Abstract
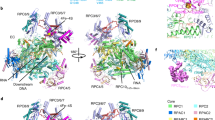
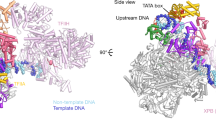
RNA polymerase (Pol) II produces messenger RNA during transcription of protein-coding genes in all eukaryotic cells. The Pol II structure is known at high resolution from X-ray crystallography for two yeast species1,2,3. Structural studies of mammalian Pol II, however, remain limited to low-resolution electron microscopy analysis of human Pol II and its complexes with various proteins4,5,6,7,8,9,10. Here we report the 3.4 Å resolution cryo-electron microscopy structure of mammalian Pol II in the form of a transcribing complex comprising DNA template and RNA transcript. We use bovine Pol II, which is identical to the human enzyme except for seven amino-acid residues. The obtained atomic model closely resembles its yeast counterpart, but also reveals unknown features. Binding of nucleic acids to the polymerase involves ‘induced fit’ of the mobile Pol II clamp and active centre region. DNA downstream of the transcription bubble contacts a conserved ‘TPSA motif’ in the jaw domain of the Pol II subunit RPB5, an interaction that is apparently already established during transcription initiation7. Upstream DNA emanates from the active centre cleft at an angle of approximately 105° with respect to downstream DNA. This position of upstream DNA allows for binding of the general transcription elongation factor DSIF (SPT4–SPT5) that we localize over the active centre cleft in a conserved position on the clamp domain of Pol II. Our results define the structure of mammalian Pol II in its functional state, indicate that previous crystallographic analysis of yeast Pol II is relevant for understanding gene transcription in all eukaryotes, and provide a starting point for a mechanistic analysis of human transcription.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Primary accessions
Electron Microscopy Data Bank
Protein Data Bank
Data deposits
Cryo-electron microscopy densities have been deposited in the Electron Microscopy Data Bank under accession numbers EMD-3218 (EC1), EMD-3219 (EC2), and EMD-3220 (EC3). Model coordinates have been deposited in the Protein Data Bank (PDB) under accession number 5FLM.
References
Cramer, P. et al. Architecture of RNA polymerase II and implications for the transcription mechanism. Science 288, 640–649 (2000)
Cramer, P., Bushnell, D. A. & Kornberg, R. D. Structural basis of transcription: RNA polymerase II at 2.8 angstrom resolution. Science 292, 1863–1876 (2001)
Spåhr, H., Calero, G., Bushnell, D. A. & Kornberg, R. D. Schizosacharomyces pombe RNA polymerase II at 3.6-Å resolution. Proc. Natl Acad. Sci. USA 106, 9185–9190 (2009)
Kostek, S. A. et al. Molecular architecture and conformational flexibility of human RNA polymerase II. Structure 14, 1691–1700 (2006)
Kassube, S. A., Jinek, M., Fang, J., Tsutakawa, S. & Nogales, E. Structural mimicry in transcription regulation of human RNA polymerase II by the DNA helicase RECQL5. Nature Struct. Mol. Biol. 20, 892–899 (2013)
Kassube, S. A. et al. Structural insights into transcriptional repression by noncoding RNAs that bind to human Pol II. J. Mol. Biol. 425, 3639–3648 (2013)
He, Y., Fang, J., Taatjes, D. J. & Nogales, E. Structural visualization of key steps in human transcription initiation. Nature 495, 481–486 (2013)
Bernecky, C., Grob, P., Ebmeier, C. C., Nogales, E. & Taatjes, D. J. Molecular architecture of the human Mediator–RNA polymerase II–TFIIF assembly. PLoS Biol. 9, e1000603 (2011)
Bernecky, C. & Taatjes, D. J. Activator–mediator binding stabilizes RNA polymerase II orientation within the human mediator–RNA polymerase II–TFIIF assembly. J. Mol. Biol. 417, 387–394 (2012)
Wu, Y. M. et al. Regulation of mammalian transcription by Gdown1 through a novel steric crosstalk revealed by cryo-EM. EMBO J. 31, 3575–3587 (2012)
