Abstract
Normal epithelial cells often exert anti-tumour effects against nearby oncogenic cells. In the Drosophila imaginal epithelium, clones of oncogenic cells with loss-of-function mutations in the apico-basal polarity genes scribble or discs large are actively eliminated by cell competition when surrounded by wild-type cells1,2,3,4,5. Although c-Jun N-terminal kinase (JNK) signalling plays a crucial role in this cell elimination1,2,3,4,5, the initial event, which occurs at the interface between normal cells and polarity-deficient cells, has not previously been identified. Here, through a genetic screen in Drosophila, we identify the ligand Sas and the receptor-type tyrosine phosphatase PTP10D as the cell-surface ligand–receptor system that drives tumour-suppressive cell competition. At the interface between the wild-type ‘winner’ and the polarity-deficient ‘loser’ clones, winner cells relocalize Sas to the lateral cell surface, whereas loser cells relocalize PTP10D there. This leads to the trans-activation of Sas–PTP10D signalling in loser cells, which restrains EGFR signalling and thereby enables elevated JNK signalling in loser cells, triggering cell elimination. In the absence of Sas–PTP10D, elevated EGFR signalling in loser cells switches the role of JNK from pro-apoptotic to pro-proliferative by inactivating the Hippo pathway, thereby driving the overgrowth of polarity-deficient cells. These findings uncover the mechanism by which normal epithelial cells recognize oncogenic polarity-deficient neighbours to drive cell competition.
Similar content being viewed by others
Main
Normal epithelial cells possess an intrinsic tumour-suppression mechanism against oncogenic neighbours. For instance, in canine kidney cell cultures and zebrafish embryos, oncogenic cells that activate Ras or Src are eliminated from an epithelial monolayer when surrounded by normal cells6,7. Similarly, in the Drosophila imaginal epithelium, oncogenic polarity-deficient cells mutant for scribble (scrib) or discs large (dlg1; hereafter dlg) are eliminated from the tissue when surrounded by wild-type cells1,2,4. The removal of these surrounding wild-type cells abolishes cell elimination and allows scrib−/− or dlg−/− loss-of-function mutant cells to overproliferate1; this context-dependent cell elimination is therefore considered to be cell competition4,8. Genetic studies in Drosophila have revealed that this tumour-suppressive cell competition is driven by JNK-dependent cell death, triggered by the Drosophila tumour necrosis factor (TNF) Eiger1,2,3. However, the initial mechanism by which normal epithelial cells recognize nearby polarity-deficient cells to drive cell competition have remained unknown.
To explore the initial event, which occurs at the interface between normal cells and oncogenic polarity-deficient cells, we conducted an ethyl methanesulfonate (EMS)-based genetic screen in Drosophila for genes required for wild-type ‘winners’ to eliminate neighbouring polarity-deficient ‘losers’ (Extended Data Fig. 1a). In the eye imaginal epithelium, clones of homozygous mutant scrib−/− cells are eliminated when surrounded by wild-type tissue1,2 (Fig. 1a, b). The elimination of scrib−/− clones is also evident in adult eyes (Fig. 1e, f). Using the FLP/FRT-mediated genetic mosaic technique9, we introduced EMS-induced homozygous mutations only in wild-type winners (see Methods for details) and screened for mutations that caused an elimination-defective (eld) phenotype in neighbouring scrib−/− losers. Among 7,490 mutant strains generated, we isolated four elimination-defective mutants (eld-4, eld-6, eld-7, and eld-8) that fell into the same lethal complementation group. Clones of scrib−/− surrounded by eld-4 clones were no longer eliminated but instead grew robustly in the eye disc (Fig. 1b, c, i) and survived into adult tissue, causing a characteristic melanization phenotype (Fig. 1f, g). Notably, clones of eld-4, eld-6, eld-7, or eld-8 cells showed neither a growth disadvantage of their own nor a suppressive effect on the growth of neighbouring wild-type tissue (Fig. 1d, h and data not shown). Thus, the complementation group eld-4/6/7/8 possesses mutations in a gene required for elimination of neighbouring scrib−/− clones.
a–d, Eye discs of ey-FLP-MARCM-induced mosaics of wild-type (WT; a), scrib−/− (b), eld4−/−//scrib−/− (c), or eld4−/− (d). Clones are marked by the presence or absence of GFP (indicated by upper bars). e–h, Adult eye of ey-FLP-induced mosaics of wild-type (e), scrib−/− (f), eld4−/−//scrib−/− (g), or eld4−/− (h). Clones are marked by the presence or absence of red pigment (indicated by upper bars). i, Quantification of relative size of clones in genotypes shown in b (n = 22 eye discs) and c (n = 24). Mean ± s.d.; **P < 0.01 by Mann–Whitney U-test. j, A schematic representation of the general domain structure of Sas. The eld-4 mutant chromosome harbours a nonsense mutation in the sas gene at amino acid codon 796 (CAG > TAG). k–n, Adult eye of ey-FLP-MARCM-induced mosaics of scrib−/− (k), scrib−/−//eld4−/− (l), scrib−/−//eld4−/−UAS-sas (m), or scrib−/−//sas-RNAi (n).
