Abstract
No immunogen to date has reliably elicited broadly neutralizing antibodies to HIV in humans or animal models. Advances in the design of immunogens that antigenically mimic the HIV envelope glycoprotein (Env), such as the soluble cleaved trimer BG505 SOSIP1, have improved the elicitation of potent isolate-specific antibody responses in rabbits2 and macaques3, but so far failed to induce broadly neutralizing antibodies. One possible reason for this failure is that the relevant antibody repertoires are poorly suited to target the conserved epitope regions on Env, which are somewhat occluded relative to the exposed variable epitopes. Here, to test this hypothesis, we immunized four cows with BG505 SOSIP. The antibody repertoire of cows contains long third heavy chain complementary determining regions (HCDR3) with an ultralong subset that can reach more than 70 amino acids in length4,5,6,7,8,9. Remarkably, BG505 SOSIP immunization resulted in rapid elicitation of broad and potent serum antibody responses in all four cows. Longitudinal serum analysis for one cow showed the development of neutralization breadth (20%, n = 117 cross-clade isolates) in 42 days and 96% breadth (n = 117) at 381 days. A monoclonal antibody isolated from this cow harboured an ultralong HCDR3 of 60 amino acids and neutralized 72% of cross-clade isolates (n = 117) with a potent median IC50 of 0.028 μg ml−1. Breadth was elicited with a single trimer immunogen and did not require additional envelope diversity. Immunization of cows may provide an avenue to rapidly generate antibody prophylactics and therapeutics to address disease agents that have evolved to avoid human antibody responses.
Similar content being viewed by others
Main
No immunogen to date has been able to reliably elicit broadly neutralizing antibodies (bnAbs) to HIV by vaccination. The difficulty in eliciting bnAbs has been attributed to the enormous antigenic diversity of the envelope glycoprotein and to the dense N-linked glycan coat that covers Env (the ‘glycan shield’). bnAbs isolated from chronically infected individuals have a number of unusual features that have been selected to cope with the glycan shield, including much longer than average HCDR3 loops10,11,12. HCDR3s in most vertebrates have restricted lengths averaging 12–16 amino acids9,13,14,15. Therefore, in many species, relatively few antibody precursors can be affinity-matured to HIV bnAbs. Cows, however, produce antibodies with HCDR3s that average ~26 amino acids in length with an ultralong subset (10–15% of the repertoire) that can be more than 70 amino acids long4,5,6,8,9,16. Repeated immunization over multiple years with a non-well-ordered Env trimer in cows led to some neutralization breadth in the immunoglobulin-rich colostrum17,18,19, although with relatively low potency. In this study, we immunized cows with the well-ordered BG505 trimer and monitored the longitudinal development of extensive neutralization breadth and potency. We also demonstrated that antibodies with ultralong HCDR3s are responsible for serum breadth and potency.
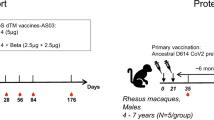
To investigate immunization with a well-ordered Env trimer in cows, we performed two experiments (Fig. 1a). The first (pilot) experiment involved immunization with JR-FL gp120 and BG505 SOSIP on each flank of two cows (nos 148 and 3441). Evaluation of sera on an indicator virus panel20 showed that both cows developed neutralization breadth and potency, with one cow (no. 3441) developing exceptionally potent responses (Fig. 1b). On the basis of these initial results, we immunized two additional cows (nos 26 and 27) with BG505 SOSIP alone (Fig. 1a). The terminal bleeds (day 381) for these two cows were also tested on the indicator virus panel and similar cross-clade neutralizing serum responses were observed (Fig. 1b).
a, Schematic of SOSIP immunization experiments in cows. b, Sera from all four cows were tested for neutralization on a 12-virus ‘global isolate’ panel (isolates listed on x axis) that serves as an indicator of breadth and potency. Presented values are neutralization ID50 titres. Murine leukaemia virus (MLV) was included as a negative control.
We next determined how quickly these responses developed. Sera from cow 26 and cow 27, from the second immunization, were sampled approximately every 7 days and tested on the same virus indicator panel (Fig. 2a and Extended Data Fig. 1). Remarkably, the results show the development of cross-neutralizing activity (8% breadth, n = 12) only 42 days after a prime and single homologous boost with BG505 SOSIP for cow 26. Autologous neutralization emerged at the same time as broad responses and potency increased over time (Extended Data Fig. 2). Cow 27 also developed broad responses, although more slowly than for cow 26 (Fig. 2a). Evaluation of the day 35 time point (14 days after the first boost) for cows 148 and 3441 (Fig. 2a) from experiment 1 also showed rapid emergence of breadth, albeit with lower potency (Extended Data Fig. 3).
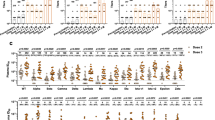
a, Sera collected longitudinally for cows 26 and 27 were tested for neutralization on the 12-virus indicator panel. Neutralization ID50 titres are presented for each virus by colour at each time point. Per cent neutralization breadth is indicated by the black line. Time in days (D) on x axis. b, Sera from selected time points were tested on a larger cross-clade 117-virus panel. Neutralization breadth and median neutralization ID50 titres are grouped by virus clade.
