Abstract
Transcription factor (TF) networks are thought to regulate embryonic stem cell (ESC) pluripotency. However, TF expression dynamics and regulatory mechanisms are poorly understood. We use reporter mouse ESC lines allowing non-invasive quantification of Nanog or Oct4 protein levels and continuous long-term single-cell tracking and quantification over many generations to reveal diverse TF protein expression dynamics. For cells with low Nanog expression, we identified two distinct colony types: one re-expressed Nanog in a mosaic pattern, and the other did not re-express Nanog over many generations. Although both expressed pluripotency markers, they exhibited differences in their TF protein correlation networks and differentiation propensities. Sister cell analysis revealed that differences in Nanog levels are not necessarily accompanied by differences in the expression of other pluripotency factors. Thus, regulatory interactions of pluripotency TFs are less stringently implemented in individual self-renewing ESCs than assumed at present.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Niwa, H., Miyazaki, J. & Smith, A. G. Quantitative expression of Oct-3/4 defines differentiation, dedifferentiation or self-renewal of ES cells. Nat. Genet. 24, 372–376 (2000).
Loh, Y.-H. et al. The Oct4 and Nanog transcription network regulates pluripotency in mouse embryonic stem cells. Nat. Genet. 38, 431–440 (2006).
Kim, J., Chu, J., Shen, X., Wang, J. & Orkin, S. H. An extended transcriptional network for pluripotency of embryonic stem cells. Cell 132, 1049–1061 (2008).
Chambers, I. & Tomlinson, S. R. The transcriptional foundation of pluripotency. Development 136, 2311–2322 (2009).
Lu, R. et al. Systems-level dynamic analyses of fate change in murine embryonic stem cells. Nature 462, 358–362 (2009).
Silva, J. et al. Nanog is the gateway to the pluripotent ground state. Cell 138, 722–737 (2009).
Som, A. et al. The PluriNetworK: an electronic representation of the network underlying pluripotency in mouse, and its applications. PLoS ONE 5, 1–13 (2010).
Van den Berg, D. L. C. et al. An Oct4-centered protein interaction network in embryonic stem cells. Cell Stem Cell 6, 369–381 (2010).
Young, R. A. Control of the embryonic stem cell state. Cell 144, 940–954 (2011).
Yeo, J.-C. & Ng, H.-H. The transcriptional regulation of pluripotency. Cell Res. 23, 20–32 (2013).
Chen, X. et al. Integration of external signaling pathways with the core transcriptional network in embryonic stem cells. Cell 133, 1106–1117 (2008).
Niwa, H., Burdon, T., Chambers, I. & Smith, A. Self-renewal of pluripotent embryonic stem cells is mediated via activation of STAT3. Genes Dev. 12, 2048–2060 (1998).
Niwa, H., Ogawa, K., Shimosato, D. & Adachi, K. A parallel circuit of LIF signalling pathways maintains pluripotency of mouse ES cells. Nature 460, 118–122 (2009).
Martello, G. et al. Esrrb is a pivotal target of the Gsk3/Tcf3 axis regulating embryonic stem cell self-renewal. Cell Stem Cell 11, 491–504 (2012).
Hall, J. et al. Oct4 and LIF/Stat3 additively induce Kruppel factors to sustain embryonic stem cell self-renewal. Cell Stem Cell 5, 597–609 (2009).
Martello, G., Bertone, P. & Smith, A. Identification of the missing pluripotency mediator downstream of leukaemia inhibitory factor. EMBO J. 32, 2561–2574 (2013).
Ang, Y. S. et al. Wdr5 mediates self-renewal and reprogramming via the embryonic stem cell core transcriptional network. Cell 145, 183–187 (2011).
Ding, J. et al. Tex10 coordinates epigenetic control of super-enhancer activity in pluripotency and reprogramming. Cell Stem Cell 16, 653–668 (2015).
Festuccia, N. et al. Esrrb is a direct Nanog target gene that can substitute for Nanog function in pluripotent cells. Cell Stem Cell 11, 477–490 (2012).
Jiang, J. et al. A core Klf circuitry regulates self-renewal of embryonic stem cells. Nat. Cell Biol. 10, 353–360 (2008).
Ye, S., Li, P., Tong, C. & Ying, Q.-L. Embryonic stem cell self-renewal pathways converge on the transcription factor Tfcp2l1. EMBO J. 32, 2548–2560 (2013).