Using a series of chromosomal-deficiency lines and subsequent cDNA sequencing, we identified a nonsense mutation in the coding region of the gene stranded at second (sas) in the eld-4 mutant strain (Fig. 1j and Extended Data Fig. 1b). Encoded by sas is a cell-surface ligand protein that has two extracellular domains—von Willebrand factor type C (VWC) and fibronectin type 3 (FN3) domains—as well as a transmembrane domain10 (Fig. 1j). Sas is required for proper axon guidance in the nervous system11, but its physiological role in epithelia is unknown. Expression of Sas was indeed lost in eld-4 clones (Extended Data Fig. 1c), but ectopic expression of Sas within eld-4 clones surrounding scrib−/− clones reversed the elimination-defective phenotype (Fig. 1k–m). Moreover, the knockdown of Sas in cells surrounding scrib−/− clones phenocopied the elimination-defective phenotype (Fig. 1n); a similar elimination-defective phenotype also occurred upon Sas knockdown in cells surrounding dlg−/− mutant eye-disc clones (Extended Data Fig. 1d–l). These data reveal that the cell-surface ligand Sas is required for normal epithelial cells to eliminate neighbouring polarity-deficient cells.
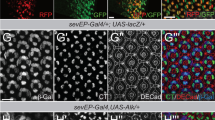
We next sought to understand the mechanism by which Sas drives the elimination of nearby cells. Sas is normally localized at the apical surface of epithelial cells10 (Extended Data Fig. 2a). Notably, however, we found that Sas relocalized to the lateral cell surface specifically at the interface between wild-type and scrib−/− or dlg−/− clones (Fig. 2a–c and Extended Data Fig. 2b). This relocalization of Sas at the clone interface was also observed between wild-type and scrib−/− sas−/− double-mutant clones (Fig. 2d–f), indicating that the Sas protein that accumulates at the clone interface is derived from surrounding wild-type cells.
a–c, Top images show xy confocal sections of eye disc bearing GFP-labelled scrib−/− clones immunostained for Sas and with DAPI; bottom images show xz cross sections. d–f, Top images show xy confocal section of eye disc bearing GFP-labelled scrib−/− sas−/− double-mutant clones immunostained with anti-Sas and DAPI; bottom shows xz cross sections. Dashed lines in the top images of c and f mark the positions of the cross-sections in the bottom images. Scale bars, 20 μm.
The fact that normal epithelial cells relocalize Sas laterally to eliminate neighbouring oncogenic cells suggests that normal cells transmit a signal to these cells through a cell-surface receptor for Sas. We therefore sought to identify the Sas receptor expressed in polarity-deficient cells. It has been reported that PTP10D, a receptor-type tyrosine phosphatase (RPTP), interacts and functions with Sas during longitudinal axon guidance in the Drosophila nervous system and that Sas–PTP10D trans-signalling occurs through glial-neuronal communication11. We therefore assumed that PTP10D and/or other RPTPs were strong candidates for the Sas receptor in the imaginal epithelium. Given that two extracellular domains of Sas, VWC and FN3, can form homophilic interactions with the same domains of other proteins and that FN3 is a domain commonly shared by RPTPs, we screened 32 RNA interference (RNAi) fly strains that target expression of Drosophila transmembrane proteins bearing either VWC or FN3 domains (Extended Data Fig. 3a). We found that only one RNAi line targeting PTP10D phenocopied the severe elimination-defective and melanization phenotypes when expressed within scrib−/− or dlg−/− mutant clones (Fig. 3b–e and Extended Data Fig. 3b, c). Like Sas, PTP10D was relocalized to the interface between scrib−/− and wild-type clones (Fig. 3f–i), whereas it normally localized at the apical surface of epithelial cells (Extended Data Fig. 2a). This lateral accumulation of PTP10D was almost eliminated when PTP10D-RNAi was expressed within scrib−/− clones (Fig. 3j–m), indicating that the PTP10D accumulating at the clone interface derives from scrib−/− mutant cells. Furthermore, immunostaining analysis of scrib−/−sas−/− double-mutant clones indicated that Sas and PTP10D are localized adjacent to each other in neighbouring cells (Extended Data Fig. 4a, b). Notably, the lateral relocalization of Sas and PTP10D at the clone interface was also observed for the neoplastic non-functional tumour-suppressor mutants vps25−/− (ref. 12), erupted−/− (ref. 13), or Rab5DN-expressing cells14, all of which are eliminated as losers of cell competition when surrounded by wild-type cells; however, such relocalization was not observed for non-neoplastic polarity stardust−/− or crumbs−/− mutants (Extended Data Fig. 5). These data suggest that in response to the emergence of neoplastic polarity-deficient cells, adjacent normal cells relocalize Sas laterally whereas nearby polarity-deficient cells relocalize PTP10D laterally, thereby driving elimination of polarity-deficient cells through trans-activated Sas–PTP10D signalling.
