Abstract
Darwin’s finches, inhabiting the Galápagos archipelago and Cocos Island, constitute an iconic model for studies of speciation and adaptive evolution. Here we report the results of whole-genome re-sequencing of 120 individuals representing all of the Darwin’s finch species and two close relatives. Phylogenetic analysis reveals important discrepancies with the phenotype-based taxonomy. We find extensive evidence for interspecific gene flow throughout the radiation. Hybridization has given rise to species of mixed ancestry. A 240 kilobase haplotype encompassing the ALX1 gene that encodes a transcription factor affecting craniofacial development is strongly associated with beak shape diversity across Darwin's finch species as well as within the medium ground finch (Geospiza fortis), a species that has undergone rapid evolution of beak shape in response to environmental changes. The ALX1 haplotype has contributed to diversification of beak shapes among the Darwin’s finches and, thereby, to an expanded utilization of food resources.
Similar content being viewed by others
Main
Adaptive radiations are particularly informative for understanding the ecological and genetic basis of biodiversity1,2. Those causes are best identified in young radiations, as they represent the early stages of diversification when phenotypic transitions between species are small and interpretable and extinctions are likely to be minimal3. Darwin’s finches are a classic example of such a young adaptive radiation3,4. They have diversified in beak sizes and shapes, feeding habits and diets in adapting to different food resources4,5 (Extended Data Table 1). The radiation is entirely intact, unlike most other radiations, none of the species having become extinct as a result of human activities4.
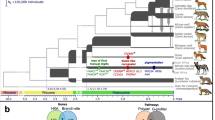
Fourteen of the currently recognized species evolved from a common ancestor in the Galápagos archipelago (Fig. 1a) in the past 1.5 million years according to mitochondrial DNA (mtDNA) dating6; a fifteenth species inhabits Cocos Island. The radiation proceeded rapidly as a result of strong isolation from the South American continent, generation of new islands by volcanic activity, climatic oscillations caused by the El Niño phenomenon, and sea level changes associated with glacial and interglacial cycles over the past million years that led to repeated alternations of island formation and coalescence7,8.
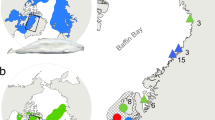
a, Geographical origin of samples; the letter after the species name is the abbreviation used for geographical origin. The map is modified from ref. 30. b, Maximum-likelihood trees based on all autosomal sites; all nodes having full local support on the basis of the Shimodaira–Hasegawa test are marked by asterisks. The colour code for groups of species applies to both panels. Taxa that showed deviations from classical taxonomy are underscored.
Traditional taxonomy of Darwin’s finches is based on morphology3, and has been largely supported by observations of breeding birds4,5 and genetic analysis6,9. However, the branching order of several recently diverged taxa is unresolved6 and genetic analysis of phylogeny has been limited to mtDNA and a few microsatellite loci. Some candidate genes for beak development are differentially expressed in species with different beak morphologies10,11,12, but the loci controlling genetic variation in beak diversity among Darwin’s finches remain to be discovered.
Here we report results from whole genome re-sequencing of 120 individuals representing all Darwin’s finch species and two closely related tanagers, Tiaris bicolor and Loxigilla noctis13. For some species we collected samples from multiple islands (Fig. 1a). We comprehensively analyse patterns of intra- and interspecific genome diversity and phylogenetic relationships among species. We find widespread evidence of interspecific gene flow that may have enhanced evolutionary diversification throughout phylogeny, and report the discovery of a locus with a major effect on beak shape.
Considerable nucleotide diversity
We generated approximately 10× sequence coverage per individual bird using 2 × 100 base-pair (bp) paired-end reads (Extended Data Fig. 1). Reads were aligned to the genome assembly of a female medium ground finch (G. fortis)14. We identified Z- and W-linked scaffolds on the basis of significant differences in read depth between males (ZZ) and females (ZW) (Supplementary Table 1) and generated a G. fortis mtDNA sequence through a combined bioinformatics and experimental approach. Stringent variant calling revealed approximately 45 million variable sites within or between populations. We found a considerable amount of genetic diversity within each population, in the range 0.3 × 10−3 to 2.2 × 10−3 (Extended Data Table 2), similar to that reported in other bird populations15 including island populations of the zebra finch16. We used these estimates of diversity to estimate effective population sizes of Darwin’s finch species within a range of 6,000–60,000 (Supplementary Text). Extensive sharing of genetic variation among populations was evident, particularly among ground and tree finches, with almost no fixed differences between species in each group (Extended Data Fig. 2).