Ying, Q. L., Nichols, J., Chambers, I. & Smith, A. BMP induction of Id proteins suppresses differentiation and sustains embryonic stem cell self-renewal in collaboration with STAT3. Cell 115, 281–292 (2003).
Ying, Q.-L. et al. The ground state of embryonic stem cell self-renewal. Nature 453, 519–523 (2008).
Berge, D. T. et al. Embryonic stem cells require Wnt proteins to prevent differentiation to epiblast stem cells. Nat. Cell Biol. 13, 1070–1075 (2011).
Krumsiek, J., Marr, C., Schroeder, T. & Theis, F. J. Hierarchical differentiation of myeloid progenitors is encoded in the transcription factor network. PLoS ONE 6, e22649 (2011).
Dunn, S.-J., Martello, G., Yordanov, B., Emmott, S. & Smith, A. G. Defining an essential transcription factor program for naïve pluripotency. Science 344, 1156–1160 (2014).
Xu, H., Ang, Y.-S., Sevilla, A., Lemischka, I. R. & Ma’ayan, A. Construction and validation of a regulatory network for pluripotency and self-renewal of mouse embryonic stem cells. PLoS Comput. Biol. 10, e1003777 (2014).
Enver, T., Pera, M., Peterson, C. & Andrews, P. W. Stem cell states, fates, and the rules of attraction. Cell Stem Cell 4, 387–397 (2009).
Chambers, I. et al. Functional expression cloning of Nanog, a pluripotency sustaining factor in embryonic stem cells. Cell 113, 643–655 (2003).
Mitsui, K. et al. The homeoprotein nanog is required for maintenance of pluripotency in mouse epiblast and ES cells. Cell 113, 631–642 (2003).
Chambers, I. et al. Nanog safeguards pluripotency and mediates germline development. Nature 450, 1230–1234 (2007).
Jaenisch, R. & Young, R. Stem cells, the molecular circuitry of pluripotency and nuclear reprogramming. Cell 132, 567–582 (2008).
Navarro, P. et al. OCT4/SOX2-independent Nanog autorepression modulates heterogeneous Nanog gene expression in mouse ES cells. EMBO J. 31, 4547–4562 (2012).
Fidalgo, M. et al. Zfp281 mediates Nanog autorepression through recruitment of the NuRD complex and inhibits somatic cell reprogramming. Proc. Natl Acad. Sci. USA 109, 16202–16207 (2012).
Morgani, S. M. et al. Totipotent embryonic stem cells arise in ground-state culture conditions. Cell Rep. 3, 1945–1957 (2013).
Hoppe, P. S., Coutu, D. L. & Schroeder, T. Single-cell technologies sharpen up mammalian stem cell research. Nat. Cell Biol. 16, 919–927 (2014).
Etzrodt, M., Endele, M. & Schroeder, T. Quantitative single-cell approaches to stem cell research. Cell Stem Cell 15, 546–558 (2014).
Schroeder, T. Long-term single-cell imaging of mammalian stem cells. Nat. Methods 8, S30–S35 (2011).
Schroeder, T. Imaging stem-cell-driven regeneration in mammals. Nature 453, 345–351 (2008).
Abranches, E. et al. Stochastic NANOG fluctuations allow mouse embryonic stem cells to explore pluripotency. Development 141, 2770–2779 (2014).
Ochiai, H., Sugawara, T. & Sakuma, T. Y. T. Stochastic promoter activation affects Nanog expression variability in mouse embryonic stem cells. Sci Rep. 4, 7125 (2014).
Singer, Z. S. et al. Dynamic heterogeneity and DNA methylation in embryonic stem cells. Mol. Cell 55, 319–331 (2014).
Kumar, R. M. et al. Deconstructing transcriptional heterogeneity in pluripotent stem cells. Nature 516, 55–61 (2014).
Glauche, I., Herberg, M. & Roeder, I. Nanog variability and pluripotency regulation of embryonic stem cells–insights from a mathematical model analysis. PLoS ONE 5, e11238 (2010).
Kalmar, T. et al. Regulated fluctuations in Nanog expression mediate cell fate decisions in embryonic stem cells. PLoS Biol. 7, e1000149 (2009).