To fully evaluate the extent of neutralization breadth, we tested sera from cow 26 on a 117 cross-clade virus panel (Fig. 2b and Supplementary Table 1) on days 42, 77, 238 and 381. The results showed higher neutralization breadth at the earliest time point (day 42, 20% breadth) than observed for the indicator panel (8%). Following a second boost (day 77), this breadth expanded to 79% of isolates. At days 238 and 381, the breadth continued to rise to 92% and 96% of isolates, respectively. Potency, determined by median ID50 titres, also increased with consecutive boosts.
Next, we attempted to isolate bnAbs from cow 26. Peripheral blood mononuclear cells (PBMCs) from days 70 and 238 were sorted with fluorophores conjugated to anti-cow IgG, and biotinylated BG505 SOSIP was used as antigen bait, as described previously21 (Extended Data Fig. 4). Sorting and screening of PBMCs at different time points (Extended Data Fig. 5) using different sort strategies (Extended Data Fig. 4) resulted in the isolation of ten mAbs named NC-Cow1 to NC-Cow10. The sequences and alignments of the heavy and light chain sequences for NC-Cow1 to NC-Cow10 are listed in Supplementary Table 2. Notably, all ten antibodies have ultralong HCDR3s (Fig. 3a)
a, HCDR3s are shown for antibodies NC-Cow1 to NC-Cow10. The sequences for the germline Vh1–7, Dh8-2 and Jh10 regions are shown at the top. HCDR3 length (L), number of cysteines within HCDR3 (#Cys), and HCDR3 numbering are also shown. Cysteines within HCDR3 are highlighted in yellow, with those conserved with the germline underlined, and the cysteine and tryptophan defining HCDR3 are highlighted in cyan. b, NC-Cow1 was analysed for neutralization breadth and potency on a 117-virus panel. c, NC-Cow1 to NC-Cow10 map to the VRC01-class epitope by competition ELISA. d, To determine whether the HCDR3 of NC-Cow1 is effective at neutralization on its own, four chimaeras (right) were tested on the 12-virus global isolates indicator panel. An IC50 value of 50 μg ml−1 was used as a cut-off for neutralization.
To understand the neutralization properties of these 10 antibodies, we evaluated them on the 12-virus indicator panel as well as several clade A viruses and found that NC-Cow2 to NC-Cow6 had narrow breadth, whereas NC-Cow1 and NC-Cow7 to NC-Cow10 had broadly neutralizing activity (Extended Data Table 1). NC-Cow1 demonstrated the greatest breadth, so this bnAb was evaluated on a larger 117-virus panel and was found to have 72% neutralization breadth with a potent median IC50 of 0.028 μg ml−1 (Fig. 3b and Supplementary Table 1). All ten monoclonal antibodies competed with VRC01-class antibodies on a stabilized BG505 SOSIP trimer termed MD3922 (Fig. 3c). With the epitope specificity of NC-Cow1 identified, we next performed serum competition experiments with NC-Cow1 and found that the main serum specificity was to the CD4 binding site (CD4bs) (Extended Data Table 2). We found relatively low serum reactivity to the BG505 V3 loop (Extended Data Table 2). Finally, we determined that NC-Cow1 was not polyreactive to human antigens using a HEp-2 assay and an enzyme-linked immunosorbent assay (ELISA) (Extended Data Fig. 6).
To further investigate the binding mechanisms of these antibodies, we used single-particle negative stain electron microscopy to visually characterize the NC-Cow1 and NC-Cow2 epitopes. In each dataset, the antigen-binding fragment (Fab) was clearly visible adjacent to the CD4bs in two-dimensional class averages (Extended Data Fig. 7). We calculated a three-dimensional reconstruction of NC-Cow1 with BG505 Env trimer, which confirmed antibody binding to the CD4bs epitope (Extended Data Fig. 7). We next compared VRC01 and NC-Cow1 binding to a panel of gp120 CD4bs alanine mutants for isolates BG505 and JR-CSF (Extended Data Fig. 7). NC-Cow 1 showed sensitivity to several known CD4bs antibody residues, including D368R, for binding to BG505 gp120 (Extended Data Fig. 7). VRC01 binding was highly dependent on N279 for BG505 and on D279 for JR-CSF, as described previously23. By contrast, NC-Cow1 binding was not dependent on D279 or N279, but was instead dependent on residues in the C2 and C5 regions of gp120.