Hu, X. et al. A Mediator-responsive form of metazoan RNA polymerase II. Proc. Natl Acad. Sci. USA 103, 9506–9511 (2006)
Gnatt, A. L., Cramer, P., Fu, J., Bushnell, D. A. & Kornberg, R. D. Structural basis of transcription: an RNA polymerase II elongation complex at 3.3 A resolution. Science 292, 1876–1882 (2001)
Armache, K. J., Kettenberger, H. & Cramer, P. Architecture of initiation-competent 12-subunit RNA polymerase II. Proc. Natl Acad. Sci. USA 100, 6964–6968 (2003)
Bushnell, D. A. & Kornberg, R. D. Complete, 12-subunit RNA polymerase II at 4.1-A resolution: implications for the initiation of transcription. Proc. Natl Acad. Sci. USA 100, 6969–6973 (2003)
Plaschka, C. et al. Architecture of the RNA polymerase II–Mediator core initiation complex. Nature 518, 376–380 (2015)
Chang, L., Zhang, Z., Yang, J., McLaughlin, S. H. & Barford, D. Atomic structure of the APC/C and its mechanism of protein ubiquitination. Nature 522, 450–454 (2015)
Liang, B. et al. Structure of the L protein of vesicular stomatitis virus from electron cryomicroscopy. Cell 162, 314–327 (2015)
Urnavicius, L. et al. The structure of the dynactin complex and its interaction with dynein. Science 347, 1441–1446 (2015)
Bai, X. C. et al. An atomic structure of human γ-secretase. Nature 525, 212–217 (2015)
Kettenberger, H., Armache, K. J. & Cramer, P. Complete RNA polymerase II elongation complex structure and its interactions with NTP and TFIIS. Mol. Cell 16, 955–965 (2004)
Martinez-Rucobo, F. W. & Cramer, P. Structural basis of transcription elongation. Biochim. Biophys. Acta 1829, 9–19 (2013)
Barnes, C. O. et al. Crystal structure of a transcribing RNA polymerase II complex reveals a complete transcription bubble. Mol. Cell 59, 258–269 (2015)
Fishburn, J., Tomko, E., Galburt, E. & Hahn, S. Double-stranded DNA translocase activity of transcription factor TFIIH and the mechanism of RNA polymerase II open complex formation. Proc. Natl Acad. Sci. USA 112, 3961–3966 (2015)
Wang, D., Bushnell, D. A., Westover, K. D., Kaplan, C. D. & Kornberg, R. D. Structural basis of transcription: role of the trigger loop in substrate specificity and catalysis. Cell 127, 941–954 (2006)
Andrecka, J. et al. Nano positioning system reveals the course of upstream and nontemplate DNA within the RNA polymerase II elongation complex. Nucleic Acids Res. 37, 5803–5809 (2009)
Martinez-Rucobo, F. W., Sainsbury, S., Cheung, A. C. & Cramer, P. Architecture of the RNA polymerase-Spt4/5 complex and basis of universal transcription processivity. EMBO J. 30, 1302–1310 (2011)
Li, W., Giles, C. & Li, S. Insights into how Spt5 functions in transcription elongation and repressing transcription coupled DNA repair. Nucleic Acids Res. 42, 7069–7083 (2014)
Hirtreiter, A. et al. Spt4/5 stimulates transcription elongation through the RNA polymerase clamp coiled-coil motif. Nucleic Acids Res. 38, 4040–4051 (2010)
Belogurov, G. A., Sevostyanova, A., Svetlov, V. & Artsimovitch, I. Functional regions of the N-terminal domain of the antiterminator RfaH. Mol. Microbiol. 76, 286–301 (2010)
Sevostyanova, A. & Artsimovitch, I. Functional analysis of Thermus thermophilus transcription factor NusG. Nucleic Acids Res. 38, 7432–7445 (2010)
Wenzel, S., Martins, B. M., Rösch, P. & Wöhrl, B. M. Crystal structure of the human transcription elongation factor DSIF hSpt4 subunit in complex with the hSpt5 dimerization interface. Biochem. J. 425, 373–380 (2010)
Korinek, A., Beck, F., Baumeister, W., Nickell, S. & Plitzko, J. M. Computer controlled cryo-electron microscopy—TOM2 a software package for high-throughput applications. J. Struct. Biol. 175, 394–405 (2011)
Li, X. et al. Electron counting and beam-induced motion correction enable near-atomic-resolution single-particle cryo-EM. Nature Methods 10, 584–590 (2013)
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012)
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015)