a, A schematic representation of the general domain structure of PTP10D. PTP10D is an RPTP bearing extracellular FN3 domains, a cytoplasmic tyrosine phosphatase (PTP) domain, and a transmembrane domain. b, c, Eye disc bearing MARCM-induced mosaics of GFP-labelled scrib−/− (b) or scrib−/−Ptp10D-RNAi (c) clones. d, Pupal eye of MARCM-induced mosaics of scrib−/−Ptp10D-RNAi clones. e, Quantification of relative size of clones in genotypes shown in b (n = 19 eye discs) and c (n = 22). Mean ± s.d.; **P < 0.01 by Mann–Whitney U-test. f–i, Top images show xy confocal section of eye disc bearing GFP-labelled scrib−/− clones immunostained with anti-PTP10D (g), anti-Sas (h), and DAPI; bottom images show xz cross sections. j–m, Top images show xy confocal section of eye disc bearing GFP-labelled scrib−/−Ptp10D-RNAi clones immunostained with anti-PTP10D (k), anti-Sas (l), and DAPI; bottom images show xz cross sections. Dashed lines in the top images of i and m mark the positions of the cross-sections in the bottom images. Note that the staining signal of lateral PTP10D in surrounding wild-type cells was decreased at the clone boundary. Scale bars, 20 μm.
We next sought to investigate the mechanism by which Sas–PTP10D signalling drives elimination of polarity-deficient cells. It has previously been shown that the activation of Eiger–JNK signalling in polarity-deficient cells is essential for their elimination1,2,5. Therefore, a possible mechanism by which PTP10D knockdown in scrib−/− clones results in an elimination-defective phenotype is through inhibition of JNK signalling. However, JNK signalling was still strongly activated in scrib−/− clones expressing PTP10D-RNAi, as assessed by the JNK target MMP1 (ref. 15) (Fig. 4a, and Extended Data Fig. 6a). This indicates that loss of PTP10D drives one or more intracellular signalling events that cause an elimination-defective phenotype in the presence of JNK activation. A strong candidate for this signalling event is activation of Ras signalling, as JNK is converted from pro-apoptotic to pro-growth in the presence of Ras activation16. Notably, it has been reported that PTP10D and its mammalian orthologue PTPRJ (also known as DEP1/CD148/SCC1/RPTPeta) negatively regulate epidermal growth factor receptor (EGFR) signalling by directly dephosphorylating the intracellular tyrosine kinase domain of EGFR17,18,19.We found that EGFR normally localizes apically in wild-type cells but relocalizes to the lateral surface together with PTP10D at the boundaries between scrib−/− and wild-type clones (Extended Data Fig. 7). More pertinently, EGFR–Ras signalling was strongly elevated in scrib−/− clones expressing PTP10D-RNAi, as assessed by downregulation of the transcription factor Capicua20 (Fig. 4b, c). Moreover, co-knockdown of EGFR and PTP10D in scrib−/− clones completely reversed the elimination-defective phenotype (Fig. 4d), with EGFR-RNAi alone having only a slight effect on the growth of normal tissue (Extended Data Fig. 6b–d). Furthermore, expression of a constitutively active form of EGFR or Ras caused overgrowth of scrib−/− clones (Extended Data Fig. 8a, b), while expression of dominant-negative form of Ras in scrib−/−PTP10D-RNAi clones strongly suppressed their growth (Extended Data Fig. 8c). Thus, scrib clones in the absence of PTP10D signalling activate both JNK and Ras signalling and overgrow in a manner dependent on EGFR signalling. The co-activation of EGFR–Ras and Eiger–JNK signalling causes hyper-accumulation of intracellular F-actin, thereby inactivating the tumour-suppressor Hippo pathway16,21. Inactivation of the Hippo pathway triggers nuclear translocation and activation of the downstream transcriptional co-activator Yorkie (Yki), which induces upregulation of various pro-growth and anti-apoptotic genes22. Indeed, scrib−/− clones expressing PTP10D-RNAi strongly accumulated intracellular F-actin (Extended Data Fig. 6f) and showed strong upregulation of the Yki target gene expanded (ex) (Fig. 4f), as well as an increased nuclear signal of Yki protein (Extended Data Fig. 6g); however, scrib mutation alone only slightly upregulated F-actin (Extended Data Fig. 6e) and ex expression (Fig. 4e). Furthermore, inhibition of Yki activity by the Yki kinase Warts (Wts) or Yki-RNAi significantly suppressed growth of scrib−/− clones in the absence of PTP10D (Fig. 4g, h and Extended Data Fig. 6h), while Wts-overexpression or Yki-RNAi alone had