multi-lineage and long-term repopulating potential arise in the AGM (aorta–gonad–mesonephros) region. These HSCs are generated from a rare and transient subset of endothelial cells, called haemogenic endothelium (HE), through an endothelial-to-haematopoietic transition (EHT). Here, we establish the absolute requirement of the transcriptional repressors GFI1 and GFI1B (growth factor independence 1 and 1B) in this unique trans-differentiation process. We first demonstrate that Gfi1 expression specifically defines the rare population of HE that generates emerging HSCs. We further establish that in the absence of GFI1 proteins, HSCs and haematopoietic progenitor cells are not produced in the AGM, revealing the critical requirement for GFI1 proteins in intra-embryonic EHT. Finally, we demonstrate that GFI1 proteins recruit the chromatin-modifying protein LSD1, a member of the CoREST repressive complex, to epigenetically silence the endothelial program in HE and allow the emergence of blood cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Costa, G., Kouskoff, V. & Lacaud, G. Origin of blood cells and HSC production in the embryo. Trends Immunol. 33, 215–223 (2012).
Palis, J., Robertson, S., Kennedy, M., Wall, C. & Keller, G. M. Development of erythroid and myeloid progenitors in the yolk sac and embryo proper of the mouse. Development 126, 5073–5084 (1999).
Frame, J. M., McGrath, K. E. & Palis, J. Erythro-myeloid progenitors: ‘definitive’ hematopoiesis in the conceptus prior to the emergence of hematopoietic stem cells. Blood Cells Mol. Dis. 51, 220–225 (2013).
McGrath, K. E. et al. Distinct sources of hematopoietic progenitors emerge before HSCs and provide functional blood cells in the mammalian embryo. Cell Rep. 11, 1892–1904 (2015).
Frame, J. M., Fegan, K. H., Conway, S. J., McGrath, K. E. & Palis, J. Definitive hematopoiesis in the yolk sac emerges from Wnt-responsive hemogenic endothelium independently of circulation and arterial identity. Stem Cells (2015)10.1002/stem.2213.
Medvinsky, A. J. & Dzierzak, E. A. Definitive hematopoiesis is autonomously initiated by the AGM region. Cell 86, 897–906 (1996).
Müller, A. M., Medvinsky, A. J., Strouboulis, J., Grosveld, F. & Dzierzak, E. A. Development of hematopoietic stem cell activity in the mouse embryo. Immunity 1, 291–301 (1994).
Jaffredo, T., Gautier, R., Eichmann, A. & Dieterlen-Lièvre, F. Intraaortic hemopoietic cells are derived from endothelial cells during ontogeny. Development 125, 4575–4583 (1998).
Zovein, A. C. et al. Fate tracing reveals the endothelial origin of hematopoietic stem cells. Cell Stem Cell 3, 625–636 (2008).
Eilken, H. M., Nishikawa, S., Nishikawa, S. & Schroeder, T. Continuous single-cell imaging of blood generation from haemogenic endothelium. Nature 457, 896–900 (2009).
Lancrin, C. et al. The haemangioblast generates haematopoietic cells through a haemogenic endothelium stage. Nature 457, 892–895 (2009).
Nishikawa, S. et al. In vitro generation of lymphohematopoietic cells from endothelial cells purified from murine embryos. Immunity 8, 761–769 (1998).
Boisset, J.-C. et al. In vivo imaging of haematopoietic cells emerging from the mouse aortic endothelium. Nature 464, 116–120 (2010).
Bertrand, J. Y. et al. Haematopoietic stem cells derive directly from aortic endothelium during development. Nature 464, 108–111 (2010).
Kissa, K. & Herbomel, P. Blood stem cells emerge from aortic endothelium by a novel type of cell transition. Nature 464, 112–115 (2010).
Lam, E. Y. N., Hall, C. J., Crosier, P. S., Crosier, K. E. & Flores, M. V. Live imaging of Runx1 expression in the dorsal aorta tracks the emergence of blood progenitors from endothelial cells. Blood 116, 909–914 (2010).
de Bruijn, M. F. T. R., Speck, N. A., Peeters, M. C. & Dzierzak, E. A. Definitive hematopoietic stem cells first develop within the major arterial regions of the mouse embryo. EMBO J. 19, 2465–2474 (2000).
