Abstract
Deciphering how neural circuits are anatomically organized with regard to input and output is instrumental in understanding how the brain processes information. For example, locus coeruleus noradrenaline (also known as norepinephrine) (LC-NE) neurons receive input from and send output to broad regions of the brain and spinal cord, and regulate diverse functions including arousal, attention, mood and sensory gating1,2,3,4,5,6,7,8. However, it is unclear how LC-NE neurons divide up their brain-wide projection patterns and whether different LC-NE neurons receive differential input. Here we developed a set of viral-genetic tools to quantitatively analyse the input–output relationship of neural circuits, and applied these tools to dissect the LC-NE circuit in mice. Rabies-virus-based input mapping indicated that LC-NE neurons receive convergent synaptic input from many regions previously identified as sending axons to the locus coeruleus, as well as from newly identified presynaptic partners, including cerebellar Purkinje cells. The ‘tracing the relationship between input and output’ method (or TRIO method) enables trans-synaptic input tracing from specific subsets of neurons based on their projection and cell type. We found that LC-NE neurons projecting to diverse output regions receive mostly similar input. Projection-based viral labelling revealed that LC-NE neurons projecting to one output region also project to all brain regions we examined. Thus, the LC-NE circuit overall integrates information from, and broadcasts to, many brain regions, consistent with its primary role in regulating brain states. At the same time, we uncovered several levels of specificity in certain LC-NE sub-circuits. These tools for mapping output architecture and input–output relationship are applicable to other neuronal circuits and organisms. More broadly, our viral-genetic approaches provide an efficient intersectional means to target neuronal populations based on cell type and projection pattern.
Similar content being viewed by others
Main
Figure 1a, b illustrates two extreme connectivity models for input–output relationships of projection neurons. At one extreme, neurons from each input A region connect to a subtype of B neurons that send output to a unique C region (Fig. 1a); this model segregates information into discrete pathways. At the other extreme, B neurons are homogeneous in receiving input from all A regions and sending output to all C regions with a common probability distribution (Fig. 1b); this model allows an overall indiscriminate integration and broadcast of information. The input–output relationship for most neural circuits is not known (Supplementary Note 1).
a, b, Schematic of two extreme connection patterns of region B neurons with inputs from A regions and outputs to C regions. c, d, Strategies for trans-synaptic input tracing from B neurons based on their outputs. TRIO (c) does not distinguish between region B cell types projecting to the selected C region (two different cell types are outlined in grey and blue). cTRIO (d) avoids labelling promiscuous projections from Cre− cells (blue). Open and filled triangles, incompatible loxP sites; open and filled half circles, incompatible FRT sites. e, Schematic of TRIO and cTRIO in mouse motor cortex. CAV was injected into contralateral motor cortex or medulla along with AAVs expressing Cre- or Flp-dependent TVA–mCherry (TC)/rabies glycoprotein (G) into motor cortex, followed by RVdG. Experiments were performed in wild-type (TRIO) or Rbp4-Cre (cTRIO) mice. f, Example coronal sections of motor cortex starter cells in TRIO and cTRIO. cMC, contralateral motor cortex; Me, medulla. Cortical layers are separated by dotted lines based on the DAPI (4′,6-diamidino-2-phenylindole) stain (blue). Starter cells (yellow, a subset indicated by arrowheads) can be distinguished from input cells labelled only with GFP from RVdG (green). TC+ cells in motor cortex spanned layers 2/3 and 5 for TRIO (left), but were restricted to L5 with cTRIO. Bottom inset, example images of input neurons from the somatosensory cortex (SC) and ventral anterior thalamus (VA), derived from larger composites (see Methods). g, Average fraction of total input neurons in Rbp4-Cre-based input tracing and cTRIO of motor cortex L5 pyramidal neurons. Values represent the average fraction of input in each category (n = 4 animals for Rbp4-Cre input tracing and contralateral motor cortex cTRIO; n = 3 animals for medulla cTRIO). Two-way ANOVA determined that inputs to Rbp4-Cre+ motor cortex starter cells generated by input tracing or cTRIO (C = cMC or C = Me) are significantly different in brain regions from which they receive input (interaction P < 0.0001). One-way ANOVA and post hoc Tukey’s multiple comparison tested the significance within each input region. Error bars, s.e.m. **P < 0.01; ***P < 0.001. Scale bars, 250 µm (f, middle row), 100 µm (f, bottom row).
To determine the input–output organization of neural circuits, we have developed a method we have termed TRIO (for tracing the relationship between input and output) and cell-type-specific TRIO (cTRIO). TRIO identifies neurons in A regions that synapse onto B neurons projecting to a specific C region. A→B connections are determined by rabies-virus-mediated retrograde trans-synaptic tracing9, which relies on monosynaptic spread of EnvA-pseudotyped, glycoprotein deleted and GFP-expressing rabies viruses (RVdG hereafter) from starter cells in the B region. Starter cells express rabies glycoprotein (G) and the TVA receptor for EnvA fused with mCherry (TC)10,11, which allow RVdG infection and complementation, enabling trans-synaptic spread to presynaptic neurons in A regions. In TRIO (Fig. 1c), expression of TVA–mCherry fusion and rabies glycoprotein depends on Cre/loxP-mediated recombination from adeno-associated viruses (AAVs) injected in region B. Cre is delivered at a specific C region by canine adenovirus type 2 (CAV hereafter) that efficiently transduces axon terminals12,13. Thus, TRIO does not distinguish between different cell types within region B. In cTRIO (Fig. 1d), TRIO is performed in transgenic mice in which Cre recombinase is expressed in a specific cell type in the B region. Expression of the TVA–mCherry fusion and rabies glycoprotein delivered in the B region depends on Flp/FRT-based recombination and Flp recombinase from CAV-FLExloxP-Flp, which is injected at a specific C region. Thus, only B neurons that express Cre and project to a specific C region can become starter cells for RVdG-mediated trans-synaptic tracing.
As a proof-of-principle, we applied TRIO and cTRIO to the mouse motor cortex (Fig. 1e). In TRIO experiments, we injected CAV-Cre in contralateral motor cortex and AAV-FLExloxP-TC/G into motor cortex of wild-type mice; this resulted in starter cells in motor cortex layers 2/3 (L2/3), L5, (Fig. 1f, left; arrowheads) and L6. In cTRIO experiments, we injected CAV-FLExloxP-Flp in contralateral motor cortex or medulla of retinol binding protein 4 (Rbp4)-Cre mice to target intracortical- or subcortical-projecting L5 neurons, respectively, and AAV-FLExFRT-TC/G into motor cortex; this resulted in starter cells that were restricted to motor cortex L5 (Fig. 1f, middle and right), as predicted by the Rbp4-Cre expression pattern14. For comparison, we also performed trans-synaptic tracing in motor cortex of Rbp4-Cre mice, where starter cells were not selected based on output projections. A large majority of the GFP+ input neurons to the motor cortex were from the cortex or thalamus (Fig. 1f, bottom). Interestingly, contralateral-motor-cortex-projecting L5 neurons received proportionally more input from the cortex, whereas medulla-projecting L5 neurons received more input from the thalamus. Almost all other inputs came from the globus pallidus, a recently identified direct input to the cortex15 (Fig. 1g; Supplementary Table 1). Control experiments indicated that rabies-mediated labelling of most local and all long-range input neurons were dependent on CAV-Cre for TRIO, and CAV-FLExloxP-Flp and Rbp4-Cre for cTRIO (Extended Data Fig. 1). These experiments demonstrated that cTRIO can restrict starter cells to a more specific population than TRIO, and suggested that subcortical- and callosal-projecting L5 neurons receive differential thalamic versus cortical input. TRIO also identified presynaptic partners of callosal- or striatal-projecting motor cortex neurons in rat (Extended Data Fig. 2).
As the precision of TRIO/cTRIO analysis is defined by CAV-mediated transduction from axons and presynaptic terminals, we characterized its spread by injecting CAV-Cre and retrobeads into the Ai14 Cre-reporter mice16 in the piriform cortex at varied distances from the mitral cell axon layer and quantifying labelled mitral cells. We found that CAV spread mostly within 200 µm from the injection site and could also infect axons in passage (Extended Data Fig. 3; Supplementary Note 2). CAV tropism was examined by injecting CAV-Cre into five diverse brain regions of Ai14 mice. Neurons known to project to these brain regions, some of which are >1 cm away from the injection sites, were efficiently labelled (Extended Data Fig. 4). Thus, CAV can infect axons and terminals of diverse neuronal types across long distances.
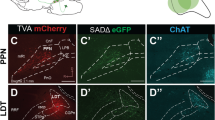
We next applied our viral-genetic tools to the noradrenaline neurons in the locus coeruleus, a small bilateral nucleus in the brainstem (∼1,500 noradrenaline neurons per locus coeruleus) that collectively project axons throughout the brain1,2,3,4. It is unclear whether different LC-NE neurons receive differential input, how LC-NE neurons divide up their brain-wide projection patterns17, and what their input–output relationships are. We first identified synaptic inputs received by these neurons using RVdG-mediated retrograde trans-synaptic tracing in dopamine β-hydroxylase (Dbh)-Cre mice (Fig. 2a), where Cre-dependent TVA–mCherry fusion and rabies glycoprotein expression was restricted to LC-NE neurons that express the noradrenaline biosynthetic enzyme Dbh (Fig. 2b). Control experiments validated that Cre recombination occurred almost exclusively in locus coeruleus neurons expressing tyrosine hydroxylase (TH), another LC-NE neuron marker, and that long-range trans-synaptic tracing depended on Dbh-Cre and AAV-delivered rabies glycoprotein (Extended Data Fig. 5).