little effect on tissue growth (Extended Data Fig. 6d, i, j). Similar upregulations of EGFR signalling and Yki activity were observed in scrib−/− clones when surrounded by sas−/− eld-4 clones (Extended Data Fig. 9a–f). Finally, we found that the number of dying cells at the boundaries between scrib−/− and wild-type clones was significantly reduced by PTP10D-knockdown (Fig. 4i–l), whereas cell proliferation was significantly increased in scrib−/− clones expressing PTP10D-RNAi (Fig. 4m–p). Together, these data indicate that when neoplastic polarity-deficient cells emerge in the epithelium, neighbouring non-neoplastic cells restrain EGFR signalling of nearby polarity-deficient cells through a Sas–PTP10D trans-interaction, which enables JNK signalling activated in polarity-deficient cells to drive cell elimination (Fig. 4q). In the absence of Sas–PTP10D, elevated EGFR–Ras signalling in polarity-deficient cells cooperates with JNK signalling to cause Yki activation, thereby leading to an elimination defect and overgrowth of polarity-deficient cells (Fig. 4r).
a–c, Right, eye disc bearing GFP-labelled scrib−/−Ptp10D-RNAi (a, c) and scrib−/− (b), or left, immunostained for MMP1 (a) or Capicua (b, c). d, Eye disc with GFP-labelled scrib−/−Ptp10D-RNAi Egfr-RNAi. e, f, Left, eye disc of an ex-lacZ/+ fly bearing GFP-labelled scrib−/− (e) or scrib−/−Ptp10D-RNAi (f) clones. Right, the same images immunostained for β-galactosidase. Endogenous ex–lacZ expression is indicated by asterisks. g, Eye disc bearing GFP-labelled scrib−/−Ptp10D-RNAi Wts clones. h, Quantification of relative size of GFP-labelled clones in genotypes shown in b (n = 19, number of eye discs), a (n = 22), d (n = 16), and g (n = 13). Mean ± s.d.; **P < 0.01 by Mann–Whitney U-test. i–k, Eye disc bearing GFP-labelled wild-type (i), scrib−/− (j), or scrib−/−Ptp10D-RNAi (k) clones immunostained for Dcp-1. Arrows indicate representative apoptotic cells at the clone boundaries. l, Quantification of the number of dying cells per clone perimeter in genotypes shown in i (n = 20 clones), j (n = 19), and k (n = 21). Mean ± s.d.; **P < 0.01 by Mann–Whitney U-test. m–o, Eye disc bearing GFP-labelled wild-type (m), scrib−/− (n), or scrib−/−Ptp10D-RNAi (o) clones immunostained with EdU-labelling and anti-GFP. p, Quantification of the number of EdU-labelled cells per clone in genotypes shown in m (n = 21 clones), (n) (n = 30), and (o) (n = 30). Mean ± s.d.; **P < 0.01 by Mann–Whitney U-test. q, r, A model for tumour-suppressive cell competition that eliminates oncogenic polarity-deficient cells via trans-interaction of Sas–PTP10D. Scale bars, 20 μm.
Our data indicate that in response to the emergence of oncogenic polarity-deficient cells, Sas and PTP10D relocalize specifically at the clone interface to the respective lateral surfaces of normal or polarity-deficient cells, enabling the ligand and receptor to interact with each other in trans. Thus, Sas–PTP10D acts as a fail-safe system for epithelial tissue, a system that protects against neoplastic development and is normally latent but activates upon oncogenic cell emergence. Notably, the Sas–PTP10D system was not required for other types of cell competition triggered by Minute8, Mahjong23, Myc24,25 or Yki26 (data not shown). Although the mechanism by which Sas and PTP10D relocalize to the clone interface is currently unknown, we found that the apical proteins Bazooka, Patj, and aPKC and the sub-apical protein E-cadherin also relocalize to the lateral surface of the clone boundary (Extended Data Fig. 10). This suggests that the apical cell surface expands to the lateral region at the clone boundary, meaning that Sas and PTP10D meet each other in trans at the clone interface.
Our genetic data reveal that Sas and PTP10D act together as tumour suppressors during cell competition. Previous studies have reported that PTPRJ, the mammalian homologue of PTP10D, also acts as a tumour suppressor27 and negatively regulates EGFR signalling19. Although no obvious homologues of Sas have been identified in mammals, thrombospondin-1 and syndecan-2 have been reported to act as ligands for PTPRJ28,29. Given that elimination of scrib-deficient cells by cell competition also occurs in mammalian systems30, and that the signalling mechanisms identified in Drosophila are evolutionarily conserved, similar cell–cell recognition mechanisms may help to safeguard human tissues against tumorigenesis.