Genome-based phylogeny
According to the classical taxonomy of Darwin’s finches, supported by morphological and mitochondrial (cytochrome b) data, warbler finches were the first to branch off, and ground and tree finches constitute the most recent major split3,6,9. Our maximum-likelihood phylogenetic tree based on autosomal genome sequences is generally consistent with current taxonomy, but shows several interesting deviations (Fig. 1b). First, Geospiza difficilis occurring on six different islands forms a polyphyletic group separated into three distinct groups: (1) populations occupying the highlands of Pinta, Santiago and Fernandina, (2) populations occupying the low islands of Wolf and Darwin in the northwest3,6,9 and (3) the population on Genovesa in the northeast. This is consistent with an earlier version of the taxonomy, in which these three groups were classified as distinct species on the basis of morphological differences17,18.
Second, Geospiza conirostris on Española showed the highest genetic similarity to another species, Geospiza magnirostris, whereas G. conirostris on Genovesa clustered with Geospiza scandens (Fig. 1b). Here, phenotypic similarity parallels genetic similarity; G. conirostris on Genovesa have a pointed beak similar to G. scandens, whereas those on Española have a blunt beak more similar to the beaks of G. magnirostris (Extended Data Fig. 3).
A network constructed from autosomal genome sequences indicates conflicting signals in the internal branches of ground and tree finches that may reflect incomplete lineage sorting and/or gene flow (Extended Data Fig. 3). The exact branching order of the most recently evolved ground and tree finches should be interpreted with caution as it may change with additional sampling. Since our data revealed some important discrepancies with the phenotype-based taxonomy, we propose a revised taxonomy for the sharp-beaked ground finch (G. difficilis) and the large cactus finch (G. conirostris) (Supplementary Text and Extended Data Fig. 4), but will use the current names in the text.
We dated phylogenetic splits on the basis of genome divergence (Fig. 2a), and compared these estimates with those obtained using mtDNA (Extended Data Fig. 5a and Supplementary Text). We infer that the most basal split, between warbler finches (Certhidea sp.) and other finches, occurred about 900,000 years ago. The rapid radiations of ground and tree finches began around 100,000–300,000 years ago. Although these estimates are based on whole-genome data, they should be considered minimum times, as they do not take into account gene flow.
a, Dating the nodes (in thousands of years) with confidence intervals (when applicable) in the phylogeny on the basis of divergence corrected for coalescence in ancestral populations; the topology is the representation of the inferred species tree from Fig. 1b. b, ABBA–BABA analysis of G. magnirostris, G. difficilis from Wolf and Pinta, and L. noctis. Number of sites supporting different trees is indicated both as a percentage and as actual numbers. The D statistic and corresponding Holm–Bonferroni-corrected P value are given for testing the null hypothesis of symmetry in genetic relationships. Finch heads are reproduced from ref. 5. How and Why Species Multiply: The Radiation of Darwin's Finches by Peter R. Grant & B. Rosemary Grant. Copyright © 2008 Princeton University Press. Reprinted by permission.
Extensive interspecies gene flow
The discrepancies between phylogenies based on morphology and genome sequences may be due to convergent evolution and/or interspecies gene flow. We found evidence of introgression from three sources: ABBA–BABA tests, discrepancies between phylogenetic trees based on autosomal and sex-linked loci, and mtDNA (Supplementary Text and Extended Data Fig. 5a).