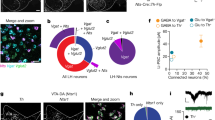
a, Strategy for trans-synaptic tracing of input to LC-NE neurons. b, Coronal section of a mouse brain at the locus coeruleus (dotted square) stained with DAPI (blue). A region within the square is magnified in the inset. LC-NE starter cells (yellow) can be distinguished from cells receiving only TC from AAV (red) or only GFP from RVdG (green) at the injection site. c–g, Coronal sections showing representative input neurons in diverse brain regions. h, Sagittal section of the cerebellum showing trans-synaptically labelled Purkinje cells. Images in b–h were derived from larger composites. i, Schematic summary of brain regions that provide the largest average fractional inputs to LC-NE neurons (n = 9 animals). Scale bars, 1 mm (b), 50 µm (b, inset; c–h). BNST, bed nucleus of the stria terminalis; CeA, central amygdala; DCN, deep cerebellar nuclei; IRN, intermediate reticular nucleus; LC, locus coeruleus; LH, lateral hypothalamus; LRN, lateral reticular nucleus; MRN, midbrain reticular nucleus; PAG, periaqueductal grey; PC, Purkinje cells; PGRN/GRN, paragigantocellular/gigantocelluar nucleus; POA, preoptic area; PRN, pontine reticular nucleus; PVH, paraventricular hypothalamic nucleus; SuC, superior colliculus; SVN, spinal vestibular nucleus; ZI, zona incerta.
We counted all input neurons to LC-NE starter cells from the anterior forebrain to posterior medulla (Fig. 2c–h) except in coronal sections immediately surrounding the locus coeruleus, as non-specific viral labelling of neurons can occur locally at the AAV/RVdG injection site (Extended Data Fig. 5c–f). We assigned each input neuron to one of 111 brain regions according to the Allen Brain Atlas (http://mouse.brain-map.org/static/atlas) to categorize brain regions ipsi- or contralateral to the injected locus coeruleus (Supplementary Table 2). Regions that contributed more than 1% of total input from nine Dbh-Cre tracing brains are summarized in Fig. 2i. Although most brain regions we identified as presynaptic to LC-NE neurons are consistent with previous retrograde tracing studies6,7,8, our experiment validated that these neurons directly synapse onto LC-NE neurons rather than just projecting axons to the locus coeruleus. We also found that deep cerebellar nuclei and cerebellar Purkinje cells contributed a notable fraction of direct synaptic input to LC-NE neurons (Fig. 2h, i), which (to our knowledge) has not been previously reported. Labelled Purkinje cells were enriched in the ipsilateral medial zones throughout the cerebellum, at distances up to 2.5 mm away from the locus coeruleus (Extended Data Fig. 6a). Consistent with a direct connection between Purkinje cells and LC-NE neurons, we found that an inhibitory postsynaptic marker, gephyrin, was present in TH+ LC-NE dendrites apposing GABAergic Purkinje cell axons (Extended Data Fig. 6b, c).
We next applied TRIO and cTRIO to test if populations of LC-NE neurons, defined by their output targets, received distinct input. We selected five diverse brain regions known to receive LC-NE projections: the olfactory bulb, auditory cortex, hippocampus, cerebellum and medulla (Fig. 3a). CAV-Cre injection into these regions in Ai14 mice confirmed labelling of noradrenaline neurons throughout the locus coeruleus (Extended Data Fig. 7a). We did not observe significant differences in the spatial distribution along the anterior–posterior or medial–lateral axes for LC-NE neurons that projected to these brain regions. However, forebrain-projecting LC-NE neurons were more dorsally biased compared to the hindbrain-projecting ones (Extended Data Fig. 7b–f), consistent with a previous observation in the rat18. We applied TRIO to olfactory bulb, auditory cortex and hippocampus, and cTRIO to cerebellum and medulla, as locus coeruleus projections to the former group predominately came from TH+ neurons, whereas the latter group contained TH− neurons (Extended Data Fig. 7a). Control experiments indicated that the labelling of input neurons depended on CAV-Cre in the case of TRIO (Extended Data Fig. 5c, e), and on both Dbh-Cre and CAV-FLExloxP-Flp in the case of cTRIO (Extended Data Fig. 8).
a, Schematic of CAV injections into locus coeruleus output regions for TRIO and cTRIO. AC, auditory cortex; Cb, cerebellum; Hi, hippocampus; Me, medulla; OB, olfactory bulb. b–d, Average fractional inputs in Dbh-Cre-based input tracing (grey, n = 9 animals), TRIO (Hi, purple; OB, red; AC, blue; n = 4 animals each), and cTRIO (Cb, green; Me, orange; n = 4 animals each). Input neurons were grouped into 16 broader categories. Magnified insets highlight the average fraction of input from striatum-like amygdala (>98% from the central amygdala) (c) or from Purkinje cells (d) to LC-NE neurons that project to the 5 output regions or in Dbh-Cre-based input tracing. Error bars, s.e.m.
We analysed inputs for the TRIO and cTRIO experiments analogous to Dbh-Cre-based input tracing (Supplementary Table 2). We observed that LC-NE neurons received inputs from all input regions regardless of their diverse output, with a grossly similar proportional distribution (Fig. 3b). These data suggest that the LC-NE circuit is largely indiscriminate with respect to its input–output relationship. However, region-by-region one-way ANOVA (Supplementary Table 3, top) rejected the overall null hypothesis that input distribution is independent of output conditions (combined P = 0.002), indicating that the input–output relationships were not entirely homogeneous. Of the individual input that exhibited the smallest P values (Supplementary Table 3, bottom), LC-NE neurons projecting to the medulla received less input from the central amygdala (Fig. 3c). In addition, the fraction of Purkinje cell inputs in Dbh-Cre based tracing was higher than any of the TRIO/cTRIO conditions (Fig. 3d), suggesting that Purkinje cells contribute input to an LC-NE population that do not project axons extensively to any of the output sites we examined.
The largely indiscriminate input–output relationship revealed by TRIO can in principle be accounted for by input convergence (Fig. 1b, left), output divergence (Fig. 1b, right), or both. A simulation analysis of the two sparsest input tracing samples (Supplementary Table 2) suggested that individual LC-NE neurons must receive input from more than 15 or 9 brain regions, respectively (Extended Data Fig. 9). This is most likely a lower bound as rabies tracing efficiency is far from 100%. However, such extensive integration is not entirely homogenous (Supplementary Table 4). Thus, individual LC-NE neurons integrate inputs from many regions, yet exhibit heterogeneity with respect to brain regions from which they receive input.
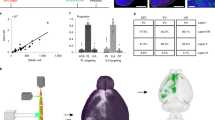
We next explored the output architecture of LC-NE neurons (Fig. 1a, b, right). Previous dual-retrograde-tracer experiments indicated that individual LC-NE neurons could project to two brain regions far apart19,20,21, but did not examine collateralization between more than two output regions in a given experiment. We devised a general method for tracing output divergence of specific neuronal populations based on their projection to one output site (Fig. 4a, b). We found that populations of LC-NE neurons projecting to the olfactory bulb, auditory cortex, hippocampus or medulla also projected to all seven additional brain regions analysed (Fig. 4c, d and Extended Data Fig. 10). Thus, the output of LC-NE neuronal populations is highly divergent, resembling the broadcast model (Fig. 1b, right) much more than the discrete output model (Fig. 1a, right). We nevertheless found a general trend of increased axon density in the output region where labelling was initiated compared to labelling initiated from the locus coeruleus, with the bias from olfactory bulb- or medulla-initiated labelling reaching statistical significance (Fig. 4e). This suggests that LC-NE neurons projecting to the olfactory bulb or medulla contain populations with biased output to these regions, consistent with the observation that LC-NE neurons projecting to these regions have a biased distribution along the dorsoventral axis in the locus coeruleus (Extended Data Fig. 7a).
a, In this strategy, neurons in region B projecting to the C region where CAV-Cre is delivered are labelled, including their collaterals to other output regions (for example, blue neurons to C2). b, In this strategy, only Cre+ neurons in region B projecting to the C region where CAV-FLExloxP-Flp is delivered are labelled. c, Schematic for data in (d). In this example, CAV was injected in the olfactory bulb and TC was injected in the locus coeruleus. TC+ LC-NE axons were imaged in the designated brain regions. All TC+ axons were co-stained with anti-noradrenaline transporter (NET; inset), confirming their noradrenaline identity. d, Average normalized fraction of TC+ LC-NE axons in each brain region when CAV was injected into four output sites, or Cre-dependent TC was injected directly into the locus coeruleus of Dbh-Cre mice (colour code on top right). LC: n = 4 animals (Dbh-Cre); Hi: n = 4 animals (Dbh-Cre); AC: n = 4 animals (2 wild-type, 2 Dbh-Cre); OB: n = 5 animals (3 wild-type, 2 Dbh-Cre); Me: n = 4 animals (Dbh-Cre). The average number of LC-NE neurons labelled in each condition was: 855 ± 102 (mean ± s.e.m, in locus coeruleus, n = 4 animals); 235 ± 35 (Hi, n = 4 animals); 80 ± 31 (OB, n = 5 animals); 114 ± 31 (AC, n = 4 animals); 202 ± 63 (Me, n = 4 animals). e, Comparison of the fraction of TC+ axons at CAV injection sites between projection-based and direct locus coeruleus labelling methods. Unpaired two-tail t-tests. *P < 0.05. Error bars, s.e.m. Scale bar, 10 µm. Abbreviations: AC, auditory cortex; Cb, cerebellum; CC, cingulate cortex; Hi, hippocampus; Hy, hypothalamus; LC, locus coeruleus; Me, medulla; OB, olfactory bulb; SC, somatosensory cortex.