Methods
Fly strains and generation of clones
Fluorescently labelled mitotic clones9,31 were produced in larval imaginal discs using the following strains: y,w, eyFLP1; FRT82B, Ubi-GFP, scrib1/TM6B (CC tester), y,w, eyFLP1;Act>y+>Gal4, UAS–GFP; FRT82B, Tub-Gal80 (82B tester-1), eyFLP5, Act>y+>Gal4, UAS-GFP; FRT82B, TubGal80 (82B tester-2), UAS-Dicer2; eyFLP5, Act>y+>Gal4, UAS-GFP; FRT82B, TubGal80 (82B tester-3), w, Tub-Gal80, FRT19A; eyFLP5, Act>y+>Gal4, UAS-GFP (19A tester). Additional strains used are as follows: scrib1 (from D. Bilder)32, dlgm52 (from S. Goode)33, UAS-sas (from K. Zinn)11, UAS-wts (from T. Xu), UAS-Ptp10D-RNAi39001 and ex697 (ex-lacZ) (Bloomington Stock Center), and UAS-sas-RNAi39086, UAS-yki-RNAi104523, and UAS-EGFR-RNAi43268 (Vienna Drosophila Resource Center). To measure clone size, ImageJ (Fiji) software was used to automatically or manually determine the threshold of the fluorescence. Relative sizes of clones in the eye-antennal disc were calculated using ImageJ and Microsoft Excel software. At least 12 individual eye-antennal discs of each genotype were analysed.
Genetic mosaic screen for elimination-defective mutants
Male flies carrying an isogenized FRT82B chromosome (w/Y; FRT82B) were fed 25 mM EMS and then mated to w; Sb/TM6B females. Single F1 males of the genotype FRT82B*/TM6B or Sb were each crossed to 4–5 females of the genotype y,w, eyFLP1; FRT82B, Ubi-GFP, scrib1/TM6B (CC tester). Clone areas of red-pigmented scrib−/− cells compared to that of white wild-type cells in the F2 eyes (non-TM6B and non-Sb adult flies) were analysed for cell competition. Mutant chromosomes were then recovered by crossing F2 males of the genotype y,w, eyFLP1/Y; FRT82B*/TM6B to w; Sb/TM6B females.
Histology
Larval tissues were stained using standard immunohistochemical procedures with rabbit anti-Sas polyclonal antibody (1:1,000; from D. Cavener), mouse anti-PTP10D monoclonal antibody (1:500, Developmental Studies Hybridoma Bank (DSHB)), mouse anti-MMP1 (1:100, from cocktail of 3A6B4, 3B8D12 and 5H7B11, DSHB), guinea-pig anti-Capicua polyclonal antibody (1:300, from I. Hariharan), rabbit anti-β-galactosidase antibody (1:1,000, MP Biomedicals), rabbit anti-Yki antibody (1:1,000; from K. Irvine), anti-cleaved Drosophila Dcp-1 (Asp216) antibody (1:100, Cell Signaling Technology), Alexa 546- or 647- conjugated phalloidin (1:50, Molecular Probes), and were mounted with 49,6-diamidino-2-phenylindole (DAPI)-containing SlowFade Gold Antifade Reagent (Molecular Probes). Images were taken with a Leica SP5 or SP8 confocal microscope.
Statistical analysis
No statistical methods were used to predetermine sample size. All n numbers represent biological replicates. Each experiment was independently performed at least three times. Experiments were not randomized or blinded. Mann–Whitney non-parametric tests were used for all experiments and differences were considered significant at P < 0.05. All data in bar graphs are expressed as mean ± s.d.
Data availability
Source data for Fig. 1i, 3e, 4h, 4l and 4p are provided with the paper. All data are available from the authors upon reasonable request.