Brock, A., Chang, H. & Huang, S. Non-genetic heterogeneity–a mutation-independent driving force for the somatic evolution of tumours. Nat. Rev. Genet. 10, 336–342 (2009).
Herberg, M., Kalkan, T., Glauche, I., Smith, A. & Roeder, I. A model-based analysis of culture-dependent phenotypes of mESCs. PLoS ONE 9, e92496 (2014).
Filipczyk, A. et al. Biallelic expression of nanog protein in mouse embryonic stem cells. Cell Stem Cell 13, 12–13 (2013).
Karwacki-Neisius, V. et al. Reduced Oct4 expression directs a robust pluripotent state with distinct signaling activity and increased enhancer occupancy by Oct4 and Nanog. Cell Stem Cell 12, 531–545 (2013).
Faddah, D. A. et al. Single-cell analysis reveals that expression of nanog is biallelic and equally variable as that of other pluripotency factors in mouse ESCs. Cell Stem Cell 13, 23–29 (2013).
Eilken, H. M., Nishikawa, S.-I. & Schroeder, T. Continuous single-cell imaging of blood generation from haemogenic endothelium. Nature 457, 896–900 (2009).
Rieger, M. A., Hoppe, P. S., Smejkal, B. M., Eitelhuber, A. C. & Schroeder, T. Hematopoietic cytokines can instruct lineage choice. Science 325, 217–218 (2009).
Schwarzfischer, M. et al. Efficient fluorescence image normalization for time lapse movies. In Proc. Microscopic Image Analysis with Applications in Biology (2011).
Garcia-Ojalvo, J. & Martinez Arias, A. Towards a statistical mechanics of cell fate decisions. Curr. Opin. Genet. Dev. 22, 619–626 (2012).
Dietrich, J.-E. & Hiiragi, T. Stochastic patterning in the mouse pre-implantation embryo. Development 134, 4219–4231 (2007).
Chan, K. K. K. et al. KLF4 and PBX1 directly regulate NANOG expression in human embryonic stem cells. Stem Cells 27, 2114–2125 (2009).
Zhang, P., Andrianakos, R., Yang, Y., Liu, C. & Lu, W. Kruppel-like factor 4 (Klf4) prevents embryonic stem (ES) cell differentiation by regulating Nanog gene expression. J. Biol. Chem. 285, 9180–9189 (2010).
Lauritzen, S. L. Graphical Models (Oxford Univ. Press, 1996).
Thomson, M. et al. Pluripotency factors in embryonic stem cells regulate differentiation into germ layers. Cell 145, 875–889 (2011).
Descalzo, S. M., Rué, P., Garcia-Ojalvo, J. & Arias, A. M. Correlations between the levels of Oct4 and Nanog as a signature for naïve pluripotency in mouse embryonic stem cells. Stem Cells 30, 2683–2691 (2012).
Feigelman, J., Theis, F. J. & Marr, C. MCA: multiresolution Correlation Analysis, a graphical tool for subpopulation identification in single-cell gene expression data. BMC Bioinform. 15, 240 (2014).
Burtscher, I. & Lickert, H. Foxa2 regulates polarity and epithelialization in the endoderm germ layer of the mouse embryo. Development 136, 1029–1038 (2009).
Wood, H. B. & Episkopou, V. Comparative expression of the mouse Sox1, Sox2 and Sox3 genes from pre-gastrulation to early somite stages. Mech. Dev. 86, 197–201 (1999).
MacArthur, B. D. et al. Nanog-dependent feedback loops regulate murine embryonic stem cell heterogeneity. Nat. Cell Biol. 14, 1139–1147 (2012).
Macarthur, B. D., Ma’ayan, A. & Lemischka, I. R. Systems biology of stem cell fate and cellular reprogramming. Nat. Rev. Mol. Cell Biol. 10, 672–681 (2009).
Trott, J., Hayashi, K., Surani, A., Babu, M. M. & Martinez-Arias, A. Dissecting ensemble networks in ES cell populations reveals micro-heterogeneity underlying pluripotency. Mol. BioSystems 8, 744 (2012).
Zhang, Y., Buchholz, F., Muyrers, J. P. & Stewart, A. F. A new logic for DNA engineering using recombination in Escherichia coli. Nat. Genet. 20, 123–128 (1998).