First, the D statistic19 associated with the ABBA–BABA test was used to compare two populations of G. difficilis from Pinta and Wolf, and G. magnirostris from Genovesa, using L. noctis as outgroup; G. magnirostris also occurs on Wolf but we lacked samples from that population. The analysis confirmed that G. difficilis on Wolf has a closer genetic relationship with G. magnirostris than with G. difficilis on Pinta (Fig. 2b). But there is evidence of gene flow between G. difficilis on Wolf and Pinta (P = 5 × 10−113), because the substantial asymmetry in genetic relationships cannot be explained by incomplete lineage sorting. However, the D statistic does not distinguish admixture from ancestral subdivision19. We conclude that the closely related populations of G. difficilis on Wolf and Darwin are a species of mixed ancestry where most of the genome originates from G. magnirostris or a close relative (Supplementary Table 2), whereas a considerable proportion of the genome, possibly including genetic variants affecting phenotypic characters, is derived from G. difficilis. Similarly, G. difficilis on Genovesa shows a closer genetic relationship to the other ground and tree finches than to G. difficilis on Pinta, but we also found evidence for gene flow between the two groups previously classified as G. difficilis (P = 3 × 10−87; Supplementary Table 2).
We next investigated gene flow involving the populations of G. conirostris on Genovesa and Española, which appear as separate species in our phylogenetic analysis. The ABBA–BABA analysis confirmed that G. conirostris on Española shows a closer genetic relationship to G. magnirostris than to G. conirostris on Genovesa (Extended Data Fig. 6a), but also provided evidence for gene flow between G. conirostris on Española and G. conirostris on Genovesa, which may explain some of their phenotypic similarities and their previous classification as a single species.
Given the evidence of relatively recent hybridization, we explored the possibility of more ancient hybridization between warbler finches (Certhidea fusca and Certhidea olivacea) and other finches. ABBA–BABA analysis provided evidence for gene flow between C. fusca and the other finches (P = 7 × 10−199; Extended Data Fig. 6b). This pattern of gene flow was apparent for all non-warbler finches, implying that it occurred before the radiation of the non-warbler finches (Supplementary Table 2).
The trees based on autosomal (Fig. 1b) and Z-linked sites (Extended Data Fig. 5b) are not completely congruent. The tree based on Z-linked polymorphisms indicated that G. difficilis present on the highlands of Pinta, Fernandina and Santiago is more closely related to Platyspiza crassirostris and emerged before the Cocos finch split off from the ground and tree finches, whereas the autosomal tree indicates a reversed order for the emergence of the two species. This discrepancy can potentially be explained by gene flow between G. difficilis and tree and ground finches after the Cocos finch became reproductively isolated from the finches on the Galápagos, which affected Z-linked and autosomal loci to different degrees. It is a common observation in closely related species that there is more interspecies sharing of sequence polymorphisms at autosomal loci than at sex-linked loci20. This interpretation of the phylogenetic status of G. difficilis (highland group) is supported by the trees based on both mtDNA and W (Extended Data Fig. 5), which suggest that G. difficilis diverged from the ancestor of other ground and tree finches before the emergence of the Cocos finch.
Finally, our analysis of demographic history using the pairwise sequentially Markovian coalescent (PSMC) model21 was consistent with extensive interspecies gene flow among the ground finches, as they have maintained larger effective population sizes than the other species (Supplementary Text and Extended Data Fig. 6c, d).
A major locus controlling beak shape
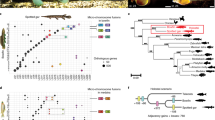
The most striking morphological difference among Darwin’s finches concerns beak shape (Extended Data Fig. 3). We performed a genome-wide scan on the basis of populations that are closely related but show different beak morphology: G. magnirostris and G. conirostris on Española have blunt beaks, whereas G. conirostris on Genovesa and G. difficilis on Wolf have pointed beaks. We used non-overlapping 15-kilobase (kb) windows to identify regions with the highest fixation indices (FST) between groups. The FST distribution was Z-transformed (ZFST) and regions with striking ZFST values were identified (Fig. 3a). Among the 15 most significant regions, six harboured genes previously associated with craniofacial and/or beak development in mammals or birds including calmodulin (CALM)11, goosecoid homeobox (GSC)22, retinol dehydrogenase 14 (RDH14)23, ALX homeobox 1 (ALX1)24,25, fibroblast growth factor 10 (FGF10)26 and forkhead box C1 (FOXC1)27. A previous study demonstrated differential expression of CALM between finches with different beak types11. Two other studies reported differential expression of bone morphogenetic protein 4 (BMP4)10,12, but we did not observe any elevated ZFST values in the vicinity of this locus, suggesting that differential expression is controlled by other loci.