References
Brumby, A. M. & Richardson, H. E. scribble mutants cooperate with oncogenic Ras or Notch to cause neoplastic overgrowth in Drosophila. EMBO J. 22, 5769–5779 (2003)
Igaki, T., Pastor-Pareja, J. C., Aonuma, H., Miura, M. & Xu, T. Intrinsic tumor suppression and epithelial maintenance by endocytic activation of Eiger/TNF signaling in Drosophila. Dev. Cell 16, 458–465 (2009)
Ohsawa, S. et al. Elimination of oncogenic neighbors by JNK-mediated engulfment in Drosophila. Dev. Cell 20, 315–328 (2011)
Amoyel, M. & Bach, E. A. Cell competition: how to eliminate your neighbours. Development 141, 988–1000 (2014)
Andersen, D. S. et al. The Drosophila TNF receptor Grindelwald couples loss of cell polarity and neoplastic growth. Nature 522, 482–486 (2015)
Hogan, C. et al. Characterization of the interface between normal and transformed epithelial cells. Nat. Cell Biol. 11, 460–467 (2009)
Kajita, M. et al. Interaction with surrounding normal epithelial cells influences signalling pathways and behaviour of Src-transformed cells. J. Cell Sci. 123, 171–180 (2010)
Morata, G. & Ripoll, P. Minutes: mutants of Drosophila autonomously affecting cell division rate. Dev. Biol. 42, 211–221 (1975)
Xu, T. & Rubin, G. M. Analysis of genetic mosaics in developing and adult Drosophila tissues. Development 117, 1223–1237 (1993)
Schonbaum, C. P., Organ, E. L., Qu, S. & Cavener, D. R. The Drosophila melanogaster stranded at second (sas) gene encodes a putative epidermal cell surface receptor required for larval development. Dev. Biol. 151, 431–445 (1992)
Lee, H. K., Cording, A., Vielmetter, J. & Zinn, K. Interactions between a receptor tyrosine phosphatase and a cell surface ligand regulate axon guidance and glial-neuronal communication. Neuron 78, 813–826 (2013)
Thompson, B. J. et al. Tumor suppressor properties of the ESCRT-II complex component Vps25 in Drosophila. Dev. Cell 9, 711–720 (2005)
Moberg, K. H., Schelble, S., Burdick, S. K. & Hariharan, I. K. Mutations in erupted, the Drosophila ortholog of mammalian tumor susceptibility gene 101, elicit non-cell-autonomous overgrowth. Dev. Cell 9, 699–710 (2005)
Ballesteros-Arias, L., Saavedra, V. & Morata, G. Cell competition may function either as tumour-suppressing or as tumour-stimulating factor in Drosophila. Oncogene 33, 4377–4384 (2014)
Uhlirova, M. & Bohmann, D. JNK- and Fos-regulated Mmp1 expression cooperates with Ras to induce invasive tumors in Drosophila. EMBO J. 25, 5294–5304 (2006)
Enomoto, M., Kizawa, D., Ohsawa, S. & Igaki, T. JNK signaling is converted from anti- to pro-tumor pathway by Ras-mediated switch of Warts activity. Dev. Biol. 403, 162–171 (2015)
Jeon, M. & Zinn, K. Receptor tyrosine phosphatases control tracheal tube geometries through negative regulation of Egfr signaling. Development 136, 3121–3129 (2009)
Jeon, M. & Zinn, K. R3 receptor tyrosine phosphatases: conserved regulators of receptor tyrosine kinase signaling and tubular organ development. Semin. Cell Dev. Biol. 37, 119–126 (2015)
Tarcic, G. et al. An unbiased screen identifies DEP-1 tumor suppressor as a phosphatase controlling EGFR endocytosis. Curr. Biol. 19, 1788–1798 (2009)
Tseng, A. S. et al. Capicua regulates cell proliferation downstream of the receptor tyrosine kinase/Ras signaling pathway. Curr. Biol. 17, 728–733 (2007)
Ohsawa, S. et al. Mitochondrial defect drives non-autonomous tumour progression through Hippo signalling in Drosophila. Nature 490, 547–551 (2012)
Pan, D. The Hippo signaling pathway in development and cancer. Dev. Cell 19, 491–505 (2010)
Tamori, Y. et al. Involvement of Lgl and Mahjong/VprBP in cell competition. PLoS Biol. 8, e1000422 (2010)
de la Cova, C., Abril, M., Bellosta, P., Gallant, P. & Johnston, L. A. Drosophila Myc regulates organ size by inducing cell competition. Cell 117, 107–116 (2004)
Moreno, E. & Basler, K. dMyc transforms cells into super-competitors. Cell 117, 117–129 (2004)
Tyler, D. M., Li, W., Zhuo, N., Pellock, B. & Baker, N. E. Genes affecting cell competition in Drosophila. Genetics 175, 643–657 (2007)
Hendriks, W. J. A. J. & Pulido, R. Protein tyrosine phosphatase variants in human hereditary disorders and disease susceptibilities. Biochim. Biophys. Acta 1832, 1673–1696 (2013)
Takahashi, K. et al. Thrombospondin-1 acts as a ligand for CD148 tyrosine phosphatase. Proc. Natl Acad. Sci. USA 109, 1985–1990 (2012)