Tang, G. et al. EMAN2: an extensible image processing suite for electron microscopy. J. Struct. Biol. 157, 38–46 (2007)
Scheres, S. H. Semi-automated selection of cryo-EM particles in RELION-1.3. J. Struct. Biol. 189, 114–122 (2015)
Scheres, S. H. Beam-induced motion correction for sub-megadalton cryo-EM particles. eLife 3, e03665 (2014)
Rosenthal, P. B. & Henderson, R. Optimal determination of particle orientation, absolute hand, and contrast loss in single-particle electron cryomicroscopy. J. Mol. Biol. 333, 721–745 (2003)
Scheres, S. H. & Chen, S. Prevention of overfitting in cryo-EM structure determination. Nature Methods 9, 853–854 (2012)
Chen, S. et al. High-resolution noise substitution to measure overfitting and validate resolution in 3D structure determination by single particle electron cryomicroscopy. Ultramicroscopy 135, 24–35 (2013)
Cardone, G., Heymann, J. B. & Steven, A. C. One number does not fit all: mapping local variations in resolution in cryo-EM reconstructions. J. Struct. Biol. 184, 226–236 (2013)
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004)
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539 (2011)
Sainsbury, S., Niesser, J. & Cramer, P. Structure and function of the initially transcribing RNA polymerase II–TFIIB complex. Nature 493, 437–440 (2013)
Winn, M. D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D 67, 235–242 (2011)
Stein, N. CHAINSAW: a program for mutating pdb files used as templates in molecular replacement. J. Appl. Crystallogr. 41, 641–643 (2008)
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010)
Meka, H., Werner, F., Cordell, S. C., Onesti, S. & Brick, P. Crystal structure and RNA binding of the Rpb4/Rpb7 subunits of human RNA polymerase II. Nucleic Acids Res. 33, 6435–6444 (2005)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010)
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010)
Barad, B. A. et al. EMRinger: side chain-directed model and map validation for 3D cryo-electron microscopy. Nature Methods 12, 943–946 (2015)
Cheng, B. & Price, D. H. Analysis of factor interactions with RNA polymerase II elongation complexes using a new electrophoretic mobility shift assay. Nucleic Acids Res. 36, e135 (2008)
Martinez-Rucobo, F. W. et al. Molecular basis of transcription-coupled pre-mRNA capping. Mol. Cell 58, 1079–1089 (2015)
Herzog, F. et al. Structural probing of a protein phosphatase 2A network by chemical cross-linking and mass spectrometry. Science 337, 1348–1352 (2012)
Missra, A. & Gilmour, D. S. Interactions between DSIF (DRB sensitivity inducing factor), NELF (negative elongation factor), and the Drosophila RNA polymerase II transcription elongation complex. Proc. Natl Acad. Sci. USA 107, 11301–11306 (2010)
Brueckner, F., Hennecke, U., Carell, T. & Cramer, P. CPD damage recognition by transcribing RNA polymerase II. Science 315, 859–862 (2007)
Brueckner, F. & Cramer, P. Structural basis of transcription inhibition by α-amanitin and implications for RNA polymerase II translocation. Nature Struct. Mol. Biol. 15, 811–818 (2008)
Cheung, A. C. & Cramer, P. Structural basis of RNA polymerase II backtracking, arrest and reactivation. Nature 471, 249–253 (2011)
Acknowledgements
We thank E. Wolf for providing bovine thymus. We thank D. Pöllmann for preparation of human DSIF, and M. Hemann for generation of peptides for mass spectrometry. We thank J. Schuller and C. Plaschka for assistance with EM data collection. We thank D. Tegunov and C. Plaschka for discussions about EM data processing. We thank D. Haselbach and W. Liu for assistance with cryo-EM grid preparation. We thank O. Berninghausen for providing access to a Tecnai Spirit microscope. C.B. was supported by the Alexander von Humboldt Foundation. P.C. was supported by the Deutsche Forschungsgemeinschaft (SFB860), the Advanced Grant ‘TRANSIT’ of the European Research Council, and the Volkswagen Foundation.