Hofemeister, H. et al. Recombineering, transfection, Western, IP and ChIP methods for protein tagging via gene targeting or BAC transgenesis. Methods 53, 437–452 (2011).
Ciotta, G. et al. Recombineering BAC transgenes for protein tagging. Methods 53, 113–119 (2011).
Klein, S., Pluim, J. P. W., Staring, M. & Viergever, M. A. Adaptive stochastic gradient descent optimisation for image registration. Int. J. Comput. Vision 81, 227–239 (2009).
Acknowledgements
We thank C. Raithel, A. Ziegler, S. Ammersdörfer and B. Vogel for technical support, M. Strasser and F. Buggenthin for helpful discussions and T. Hilger for statistical advice. This work was supported by the German Federal Ministry of Education and Research (BMBF), the European Research Council starting grant (Latent Causes), the BioSysNet (Bavarian Research Network for Molecular Biosystems), the International Human Frontier Science Program Organization, and by the German Research Foundation (DFG) within the SPPs 1395 and 1356. S.S. and O.H. acknowledge financial support for this project from SystemsX.ch.
Author information
Authors and Affiliations
Contributions
A.F. and S.H. performed experiments and analysed data with C.M., J.F. and M.S. M.S. developed QTFy and performed protein number estimation with J.H. J.F. performed dynamical modelling and parameter estimation with J.H. P.S.H. established quantitative TF imaging and maintained and advised on FACS procedures with M.E. K.A. produced reporter ESC lines. F.J.T. designed and supervised data analysis, modelling and QTFy software development, and commented on the manuscript. T.S. designed and supervised the study, developed and maintained long-term bioimaging with D.L. and K.D.K., and single-cell tracking and analysis software with B.S., O.H. and S.S., and wrote the manuscript with A.F., C.M. and S.H.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 3 Related to Fig. 1.
(a) Normal Proliferation of NanogVENUS and Oct4VENUS ESC lines. Both the untargeted parental R1 wild-type ESC line and the protein fusion reporter lines NanogVENUS and Oct4VENUS were passaged every two days and the number of total cells counted at each passage interval for a duration of 14 days. The resulting growth curves (mean ± SD, n = 3 independent experiments) show that both protein fusion lines proliferate as their wild-type counterpart. (b) Down-regulation of NanogVENUS and Oct4VENUS during differentiation. NanogVENUS and Oct4VENUS protein fusion reporter lines were passaged for 6 days in the absence of LIF. The proportion of VENUS positive cells in NanogVENUS and Oct4VENUS protein fusion lines was reduced significantly over the time course as measured by FACS (mean ± SD, n = 3 independent experiments), indicating the lines are sensitive reporters at the protein level for Nanog and Oct4, respectively. (c) Endogenous Nanog levels strongly correlate to NanogVENUS levels (R2 = 0.74, 492 cells). NanogVENUS ESCs were immunostained with an antibody against Nanog, imaged, background corrected, segmented, and quantified in both fluorescence channels. One representative experiment of 3 independent experiments shown. (d) Oct4GFP and Oct4VENUS are expressed similarly, regardless of genetic background and fluorescent protein (R1 in VENUS line and E14 in GFP line). (e) ESC culture on E-Cadherin does not change Nanog expression. The NanogVENUS ESC line was passaged for 14 days on gelatin or E-cadherin substrate and subsequently analysed by FACS. The proportion of NanogVENUS positive cells grown on either substrate was unchanged (mean ± SD, n = 3 independent experiments).
Supplementary Figure 4 Related to Fig. 2.
(a) NanogVENUS intensity is strongly correlated (Pearson correlation coefficient R = 0.94, coefficient of determination R2 = 0.89, n = 457 cells) to Nanog staining intensity. Shown is one representative experiment out of four independent experiments. On average, 74 ± 12% (coefficient of determination R2, mean ± SD, n = 4 independent experiments) of the variance in the Nanog staining intensity is explained by NanogVENUS intensity. (b) A scatter plot of the percentiles of each cell’s intensity in the respective NanogVENUS and Nanog staining distribution reveals that less than 3.5% of the cells deviate by more than 25% (points outside the gray shaded region) in the same experiment as shown in (a). (c) On average, less than 10 ± 6% (mean ± SD, n = 4 independent experiments) of cells exceed a percentile difference of 25%. (d) Monolayer colonies (representative examples shown at 30 h and 45 h after movie start) can be reliably tracked and quantified throughout the whole movie. Scale bars are 20 μm. (e) Approximately 10% of ESC colonies grow in 3D on E-cadherin coated dishes (example colony shown at 120 h) and cannot be tracked reliably. In ∼50% of these cases, colonies flatten out later (flattened out colony shown at 135.5 h). (f) NanogVENUS intensity does not differ significantly (p = 0.26, two sample t-test, n = 429) between the reference NanogVENUS distribution and cells grown in 3D colonies . Colonies with 3D growth can therefore be excluded from analyses without introducing bias.