Whiteford, J. R. et al. Syndecan-2 is a novel ligand for the protein tyrosine phosphatase receptor CD148. Mol. Biol. Cell 22, 3609–3624 (2011)
Norman, M. et al. Loss of Scribble causes cell competition in mammalian cells. J. Cell Sci. 125, 59–66 (2012)
Lee, T. & Luo, L. Mosaic analysis with a repressible cell marker for studies of gene function in neuronal morphogenesis. Neuron 22, 451–461 (1999)
Bilder, D., Li, M. & Perrimon, N. Cooperative regulation of cell polarity and growth by Drosophila tumor suppressors. Science 289, 113–116 (2000)
Goode, S. & Perrimon, N. Inhibition of patterned cell shape change and cell invasion by Discs large during Drosophila oogenesis. Genes Dev. 11, 2532–2544 (1997)
Acknowledgements
We thank J. Vaughen for comments on the manuscript; Y. Matsumoto, T. Sawada, K. Takino, M. Katsukawa, and C. Iida for technical support; D. Cavener, I. Hariharan, and K. Irvine for antibodies; D. Bilder, S. Goode, T. Xu, K. Zinn, the Bloomington Drosophila Stock Center (Indiana, USA), the Vienna Drosophila Resource Center (Vienna, Austria), the NIG Stock Center (Mishima, Japan), and the Drosophila Genomics and Genetic Resources (Kyoto Stock Center, Japan) for fly stocks. This work was supported in part by grants from the MEXT/JSPS KAKENHI (grant number 26114002, 25112710, 15H05862, and 23127508) to S.O. and T.I., the Nakajima Foundation to S.O., the Inoue Science Research Award to S.O, the Naito Foundation to T.I, the Takeda Science Foundation to S.O. and T.I., and the JST to T.I. M.Y. was supported by the Platform for Dynamic Approaches to Living System from AMED.
Author information
Authors and Affiliations
Contributions
T.I. designed the screen; S.O. and K.K. conducted the screen; M.Y., S.O., K.K. and T.I. designed subsequent experiments; M.Y., S.O. and K.K. performed all experiments; M.Y., S.O., K.K. and T.I. analysed the data; and M.Y., S.O. and T.I. wrote the manuscript.
Corresponding author
Additional information
Reviewer Information Nature thanks G. Morata, K. Zinn and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Figure 1 Sas is required for normal cells to eliminate polarity-deficient neighbours.
a, A ‘non-autonomous’ genetic mosaic screen for mutants that fail to eliminate polarity-deficient cells from the eye imaginal epithelium. scrib−/− clones are normally eliminated by cell competition when surrounded by wild-type tissue. Using the genetic mosaic technique, EMS-induced mutations were introduced only in wild-type cells in scrib−/− mosaic eye discs (see Methods for details). The elimination-defective phenotype was assessed by red-pigmented clones or melanization generated in the adult eye. b, Mapping of a mutation causing the elimination-defective phenotype. Using a series of chromosomal deficiency fly lines, the responsible genomic region was narrowed down to 84C8–84D2 based on lethality with eld-4. c, Eye-antennal-disc-bearing GFP-labelled eld-4 clones immunostained for Sas and DAPI. d–g, Adult eyes of MARCM-induced mosaics of wt//wt (d), wt//dlg−/− (e), dlg−/−//sas-RNAi (f), and wt//sas-RNAi (g) clones. h–k, Eye discs of MARCM-induced mosaics of wt//wt (g), wt//dlg−/− (h), dlg−/−//sas-RNAi (i), and wt//sas-RNAi (j) clones. l, Quantification of relative size of clones in genotypes shown in h (n = 13, number of eye discs), i (n = 14), j (n = 13), k (n = 15). Error bars show s.d.; ns, not significant (P > 0.05); **P < 0.01 by Mann–Whitney U-test. Scale bars, 20 μm.
Extended Data Figure 2 Sas accumulates at the interface between normal and dlg−/− cells.
a, b, xy (upper panels) or xz (lower panels) confocal sections of eye disc bearing GFP-labelled wild-type (a) or dlg−/− (b) clones immunostained for Sas (a’, b’) and anti-PTP10D (a’’, b’’). Scale bars, 20 μm.
Extended Data Figure 3 In vivo RNAi screen to identify the Sas receptor required for elimination of polarity-deficient cells.
a, In vivo RNAi screen schematic for identifying the Sas receptor. As the VWC and FN3 domains of Sas can bind homophilically, RNAi against 32 Drosophila transmembrane proteins containing either VWC or FN3 domains was driven individually inside scrib−/− clones and assessed for overgrowth in adult eyes. b, c, Adult eyes of MARCM-induced GFP-labelled mosaics of scrib−/− clones (b, as a control) simultaneously expressing each candidate RNAi (c).