a, Genome-wide FST screen comparing G. magnirostris and G. conirostris (Española) having blunt beaks with G. conirostris (Genovesa) and G. difficilis (Wolf) having pointed beaks. The y axis represents ZFST values. b, Nucleotide diversities in the ALX1 region. The 240-kb region showing high homozygosity in blunt-beaked species is highlighted. Red and blue colours in b–d refer to blunt and pointed beak haplotypes, respectively. c, Neighbour-joining haplotype tree of ALX1 region. Haplotypes originating from heterozygous birds (see text) are indicated in yellow. Estimated time since divergence (± confidence interval) of blunt and pointed beak haplotypes are given in thousands of years. d, Upper panel: genotypes at 335 SNPs showing complete fixation between ALX1 haplotypes associated with blunt (B) and pointed (P) beaks. d, Middle panel: classification of alleles associated with blunt beaks at the 335 SNPs as derived or ancestral on the basis of allelic state in the outgroup. d, Lower panel: PhastCons35 scores (on the basis of human, mouse and finch alignments) for the 335 SNP sites. TFBS, transcription factor binding sites. e, Linear regression analysis of beak-shape scores among G. fortis individuals on Daphne Major Island classified according to ALX1 genotype; distribution of pointedness in each class is shown as a boxplot; n = 62; F = 17.7, adjusted R2 = 0.22. Differences in six individual body and beak size traits were not significant (all P > 0.05).
The most striking finding was a 240-kb region with high ZFST values, including the window with the highest ZFST score (9.46) overall (Fig. 3a, b). The region overlaps part of LRRIQ1 (leucine-rich repeats and IQ motif containing 1), the entire ALX1 gene and about 130 kb downstream of ALX1. No previous report indicates that LRRIQ1 has a role during development in vertebrates. By contrast, ALX1 is an excellent candidate for variation in beak morphology. It encodes a paired-type homeodomain protein that plays a crucial role in development of structures derived from craniofacial mesenchyme, the first branchial arch and the limb bud24, and on migration of cranial neural crest cells, highly relevant to beak development25. Loss of ALX1 in humans causes disruption of early craniofacial development24.
All individuals in the blunt beak category were homozygous for a blunt beak-associated haplotype (denoted B), except one heterozygous G. conirostris individual from Española. Furthermore, except for one heterozygous bird from Genovesa, all 19 G. difficilis individuals not included in the FST scan were homozygous for a pointed beak haplotype (P), consistent with their phenotypic appearance (sharp-beaked ground finches). This is notable because genome-wide, G. difficilis on Wolf, Darwin and Genovesa are all more closely related to the blunt-beaked G. magnirostris than to the pointed-beaked G. difficilis from Pinta (Fig. 2b).
A phylogenetic tree based on this region revealed a deep divergence between the B and P haplotypes that must have occurred soon after the split between warbler finches and other Darwin’s finches (Fig. 3c). Apart from the blunt-beaked G. magnirostris and G. conirostris on Española, all individuals except three were homozygous for P haplotypes, the remaining three being heterozygous. The two G. fortis from Daphne Major Island were both homozygous, but for different haplotypes (BB and PP; Fig. 3c). The short branch lengths among B haplotypes are consistent with a selective sweep. There were 335 fixed differences between the B and P haplotypes (Fig. 3d, upper panel), which we assigned as derived or ancestral on the basis of comparison with the outgroup sequence (L. noctis). Derived alleles on the B haplotype were aggregated in the vicinity of ALX1, including the downstream region (Fig. 3d, middle panel). Furthermore, 8 of these 335 fixed differences occurred at conserved sites, and the B haplotype carried the derived allele at seven of them (Fig. 3d, lower panel). Four derived alleles occurred at sites corresponding to transcription factor binding sites in the human genome28. Two other changes constitute missense mutations (L112P and I208V) at ALX1 amino-acid residues that are highly conserved among birds and mammals (Extended Data Fig. 7), and ‘Sorting Intolerant From Tolerant’ (SIFT)29 analysis classified both as damaging (score 0.03 for both). The ratio of non-synonymous to synonymous substitutions between the P and B alleles is high (2/1 = 2.00) compared with the ratio observed between the ancestral P allele and orthologous zebra finch (2/14 = 0.14) and human (21/122 = 0.17) sequences, suggesting that one or both of these missense mutations are non-neutral.