We next investigated whether the HCDR3 knob is functional on its own and whether the antibody retains its function when reverted to its inferred germline. We evaluated NC-Cow1 for neutralization on the 12-virus indicator panel, which demonstrated 100% breadth at a potent median IC50 of 0.007 μg ml−1 (Fig. 3d). Compared to this affinity-matured antibody, partial or fully reverted antibody variants showed no or little decrease in neutralization breadth and potency (Fig. 3d). We next transplanted the 60-amino-acid HCDR3 of NC-Cow1 to a germline-reverted variant of HIV bnAb PG9 and tested for neutralization on the 12-virus indicator panel. We observed a moderate drop to 88% neutralization breadth with a reduction in potency to a median IC50 of 0.054 μg ml−1. Although less active than the original antibody, the neutralization breadth and potency of a 60-amino acid HCDR3 knob transplanted onto a germline antibody is remarkable, and may have implications for therapeutic designs.
A number of HIV bnAbs have been or are being tested as potential microbicides to prevent mucosal HIV acquisition24. Notably, VRC01-class antibodies rely on a critical salt bridge interaction with an aspartic acid residue at position 368 of gp12023 that is disrupted at low pH (Extended Data Fig. 8), which is characteristic of conditions found in the vagina. NC-Cow1, however, retained affinity for gp120 in simulated vaginal fluid at pH 4.5, suggesting that the antibody might be useful as a potential microbicide, although this application warrants further exploration (Extended Data Fig. 8).
Here we have shown that immunization with a well-ordered Env trimer in cows reliably and rapidly elicits broad and potent neutralizing serum responses, in contrast to previous experiments in other animals. As far as we are aware, the only previous example of elicitation of broad serum neutralization and subsequent isolation of bnAbs was in llamas, when seven immunizations with an Env trimer (not well-ordered) over 4 months resulted in poor serum neutralization breadth and potency and subsequent screening of more than 2,800 unique camelid-specific variable region (VHH) fragments identified only one with broad and potent neutralizing activity25,26.
The results described here support the idea that bnAb epitopes on HIV Env are immunoquiescent only in a repertoire-dependent fashion. The CD4bs is recessed and occluded on the native trimer, which greatly hinders access by human neutralizing antibodies and thereby renders the CD4bs effectively immunoquiescent in most humans. The long HCDR3 of cow antibodies can nonetheless easily access the CD4bs on the trimer, thereby rendering this region immunogenic in the context of the cow antibody repertoire.
Importantly, different trimer isolates were not required to elicit breadth, indicating that trimer diversity might not be required provided that the conserved epitopes are accessible and immunogenic. The speed of developing a bnAb to the CD4bs of HIV Env in cows is remarkable when contrasted with the length of time required to elicit similar antibodies in humans through natural infection (more than 5 years). One lesson from these studies for HIV vaccine design is that immunization-priming steps that favour the selection of long HCDR3s may accelerate the development of bnAbs.
Finally, the rapid elicitation of functional responses against a difficult target such as HIV Env suggests that cow immunization should be explored for rapid generation of antibodies against other pathogens that have evolved to avoid human antibody responses. Such antibodies, as well as their intrinsic prophylactic and therapeutic value, could help to define targets for vaccine and drug design.
Methods
Pseudovirus neutralization assays
Plasmids encoding HIV Env were co-transfected into HEK293T cells (ATCC) with pSG3ΔEnv, an Env-deficient genomic backbone plasmid, in a 1:2 ratio using X-tremeGENE HP (Roche) as transfection reagent. Cell culture supernatants were harvested 3 days after transfection and sterile filtered through a 0.22-μm filter. Neutralizing activity was measured by incubating monoclonal antibodies or sera with replication-incompetent pseudovirus for 1 h at 37 °C before transferring onto TZM-bl target cells (https://www.aidsreagent.org/) as described previously10.
BG505 SOSIP trimer expression and purification
BG505 SOSIP.664 gp140, BG505 SOSIP.664-His gp140, and BG505 SOSIP.664-avi gp140 were expressed in HEK293F cells (Invitrogen) as described previously21. In brief, HEK293F cells were maintained in FreeStyle medium (Invitrogen). For gp140 trimer production, HEK293F cells were seeded at a density of 0.5 × 106 per ml. After 24 h, cells were transfected with 1 mg 293Fectin (Invitrogen) with 300 μg Env plasmid and 75 μg furin plasmid in OPTI-MEM according to the manufacturer’s protocol. Supernatants were purified using a Galanthus nivalis lectin (Vector Labs) column and protein was eluted with 1 M methyl-α-d-mannopyranoside (MMP, Sigma). Following buffer exchange into PBS, only trimers with AviTags were in vitro biotinylated using the BirA enzyme according to the manufacturer’s protocol (Avidity). The affinity-purified Env proteins were further purified to size homogeneity using size exclusion chromatography (SEC) on a Superose 6 10/300 GL column (GE Healthcare) in PBS. The trimer fractions were collected and pooled and protein concentrations were determined using either a bicinchonic acid-based assay (Thermo Scientific) or UV280 absorbance using theoretical extinction coefficients.