Extended Data Figure 4 Sas and PTP10D are localized adjacent to each other in neighbouring cells.
a, b, xz sections of the confocal image of GFP-labelled scrlb−/− mutant clones (a, cyan) or scrib−/− sas−/− double mutant clones (b, cyan) immunostained for PTP10D (green) and Sas (magenta). Fluorescent intensities of Sas and PTP10D are measured by imageJ software at the yellow lines (a’ and b’).
Extended Data Figure 5 Sas and PTP10D localize at the lateral interface between normal and neoplastic tumour-suppressor mutants but not non-oncogenic polarity mutants.
a–e, xz sections of the confocal image of GFP-labelled crb−/− (a), sdt−/− (b), RabDN (c), ept−/− (d), and vps25−/− (e) clones immunostained with anti-Sas (a’–e’), anti-PTP10D (a’’–e’’), and merged image (a’’’–e’’’) are shown.
Extended Data Figure 6 Sas–PTP10D drives elimination of polarity-deficient cells by modulating EGFR and Hippo signalling.
a–c, Eye disc bearing GFP-labelled scrib−/− (a), wild-type (b), or Egfr-RNAi (c) clones; d, Quantification of relative size of clones in genotypes shown in b (n = 15, number of eye discs), c (n = 12), i (n = 14), j (n = 14). Error bars show s.d.; ns, not significant (P > 0.05); *P < 0.05 by Mann–Whitney U-test. e, f, Eye disc bearing GFP-labelled scrib−/− (e) or scrib−/− and Ptp10D-RNAi (f) clones stained with phalloidin. g, Eye disc bearing GFP-labelled scrib−/− + Ptp10D-RNAi clones stained for Yki and with DAPI. h–j, Eye disc bearing GFP-labelled scrib−/− + Ptp10D-RNAi + yki-RNAi (h), yki-RNAi (i), or Wts-overexpressing (j) clones. Egfr-RNAi or yki-RNAi did not reduce clone size, perhaps owing to incomplete knockdown. Scale bars, 20 μm.
Extended Data Figure 7 EGFR and PTP10D relocalize from the apical to lateral cell surface at the boundary between scrib clones and wild-type clones.
a–d, xz section of the confocal image of GFP-labelled scrlb−/− clones (a) immunostained for EGFR (b), PTP10D (c), and merged image (d) are shown.
Extended Data Figure 8 Expression of EGFRCA or RasV12 abolishes scrib cell elimination while expression of RasDN in scrib + PTP10D-RNAi clones suppresses their growth.
a–c, Eye disc bearing GFP-labelled MARCM clones with scrib−/− + EGFRCA(a), scrib−/− + RasV12 (b), or scrib−/− + PTP10D-RNAi + UAS-RasDN (c) is shown.
Extended Data Figure 9 scrib clones surrounded by sas clones activate EGFR–Ras signalling and Yki.
a, b, Eye disc bearing GFP-labelled scrlb−/− clones surrounded by GFP-negative wild-type (a–a’’) or sas−/− eld-4 (b–b’’) clones stained with anti-Capicua. Arrows represent typical scrib mutant clones that downregulate Cic (b-b’’) or upregulate ex-lacZ (d-d’’) expression. c, d, Eye disc bearing GFP-labelled scrlb−/− clones surrounded by GFP-negative wild-type (c–c’’) or sas−/− eld-4 (d–d’’) clones immunostained for β-galactosidase to label the expanded (ex)–lacZ reporter. e, f, Eye disc bearing GFP-labelled scrlb−/− clones surrounded by GFP-negative wild-type (e–e’’) or sas−/− eld-4 (f–f’’) clones stained with anti-Yorkie (Yki).
Extended Data Figure 10 Apical and sub-apical proteins are localized at the lateral interface between wild-type and scrib−/− clones.
xz sections of the confocal image of GFP-labelled scrlb−/− clones immunostained for Baz (a), Patj (b), aPKC/Sas (c), and E-cadherin/Sas (d).
Supplementary information
Supplementary Information
This file contains a list of detailed genotypes. (PDF 162 kb)
Rights and permissions
About this article
Cite this article
Yamamoto, M., Ohsawa, S., Kunimasa, K. et al. The ligand Sas and its receptor PTP10D drive tumour-suppressive cell competition. Nature 542, 246–250 (2017). https://doi.org/10.1038/nature21033
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature21033
This article is cited by
-
Cell competition and cancer from Drosophila to mammals
Oncogenesis (2024)
-
Cell competition in development, homeostasis and cancer
Nature Reviews Molecular Cell Biology (2023)
-
A mathematical model with aberrant growth correction in tissue homeostasis and tumor cell growth
Journal of Mathematical Biology (2023)
-
Pleiotropic effects of cell competition between normal and transformed cells in mammalian cancers
Journal of Cancer Research and Clinical Oncology (2023)
-
WASH activation controls endosomal recycling and EGFR and Hippo signaling during tumor-suppressive cell competition
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.