That ALX1 is polymorphic in G. fortis (Fig. 3c, d, upper panel) is particularly interesting, because field observations have shown there is considerable diversity in beak shape in this species5,30. We genotyped an additional 62 G. fortis birds from Daphne Major Island for a diagnostic single nucleotide polymorphism (SNP), and observed a significant association with beak shape (P = 8.8 × 10−5, Fig. 3e). PP homozygotes tended to have proportionately long, pointed beaks, BB homozygotes had proportionately deep, blunt beaks, whereas heterozygotes (BP) had intermediate beak shapes. We also compared haplotype frequencies among G. fortis individuals on Daphne Major Island with those on Santa Cruz, which have a larger and blunter beak on average31, possibly as a result of introgressive hybridization with G. magnirostris4,5. We found the B haplotype to be more frequent on Santa Cruz than on Daphne Major (0.74, n = 21 versus 0.49, n = 62; P = 0.007, Fisher’s exact test).
Natural selection on beak size and shape of G. fortis on Daphne Major Island has led to evolutionary change in the past few decades5,30. Moreover, genetic variation in beak shape has been increased through introgressive hybridization5,30 with two species of Geospiza, scandens and fuliginosa, that have relatively pointed beaks. Therefore we expect hybrids and backcrosses in the G. fortis population to have a relatively high frequency of the P haplotype. We genotyped an additional 25 G. fortis at ALX1, added them to the sample of 62 (Methods) and compared the haplotype frequencies in eight hybrids (including backcrosses) and 79 non-hybrids. ALX1-P had a frequency of 0.75 among hybrids, and 0.44 among the others, which is statistically significant in the expected direction (P = 0.03, Fisher’s exact test). Thus, ALX1-P alleles introduced by introgressive hybridization most probably contributed to evolution of more pointed beaks in 1987 following natural selection as a result of a change in food supply in the 1985–86 drought30.
Discussion
Our revised and dated phylogeny of Darwin’s finches shows that the adaptive radiation took place in the past million years, with a rapid accumulation of species recently (Supplementary Text). We have genomically characterized the entire radiation, which has revealed a striking connection between past and present evolution. Evidence of introgressive hybridization, which has been documented as a contemporary process, is found throughout the radiation. Hybridization has given rise to species of mixed ancestry, in the past (this study) and the present30. It has influenced the evolution of a key phenotypic trait: beak shape. Similar introgressive hybridization affecting an adaptive trait (mimicry) has been described in Heliconius butterflies32. The degree of continuity between historical and contemporary evolution is unexpected because introgressive hybridization plays no part in traditional accounts of adaptive radiations of animals1,2. For young radiations it complements the better-known role of natural selection.
Charles Darwin first noted the diversity in beak shapes among the finches on Galápagos. Our genomic study has now revealed some of the underlying genetic variation explaining this diversity. A polygenic basis for beak diversity is indicated by our discovery of about 15 regions with strong genetic differentiation between groups of finches with blunt or pointed beaks. We present evidence that the ALX1 locus contributes to beak diversity, within and among species. The derived ALX1-B haplotype associated with blunt beaks has a long evolutionary history (hundreds of thousands of years), because its origin predates the radiation of vegetarian, tree and ground finches (Fig. 3c). This haplotype is fixed or nearly fixed in two ground finches with blunt beaks, G. magnirostris and G. conirostris on Española, and it co-segregates with variation in beak shape in G. fortis. As previously documented in domestic animals33 and natural populations34, the haplotype might have evolved by accumulating both coding and regulatory changes affecting ALX1 function. Natural selection and introgression affecting this locus have contributed to the diversification of beak shapes among Darwin’s finches and hence to their expanded utilization of food resources on Galápagos.