Cow immunization
Six-month-old Bos taurus calves were used to analyse the bovine immune response to HIV antigens. Animals were primed and boosted by intradermal inoculation. Two animals were selected for two different immunization experiments as a pilot study, yielding a sample size of four animals in total. All four animals were bled from the jugular vein as often as once per week for serum and for the isolation of mononuclear cells from peripheral blood. Animals and subsequent analyses were not randomized or blinded. No statistical methods were used to predetermine sample size. Heifers 148 and 3441 were Angus cross breeds immunized with 200 μg BG505 SOSIP trimer (200 μl antigen/800 μl adjuvant inoculations were divided into five 200-μl injections with Iscomatrix adjuvant for cow 148 and RIBI for cow 3441) on one side and 200 μg JRFL gp120 on the other (200 μl antigen/800 μl adjuvant inoculations were split into five 200-μl injections with RIBI adjuvant). Both heifers 148 and 3441 were boosted on Day 21 with 200 μg of the same antigen on the same side of the neck as previously received, and once more on Day 78 with the same delivery of immunogen as the first boost. All boosts of these two heifers employed RIBI as adjuvant. Holstein steers 26 and 27 received immunizations of 200 μg BG505 SOSIP trimer (total spread over five sites on each side of neck) emulsified in an equal volume of ENABL C1 (VaxLiant) adjuvant. Boosts of equal amounts of antigen were administered on Days 36, 64, 99, 148, 211 and 360. Exceptions were that on Day 99 the boost was administered with RIBI adjuvant instead of ENABL C1. These protocols were approved by the Texas A&M Institutional Animal Care and Use Committee for M.F.C. as AUP 2015-078.
Single-particle negative stain electron microscopy
BG505 SOSIP.664 + NC-Cow1 complexes were placed on glow-discharged carbon-coated copper mesh grids and stained with 2% uranyl formate. Grids were screened for appropriate stain thickness and particle distribution using an FEI Morgagni (80 keV) electron microscope. The final dataset was collected on an FEI Tecnai Spirit T12 (120 keV) electron microscope with a Tietz TVIPS CMOS (4K by 4K) camera controlled with Leginon automated imaging software27. Images were collected at 52,000× magnification with a −1.3 μm defocus for a final magnified pixel size of 2.05 Å per pixel. Both automated particle picking performed with DoG-Picker28 and reference-free 2D classification with iterative MRA-MSA (multi-reference alignment, multivariate statistical analysis)29 were executed through the Appion database30. For 3D analysis, micrographs were first contrast transfer function (CTF) estimated with GCTF then particles were extracted, phase-flipped, and subjected to reference-based 3D classification and refinement in Relion version 2.031. The final 3D reconstruction contained ~3.5k particles out of the initial ~13.5k that went into 3D classification. UCSF Chimera was used to generate figures32.
Single-cell sorting of cow PBMCs using flow cytometry
Sorting of cow PBMCs was performed as described previously with some modifications21. Cow PBMCs were stained with primary fluorophore-conjugated antibodies binding cow IgG (AbCam) and biotinylated BG505 SOSIP.664-avi gp140 coupled to streptavidin-APC and PE (Life Technologies) in equimolar ratios. Cells were stained for 1 h at 4 °C in PBS containing 1 mM EDTA and 1% FBS and 50 nM of conjugated bait. Cells were sorted for IgG+/BG505 SOSIP.664-avi-PE+/BG505 SOSIP.664-avi-APC+ events. Target cells were single-cell sorted into 96-well plates containing lysis buffer on a BD Fusion sorter and were immediately frozen on dry ice.
Single-cell PCR amplification and cloning of antibody variable genes
cDNA synthesis from mRNA and subsequent rounds of PCR amplification of antibody variable genes were performed as previously described, but using primers for cow immunoglobulin (Supplementary Table 3). PCR reactions were set up in 25 μl volume with 2.5 μl cDNA or PCR1 product using HotStarTaq Master Mix (Qiagen). Heavy and light chain pairs retrieved from single sorted cells were cloned into human antibody expression vectors as described previously33.
Antibody production and purification
Antibody plasmids containing heavy chain and light chain genes were co-transfected (1:1 ratio) in either HEK293T or HEK293F cells using X-tremeGENE (Roche) or 293Fectin (Invitrogen) as transfection reagents, respectively. Antibody-containing supernatants were harvested 4 days after transfection and 0.22-μm sterile filtered. Antibodies produced in HEK293T cells were quantified by anti-Fc ELISA and used directly in neutralization assays for screening purposes. Antibody supernatants produced in HEK293F cells were purified over Protein A Sepharose 4 Fast Flow (GE healthcare) columns as described previously18.