Taoudi, S. & Medvinsky, A. J. Functional identification of the hematopoietic stem cell niche in the ventral domain of the embryonic dorsal aorta. Proc. Natl Acad. Sci. USA 104, 9399–9403 (2007).
Boisset, J.-C. et al. Progressive maturation toward hematopoietic stem cells in the mouse embryo aorta. Blood 125, 465–469 (2015).
North, T. et al. Cbfa2 is required for the formation of intra-aortic hematopoietic clusters. Development 126, 2563–2575 (1999).
Sroczynska, P., Lancrin, C., Kouskoff, V. & Lacaud, G. The differential activities of Runx1 promoters define milestones during embryonic hematopoiesis. Blood 114, 5279–5289 (2009).
Lacaud, G. et al. Runx1 is essential for hematopoietic commitment at the hemangioblast stage of development in vitro. Blood 100, 458–466 (2002).
North, T. E. et al. Runx1 expression marks long-term repopulating hematopoietic stem cells in the midgestation mouse embryo. Immunity 16, 661–672 (2002).
North, T. E., Stacy, T., Matheny, C. J., Speck, N. A. & de Bruijn, M. F. T. R. Runx1 is expressed in adult mouse hematopoietic stem cells and differentiating myeloid and lymphoid cells, but not in maturing erythroid cells. Stem Cells 22, 158–168 (2004).
Okuda, T., Hiebert, S. W. & Downing, J. R. AML1, the target of multiple chromosomal translocations in human leukemia, is essential for normal fetal liver hematopoiesis. Cell 84, 321–330 (1996).
Wang, Q. et al. Disruption of the Cbfa2 gene causes necrosis and hemorrhaging in the central nervous system and blocks definitive hematopoiesis. Proc. Natl Acad. Sci. USA 93, 3444–3449 (1996).
Chen, M. J., Yokomizo, T., Zeigler, B. M., Dzierzak, E. A. & Speck, N. A. Runx1 is required for the endothelial to haematopoietic cell transition but not thereafter. Nature 457, 887–891 (2009).
Lancrin, C. et al. GFI1 and GFI1B control the loss of endothelial identity of hemogenic endothelium during hematopoietic commitment. Blood 120, 314–322 (2012).
Vassen, L., Okayama, T. & Möröy, T. Gfi1b: green fluorescent protein knock-in mice reveal a dynamic expression pattern of Gfi1b during hematopoiesis that is largely complementary to Gfi1. Blood 109, 2356–2364 (2007).
Rybtsov, S. et al. Hierarchical organization and early hematopoietic specification of the developing HSC lineage in the AGM region. J. Exp. Med. 208, 1305–1315 (2011).
Taoudi, S. et al. Extensive hematopoietic stem cell generation in the AGM region via maturation of VE-cadherin + CD45+ pre-definitive HSCs. Cell Stem Cell 3, 99–108 (2008).
Yokomizo, T. & Dzierzak, E. A. Three-dimensional cartography of hematopoietic clusters in the vasculature of whole mouse embryos. Development 137, 3651–3661 (2010).
Fiolka, K. et al. Gfi1 and Gfi1b act equivalently in haematopoiesis, but have distinct, non-overlapping functions in inner ear development. EMBO Rep. 7, 326–333 (2006).
Saleque, S., Orkin, S., Kim, J. & Rooke, H. M. Epigenetic regulation of hematopoietic differentiation by Gfi-1 and Gfi-1b is mediated by the cofactors CoREST and LSD1. Mol. Cell 27, 562–572 (2007).
Harris, W. J. et al. The histone demethylase KDM1A sustains the oncogenic potential of MLL-AF9 leukemia stem cells. Cancer Cell 21, 473–487 (2012).
Lancrin, C. et al. Blood cell generation from the hemangioblast. J. Mol. Med. 88, 167–172 (2010).
Foster, C. T. et al. Lysine-specific demethylase 1 regulates the embryonic transcriptome and CoREST stability. Mol. Cell. Biol. 30, 4851–4863 (2010).
Wang, J. et al. The lysine demethylase LSD1 (KDM1) is required for maintenance of global DNA methylation. Nat. Genet. 41, 125–129 (2009).