Our study provides the first whole-brain quantitative analysis of synaptic input onto LC-NE neurons (Figs 2i and 3b–d and Supplementary Table 2). Although the total input encompasses brain regions that control cognitive, autonomic, endocrine and somatic motor activities22, LC-NE neurons receive abundant input from motor-related nuclei in the midbrain, pons, medulla and cerebellum (Fig. 3b). Our viral-genetic tools also revealed a highly extensive output divergence (Fig. 4d). Together with input convergence, this probably explains the largely indiscriminate input–output relationship of LC-NE neurons (Fig. 3b). This property fits well with a primary function of the LC-NE neurons in regulating states of the entire brain during sleep/wake cycles and arousal3,23,24,25.
Despite the overall integrative nature, however, our data also revealed specificity in the input–output relationship of LC-NE sub-circuits. Medulla-projecting LC-NE neurons receive disproportionally smaller input from the central amygdala than LC-NE neurons projecting to other regions (Fig. 3b, c). Input from the central amygdala to the locus coeruleus is an important component for initiating stress response26. Our observation implies that modulation of medulla by LC-NE neurons is preferentially immune to this type of stress input. Although our output studies demonstrated the broad projection pattern of LC-NE neurons, they also highlighted specificities both in regard to biased cell body distribution within the locus coeruleus (Extended Data Fig. 7f) and biased projections (Fig. 4e). The existence of such input–output specificity, along with differential distribution of adrenergic receptors in target neuronal populations4, enables the LC-NE circuit to selectively modulate specific targets.
The viral-genetic tools we described here can be applied to other circuits in the mammalian brain, such as motor cortex (Fig. 1 and Extended Data Fig. 2). TRIO and particularly cTRIO have extended previous projection-selective targeting methods27,28,29 for analysing complex circuits in the central nervous system. Furthermore, intersecting projection and cell type using CAV-FLExloxP-Flp and numerous Cre transgenic mice can refine genetic access to specific populations of neurons to record and functionally manipulate their activity30.
Methods
Animals
Dbh-Cre mice31 and Rbp4-Cre transgenic mice14 were obtained from the Mutant Mouse Regional Resource Center. Purkinje cell protein 2 (Pcp2)-Cre32 and the ROSA26Ai14 Cre-dependent tdTomato reporter (Ai14)16 were obtained from the Jackson Laboratories. Mice were housed on a 12-h light/dark cycle with food and water ad libitum. Transgenic mice were of a mixed genetic background, and there was a similar distribution of male and female mice included in all experiments. Wister Rats were purchased from Japan SLC (Hamamatsu, Japan). All rats used for experiments were female. All procedures for mice followed animal care guidelines approved by Stanford University’s Administrative Panel on Laboratory Animal Care (APLAC). All rat experiments were performed in accordance with the animal care and use committee guidelines of the University of Tokyo.
DNA constructs
CAG-FLExloxP-G and CAG-FLExloxP-TC (same as CAG-FLEx-TCB) have been described previously10. CAG-FLExFRT-G and CAG-FLExFRT-TC were constructed using standard molecular cloning methods with enzymes from New England Biolabs (Ipswich, USA). The custom DNA fragment (389 base pairs) shown below was synthesized by DNA 2.0 (Menlo Park, USA). This DNA fragment contained restriction enzyme sites and two heterospecific pairs of FRT (shown by the underline) and FRT5 (shown by bold italic) in the following order: 5′-NotI-MluI-KpnI-FRT-FRT5-SalI-AscI-FRT(complementary)-FRT5(complementary)-HindIII-SpeI-NotI, with the sequence as follows: 5′-GCGGCCGCACGCGTACGTGGTACCGAAGTTCCTATTCCGAAGTTCCTATTCTCTAGAAAGTATA GGAACTTCATCAAAATAGGAAGACCAATGCTTCACCATCGACCCGAATTGCCAAGCATCACCATCGACAGAA GTTCCTATTCCGAAGTTCCTATTCTTCAAAAGGTATAGGAACTTCGTCGACAATTGGCGCGCCGAAGTTCCTA TACTTTCTAGAGAATAGGAACTTCGGAATAGGAACTTCCGTTGGGATTCTTCCTATTTTGATCCAAGCATCACCATCGACCCTCTAGTCCAGATCTCACCATCGACCCGAAGTTCCTATACCTTTTGAAGAATAGGAACTTCGGAATA GGAACTTCAAGCTTAATTACTAGTGCGGCCGC.
This DNA fragment was cloned into the modified pBluescript II SK vector that contains only a NotI recognition sequence in the cloning site. We serially inserted into this pBluescript the following DNA fragments by using the unique restriction sites. (1) HindIII/SpeI fragment containing the WPRE and human growth hormone polyA signal obtained from pAAV-TRE-HTG (Addgene number 27437)33. (2) MluI/KpnI fragment containing the CAG promoter. To make this fragment, we sub-cloned PstI–XmaI flanked CAG promoter from pCA-T-int-G (Addgene number 36887)34 into a modified pBluescript II SK vector that only contains MluI-PstI-XmaI-KpnI recognition sequence in the cloning site. (3) AscI/SalI fragment containing coding sequence of G or TC cassette, obtained from pAAV CAG-FLExloxP-G (Addgene number 48333) or pAAV CAG-FLExloxP-TC (Addgene number 48332)10. The assembled cassettes were sub-cloned into pAAV-MCS (AAV helper free system, Stratagene, catalogue number 240071-12) using the NotI sites. Flp-dependent mCherry expression from CAG-FLExFRT-TC was confirmed by transient transfection into cultured HEK293 cells by using a Flp-expressing plasmid (data not shown). To generate CAV-FLExloxP-Flp, we first constructed the CAV targeting vector pCAV-FLExloxP-Flp. AscI-SalI flanking Flpo coding sequence was PCR amplified by using pBT340 (Addgene number 52549) as a template and sub-cloned into SalI–AscI site of a precursor of pAAV CAG-FLExloxP-TCB (ref. 10) that does not contain WPRE-polyA signal. XmaI–NotI fragment containing FLExloxP-Flp was then subcloned into KpnI–NotI site of pCL20c lentivirus vector35 by using blunt-end ligation. EcoRI–SpeI fragment containing FLExloxP-Flp was then subcloned into EcoRI–EcoRV site of pTCAV-12vk (description available upon request to E.J.K.) by using one-sided (SpeI site) blunt-end ligation, resulting in pCAV-FLExloxP-Flp.
Virus preparations
All viral procedures followed the Biosafety Guidelines approved by the Stanford University Administrative Panel on Laboratory Animal Care (A-PLAC), Administrative Panel of Biosafety (APB), and equivalent committees of the University of Tokyo. Recombinant AAV vectors (serotype 5 for TVA receptor fused with mCherry and serotype 8 for rabies glycoprotein) were produced in the Stanford University or University of North Carolina Viral Core. The AAV titre was estimated to be 2.6 and 1.3 × 1012 viral particles per ml for CAG-FLExFRT-TC and CAG-FLExFRT-G, based on quantitative PCR analysis. RVdG was prepared as previously described36. The pseudotyped RVdG titer was estimated to be ∼5 × 109 infectious particles per ml based on serial dilutions of the virus stock followed by infection of the 293-TVA800 cell line. The recombinant CAV-Cre and CAV-FLExloxP-Flp were generated, expanded and purified by previously described methods37. The final titre of CAV-Cre and CAV-FLExloxP-Flp were 2.5 × 1012 and 5 × 1012 viral particles per ml, respectively. All handling of CAV and rabies virus followed procedures approved by Stanford University’s Administrative Panel on Biosafety (APB) for biosafety level 2, and the equivalent committees of the University of Tokyo (P2/P2A).
Locus coeruleus trans-synaptic input tracing
Experiments in Fig. 2 were performed in Dbh-Cre mice at 8–12 weeks of age following procedure as described previously10,38. Mice were anaesthetized with 65 mg per kg ketamine and 13 mg per kg xylazine (Vedco/Lloyd Laboratories) via intraperitoneal injection. Then ∼0.5 μl of a 1:1 mixture of AAV8 CAG-FLExloxP-G and AAV5 CAG-FLExloxP-TC were injected into the left locus coeruleus of the mouse using stereotaxic equipment (Kopf). The coordinates were 0.8 mm lateral from midline, 0.8 mm posterior from lambda, and 3.2 mm ventral from the surface of the brain. Two weeks later, 0.3–0.5 μl RVdG was injected into the same area of the locus coeruleus using the procedure described above. After recovery, mice were housed in a biosafety level 2 (BSL2) facility for 4 days before euthanasia.