ELISA assays
ELISA plates were first coated with an anti-C5 gp120 antibody at 4 °C in 1 × PBS overnight. Plates were then washed five times with PBS + 0.05% Tween and blocked with 3% BSA in 1 × PBS at room temperature for 1 h. Mutant pseudovirus supernatants were lysed with 1% NP40 and then captured on ELISA plates at 37 °C for 2 h using antibody D7324. Plates were washed five times with PBS + 0.05% Tween and then serial dilutions of monoclonal antibodies were added to the wells and plates were incubated at room temperature for 1 h. Plates were washed five times with PBS + 0.05% Tween and then goat anti-human IgG F(ab′)2 conjugated to alkaline phosphatase (Pierce) was diluted 1:1,000 in PBS containing 1% BSA and 0.05% Tween and added to the wells. The plate was incubated at room temperature for 1 h and washed five times with PBS + 0.05% Tween. Plates were then developed by adding 50 μl alkaline phosphatase substrate (Sigma) dissolved in alkaline phosphatase staining buffer (pH 9.8), according to the manufacturer’s instructions. The optical density at 405 nm was read on a microplate reader (Molecular Devices). For experiments involving antibody binding to antigen at different pH values, serial dilutions of monoclonal antibodies were incubated in PBS at different pHs or in simulated vaginal fluid34 (SVF) for 1 h at room temperature and then washed five times with PBS + 0.05% Tween before addition of goat anti-human IgG F(ab′)2 conjugated to alkaline phosphatase secondary. SVF was made with citric acid instead of lactic acid to improve buffering at higher pH levels.
Competition ELISA
For competition ELISA experiments, competing antibodies were biotinylated using an antibody biotinylation kit (Thermo Scientific). Plates were coated with an anti-His antibody (Roche) at 5 μg/ml overnight. Following washing, plates were blocked with 3% BSA for 1 h at room temperature. The stabilized BG505 SOSIP construct MD3922 was then captured at 2.5 μg/ml in PBS (50 μl/well) for 2 h at 37 °C. After washing, serially diluted antibodies in PBS/1% BSA were added for 30 min. To this was added biotinylated antibody at a constant EC70 concentration for 1 h. Plates were washed and detection was measured using alkaline phosphatase-conjugated streptavidin (Pierce) at 1:1,000 for 1 h at room temperature. Absorption was measured at 405 nm.
HEp-2 cell staining assay
The HEp-2 cell-staining assay was performed using kits purchased from Aesku Diagnostics according to the manufacturer’s instructions. These Aesku slides use optimally fixed human epithelial (HEp-2) cells (ATCC) as a substrate and affinity-purified, FITC-conjugated goat anti-human IgG for detection. In brief, 2.5 μg or 25 μl of 100 μg/ml monoclonal antibodies or controls were added to wells and incubated on HEp-2 slides in a moist chamber at room temperature for 30 min. Slides were then rinsed and submerged in PBS and 25 μl FITC-conjugated goat anti-human IgG was immediately applied to each well. Slides were allowed to incubate at room temperature in a moist chamber for another 30 min. Slides were then washed in the same manner as above and then mounted on coverslips using the provided mounting medium. Slides were viewed at 20× magnification and photographed on an EVOS f1 fluorescence microscope at 250 ms exposure with 100% intensity. Positive and negative control sera were provided by the vendor. Samples showing fluorescence greater than the negative control were considered positive for HEp-2 staining.
Polyspecificity reagent (PSR) binding assay
Monoclonal antibodies were screened for reactivity with preparations of solubilized membrane proteins (SMP) and cytosolic proteins (SCP) as described previously with a small modification35. In brief, SMP and SCP were extracted from CHO cells (ATCC). The protein concentration was determined using the Dc-protein assay kit (BioRad). SMP and SCP were then immobilized on ELISA plates for monoclonal antibody screening. The results were established by reading the absorbance at 450 nm of the examined samples.
Single autoantigen reactivity
Single-antigen ELISAs for SSA/Ro, SS-B/La, Sm, ribonucleoprotein (RNP), Jo-1, double-stranded DNA, centromere B, and histones were purchased from Aesku Diagnostics. The 96 wells were separately coated with these eight cellular and nuclear antigens for the qualitative detection of monoclonal antibody reactivity. A cut-off calibrator was provided by the manufacturer. The negative control was diluted human serum.
Data availability
Gene sequences of the reported antibodies have been deposited in GenBank under accession numbers MF167436– MF167455. Source Data for Figs 1, 2, 3 and Extended Data Figs 2, 5, 8 are provided with the online version of the paper. Other data are available on request from the corresponding authors.