Whyte, W. A. et al. Enhancer decommissioning by LSD1 during embryonic stem cell differentiation. Nature 482, 221–225 (2012).
Vogel, M. J., Peric-Hupkes, D. & van Steensel, B. Detection of in vivo protein-DNA interactions using DamID in mammalian cells. Nat. Protoc. 2, 1467–1478 (2007).
Lie-A-Ling, M. et al. RUNX1 positively regulates a cell adhesion and migration program in murine hemogenic endothelium prior to blood emergence. Blood 124, e11–e20 (2014).
Lichtinger, M. et al. RUNX1 reshapes the epigenetic landscape at the onset of haematopoiesis. EMBO J. 31, 4318–4333 (2012).
Wang, L. & Dudek, S. M. Regulation of vascular permeability by sphingosine 1-phosphate. Microvasc. Res. 77, 39–45 (2009).
Chen, M. J. et al. Erythroid/myeloid progenitors and hematopoietic stem cells originate from distinct populations of endothelial cells. Stem Cell 9, 541–552 (2011).
Hadland, B. K. et al. A requirement for Notch1 distinguishes 2 phases of definitive hematopoiesis during development. Blood 104, 3097–3105 (2004).
Kumano, K. et al. Notch1 but not Notch2 is essential for generating hematopoietic stem cells from endothelial cells. Immunity 18, 699–711 (2003).
Bigas, A. & Robert-Moreno, L. The Notch pathway in the developing hematopoietic system. Int. J. Dev. Biol. 54, 1175–1188 (2010).
Guiu, J. et al. Hes repressors are essential regulators of hematopoietic stem cell development downstream of Notch signaling. J. Exp. Med. 210, 71–84 (2013).
Costa, G. et al. SOX7 regulates the expression of VE-cadherin in the haemogenic endothelium at the onset of haematopoietic development. Development 139, 1587–1598 (2012).
Clarke, R. L. et al. The expression of Sox17 identifies and regulates haemogenic endothelium. Nat. Cell Biol. 15, 1–10 (2013).
de Bruijn, M. F. T. R., Robin, C., Ottersbach, K., Sanchez, M.-J. & Dzierzak, E. A. Hematopoietic stem cells localize to the endothelial cell layer in the midgestation mouse aorta. Immunity 16, 673–683 (2002).
Swiers, G. et al. Early dynamic fate changes in haemogenic endothelium characterized at the single-cell level. Nat. Commun. 4, 2924 (2013).
Kim, W., Klarmann, K. D. & Keller, J. R. Gfi-1 regulates the erythroid transcription factor network through Id2 repression in murine hematopoietic progenitor cells. Blood 124, 1586–1596 (2014).
Möröy, T. & Khandanpour, C. Growth factor independence 1 (Gfi1) as a regulator of lymphocyte development and activation. Semin. Immunol. 23, 368–378 (2011).
Pearson, S., Cuvertino, S., Fleury, M., Lacaud, G. & Kouskoff, V. In vivo repopulating activity emerges at the onset of hematopoietic specification during embryonic stem cell differentiation. Stem Cell Rep. 4, 431–444 (2015).
Ledran, M. H. et al. Efficient hematopoietic differentiation of human embryonic stem cells on stromal cells derived from hematopoietic niches. Cell Stem Cell 3, 85–98 (2008).
Wang, L. et al. Generation of hematopoietic repopulating cells from human embryonic stem cells independent of ectopic HOXB4 expression. J. Exp. Med. 201, 1603–1614 (2005).
Kyba, M., Perlingeiro, R. C. R. & Daley, G. Q. HoxB4 confers definitive lymphoid-myeloid engraftment potential on embryonic stem cell and yolk sac hematopoietic progenitors. Cell 109, 29–37 (2002).
Pereira, C. F. et al. Induction of a hemogenic program in mouse fibroblasts. Cell Stem Cell 13, 205–218 (2013).
Sandler, V. M. et al. Reprogramming human endothelial cells to haematopoietic cells requires vascular induction. Nature 511, 312–318 (2014).