Assessment of locus coeruleus terminal infectivity by CAV-Cre
Experiments in Extended Data Figs 4 and 7 were performed in Ai14 mice 6–8 weeks of age. Mice were anaesthetized and injected, as described above, with 0.25–0.5 μl CAV-Cre plus 0.02 μl green retrobeads (Lumofluor, USA) into predicted LC output sites: olfactory bulb (OB), auditory cortex (AC), hippocampus (Hi), medulla (Me), or cerebellum (Cb). The coordinates used for CAV-Cre injection sites are listed as measurements from bregma for OB, AC and Hi, and from lambda for Me and Cb. Ventral measurements are from the surface of the brain. OB: 0.75 mm lateral, 4.0 mm anterior, 1 mm ventral; AC: 4.2 mm lateral, 2.5 mm posterior, 0.8 mm ventral; Hi: 1.5 mm lateral, 2 mm posterior, 1.5 mm ventral; Me: 0.75 mm lateral, 3.3 mm posterior, 3.5 mm ventral; Cb: 2.0 mm lateral, 3.0 mm posterior, 1.5 mm ventral. After recovery, mice were housed in a BSL2 facility for 5–7 days before euthanasia.
Locus coeruleus projection-based viral labelling
Experiments in Fig. 4 and Extended Data Fig. 10 were performed in wild-type or Dbh-Cre mice 8–12 weeks of age. Mice were anaesthetized and injected with ∼0.25 μl AAV5-expressing TC (CAG-FLExloxP-TC for wild-type mice or CAG-FLExFRT-TC for Dbh-Cre mice) into the left locus coeruleus as described above. Mice were also injected with ∼0.5 μl CAV-Cre (wild-type mice) or CAV-FLExloxP-Flp (Dbh-Cre mice) at locus coeruleus output sites in the ipsilateral hemisphere (OB, AC, Hi, Me; coordinates listed above). After recovery, mice were housed in a BSL2 facility for 3–4 weeks before euthanasia.
Locus coeruleus TRIO and cTRIO
Experiments shown in Fig. 3 were performed in wild-type (TRIO) or Dbh-Cre (cTRIO) mice at 8–12 weeks of age. For TRIO, mice were anaesthetized and injected with ∼0.5 μl of a 1:1 mixture of AAV8 CAG-FLExloxP-G and AAV5 CAG-FLExloxP-TC into the left locus coeruleus, and also injected with ∼0.5 μl CAV-Cre into ipsilateral OB, Hi, or AC using coordinates described above. For cTRIO, mice were anaesthetized and injected with ∼0.5 μl of a 1:1 mixture of AAV8 CAG-FLExFRT-G and AAV5 CAG-FLExFRT-TC into the left locus coeruleus, and also injected with ∼0.5 μl CAV-FLExloxP-Flp into either ipsilateral Cb or Me using coordinates described above. After recovery, mice were housed in a BSL2 facility. Two weeks later, 0.3–0.5 μl RVdG was injected into the locus coeruleus using the procedure described above. After recovery, mice were housed in a BSL2 facility for 4 days before euthanasia.
Motor cortex TRIO and cTRIO
Experiments in Fig. 1 were performed in wild-type (TRIO) or Rbp4-Cre (cTRIO) mice at 8–12 weeks of age. For TRIO, mice were anaesthetized and injected with ∼0.5 μl of a 1:1 mixture of AAV8 CAG-FLExloxP-G and AAV5 CAG-FLExloxP-TC into the left motor cortex (MC), and also injected with ∼0.5 μl CAV-Cre into contralateral MC (cMC) using the following coordinates from bregma: 1.5 mm lateral, 1.5 mm anterior, 0.8 mm ventral from the surface of the brain. For cTRIO, Rbp4-Cre mice were anaesthetized and injected with ∼0.5 μl of a 1:1 mixture of AAV8 CAG-FLExFRT-G and AAV5 CAG-FLExFRT-TC into the left motor cortex, and also injected with ∼0.5 μl CAV-FLExloxP-Flp into the coordinates described above (for cMC), or the following coordinates for medulla (from lambda): 1 mm lateral, 3 mm posterior, 4 mm ventral from the surface of the brain. After recovery, mice were housed in a BSL2 facility. Two weeks later, 0.3–0.5 μl RVdG was injected into motor cortex. After recovery, mice were housed in a BSL2 facility for 4 days before euthanasia.
Rat motor cortex TRIO
For TRIO experiments in rat (Extended Data Fig. 2), ∼0.4 μl of a 1:1 mixture of AAV2 CAG-FLExloxP-G and AAV2 CAG-FLExloxP-TC was injected into the brain of ∼5-week-old Wister rat using stereotaxic equipment (Narishige, Japan). During surgery, animals were anaesthetized with 65 mg per kg ketamine and 13 mg per kg xylazine. For motor cortex injections, the needle was placed 2.5 mm anterior and 2.3 mm lateral from the bregma, and 0.9 mm ventral from the brain surface. ∼0.5 μl CAV-Cre was injected subsequently into either the ipsilateral striatum (1.0 mm posterior and 4.0 mm lateral from the bregma, and 4.0 mm ventral from the brain surface) or the contralateral motor cortex. After recovery, animals were housed in a BSL2 room. Two weeks later, 0.3 μl RVdG was injected into the AAV injection site under anaesthesia. After recovery, animals were housed in a BSL2 room for 4 days before euthanasia.
Control TRIOs
Control experiments (Extended Data Figs 1, 2, 5 and 8) were performed using conditions described above for locus coeruleus and motor cortex experiments.
Characterizing CAV-Cre spread in the piriform cortex
To test the extent of local spread of CAV-Cre in the injection site (Extended Data Fig. 3), ∼0.3 μl CAV-Cre plus ∼0.025 μl green retrobeads (Lumofluor, USA) were injected into the anterior piriform cortex or surrounding areas of adult mice heterozygous for the Ai14 Cre reporter. During surgery, animals were anaesthetized with 65 mg per kg ketamine and 13 mg per kg xylazine. The stereotactic coordinates were anterior 1.7 mm from the bregma; lateral 1.7–2.8 mm from the midline; ventral 2.5–4.0 mm from the surface of the brain. One week after the injection, animals were perfused and brain tissue was processed and sectioned as described in the histology and imaging section. Then, 60-μm coronal sections through the olfactory bulb until the end of anterior piriform cortex (APC) were collected. The needle location was visualized by the presence of concentrated retrobeads. The distance between the needle tip and the nearest layer 1a of anterior piriform cortex or lateral olfactory tract was measured. In the main and accessory olfactory bulb, the number of tdTomato+ cells was counted either in every section (when total number of labelling was less than ∼1,000 cells) or in every other section (when total the number of labelling was greater than ∼1,000 cells).
Histology and imaging
For all tracing analyses and CAV-Cre infectivity analyses (Figs 1, 2, 3, 4 and Extended Data Figs 1, 2, 3, 4, 5, 6a, 7, 8 and 10), animals were perfused transcardially with phosphate-buffered saline (PBS) followed by 4% paraformaldehyde (PFA) in PBS. Brains were dissected, post-fixed in 4% PFA for 24 h, and placed in 30% sucrose in PBS for 24–48 h. After embedding in Optimum Cutting Temperature (OCT, Tissue Tek), samples were stored at −80 °C until sectioning. For Figs 1, 2, 3 and Extended Data Figs 1, 2, 3, 4, 5, 6a, and 8, consecutive 60-μm coronal sections were collected onto Superfrost Plus slides, washed 2 × 20 min with PBS, and stained with DAPI (1:10,000 of 5 mg ml−1, Sigma-Aldrich), which was included in the last PBS wash. Slides were coverslipped with Fluorogel (Electron Microscopy Sciences). Samples were imaged using a Leica Ariol slide scanner with the SL200 slide loader. Briefly, the scanner first imaged slides using a 1.25× objective and the ‘TissueFind’ function to generate composite brightfield images of the entire slide. Then, the scanner automatically detected individual tissue sections from these brightfield images (∼15–20 coronal tissue sections per slide), and performed automated tiled imaging of each tissue section on the slide in two channels (DAPI and Spectrum Green filters) using a 5× objective. Each tile was approximately 1.2 mm by 1.2 mm, and included ∼20-μm overlap between tiles. Leica Ariol software automatically stitched together individual tiles during image collection to generate a composite SCN file of the entire slide. For analysis of locus coeruleus output (Fig. 4 and Extended Data Fig. 10), every 50-μm sagittal section within the brain regions designated for analysis were collected sequentially into PBS. Sections were washed 2 × 10 min in PBS and blocked for 2–3 h at room temperature (RT) in 10% normal donkey serum (NDS) in PBS with 0.3% Triton-X100 (PBST). Primary antibodies (mouse anti-noradrenaline transporter (NET), PhosphoSolutions, 1447-NET, 1:10,000; rat anti-mCherry, M11217, Invitrogen, 1:2,000) were diluted in 5% NDS in PBST and incubated for four nights at 4 °C. After 3 × 10 min washes in PBST, secondary antibodies were applied for 2–3 h at room temperature (donkey anti-mouse, Alexa-488, and donkey anti-rat Cy3, Jackson ImmunoResearch), followed by 3 × 10 min washes in PBST. Sections were additionally stained with DAPI. For immunostaining of locus coeruleus neuron cell bodies (Extended Data Figs 5 and 7), 50-μm coronal sections through the locus coeruleus were collected into PBS. Sections were washed, immunostained, and mounted as described above, using a primary antibody for tyrosine hydroxylase (rabbit anti-tyrosine hydroxylase (TH), Millipore, AB152, 1:2,000). All images were processed using NIH ImageJ software. For gephyrin immunostaining (Extended Data Fig. 6), fresh tissue was processed and 14-μm horizontal sections were collected through the locus coeruleus following ‘Method B’ in a previously published protocol39. Sections were immunostained with primary antibodies for gephyrin (mouse anti-gephyrin, Synaptic Systems, 147011, 1:700), and tyrosine hydroxylase (Millipore, 1:2,000). Representative images in Fig. 1f (top and middle) and Extended Data Figs 1, 4, 5c, d, 6b, c (left), 7a, c, 8a, b (bottom) and Extended Data Fig. 10a were obtained on a Zeiss epifluorescence microscope with a Nikon CCD camera. Representative images in Figs 2b (inset), 4c (inset), Extended Data Figs 5a, 6b (right), c (middle and right), and 7a (inset) were obtained on a Zeiss LSM 780 confocal microscope. Representative images in Figs 1f (bottom), 2b–h, and Extended Data Fig. 6a were obtained on a Leica Ariol slide scanner with the SL200 slide loader. Representative images in Extended Data Figs 2 and 3 were obtained by cooled CCD camera (ORCA-R2, Hamamatsu Photonics) connected with a upright fluorescent microscope (4× or 10× objective, BX53, Olympus).