References
Sanders, R. W. et al. A next-generation cleaved, soluble HIV-1 Env trimer, BG505 SOSIP.664 gp140, expresses multiple epitopes for broadly neutralizing but not non-neutralizing antibodies. PLoS Pathog. 9, e1003618 (2013)
McCoy, L. E. et al. Holes in the glycan shield of the native HIV envelope are a target of trimer-elicited neutralizing antibodies. Cell Reports 16, 2327–2338 (2016)
Sanders, R. W . et al. HIV-1 neutralizing antibodies induced by native-like envelope trimers. Science 349, aac4223 (2015)
Berens, S. J., Wylie, D. E. & Lopez, O. J. Use of a single VH family and long CDR3s in the variable region of cattle Ig heavy chains. Int. Immunol. 9, 189–199 (1997)
Lopez, O., Perez, C. & Wylie, D. A single VH family and long CDR3s are the targets for hypermutation in bovine immunoglobulin heavy chains. Immunol. Rev. 162, 55–66 (1998)
Saini, S. S., Allore, B., Jacobs, R. M. & Kaushik, A. Exceptionally long CDR3H region with multiple cysteine residues in functional bovine IgM antibodies. Eur. J. Immunol. 29, 2420–2426 (1999)
Saini, S. S. & Kaushik, A. Extensive CDR3H length heterogeneity exists in bovine foetal VDJ rearrangements. Scand. J. Immunol. 55, 140–148 (2002)
de los Rios, M., Criscitiello, M. F. & Smider, V. V. Structural and genetic diversity in antibody repertoires from diverse species. Curr. Opin. Struct. Biol. 33, 27–41 (2015)
Wang, F. et al. Reshaping antibody diversity. Cell 153, 1379–1393 (2013)
Walker, L. M. et al. Broad neutralization coverage of HIV by multiple highly potent antibodies. Nature 477, 466–470 (2011)
Doria-Rose, N. A. et al. Developmental pathway for potent V1V2-directed HIV-neutralizing antibodies. Nature 509, 55–62 (2014)
Bonsignori, M. et al. Analysis of a clonal lineage of HIV-1 envelope V2/V3 conformational epitope-specific broadly neutralizing antibodies and their inferred unmutated common ancestors. J. Virol. 85, 9998–10009 (2011)
Shi, B. et al. Comparative analysis of human and mouse immunoglobulin variable heavy regions from IMGT/LIGM-DB with IMGT/HighV-QUEST. Theor. Biol. Med. Model. 11, 30 (2014)
Lee, E.-C. et al. Complete humanization of the mouse immunoglobulin loci enables efficient therapeutic antibody discovery. Nat. Biotechnol. 32, 356–363 (2014)
Kodangattil, S. et al. The functional repertoire of rabbit antibodies and antibody discovery via next-generation sequencing. MAbs 6, 628–636 (2014)
Saini, S. S., Farrugia, W., Ramsland, P. A. & Kaushik, A. K. Bovine IgM antibodies with exceptionally long complementarity-determining region 3 of the heavy chain share unique structural properties conferring restricted VH + Vlambda pairings. Int. Immunol. 15, 845–853 (2003)
Heydarchi, B. et al. Trimeric gp120-specific bovine monoclonal antibodies require cysteine and aromatic residues in CDRH3 for high affinity binding to HIV Env. MAbs 9, 550–566 (2016)
Heydarchi, B. et al. Repeated vaccination of cows with HIV Env gp140 during subsequent pregnancies elicits and sustains an enduring strong Env-binding and neutralising antibody response. PLoS ONE 11, e0157353 (2016)
Kramski, M. et al. Hyperimmune bovine colostrum as a low-cost, large-scale source of antibodies with broad neutralizing activity for HIV-1 envelope with potential use in microbicides. Antimicrob. Agents Chemother. 56, 4310–4319 (2012)
deCamp, A. et al. Global panel of HIV-1 Env reference strains for standardized assessments of vaccine-elicited neutralizing antibodies. J. Virol. 88, 2489–2507 (2014)
Sok, D. et al. Recombinant HIV envelope trimer selects for quaternary-dependent antibodies targeting the trimer apex. Proc. Natl Acad. Sci. USA 111, 17624–17629 (2014)
Steichen, J. M. et al. HIV vaccine design to target germline precursors of glycan-dependent broadly neutralizing antibodies. Immunity 45, 483–496 (2016)
Zhou, T. et al. Structural basis for broad and potent neutralization of HIV-1 by antibody VRC01. Science 329, 811–817 (2010)
Veselinovic, M., Neff, C. P., Mulder, L. R. & Akkina, R. Topical gel formulation of broadly neutralizing anti-HIV-1 monoclonal antibody VRC01 confers protection against HIV-1 vaginal challenge in a humanized mouse model. Virology 432, 505–510 (2012)
McCoy, L. E. et al. Molecular evolution of broadly neutralizing Llama antibodies to the CD4-binding site of HIV-1. PLoS Pathog. 10, e1004552 (2014)
McCoy, L. E. et al. Potent and broad neutralization of HIV-1 by a llama antibody elicited by immunization. J. Exp. Med. 209, 1091–1103 (2012)
Suloway, C. et al. Automated molecular microscopy: the new Leginon system. J. Struct. Biol. 151, 41–60 (2005)
Voss, N. R., Yoshioka, C. K., Radermacher, M. & Potter, C. S. & Carragher, B. DoG Picker and TiltPicker: software tools to facilitate particle selection in single particle electron microscopy. J. Struct. Biol. 166, 205–213 (2009)