Batta, K., Florkowska, M., Kouskoff, V. & Lacaud, G. Direct reprogramming of murine fibroblasts to hematopoietic progenitor cells. Cell Rep. 9, 1871–1884 (2014).
Wilson, N. K. et al. Gfi1 expression is controlled by five distinct regulatory regions spread over 100 kilobases, with Scl/Tal1, Gata2, PU.1, Erg, Meis1, and Runx1 acting as upstream regulators in early hematopoietic cells. Mol. Cell. Biol. 30, 3853–3863 (2010).
Stefanska, M., Costa, G., Lie-A-Ling, M., Kouskoff, V. & Lacaud, G. Smooth muscle cells largely develop independently of functional hemogenic endothelium. Stem Cell Res. 12, 222–232 (2014).
Yücel, R., Kosan, C., Heyd, F. & Möröy, T. Gfi1:green fluorescent protein knock-in mutant reveals differential expression and autoregulation of the growth factor independence 1 (Gfi1) gene during lymphocyte development. J. Biol. Chem. 279, 40906–40917 (2004).
Kumaravelu, P. et al. Quantitative developmental anatomy of definitive haematopoietic stem cells/long-term repopulating units (HSC/RUs): role of the aorta-gonad-mesonephros (AGM) region and the yolk sac in colonisation of the mouse embryonic liver. Development 129, 4891–4899 (2002).
Nichols, J. et al. Validated germline-competent embryonic stem cell lines from nonobese diabetic mice. Nat. Med. 15, 814–818 (2009).
Fehling, H. J. et al. Tracking mesoderm induction and its specification to the hemangioblast during embryonic stem cell differentiation. Development 130, 4217–4227 (2003).
Sroczynska, P., Lancrin, C., Pearson, S., Kouskoff, V. & Lacaud, G. In vitro differentiation of mouse embryonic stem cells as a model of early hematopoietic development. Methods Mol. Biol. 538, 317–334 (2009).
Moignard, V. et al. Characterization of transcriptional networks in blood stem and progenitor cells using high-throughput single-cell gene expression analysis. Nat. Cell Biol. 15, 363–372 (2013).
Acknowledgements
We thank the staff at the Advanced Imaging, animal facility, Molecular Biology Core facilities and Flow Cytometry of CRUK Manchester Institute for technical support and M. Lie-A-Ling for help with initiating the DamID-PIP bioinformatics project. We thank members of the Stem Cell Biology group, the Stem Cell Haematopoiesis groups and M. Gering for valuable advice and critical reading of the manuscript. Work in our laboratory is supported by the Leukaemia and Lymphoma Research Foundation (LLR), Cancer Research UK (CRUK) and the Biotechnology and Biological Sciences Research Council (BBSRC). S.C. is the recipient of an MRC senior fellowship (MR/J009202/1).
Author information
Authors and Affiliations
Contributions
R.T. designed and performed most of the experiments, analysed the data and wrote the manuscript, M.M. initiated the project, designed performed experiments and analysed the data. R.P., V.M., M.S. and C.L. designed and performed experiments. E.M. and Y.L. performed bioinformatics analysis on the sequencing data and microarray. T.C., T.M., C.R., C.M., S.C. and B.G. contributed valuable tools and protocols. V.K. and G.L. designed and supervised the research project, analysed the data, and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Gfi1tomato mouse line.
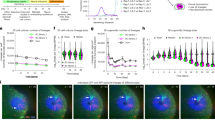
(a) FACS analysis of Gfi1tomato and Gfi1GFP/+ bone marrow cells after staining for lymphoid (CD4/CD8) and myeloid (GR-1/MAC-1) markers. Bone marrow populations were gated into GFP− and GFP+ for Gfi1GFP/+ (top panel) and TOM− and TOM+ for Gfi1tomato (lower panel) fractions (representative FACS plot of one independent experiment). (b) IHC for CD31 (red), GFI1 (cyan) and GFI1B (green) on E10.5 AGM section counterstained with DAPI (representative images of 3 independent experiments). (c) Frequencies of GFI1+/GFI1− and GFI1+/GFI1+ cells in the CDH5 compartment of E10.5 and E11.5 AGMs (AGMs from 11 independent litters, n = 6 independent experiments for E10.5 and n = 4 independent experiments for E11.5 were analysed, p-values were determined by a two-tailed Student’s t-test. Error bars represent standard error of the mean (S.E.M.). The source data used to determine the statistical significance can be found in supplementary table 3. (d) FACS analysis of GFI1 and GFI1B cells in the CDH5 compartment of E11.5 AGMs (representative FACS plot from one independent experiment). Scale bar = 10 μm.