Methods
Study samples
No statistical methods were used to predetermine sample size. Blood samples from a total of 200 individuals of Darwin’s finches, captured in mist nets and then released, were collected on FTA papers and stored at −70 °C until DNA preparation. These included all 15 species of Darwin’s finches currently present on the Galápagos and Cocos Island, and two closely related tanagers from Barbados used as outgroups13. Details on the name of each species, the specific island where they were sampled and the total number of individuals sampled from each species are in Extended Data Table 2 and phenotype descriptions of each species are in Extended Data Table 1.
Whole-genome sequencing
DNA was isolated from pieces of FTA papers using DNeasy tissue kit (QIAGEN). Each DNA sample was uniquely tagged with a sequence index during multiplexing library preparation protocol. The libraries (average fragment size about 400 bp) were sequenced using Illumina Hiseq2000 sequencers and 2 × 100 bp paired-end reads were generated. The amount of sequence per bird was targeted to approximately 10× coverage.
Reference genome assembly
Sequence reads were aligned to the genome assembly of a female medium ground finch (G. fortis)14. This draft genome assembly has a size of ∼1.07 Gb with scaffold N50 size of ∼5.2 Mb and contig N50 size of ∼30 kb. The annotation of the genome included a total of 16,286 protein-coding genes.
In addition, as the complete sequence for mtDNA was not previously available for any of the Darwin’s finches, we also generated an assembly of the mtDNA genome sequence. For this, we first mapped all reads from one G. fortis individual against the zebra finch (Taeniopygia guttata) mtDNA. All the aligned reads were locally reassembled using SOAP DENOVO36, and then the gaps between the contigs were filled using Sanger sequencing to generate a single mtDNA genome sequence of 16.8 kb in length.
Sequence alignment and variant calling
The short sequence reads (2 × 100 bp) were quality checked using FASTQC (http://www.bioinformatics.babraham.ac.uk/projects/fastqc/). Then we used BWA37 (version 0.6.2) with default parameters to map the genomic reads from each individual against the reference genome assembly. The alignments were further checked for PCR duplicates using PICARD (http://picard.sourceforge.net/). We used Genome Analysis Toolkit (GATK)38 for base quality recalibrations, insertion/deletion (INDEL) realignment, SNP and INDEL discovery and genotyping across all 120 samples simultaneously according to GATK best practice recommendations39,40.
Quality filtering of the raw variant calls was done according to an in-house filtering pipeline that excluded a variant as low quality if it did not satisfy the following cut offs for filtering: SNP quality > 100, base quality > 30, mapping quality > 50, haplotype score < 10, Fisher strand bias <60, mapping quality rank sum > −4.0, read position rank sum > −2.0, quality by depth > 2.0, minimum depth (summing all 120 samples) > 125, and maximum depth (summing all 120 samples) < 1,875. These parameters are explained in detail in the GATK user manual39. The cut-offs were chosen on the basis of the distribution of each of these parameters from the raw variant calls generated by the GATK UnifiedGenotyper module. The missing and low quality genotypes from the call set were inferred separately for each population using BEAGLE (version 3.3.2)41. Finally, we retained 44,753,624 variable sites in the data set. The variant calling in mtDNA was also performed using a similar BWA and GATK pipeline as described above. We identified 1,429 mtDNA variable sites in mtDNA. We calculated the average nucleotide diversity for autosomes, chromosomes Z and W, and in the mtDNA genome separately to estimate the amount of genetic variation in each population in different parts of the genome.