Data analysis for trans-synaptic tracing and TRIO
Because each brain differed in total numbers of input neurons, we normalized neuronal number in each region by the total number of input neurons counted in the same brain. For trans-synaptic tracing and TRIO/cTRIO analyses (Figs 1, 2, 3, and Supplementary Tables 1 and 2), GFP+ input neurons were manually counted from every 60-μm section through the entire brain, except near the starter cell location (motor cortex or locus coeruleus), as specified in Extended Data Figs 1 and 5. GFP+ input neurons were assigned to specific brain regions based on classifications of the Allen Brain Atlas (http://mouse.brain-map.org/static/atlas), using anatomical landmarks in the sections visualized by DAPI counterstaining and autofluorescence of the tissue itself. In a small minority of cases, assignment of input neurons to specific brain nuclei may be approximate if GFP+ cell bodies were located on borders between regions, or when anatomical markers were lacking between directly adjacent regions (such as hypoglossal nucleus/nucleus prepositus or lateral reticular nucleus/gigantocellular reticular nucleus). However, quantitative analyses of input tracing results (Figs 1, 2, 3) were performed on anatomical classifications (specified by the Allen Brain Atlas) that were at least one hierarchical level broader than the discrete brain regions in which GFP+ cells were originally assigned to. For instance, the fraction of GFP+ cells designated to ‘paraventricular hypothalamic nucleus’ and ‘lateral hypothalamic nucleus’ were grouped into a broader category of ‘hypothalamus’. In almost all cases where individual GFP+ cells were difficult to classify, their location was within brain regions that belonged to the same broad group. Also, some brain regions partially overlapped with the region excluded from analysis, such as motor and somatosensory cortex for the motor cortex tracing/cTRIO analyses (Fig. 1g, Supplementary Table 1), and the dorsal raphe, periaqueductal grey, and pontine reticular nucleus for the locus coeruleus tracing/TRIO/cTRIO analyses (Figs 2i, 3b, and Supplementary Table 2). Therefore, the inputs reported for these regions are likely under-representations of their contribution to motor cortex or locus coeruleus input. We did not adjust for the possibility of double-counting cells in any of our quantifications, which likely results in slight over-estimates, with the amount of over-estimation depending on the size of the cell in each region quantified. Nearly all starter cells were TH+ for TRIO experiments with the CAV injection site being hippocampus (98.6% ± 0.8%, n = 4 animals), olfactory bulb (96.0% ± 2.7%, n = 4 animals), or auditory cortex (98.8% ± 1.2%, n = 4 animals), consistent with our observation that cells at the locus coeruleus projecting to these target areas are predominantly noradrenaline neurons (Extended Data Fig. 7a). All starter cells were confirmed to be TH+ in cTRIO experiments with the CAV injection site in Cb (100%, n = 4 animals) or Me (100%, n = 4 animals).
Data analysis for locus coeruleus output
For locus coeruleus output quantification (Fig. 4 and Extended Data Fig. 10), images were taken from 5 consecutive sections in each of the 8 brain regions (OB, AC, CC, SC, Hi, Hy, Cb, Me) on a Zeiss epifluorescence microscope with a 10× objective. We attempted to image from identical volumes within each brain region between samples, based on pre-determined coordinates for these regions and the section number, as sections were collected sequentially and kept in order. The field of view for each image was located based solely on DAPI staining so the experimenter was blind to the level of TC+ locus coeruleus axons before imaging. An image was also taken of the noradrenaline transporter (NET) immunostaining in the same field of view to confirm that all TC+ axons were also NET+. For analysis, the TC channels for each image were made binary in ImageJ, after performing background subtraction and thresholding to a value ∼4× greater than mean background intensity. The pixel densities of these binary images were measured and averaged (5 images per brain region) to determine the fraction of TC+ axons resulting from LC-NE neurons in each brain region, as a fraction of the total TC+ axons quantified from all of the imaged regions in that brain.
Analysis of spatial distribution in the locus coeruleus
To assess the distribution of discrete LC-NE neurons within the locus coeruleus dependent on their output (Extended Data Fig. 7), 50-μm coronal sections were collected in order through the locus coeruleus of experimental mice and processed as described above in the histology and imaging section. The experimenter was blind to the location of the output injection site when imaging and quantifying locus coeruleus sections. The outlines of the digital model were manually drawn using TH immunostaining of LC-NE cell bodies as a guide. A cross (+) marks the approximate centre of each locus coeruleus section following the procedure below. (1) measure the maximal height (Hmax) of the locus coeruleus (based on the shape of TH immunostaining); (2) measure the maximum width (Wmax) of the locus coeruleus at the 0.5 Hmax height level; (3) place a cross at the 0.5 Wmax position. This cross was used to align each locus coeruleus image with its corresponding digital locus coeruleus section, before designating the location of the tdTomato+ LC-NE cell bodies with coloured dots on the digital section. Before quantifying experimental sections, two independent sets of TH-immunostained locus coeruleus sections were found to fit within the boundaries of the digital model using this method of alignment. The distribution of tdTomato+ LC-NE dots from each digital section were counted and assigned to dorsal/ventral and medial/lateral sub-regions using horizontal and vertical lines drawn through the centre cross of each digital section, respectively (Extended Data Fig. 7).
Input simulation
We simulated the number of input areas for the two sparsest Dbh-Cre tracing samples with Matlab. For each starter cell, when assuming it receives inputs from n areas, we randomly sampled n areas from the 111 input areas without replacement, weighted by the total counts of cells in each area derived from all the Dbh-Cre brains. The simulated input areas to the 4 (sparsest sample) or 22 (second sparsest sample) starter cells were then consolidated to generate the final number of input areas. Ten thousand rounds of simulations were performed for each n between 11 and 30 (sparsest sample) or 3 and 22 (second sparsest sample).
Statistical methods
No statistical methods were used to predetermine sample size. Animals were excluded from certain experiments using the following pre-established criteria. For all trans-synaptic tracing, TRIO, and cTRIO experiments in motor cortex and locus coeruleus, samples were excluded if less than 50 GFP+ neurons were observed in the brain outside of the area designated as local background. For projection-based viral-genetic labelling, samples were excluded if less than 10 LC-NE neurons were observed to be TC+. No method of randomization was used in any of the experiments. For ANOVA analyses, the variances were similar as determined by Brown–Forsythe test.