Supplementary Figure 2 GFI1+/GFI1B− cells gain GFI1B expression in vitro.
(a) CDH5+/GFI1+/GFI1B− cells were FACS sorted from Gfi1tomatoGfi1GFP/+ E10.5 AGM and cultured on OP-9 stromal cells. Images at day 2 and 7 of in vitro cultures (representative images of n = 4 independent experiments). (b) Day 7 cultures were analysed by FACS and indicated gates were used for sorting. (c) The initial sorted E10.5 CDH5+/GFI1+/GFI1B− AGM cells and the different cell populations isolated after 7 days of their cultures (GFI1+/GFI1B−, GFI1+/GFI1B+, GFI1−/GFI1B+) were analysed for Gfi1b expression level by q-PCR (representative data from one experiment).
Supplementary Figure 3 Heatmap of all cells and genes analysed by single cell PCR.
Heat map depicting the clustering all the single cells and genes analysed.
Supplementary Figure 4 E11.5 CDH5+/GFI1+ cells contribution to recipients.
(a) GFI1 and GFI1B expression mark all hematopoietic cells in the E10.5 AGM (i) schematic representation of the experimental design (ii and iii) E10.5 AGM cells were FACS sorted and re-plated into haematopoietic assay either before (one independent experiment with embryos from one litter) (ii) or after (one independent experiment with embryos from one litter) (iii) co-culture step with OP-9. Colonies were scored 9-11 days later. (b) FACS analysis of recipients transplanted with either CDH5+/GFI1+ or CDH5+/GFI1− E11.5 AGM cells (CD45.1/CD45.2 double positives) at 17 weeks after transplantation. Bone marrow LSK compartments of recipients (CD45.1) were analysed for CD45.1 and CD45.2 (representative FACS plots of one independent experiment). (c) Donor contribution to haematopoietic lineages determined by sub-gating on CD45.1/CD45.2 double positive population in the recipient. Donor contribution to T cells (CD4/CD8) in the spleen and thymus, B cells (IgM/B220), myeloid cells (GR-1/MAC-1) and erythroid cells (CD71/TER119) in different haematopoietic organs are shown (representative FACS plots of one independent experiment).
Supplementary Figure 5 GFI1 or GFI1B single knock out embryos can generate IAHC.
(a) IHC on E10.5 GFI1KOGFI1B+/+ and GFI1+/+GFI1BKO AGM section for CD31, GFP and c-KIT (counterstained with DAPI) (representative images of one independent experiment with embryos form different litters). (b) In situ hybridisation for Gfi1b (red dots) and IHC for CD31 (brown) on GFI1GFP/+GFI1BGFP/+ and GFI1KOGFI1BGFP/+ E10.5 embryo sections. (c) qPCRs on E10.5 AGM cells. (i) qPCR for Gfi1 and Gfi1b expression in AGM cell lysate of GFI1GFP/+GFI1BGFP/+ (het/het), GFI1GFP/GFPGFI1BGFP/+ (KO/het) and GFI1GFP/+GFI1BGFP/GFP (het/KO) embryos (representative data from one independent experiment with embryos from the same litter). (ii) PCR for Gfi1 and Gfi1b on sorted CDH5+/GFP+/c-KIT− HE cells of GFI1GFP/+GFI1BGFP/+ (het/het) and GFI1GFP/GFPGFI1BGFP/+ (KO/het) embryos (one independent experiment with embryos from the same litter). Scale bar = 10 μm.
Supplementary Figure 6 Conditional Lsd1 knock-out recapitulates LSD1 inhibition phenotype and leads to decrease in proliferation and apoptosis.