Author information
Authors and Affiliations
Contributions
C.B. designed and performed all experiments and data analysis. F.H. supervised mass spectrometry. W.B. and J.P. provided access to a high-end EM facility. J.P. supervised EM data collection. P.C. designed and supervised research. C.B. and P.C. prepared the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
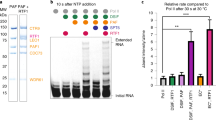
Extended Data Figure 1 Bovine Pol II purification and transcription activity.
a, SDS–polyacrylamide gel electrophoresis analysis of size-exclusion-purified 12-subunit bovine Pol II, recombinant Gdown1, and reconstituted complete Pol II with all 13 subunits (right). b, Extension of RNA by bovine Pol II. Activity assay was performed on Pol II ECs assembled on fully complementary DNA. Complexes were assembled with 12-subunit Pol II (Pol II) or 12-subunit Pol II in complex with Gdown1 (Pol II (G)). Transcription was allowed to proceed for 10 min at 30 °C in the presence of the indicated concentration of NTPs. Purified endogenous bovine Pol II is active in transcription. Approximately 40% of the RNA appears to be extended in the presence of Gdown1. c, Extension of RNA by bovine Pol II–human Gdown1 on the bubble scaffold used for structural studies. Uncrosslinked Pol II EC was incubated with the indicated amount of UTP at 30 °C for 10 min to allow the incorporation of two additional nucleotides. Approximately 30% of RNA is extended. d, Observation of individual Pol II ECs of the expected size under cryo-EM conditions with a K2 direct electron detection device. e, Fifty representative 2D class averages resulting from classification of the full Pol II EC data set.
Extended Data Figure 2 Computational sorting of cryo-EM particle images.
Particle images were sorted by 3D classification in RELION to reveal additional features of the Pol II EC. To reduce computational costs, all classifications were performed without image alignment. For global classifications, a mask encompassing the entire Pol II EC was used, whereas for local classifications masks encompassed only the area of interest as indicated in the diagram. The percentage of total data included in each class is indicated above each panel. Density was coloured to assist in visualization. For global classifications (upper left), density was coloured according to observed mobile regions. Green: clamp, anchor, part of the RPB1 cleft, part of the hybrid-binding region, and part of the active site. Orange: RPB9, RPB1 upper jaw, and part of the RPB1 cleft. Purple: RPB4–RPB7 stalk. Blue: nucleic acids. For focused classifications, density for the region of interest was coloured. This corresponded to the region within the classification mask, except for the classification of upstream DNA, in which case the density in the region of the entire nucleic-acid scaffold was coloured. The three reconstructions used for modelling the Pol II EC are shown in yellow, red, and blue boxes. The EC class displaying a mobile jaw-lobe region is highlighted with an orange box, and the EC class displaying a mobile clamp-active site region is highlighted with a green box.
Extended Data Figure 3 Quality of the cryo-EM reconstructions.
a, Angular distribution of particle images for the Pol II EC1, EC2, and EC3 reconstructions. Black shading indicates the number of particles assigned to a given view, while red dots indicate represented views (at least one particle was assigned within 1° of the point). b, FSC plots for the reconstructions described in a, as well as the FSC plot for the Pol II EC model versus the unsharpened Pol II EC1 map. The first data point after phase randomization is omitted. c, Top–front view and middle slice of the B-factor sharpened Pol II EC1 reconstruction coloured by local resolution. The top–front view (also shown in Extended Data Figure 2) is a rotation of either the top view or front view by 45°. d, Top–front view and middle slice of the unsharpened Pol II EC2 reconstruction coloured by local resolution. e, Top–front view and middle slice of the unsharpened Pol II EC3 reconstruction coloured by local resolution. f, RPB4–RPB7 stalk density for the unsharpened Pol II EC1, Pol II EC2, and stalk focused refinements shown at the same threshold level and filtered to the nominal resolutions (3.4 Å, 3.6 Å, and 4.2 Å). Continuous density for the stalk is visible in the lower resolution focused refinement. g, Density for the protrusion in the unsharpened Pol II EC1 and protrusion-focused refinements shown at the same threshold level and filtered to the nominal resolutions (3.4 Å and 3.8 Å). Density for the protrusion tip is more visible in the focused refinement map.