Ogura, T., Iwasaki, K. & Sato, C. Topology representing network enables highly accurate classification of protein images taken by cryo electron-microscope without masking. J. Struct. Biol. 143, 185–200 (2003)
Lander, G. C. et al. Appion: an integrated, database-driven pipeline to facilitate EM image processing. J. Struct. Biol. 166, 95–102 (2009)
Kimanius, D ., Forsberg, B. O ., Scheres, S. H . & Lindahl, E. Accelerated cryo-EM structure determination with parallelisation using GPUs in RELION-2. eLife 5, 19 (2016)
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004)
Tiller, T. et al. Efficient generation of monoclonal antibodies from single human B cells by single cell RT–PCR and expression vector cloning. J. Immunol. Methods 329, 112–124 (2008)
Owen, D. et al. A vaginal fluid simulant. Contraception 59, 91–95 (1999)
Jardine, J. et al. Minimally mutated HIV-1 broadly neutralizing antibodies to guide reductionist vaccine design. PLoS Pathog. 12, e1005815 (2016)
Kearse, M. et al. Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28, 1647–1649 (2012)
Acknowledgements
This work was supported by the International AIDS Vaccine Initiative Neutralizing Antibody Consortium through the Collaboration for AIDS Vaccine Discovery grant OPP1084519 (D.R.B., I.A.W., A.B.W.), NIH grants R21 AI120791 (V.V.S.), R01 GM105826 (V.V.S.), Center for HIV/AIDS Vaccine Immunology and Immunogen Discovery Grant UM1AI100663 (D.R.B., I.A.W., A.B.W.), IOS 1257829 (M.F.C.), and USDA-NIFA grant number CSREES 2008-35204 (W.M.). D.R.B. acknowledges the support of the James and Jessie Minor Chair in Immunology. We thank B. Schief and S. Menis for providing MD39 for competition experiments. This work was funded in part by IAVI and made possible by the support of many donors, including: the Bill & Melinda Gates Foundation, the Ministry of Foreign Affairs of Denmark, Irish Aid, the Ministry of Finance of Japan in partnership with The World Bank, the Ministry of Foreign Affairs of the Netherlands, the Norwegian Agency for Development Cooperation (NORAD), the United Kingdom Department for International Development (DFID), and the United States Agency for International Development (USAID). The full list of IAVI donors is available at http://www.iavi.org. The contents of this manuscript do not necessarily reflect the views of USAID or the US Government.
Author information
Authors and Affiliations
Contributions
D.S., K.M.L., J.R. and A.R. performed antigen B cell sorts. K.M.L., J.R. and A.R. performed PCR and antibody cloning, expression, and purification. K.L.S.-F., K.M.L. and J.R. performed neutralization and mutagenesis experiments. M.L.V. and D.S. designed and validated VH and VL gene-specific primers. D.S., K.M.L., M.L.V. and V.V.S. analysed V-region sequences. L.K. and I.A.W. provided protein for immunization experiments. J.L.T., Z.T.B., R.S., A.B.W. and I.A.W. performed structural analysis. J.G.J. performed pH experiments. C.-H.L. performed polyreactivity experiments. P.L.C., M.F.C. and W.M. performed cow immunization and serum ELISA experiments. P.L.C., M.F.C., W.M. and M.L.V. processed lymphocytes and produced mRNA and cDNA. D.S., V.V.S. and D.R.B. helped design and oversaw experiments. D.S., V.V.S. and D.R.B. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
D.S., D.R.B. and V.V.S. are inventors on a patent describing the NC-Cow antibodies: US provisional application filed on 14 July 2017.
Additional information
Reviewer Information Nature thanks J. Mascola and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Sera collected at different time points over the course of immunization for cows 26 and 27 were evaluated for neutralization breath and potency on the 12-virus global indicator panel.
Values represent serum ID50 against the indicated virus.
Extended Data Figure 2 Serum tested at given time points for neutralization on BG505 pseudovirus.
Neutralization against autologous virus emerged at the same time as breadth (day 42) and increased in potency over time. Values represent serum ID50.
Extended Data Figure 3 Development of neutralization breadth at 35 days post immunization from cows in experiment 1.
Serum collected at day 35 from experiment 1 was tested for neutralization breadth and potency on the 12-virus global indicator panel. Values represent serum ID50.
Extended Data Figure 4 Sorting strategy for isolating antigen-specific IgG+ B cells.
a, Cow PBMCs were sorted for IgG+ cells that bound to biotinylated BG505 SOSIP-AviTag conjugated on PE and APC fluorophores. b, To isolate epitope-specific antibodies, unliganded BG505 SOSIP (blue) and BG505 SOSIP liganded with NC-Cow1 was used to antigen-sort memory B cells. Epitope-specific B cells are defined as binding unliganded SOSIP and not binding to liganded SOSIP. This sort strategy yielded the broadly neutralizing antibodies NC-Cow7 to NC-Cow10.