(a) FLK1+ cells from the conditional Lsd1Δ/lox line were isolated from day 3 EBs and cultured as monolayer. FACS analysis on 3 consecutive days with staining for TIE-2, CDH5 and CD41 are shown. Lsd1Δ/lox FLK1 + cells were either cultured with control ETOH or with 1 uM of 4OHT (in ETOH) to induce the activity of Cre-ERT2 and generate Lsd1Δ/Δ (representative FACS plots of 3 independent experiments). (b) EdU assay on Day 3 control (Lsd1Δ/lox) or Lsd1 deleted (Lsd1Δ/Δ). Blast cultures (Representative FACS plots of one independent experiment). (c) Annexin5/7-AAD staining on Day 3 control (Lsd1Δ/lox) or Lsd1 deleted (Lsd1Δ/Δ) Li-Blast cultures. Additionally, the cultures were stained with CD41 to differentiate between HE and non-HE cells.
Supplementary Figure 7 LSD1 is ubiquitously expressed and its inhibition leads to loss of EHT in E10.5 AGM.
(a) IHC for CD31, tomato and LSD1 on E10.5 AGM sections of Gfi1tomato embryos (counterstained with DAPI). (b) IHC for CDH5, GFI1 and CD45 on E9.5 P-Sp explants. (c) Single channel control images from live imagining of E10.5 Gfi1tomatoGfi1GFP/+ AGM slices wither treated with DMSO (ctrl) or 500nM of the LSD1 inhibitor. (di–iii) Screen shots of the Oit-3, Gata-2 and Hey-2 locus from the UCSC browser showing examples of GFI1:DAM (Cyan) and GFI1B:DAM (green) binding enrichment after sequencing. Region of interest is marked with a black box and specific peaks detected are highlighted with black bars below. Scale bar = 10 μm.
Supplementary information
Supplementary Information
Supplementary Information (PDF 3014 kb)
Supplementary Table 1
Supplementary Information (XLSX 12 kb)
Supplementary Table 2
Supplementary Information (XLS 85 kb)
Supplementary Table 3
Supplementary Information (XLSX 25 kb)
41556_2016_BFncb3276_MOESM28_ESM.mov
Time-lapse imaging of FLK1+ cells blast monolayer cultures treated with 300 nM of the LSD1 inhibitor (LSD1 inhib). (MOV 4541 kb)
41556_2016_BFncb3276_MOESM29_ESM.mov
3D re-construction of z-stacks (4.2 um) of E9.5 P-Sp explants (2 days cultures) treated with DMSO and stained for CD31 (red), GFI1 (cyan) and GFI1B (green). (MOV 1359 kb)
41556_2016_BFncb3276_MOESM30_ESM.mov
3D re-construction of z-stacks (4.1 um) of E9.5 P-Sp explants (2 days cultures) treated with the LSD1 inhibitor (300 nM) and stained for CD31 (red), GFI1 (cyan) and GFI1B (green). (MOV 2698 kb)
41556_2016_BFncb3276_MOESM31_ESM.mov
Time-lapse imaging of FLK1+ cells from the cLsd1 ES line in Liquid Blast culture treated with EtOH (Lsd1Δ/lox). (MOV 9637 kb)
41556_2016_BFncb3276_MOESM32_ESM.mov
Time-lapse imaging of FLK1+ cells from the cLsd1 ES line in Liquid Blast culture treated with 4-OHT (Lsd1Δ/Δ). (MOV 6479 kb)
Rights and permissions
About this article
Cite this article
Thambyrajah, R., Mazan, M., Patel, R. et al. GFI1 proteins orchestrate the emergence of haematopoietic stem cells through recruitment of LSD1. Nat Cell Biol 18, 21–32 (2016). https://doi.org/10.1038/ncb3276
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3276
This article is cited by
-
Activation of lineage competence in hemogenic endothelium precedes the formation of hematopoietic stem cell heterogeneity
Cell Research (2023)
-
Meis1 establishes the pre-hemogenic endothelial state prior to Runx1 expression
Nature Communications (2023)
-
A genome-wide relay of signalling-responsive enhancers drives hematopoietic specification
Nature Communications (2023)
-
Histone demethylase Lsd1 is required for the differentiation of neural cells in Nematostella vectensis
Nature Communications (2022)
-
GFI1 regulates hair cell differentiation by acting as an off-DNA transcriptional co-activator of ATOH1, and a DNA-binding repressor
Scientific Reports (2022)