References
Dahlström, A. & Fuxe, K. Evidence for the existence of monoamine containing neurons in the central nervous system. I. Demonstration of monoamines in the cell bodies of brain stem neurons. Acta Physiol. Scand. 232 (Suppl.) 1–55 (1964)
Swanson, L. W. & Hartman, B. K. The central adrenergic system. An immunofluorescence study of the location of cell bodies and their efferent connections in the rat utilizing dopamine-beta-hydroxylase as a marker. J. Comp. Neurol. 163, 467–505 (1975)
Sara, S. J. & Bouret, S. Orienting and reorienting: the locus coeruleus mediates cognition through arousal. Neuron 76, 130–141 (2012)
Szabadi, E. Functional neuroanatomy of the central noradrenergic system. J. Psychopharmacol. 27, 659–693 (2013)
Robertson, S. D., Plummer, N. W., de Marchena, J. & Jensen, P. Developmental origins of central norepinephrine neuron diversity. Nature Neurosci. 16, 1016–1023 (2013)
Cedarbaum, J. M. & Aghajanian, G. K. Afferent projections to the rat locus coeruleus as determined by a retrograde tracing technique. J. Comp. Neurol. 178, 1–16 (1978)
Aston-Jones, G. et al. Afferent regulation of locus coeruleus neurons: anatomy, physiology and pharmacology. Prog. Brain Res. 88, 47–75 (1991)
Luppi, P. H., Aston-Jones, G., Akaoka, H., Chouvet, G. & Jouvet, M. Afferent projections to the rat locus coeruleus demonstrated by retrograde and anterograde tracing with cholera-toxin B subunit and Phaseolus vulgaris leucoagglutinin. Neuroscience 65, 119–160 (1995)
Wickersham, I. R. et al. Monosynaptic restriction of transsynaptic tracing from single, genetically targeted neurons. Neuron 53, 639–647 (2007)
Miyamichi, K. et al. Dissecting local circuits: parvalbumin interneurons underlie broad feedback control of olfactory bulb output. Neuron 80, 1232–1245 (2013)
Watabe-Uchida, M., Zhu, L., Ogawa, S. K., Vamanrao, A. & Uchida, N. Whole-brain mapping of direct inputs to midbrain dopamine neurons. Neuron 74, 858–873 (2012)
Soudais, C., Laplace-Builhe, C., Kissa, K. & Kremer, E. J. Preferential transduction of neurons by canine adenovirus vectors and their efficient retrograde transport in vivo. FASEB J. 15, 2283–2285 (2001)
Salinas, S. et al. CAR-associated vesicular transport of an adenovirus in motor neuron axons. PLoS Pathog. 5, e1000442 (2009)
Gerfen, C. R., Paletzki, R. & Heintz, N. GENSAT BAC Cre-recombinase driver lines to study the functional organization of cerebral cortical and basal ganglia circuits. Neuron 80, 1368–1383 (2013)
Saunders, A. et al. A direct GABAergic output from the basal ganglia to frontal cortex. Nature 521, 85–89 (2015)
Madisen, L. et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nature Neurosci. 13, 133–140 (2010)
Chandler, D. J., Gao, W. J. & Waterhouse, B. D. Heterogeneous organization of the locus coeruleus projections to prefrontal and motor cortices. Proc. Natl Acad. Sci. USA 111, 6816–6821 (2014)
Mason, S. T. & Fibiger, H. C. Regional topography within noradrenergic locus coeruleus as revealed by retrograde transport of horseradish peroxidase. J. Comp. Neurol. 187, 703–724 (1979)
Room, P., Postema, F. & Korf, J. Divergent axon collaterals of rat locus coeruleus neurons: demonstration by a fluorescent double labeling technique. Brain Res. 221, 219–230 (1981)
Nagai, T., Satoh, K., Imamoto, K. & Maeda, T. Divergent projections of catecholamine neurons of the locus coeruleus as revealed by fluorescent retrograde double labeling technique. Neurosci. Lett. 23, 117–123 (1981)
Steindler, D. A. Locus coeruleus neurons have axons that branch to the forebrain and cerebellum. Brain Res. 223, 367–373 (1981)
Swanson, L. W. Brain Architecture: Understanding the Basic Plan (Oxford Univ. Press, 2011)
Aston-Jones, G. & Bloom, F. E. Activity of norepinephrine-containing locus coeruleus neurons in behaving rats anticipates fluctuations in the sleep-waking cycle. J. Neurosci. 1, 876–886 (1981)
Gompf, H. S. et al. Locus ceruleus and anterior cingulate cortex sustain wakefulness in a novel environment. J. Neurosci. 30, 14543–14551 (2010)
Carter, M. E. et al. Tuning arousal with optogenetic modulation of locus coeruleus neurons. Nature Neurosci. 13, 1526–1533 (2010)
Van Bockstaele, E. J., Colago, E. E. & Valentino, R. J. Corticotropin-releasing factor-containing axon terminals synapse onto catecholamine dendrites and may presynaptically modulate other afferents in the rostral pole of the nucleus locus coeruleus in the rat brain. J. Comp. Neurol. 364, 523–534 (1996)
Stepien, A. E., Tripodi, M. & Arber, S. Monosynaptic rabies virus reveals premotor network organization and synaptic specificity of cholinergic partition cells. Neuron 68, 456–472 (2010)
Pivetta, C., Esposito, M. S., Sigrist, M. & Arber, S. Motor-circuit communication matrix from spinal cord to brainstem neurons revealed by developmental origin. Cell 156, 537–548 (2014)
Kinoshita, M. et al. Genetic dissection of the circuit for hand dexterity in primates. Nature 487, 235–238 (2012)
Luo, L., Callaway, E. M. & Svoboda, K. Genetic dissection of neural circuits. Neuron 57, 634–660 (2008)
Gong, S. et al. Targeting Cre recombinase to specific neuron populations with bacterial artificial chromosome constructs. J. Neurosci. 27, 9817–9823 (2007)
Zhang, X. M. et al. Highly restricted expression of Cre recombinase in cerebellar Purkinje cells. Genesis 40, 45–51 (2004)
Miyamichi, K. et al. Cortical representations of olfactory input by trans-synaptic tracing. Nature 472, 191–196 (2011)
Tasic, B. et al. Site-specific integrase-mediated transgenesis in mice via pronuclear injection. Proc. Natl Acad. Sci. USA 108, 7902–7907 (2011)
Hanawa, H. et al. Efficient gene transfer into rhesus repopulating hematopoietic stem cells using a simian immunodeficiency virus-based lentiviral vector system. Blood 103, 4062–4069 (2004)
Osakada, F. & Callaway, E. M. Design and generation of recombinant rabies virus vectors. Nature Protocols 8, 1583–1601 (2013)
Kremer, E. J., Boutin, S., Chillon, M. & Danos, O. Canine adenovirus vectors: an alternative for adenovirus-mediated gene transfer. J. Virol. 74, 505–512 (2000)
Weissbourd, B. et al. Presynaptic partners of dorsal raphe serotonergic and GABAergic neurons. Neuron 83, 645–662 (2014)
Schneider Gasser, E. M. et al. Immunofluorescence in brain sections: simultaneous detection of presynaptic and postsynaptic proteins in identified neurons. Nature Protocols 1, 1887–1897 (2006)
Acknowledgements
We thank N. Makki for initiating the locus coeruleus project, Stanford and UNC Viral Cores for producing AAVs, K. Touhara for support, D. Berns for suggesting the TRIO acronym, and members of the Luo laboratory, L. de Lecea and S. Hestrin for critiques. L.A.S. is supported by a Ruth L. Kirschstein National Service Research Award from NIMH, X.J.G. is supported by a Stanford Bio-X Enlight Foundation Interdisciplinary Fellowship, B.W. is supported by a Stanford Graduate Fellowship and an NSF Graduate Research Fellowship, E.J.K. is supported by EU FP7 BrainVector (no. 286071), K.M. was a Research Specialist and L.L. is an investigator of HHMI. This work is supported by an HHMI Collaborative Innovation Award.
Author information
Authors and Affiliations
Contributions
L.A.S. performed all the experiments on locus coeruleus input, output and TRIO analysis, as well as motor cortex TRIO experiments and analysis. K.M. designed the TRIO and cTRIO methods, made all the constructs, performed proof-of-principle experiments for TRIO along with B.W. and performed rat TRIO and mitral cell experiments. X.J.G. performed statistical analysis together with L.A.S. K.T.B. participated in testing TRIO and cTRIO conditions and is co-supervised by R.C.M. and L.L. K.E.D. and J.R. provided technical support. S.I. and E.J.K. produced CAV-FLExloxP-Flp. L.L. supervised the project and wrote the paper together with L.A.S., with contributions from all authors, in particular K.M. and X.J.G.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Controls for TRIO and cTRIO at the motor cortex.
a–c, Negative control experiments omitting CAV-Cre for TRIO (a), and omitting CAV-FLExloxP-Flp (b) or the Rpb4-Cre transgene (c) for cTRIO showed only local non-specific infection of RVdG. This background labelling is likely due to Cre- or Flp-independent leaky expression of a small amount of TVA–mCherry (TC), too low for mCherry to be detected but still capable of permitting infection by EnvA-pseudotyped RVdG due to the high sensitivity of TVA10. d, Quantification for three controls (n = 4, 4, 7 animals, respectively). By comparison, 672 GFP+ neurons were counted in the same region for an experimental brain that has the lowest starter cells among the 11 brains whose data were used for quantitative analysis of motor cortex TRIO input tracing. These background cells were restricted within ∼500 μm of the injection site. Because of these observations, GFP+ cells on sections within ∼600 μm of the injection site were excluded from the input analysis in Fig. 1g. Scale bar, 100 µm. Error bars, s.e.m.