Supplementary Figure 5 Related to Fig. 3.
(a) Compartmentalized transition matrix of NanogVENUS cells. From 1,356 negative Nanog cells analysed (pooled from 3 independent experiments), 2.3 ± 1.7% (mean ± SD, n = 3 independent experiments) gave rise to Nanog high daughter cells, while 62 ± 12% remained in the negative compartment in the next generation. (b) Analysis of the power spectrum of 60 branches from an unsorted, manually curated tree shows no indication of oscillations, ruling out oscillatory behaviour as a general pattern of Nanog dynamics. (c) Cell cycle corrected NanogVENUS and Oct4VENUS distributions differ: NanogVENUS spans a wider range and extends into the negative gate, while Oct4VENUS shows a higher overall protein expression. (d) Cell cycle corrected Oct4VENUS distribution is well above background and shows a dynamic range of DR = 8.0 dB. The uncorrected distribution, where cells are sampled randomly in cell cycle, is much broader with a DR = 11.0 dB. (e) The Oct4VENUS transition kernel is less complex than for NanogVENUS (cf Fig. 3d) and shows a strong bias towards Oct4 mid cells.
Supplementary Figure 6 Related to Fig. 4.
(a) NanogVENUS intensity trajectories are modelled as arising from a simple gene regulation model. The NanogVENUS gene is assumed to be initially basally active with low mRNA production rate kr0. At the switch time t0, the gene becomes active, giving rise to constant mRNA production kr1and Nanog protein production with rate kp proportional to the mRNA quantity. mRNA and protein are both degraded at a constant rate (γr and γp, respectively). (b) NanogVENUS trajectories are optimized using a gradient-based method, assuming Gaussian observation error. Model parameters are constrained to a region reflecting prior biological knowledge. (c) We estimate the maximum rate of protein production and observe significant heterogeneity in the rapidity of NanogVENUS activation across individual trajectories. Mean maximum production rate was 45,000 proteins per hour. (d) NanogVENUS onsets occur throughout the length of the movie, with estimated residence times of up to 52 h in the Nanog negative/low state. (e) Onsets occur throughout the cell cycle, and do not show a statistically significant difference from a uniform distribution (Kolmogorov–Smirnov test, p = 0.051). (f) The rate of convergence of NanogVENUS to stably high levels differs drastically across cellular branches. The shortest observed time until half-maximum intensity was 0.35 h, and the median 5.75 h. (g, h) Colonies which contained a large proportion of differentiated or senescent looking cells have been excluded from analysis. Representative brightfield images from one out of 3 independent experiments show Nanog negative/low colonies (g) and overtly differentiated or senescent cells (h). Green lines indicate boundaries of a colony.
Supplementary Figure 7 Related to Fig. 5.
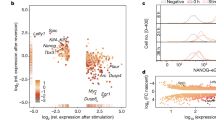
(a) No cells in the Nanog negative/low and mosaic colonies analysed were negative for Oct4 and Sox2. Around 30% of cells are negative for Klf4 in the Nanog negative/low colonies, and ∼11% in the Nanog mosaic colonies. (b) Comparison of partial correlations of transcription factor pairs reveals differences between Nanog mosaic and Nanog negative/low colonies. The most drastic difference occurs for Oct4–Klf4, where the partial correlation changes significantly from −0.14 ± 0.09 (mean ± SD, n = 3 independent experiments) in Nanog negative/low colonies to +0.09 ± 0.11 in Nanog mosaic colonies (p = 9.3 × 10−6, n = 3 independent experiments, partial correlation population test52). We show mean and 95% bootstrap confidence interval for each replicate. (c) Nanog negative/low cells from Nanog mosaic colonies that express NanogVENUS at the same level as Nanog negative colonies still exhibit altered partial correlations. Significant differences are found for Oct4–Klf4, Oct4–Sox2, and Nanog–Sox2. We show mean ± SD for each replicate. (d,e) Partial correlations were visualized using multiresolution correlation analysis plots, within which each point (w, dw) represents a cellular subpopulation whose mean NanogVENUS intensity is given by the point’s x-coordinate w and the subpopulation window size is given by the y-coordinate dw. The topmost point in the plot represents the whole population, with the mean given by the 50% quantile.