Extended Data Figure 5 Functional screening and sequencing information for isolated antibodies.
a, Amplified heavy chains were paired with universal cow light chain or with NC-Cow1 light chain and tested for expression (anti-Fc ELISA), antigen binding (BG505 SOSIP), autologous neutralization (BG505 pseudovirus), and heterologous neutralization (Q23 pseudovirus). Sequence alignments of recovered heavy chains are listed underneath. As ultralong HCDR3 antibodies have been reported to pair with a single germline light chain (V30), amplified heavy chain genes were first paired with the universal cow light chain and screened for expression, binding to BG505 SOSIP, neutralization of BG505 virus, and neutralization of a clade A heterologous virus, Q23. From this dataset, three antibodies (NC-Cow1, NC-Cow2, and NC-Cow3) were selected that showed autologous neutralization (all from day 238) and corresponding native light chains were amplified, with success for only NC-Cow1. These three antibodies were expressed and purified at a larger scale for additional characterization by maintaining native pairing for NC-Cow1 and pairing with germline V30 for NC-Cow2 and NC-Cow3. For the day 70 time point, three heavy chains were selected that showed binding to BG505 SOSIP, but no neutralization, and these antibodies were produced with their native light chain pairs (NC-Cow4 to NC-Cow6). Finally, as NC-Cow1 could neutralize isolate Q23 in the neutralization screen, an additional sort with PBMCs from day 381 was performed, but used BG505 SOSIP liganded with and without NC-Cow1 to enrich for epitope-specific antibodies. The enrichment yielded an additional five hits by neutralization screen, and four out of these five antibodies were produced at larger scale with their native light chain pairs (NC-Cow7 to NC-Cow10). Small-scale screening was also performed with all heavy chains paired with NC-Cow1 light chain and no significant increase in neutralization breadth was observed, although there were slight improvements in BG505 SOSIP affinity and autologous potency. b, Nucleotide alignment of heavy chain sequences of isolated monoclonal antibodies36.
Extended Data Figure 6 NC-Cow1 is not polyreactive to human antigens.
a, NC-Cow1 was tested for antigen reactivity in a HEp-2 assay compared to the known polyreactive antibody 4E10, and negative and positive control sera supplied by the manufacturer. b, NC-Cow1 was also tested for reactivity with a range of typical human autoantigens by ELISA as well as for binding to solubilized membrane (SMP) and cytosolic preparations (SCP) from CHO cells. Values are optical density values (OD405) at a dose of 100 μg ml−1. Black lines indicate cut-off values as indicated by the manufacturer.
Extended Data Figure 7 Epitope mapping of NC-Cow1.
a, Representative 2D class averages of cow Fabs NC-Cow1 and NC-Cow2 bound to BG505 SOSIP trimers to demonstrate CD4bs specificity. The Fabs appear at slightly different angles relative to the trimer, perhaps owing to some flexibility between the Fab and HCDR3. In the 2D class averages for NC-Cow1, the HCDR3 appears as a faint density bridging between the Fab and CD4bs. b, Top and side views of 3D reconstruction of NC-Cow1 bound to BG505 SOSIP trimer with previously published Env trimer structure (green, PDB 5CEZ) and cow Fab (orange, PDB 5IJV) docked in. The body of NC-Cow1 Fab is approximately 20 Å from the CD4bs, which is the estimated length of the ultralong HCDR3. c, NC-Cow 1 was tested by ELISA for binding to BG505 or JRCSF gp120 captured from lysed virions with PGT121 (V3-glycan epitope) and VRC01 (CD4bs epitope) included for comparison. Values are EC50 in μg ml−1. NC-Cow1 was also tested for neutralization on BG505 or JR-CSF pseudoviruses and corresponding alanine mutants with PGT121 (V3-glycan epitope) and 12A12 (CD4bs epitope) included for comparison. Values are IC50 in μg ml−1. d, NC-Cow1 was tested for binding to wild-type and D368R protein. Antibodies VRC01 (CD4bs) and 14e (V3) were included for comparison.
Extended Data Figure 8 Effects of pH on binding of NC-Cow1 and CD4bs antibodies to gp120.
a, CD4bs antibodies PGV04, 12A12, 3BNC60 and b12 were tested for binding to gp120 (isolate 92BR020) by ELISA in buffers at different pHs (3.5, 4.0, 4.5, 5.0, 5.5 and 6.0). b, NC-Cow1 and VRC01 were tested for binding to BG505 gp120 in PBS buffer (pH 7.4) compared to simulated vaginal fluid (SVF) at pH 4.5.
Supplementary information
Supplementary Tables
This file contains Supplementary Tables 1–3. (PDF 221 kb)
Rights and permissions
About this article
Cite this article
Sok, D., Le, K., Vadnais, M. et al. Rapid elicitation of broadly neutralizing antibodies to HIV by immunization in cows. Nature 548, 108–111 (2017). https://doi.org/10.1038/nature23301
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature23301
This article is cited by
-
Mechanism of glycoform specificity and in vivo protection by an anti-afucosylated IgG nanobody
Nature Communications (2023)
-
De novo genome assembly depicts the immune genomic characteristics of cattle
Nature Communications (2023)
-
Assessing immunogenicity barriers of the HIV-1 envelope trimer
npj Vaccines (2023)
-
Evolution of immunogenetic components encoding ultralong CDR H3
Immunogenetics (2023)
-
SARS CoV-2 infections in animals, two years into the pandemic
Archives of Virology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.