Extended Data Figure 2 TRIO applied to rat primary motor cortex.
a, Schematic of injection sites used for TRIO in rat motor cortex (see Fig. 1c for details of the viruses). Two different C regions were tested: striatum or contralateral motor cortex (cMC). b, c, Coronal section of rat motor cortex stained with DAPI (blue). Starter pyramidal neurons projecting to contralateral motor cortex (b) or striatum (c) (yellow, a subset indicated by arrowheads) can be distinguished from neurons receiving CAV-Cre and AAV-FLExloxP-TC (red) or GFP from RVdG (green). Bottom insets, coronal sections showing representative presynaptic GFP+ cells in somatosensory cortex (SC) or thalamus (Th). These data indicate that callosal-projecting neurons and striatum-projecting neurons in rat motor cortex both receive direct synaptic input from somatosensory cortex and thalamus (n = 2 animals for cMC C region; n = 3 animals for striatum C region). d, e, Omitting CAV-Cre for TRIO in the rat also resulted in local non-specific infection of RVdG. On average ∼200 cells were observed (n = 4 animals) within 800 µm from the injection site in these control experiments. By comparison, 1,392 GFP+ neurons were counted in the same region of a TRIO sample that has the lowest starter cells among the 5 brains analysed. Scale bars, 100 µm. Error bars, s.e.m.
Extended Data Figure 3 Evaluation of CAV-Cre spread by using the OB→APC projection.
a, Four representative 60-μm coronal sections of the CAV-Cre injection site in the anterior piriform cortex (APC) of four Ai14 Cre-reporter mice. Red, tdTomato; green, retrobeads; blue, DAPI. The location of the injection site was readily visualized by concentrated retrobeads. D, dorsal; L, lateral; M, medial; V, ventral. In each mouse, we determined the minimal distance (D) between the injection site and layer 1a of the piriform cortex, where mitral cell axons terminate, or lateral olfactory tract, where mitral cell axon bundles are present. Dashed lines represent the boundary between layer 1a and layer 1b. For each sample, we counted the number of tdTomato-labelled mitral cells (numbers below each image) from serial olfactory bulb (OB) sections. b, An example 60-μm coronal section of the OB. Both tdTomato and retrobeads signals were found to be mostly restricted to the mitral cell layer (M) of the main olfactory bulb (MOB) and accessory olfactory bulb (AOB) with minor labelling in the granule cell layer (Gra). As AOB mitral cells do not form synapses in the APC, this observation indicates that CAV-Cre can infect axons-in-passage. c, Distribution of D among 26 injections (x axis) and relationship between D and the numbers of labelled cells in the MOB (y axis). d, Histogram based on c. Dense labelling (over 1,000) was obtained only when D < 100 μm. CAV-Cre injections with D > 800 μm rarely labelled the OB (2.8 ± 1.9 cells per bulb, n = 4 animals). e, Cumulative distribution plot of MOB cell counts. A sample of the ninth smallest D (D = 200 μm) reached 90% of the labelling (indicated by vertical dotted line) detected in all 26 samples, suggesting that given our sample distribution, ∼90% of axonal transduction occurred within 200 μm from the CAV-Cre injection site. Scale bars, 100 µm. Error bars, s.e.m.
Extended Data Figure 4 Evaluation of retrograde infection by CAV-Cre.
a, Representative coronal sections of the injection sites where CAV-Cre plus retrobeads were delivered into the olfactory bulb, dorsal hippocampus, auditory cortex, cerebellum or medulla of the Ai14 Cre-reporter mice (see Methods for coordinates). Red, tdTomato; green, retrobeads; blue, DAPI. tdTomato labelling was densest at the injection site, and corresponded with the presence of retrobeads. We did not observe dense tdTomato or retrobeads labelling in other brain regions adjacent to the injection site unless these sites sent direct projections to the injection site, indicating that for our experiments, CAV-Cre was efficiently and specifically delivered to the targeted brain regions. n = 4 animals per injection site. b, Representative coronal sections of brain regions that contained tdTomato+ labelling of specific cell populations known to project to CAV-Cre injection sites. The following is a partial list: neurons projecting to olfactory bulb (first column): ipsi- and contralateral anterior olfactory nucleus (AON), piriform cortex (Pir), nucleus of the lateral olfactory tract (nLOT), but not contralateral olfactory bulb; to dorsal hippocampus (second column): lateral and medial septum (LS, MS) and entorhinal cortex (Ent); to auditory cortex (third column): somatosensory cortex (SC), entorhinal cortex (Ent), and medial geniculate nucleus (MGN); to cerebellum (fourth column): contralateral pontine nuclei (PN) and inferior olive (IF); to medulla (fifth column): insular cortex (Ins), central amygdala (CeA), and paraventricular hypothalamic nucleus (PVH). Coronal images are composites generated from overlapping tiled images. Insets show high magnification images of boxed regions. Bottom, sagittal schematic of the CAV-Cre injection sites (a) and the approximate location of the two representative coronal sections above. Scale bars, 1 mm; inset, 100 µm.
Extended Data Figure 5 Controls for Dbh-Cre-based trans-synaptic tracing and TRIO analysis in locus coeruleus.
a, A Representative coronal section of the locus coeruleus from a mouse heterozygous for Dbh-Cre and Ai14 Cre-reporter transgenes. Sections were labelled with an antibody against tyrosine hydroxylase (TH), an enzyme in the biosynthetic pathway for noradrenaline (green), while cells expressing Cre recombinase are visible by expression of tdTomato (red). b, Quantification of the number of tdTomato+ neurons in the locus coeruleus that were also labelled by TH antibody (n = 3 animals). Every 50-µm section through the locus coeruleus was collected for quantification. Qualitatively, all TH+ cells expressed tdTomato; however, we cannot determine quantitatively because we could not accurately count TH+ cells due to dense process staining. c, Top, schematic for negative control where AAVs that express Cre-dependent TVA–mCherry fusion (TC) and rabies glycoprotein (G) were injected into the locus coeruleus of wild-type mice, followed by injection of RVdG. Middle, coronal section of the locus coeruleus stained with DAPI (blue) shows a small number of GFP+ neurons at the injection site. The dotted rectangle highlights GFP+ neurons magnified in the bottom panel. d, Top, in this negative control, Dbh-Cre mice received Cre-dependent TVA–mCherry fusion (without rabies glycoprotein) via AAV injection into the locus coeruleus, followed by RVdG. Middle, a coronal section of the locus coeruleus stained with DAPI (blue) shows infection of Cre+ locus coeruleus neurons with TC (red) or TC and RVdG (yellow) at the injection site. The dotted rectangle highlights infected locus coeruleus neurons magnified in the bottom panel. Most green cells are also red. No GFP+ cells were observed outside the region immediately adjacent to the injection site, indicating that trans-synaptic tracing depends on rabies glycoprotein. e, Quantification of the number of GFP+ cells (c), or GFP+ cells that did not colocalize with TC (d), that were observed in the experiments described in (c, n = 8 animals) and (d, n = 6 animals). By comparison, 1,381 GFP+ neurons were counted in the same region for an experimental brain that has the median number of starter cells among the 9 brains. For explanation of background labelling, see Extended Data Fig. 1a–c. In either case, no GFP+ neurons were visible >800 μm away from the injection site. f, Schematic of brain regions quantified for presynaptic GFP+ neurons. Regions approximately 800 μm anterior and posterior to the centre of the locus coeruleus were excluded from analysis due to local background labelling from TVA–mCherry fusion and GFP. Scale bars, 50 µm (a), 1 mm (c, d, middle panels), 100 µm (c, d, bottom panels). Error bars, s.e.m.
Extended Data Figure 6 Purkinje cell axons contact noradrenaline processes in the locus coeruleus.
a, Coronal sections counterstained with DAPI (blue) showing representative GFP+ Purkinje cells (green) from Dbh-Cre trans-synaptic tracing experiments described in Fig. 2. Labelled Purkinje cells span the anterior–posterior axis, but are enriched in the medial portion of the ipsilateral cerebellum. b, Sagittal section through the locus coeruleus of mice heterozygous for the transgenes Pcp2-Cre and Ai14, in which tdTomato (tdT) expression was restricted to cerebellar Purkinje cells and their processes (red). Sections were labelled with DAPI (blue) and anti-TH antibody (green) to label LC-NE neurons. The right panel is a maximum-projection confocal stack taken with a 40× objective of the boxed region in the left panel. Purkinje cell axons are intermingled with TH+ locus coeruleus neurons and their processes. c, Left, Representative image of a horizontal section collected through the locus coeruleus of a mouse heterozygous for the transgenes Pcp2-Cre and Ai14. Sections were stained with anti-TH antibody (green) to label LC-NE neurons and their processes, and anti-gephyrin (geph) antibody (white) to label inhibitory post-synaptic densities. Middle, maximum-projection confocal stack taken with a 40× objective of the dashed box of the left panel showing the overlap between tdTomato+ Purkinje cell axons and TH+ LC processes. Right, high magnification of the dashed box of the middle panel, showing that several of these contact points also contained gephyrin+ puncta (arrowheads) within green processes apposing the red processes, consistent with GABAergic Purkinje cell axons forming synapses onto dendrites of TH+ LC-NE neurons. Images in a were derived from larger composite images generated by a Leica Ariol Slide Scanner. A, anterior; D, dorsal; L, lateral; M, medial; P, posterior; V, ventral. Scale bars, 1 mm (a; b and c, left), 100 μm (a, inset; b, right), 10 μm (c, middle and right).