Supplementary Figure 8 Related to Fig. 6.
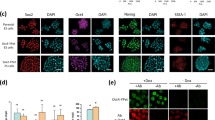
(a) Three-dimensional scatter plots show that negative/low and mosaic colonies are distinct in their potential to differentiate into a Foxa2 +, Sox1 −, small nuclear Area cell population. One representative experiment of 3 independent experiments shown. (b) Scatter plots of quantified immunostainings after differentiation of all negative/low or mosaic colonies. Two representative experiments of 3 independent experiments shown which are different replicates than the one used in Fig. 6d and Supplementary Fig. 6a. Foxa2 was not compensated for mCHERRYnucmem bleed-through in replicate 3. (c) Positive correlation of mCHERRYnucmem and Foxa2 signal intensity in a large proportion of dim Foxa2 cells in uncompensated dataset (top). Removal of bleed-through in each cell was achieved by subtraction of mCHERRYnucmem signal with a manually adjusted bleed-through factor (bottom). Two independent experiments shown. (d) Compensation of Foxa2 allowed better separation of differentiated subpopulations. Two independent experiments shown. (e) Foxa2 median ratios of NanogVENUS subpopulations are similar in uncompensated (n = 2,740 cells from negative/low colonies, n = 5,531 cells from mosaic colonies, n = 108 low cells in mosaic colonies and n = 125 high cells in mosaic colonies, pooled from 3 independent experiments) and compensated data sets (n = 2,152 cells from negative/low colonies, n = 4,027 cells from mosaic colonies, n = 69 low cells in mosaic colonies and n = 88 high cells in mosaic colonies, pooled from 2 experiments out of 3 independent experiments).
Supplementary Figure 9 Related to Fig. 7.
NanogVENUS low ESCs were imaged continuously for 70 h and then fixed before antibody staining of other TFs. Partial correlation analysis of subtrees from a Nanog negative/low colony shows variability between subtrees. Only significant (p < 0.05, partial correlation population test) edges are shown.
Supplementary Figure 10 Related to Fig. 8.
(a) Schematic of a pair of sister cells that exhibits divergent expression for Nanog may also exhibit divergence for other transcription factors. If the cells are divergent for another TF in the same manner as Nanog, the TF is labelled as congruent divergent (as for Oct4); if they are divergent, but in the opposite manner they are labelled incongruent divergent (as for Sox2). Lastly, they may also be non-divergent if the ratio of TFs between sister cells does not exceed the predefined threshold (as for Klf4). (b) The significance of congruent sisters with respect to all TF pairs is stable for thresholds smaller than the chosen fold-change threshold of 2 (Fisher’s exact test on 136 sister cell pairs pooled from 3 independent experiments). (c) Congruency patterns for all 136 sister pairs analysed.
Supplementary information
Supplementary Information
Supplementary Information (PDF 2804 kb)
Rights and permissions
About this article
Cite this article
Filipczyk, A., Marr, C., Hastreiter, S. et al. Network plasticity of pluripotency transcription factors in embryonic stem cells. Nat Cell Biol 17, 1235–1246 (2015). https://doi.org/10.1038/ncb3237
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3237
This article is cited by
-
Open-source personal pipetting robots with live-cell incubation and microscopy compatibility
Nature Communications (2022)
-
Machine learning phenomics (MLP) combining deep learning with time-lapse-microscopy for monitoring colorectal adenocarcinoma cells gene expression and drug-response
Scientific Reports (2022)
-
A cytokine screen using CRISPR-Cas9 knock-in reporter pig iPS cells reveals that Activin A regulates NANOG
Stem Cell Research & Therapy (2020)
-
In vivo tracking of single cells with PET
Nature Biomedical Engineering (2020)
-
Ontology patterns for the representation of quality changes of cells in time
Journal of Biomedical Semantics (2019)