Extended Data Figure 4 Structural comparison of bovine and yeast Pol II.
a, Structures of the bovine and yeast EC (PDB 1Y1W) were superimposed by alignment of the active site. The protein regions of the bovine EC are shown in two views, coloured by r.m.s.d. of the yeast and bovine models. Insertions and uniquely structured regions within the bovine EC are coloured purple. b, Surface conservation between yeast and bovine Pol II. Strong and weak conservation groups were assigned according to Clustal conventions. c, Structural rearrangements within the RPB5 jaw domain. Axes were drawn through the four longest helices of the bovine and yeast RPB5 jaw using UCSF Chimera. d, Comparison of the backbone path of the trigger loop to previously known trigger loop conformations. The trigger loop backbone of the bovine Pol II EC is shown in thick, dark grey; the open conformation of the yeast EC (PDB 1Y1W)20 is in blue; the closed conformation of the yeast EC in the presence of GTP (PDB 2E2H)24 is in yellow; the wedged conformation of the yeast EC in the presence of α-amanitin (PDB 2VUM)58 is in dark green; the trapped conformation of the arrested and backtracked yeast EC (PDB 3PO2)59 is in purple; the locked conformation of the yeast EC reactivation intermediate in the presence of TFIIS (PDB 3PO3)59 is in orange; and the new open position observed in the ten-subunit yeast transcription complex crystallized in the presence of TFIIF is in light green. The binding site for α-amanitin is denoted in dashed green lines. The binding site for the incoming NTP is outlined in dashed yellow lines. e, Comparison of the path of DNA in the closed initiation complex (yellow7) and the elongation complex (blue, this study). Downstream DNA was extended.
Extended Data Figure 5 Modelled position of the human SPT5 NGN domain on the bovine Pol II EC.
a, Zoom-in view of Pol II EC3 nucleic-acid density superimposed on the Pol II–SPT4–SPT5-NGN EC model reveals proximity of non-template DNA to the SPT5 NGN domain. The human SPT4–SPT5-NGN crystal structure (PDB 3H7H)31 was positioned on the Pol II EC model by alignment with the archaeal RNA polymerase clamp–SPT4–SPT5 crystal structure (PDB 3QQC)26. b, Colouring a surface representation of the X-ray structure of the SPT4–SPT5-NGN by Coulombic surface charge reveals positively charged patches in close proximity to non-template and upstream DNA. Positive charge is in blue; negative charge is in red.
Supplementary information
Supplementary Table 1
This table contains a list of observed Pol II EC-DSIF protein-protein crosslinks. Inter-protein crosslinks are listed first, followed by intra-protein crosslinks. Within these categories, crosslinks are listed in order of decreasing Id-Score. Crosslink precursor ions with unique M/z values were treated as unique crosslinks in the analysis. (PDF 46 kb)
Rights and permissions
About this article
Cite this article
Bernecky, C., Herzog, F., Baumeister, W. et al. Structure of transcribing mammalian RNA polymerase II. Nature 529, 551–554 (2016). https://doi.org/10.1038/nature16482
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature16482
This article is cited by
-
Lowering mutant huntingtin by small molecules relieves Huntington’s disease symptoms and progression
EMBO Molecular Medicine (2024)
-
The SPOC proteins DIDO3 and PHF3 co-regulate gene expression and neuronal differentiation
Nature Communications (2023)
-
Enhancer RNAs stimulate Pol II pause release by harnessing multivalent interactions to NELF
Nature Communications (2022)
-
Structural insights into nuclear transcription by eukaryotic DNA-dependent RNA polymerases
Nature Reviews Molecular Cell Biology (2022)
-
Causes and consequences of RNA polymerase II stalling during transcript elongation
Nature Reviews Molecular Cell Biology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.