Extended Data Figure 7 Spatial distribution of LC-NE neurons projecting to distinct output brain regions.
a, Representative images of individual LC-NE neurons labelled within the locus coeruleus by injection of CAV-Cre at specific output sites (see Extended Data Fig. 4a) in Ai14 Cre-reporter mice. Coronal sections through the locus coeruleus were collected in order and stained with anti-TH antibody (pseudocoloured in green). All tdTomato (tdT)+ neurons within the locus coeruleus were also TH+, and many of these cells also contained retrobeads (green in inset). Injection of CAV-Cre into the olfactory bulb, hippocampus or auditory cortex resulted in high tdTomato expression in NE+ neurons within the locus coeruleus, whereas tdTomato labelling was almost completely absent in adjacent brain regions, indicating that regions next to the locus coeruleus contribute minimal projections to these output sites. However, CAV-Cre injected into the cerebellum or medulla labelled NE+ locus coeruleus neurons as well as adjacent, NE− cell populations (a subset of which are highlighted by arrowheads). b, The locations of tdTomato+ LC-NE neurons from sequential 50-μm coronal sections collected through the entire locus coeruleus were transferred to corresponding sections of a digital locus coeruleus model and are represented by coloured dots (see Methods). c, Schematic of the dorsal/ventral and medial/lateral classifications used with tdTomato+ LC-NE neurons occurring from CAV-Cre injections into the olfactory bulb (left) or cerebellum (right) of Ai14 mice. These classifications were made by drawing horizontal and vertical lines through the cross (b) designating the middle of each locus coeruleus section. d, Quantification of the fraction of tdTomato+ LC-NE cells in each locus coeruleus section along the anterior–posterior axis of the locus coeruleus. No significant differences were observed for the anterior–posterior distribution of tdTomato+ LC-NE neurons projecting to different output sites. e, Quantification of the medial–lateral distribution of LC-NE neurons projecting to different output sites. LC-NE neurons showed no bias in the medial versus lateral portion of the locus coeruleus, regardless of where they sent projections. f, Quantification of the dorsal–ventral distribution of tdTomato+ LC-NE neurons projecting to different output sites. Although no bias was observed in the posterior locus coeruleus, significant differences were observed in the anterior and mid-LC. Specifically, LC-NE neurons projecting to the forebrain showed a dorsal bias for tdTomato+ cell labelling within the anterior locus coeruleus, whereas LC-NE neurons projecting to the cerebellum and medulla were located in more ventral portions of the anterior- and mid-LC. n = 4 animals per CAV injection site. Data in d was analysed with one-way ANOVA. Data in e, f were analysed by first performing two-way ANOVA, which did not uncover any significance in the medial/lateral bias of tdTomato+ LC-NE neurons. Two-way ANOVA determined that (1) the location of the CAV injection site contributes to the dorsal/ventral bias of tdTomato+ LC-NE neurons within the locus coeruleus (P < 0.0001), (2) there is interaction between the CAV injection site and the location (anterior, mid, posterior) of tdTomato+ NE neurons within the locus coeruleus (P = 0.0389), and (3) the locus coeruleus subdivisions themselves did not significantly contribute to the variance observed in tdTomato+ LC-NE neurons. One-way ANOVA and post hoc Tukey’s multiple comparison were then performed to test the significance of dorsal/ventral bias in each locus coeruleus region based on CAV injection sites. Scale bars, 50 µm. Error bars represent s.e.m. *P < 0.05; **P < 0.01, ***P < 0.001.
Extended Data Figure 8 Controls for locus coeruleus cTRIO.
a, Top, schematic for negative controls where AAVs expressing Flp-dependent TVA–mCherry fusion and rabies glycoprotein were injected into the locus coeruleus of Dbh-Cre mice, followed by RVdG injection into the locus coeruleus, but the CAV-FLExloxP-Flp injection was omitted. Middle, coronal section of the locus coeruleus stained with DAPI (blue) shows a small number of GFP+ neurons at the injection site. The dotted rectangle highlights GFP+ neurons magnified in the lower panel. b, Top, schematic for negative control where CAV-FLExloxP-Flp was injected into the olfactory bulb and AAVs expressing Flp-dependent TVA–mCherry fusion and rabies glycoprotein were injected into the locus coeruleus of wild-type mice, followed by RVdG injection; hence there was no Cre to mediate Flp expression in locus coeruleus cells. Middle, coronal section of the locus coeruleus stained with DAPI (blue) shows a small number of GFP+ neurons at the injection site. The dotted rectangle highlights GFP+ neurons magnified in the lower panel. c, Quantification of GFP+ background labelling in the locus coeruleus (n = 4 and 8 animals). This labelling is likely caused by leaky TVA expression as discussed in Extended Data Fig. 1. In none of these control experiments did we observe GFP+ or TC+ neurons >800 μm away from the injection site. Scale bars, 1 mm (middle panels), 100 µm (lower panels). Error bars represent s.e.m.
Extended Data Figure 9 Simulation of input convergence in Dbh-Cre tracing experiments.
In the sparsest Dbh-Cre trans-synaptic tracing brain, 4 starter cells received input from 43 distinct input regions (309 input neurons, see Supplementary Table 2, sample number 8). In the second sparsest sample, 22 starter cells received input from 66 distinct input regions (756 input neurons; see Supplementary Table 2, sample number 9). a, The relation between the number of input regions for each LC-NE starter cell and the probability of observing >42 (left) or >65 (right) input regions in simulation, assuming that each starter cell receives input from a given region with the same probability. As the number of input regions per starter cell increases, the probability of observing inputs from >42 or >65 regions also increases. Based on a threshold of P value <0.001, these simulations suggest that, to account for the total number of observed input areas in each brain sample, there must be individual LC-NE neurons that receive input from more than 15 regions for the sparsest sample (red dot, left) or more than 9 regions for second sparsest sample (red dot, right). b, Detailed view of the distribution of simulation results corresponding to the red dots in a. Assuming that each cell receives input from 15 (left) or 9 (right) distinct regions, only 5 (left) or 6 (right) out of 10,000 simulations label >42 (left) or >65 (right) input regions. Note that if the assumption that each starter cell receives input from the same number of regions does not apply, then there must be at least one cell receiving input from more regions than the number specified in the simulation.
Extended Data Figure 10 Representative images and distribution of individual samples for projection-based viral-genetic labelling experiments.
a, Representative images from sagittal sections of TC+ LC-NE axons in 8 brain regions indicated at the top of each column (the last column shows cell bodies for LC-NE neurons) resulting from CAV injections at four projection sites indicated on the left (top four rows), or AAV-FLExloxP-TC injection at the locus coeruleus of Dbh-Cre animals (bottom row). All TC+ processes were confirmed to contain noradrenaline transporter (NET, an NE neuron marker) by anti-NET immunostaining (not shown; see Fig. 4 inset). b, The normalized fraction of TC+ LC-NE axons for individual experiments for five conditions are colour coded on the top right. Filled symbols represent experiments where Dbh-Cre mice were used along with CAV-FLExloxP-Flp; open symbols represent experiments where wild-type mice were used along with CAV-Cre. The distribution of individual samples with regards to the fraction of TC+ axons observed at output sites was similar between wild-type and Dbh-Cre mice. Collectively, the samples for each condition were averaged to quantify the normalized fraction of TC+ LC-NE axons in each brain region as reported in Fig. 4d. Scale bar, 50 µm. Error bars represent s.e.m. Abbreviations: AC, auditory cortex; CC, cingulate cortex; Cb, cerebellum; Hi, hippocampus; Hy, hypothalamus; LC, locus coeruleus; Me, medulla; OB, olfactory bulb; SC, somatosensory cortex.
Supplementary information
Supplementary Information
This file contains Supplementary Notes 1-2 and additional references. (PDF 157 kb)
Supplementary Table 1
Distribution of presynaptic input to layer 5 motor cortex neurons for Rbp4-Cre input tracing and cTRIO. (XLSX 12 kb)
Supplementary Table 2
Distribution of presynaptic input to locus coeruleus norepinephrine neurons for Dbh-Cre input tracing, TRIO, and cTRIO. (XLSX 134 kb)
Supplementary Table 3
One-way ANOVA p-values for Dbh-Cre input tracing, TRIO, and cTRIO comparisons. (XLSX 17 kb)
Supplementary Table 4
Contingency table for testing input heterogeneity. (XLSX 14 kb)
Rights and permissions
About this article
Cite this article
Schwarz, L., Miyamichi, K., Gao, X. et al. Viral-genetic tracing of the input–output organization of a central noradrenaline circuit. Nature 524, 88–92 (2015). https://doi.org/10.1038/nature14600
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14600
This article is cited by
-
The crucial role of locus coeruleus noradrenergic neurons in the interaction between acute sleep disturbance and headache
The Journal of Headache and Pain (2024)
-
LC-derived excitatory synaptic transmission to dorsal raphe serotonin neurons is inhibited by activation of alpha2-adrenergic receptors
Neuropsychopharmacology (2024)
-
Identification of a novel perifornical-hypothalamic-area-projecting serotonergic system that inhibits innate panic and conditioned fear responses
Translational Psychiatry (2024)
-
A Novel Retrograde AAV Variant for Functional Manipulation of Cortical Projection Neurons in Mice and Monkeys
Neuroscience Bulletin (2024)
-
The locus coeruleus input to the rostral ventromedial medulla mediates stress-induced colorectal visceral pain
Acta Neuropathologica Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.