Identification of scaffolds from chromosomes Z and W
The medium ground-finch genome assembly contains 27,239 scaffolds unassigned to chromosomes. We used the MultiSV package to identify scaffolds that belong to chromosomes Z and W by comparing the read depth for each scaffold in 85 males and 35 females. This analysis identified 133 scaffolds, which belonged to chromosome Z with a total length of 67,176,652 bp (Supplementary Table 1a), and 662 scaffolds, which belonged to chromosome W with a total length of 643,111 bp (Supplementary Table 1b).
Estimation of genetic distance and phylogeny reconstruction
We used PLINK (version 1.07)42 to calculate genetic distance (on the basis of proportion of alleles identical by state) for all pairs of individuals separately for autosomes and the Z chromosome. We used the neighbour-net method of SplitsTree4 (http://www.splitstree.org/) to compute the phylogenetic network from genetic distances. We used FastTree to infer approximately maximum-likelihood phylogenies with standard parameters for nucleotide alignments of variable positions in the data set (http://meta.microbesonline.org/fasttree/). FastTree computes local support values with the Shimodaira–Hasegawa test.
ABBA–BABA analysis
Patterns of gene flow and the extent of admixture in populations were analysed and tested for asymmetry in the frequencies of discordant gene trees in a three-population phylogeny rooted with an outgroup using the D statistic43 as implemented for polymorphic sites19. The D statistics were transformed to Z scores by division with the standard error, which was calculated with a jackknife procedure. Blocks of 40,000 variable sites for autosomes and 10,000 for the Z chromosome were used in the jackknife to overcome the effect of linkage disequilibrium, which yielded 1,027 and 291 blocks, respectively. The Z scores were translated to two-sided P values that were Holm–Bonferroni-corrected44 for multiple testing by stepwise division of the lowest P value with the remaining number of tests performed for all 1,768 possible tests in the phylogeny and the two tests with pooled species (Supplementary Table 2).
Mutation rates
We used the following previously reported estimated mutation rates for nuclear and mtDNA: nuclear DNA, 2.04 × 10−9 per site per year estimated from the synonymous mutation rate on the Darwin’s finches’ lineage since the split from zebra finch45; mtDNA, a fossil-calibrated divergence rate of 2.1% per million years for bird cytochrome b sequences46.
Estimation of effective population size
Effective population sizes (Ne) were calculated from Watterson’s θ (ref. 47) across the whole genome and the above-mentioned mutation rate. Fluctuations in Ne were inferred using PSMC37 and with ‘64*1’ as the time interval parameter pattern. Plots were scaled assuming a mutation rate per generation of 1.02 × 10−8 and a generation time of 5 years (ref. 48).
Dating the nodes in the phylogeny and demographic history
Times of population splits were calculated with our estimates of genetic distances in the two subtrees of a node and corrected for the time to coalescence in ancestral populations49 and mutation rate. Confidence intervals were estimated from the standard deviation of genetic distances estimated from the pairwise species comparisons. We estimated the time of divergence between the blunt and pointed ALX1 haplotypes by estimating the average pairwise difference at this locus between species containing all blunt and all pointed haplotypes and correcting for mutation rate. G. fortis and heterozygous individuals were excluded. Cytochrome b sequences were used to date the mtDNA phylogeny in which the most recently evolved ground finches (that is, G. magnirostris, conirostris, scandens, fortis, fuliginosa and difficilis on Genovesa) were treated as one population, with diversities averaged across species, because they did not form monophyletic groups according to species.
To elucidate and display the demographic history of Darwin’s finches we used the pairwise sequentially Markovian coalescent (PSMC) model, which infers fluctuations in effective population size over evolutionary time from a single genome sequence21.
Signatures of selection for beak diversification
We scanned the whole genome in non-overlapping 15-kb windows to identify regions with increased genetic divergence (FST) between species with blunt and pointed beaks. We used VCFtools version 0.1.11 (ref. 50) to calculate FST. The genomic windows with high ZFST (>6) were analysed for gene content.
ALX1 genotyping in additional samples
A Taqman SNP genotyping assay (Life Technologies) was designed for one SNP (A/C at nucleotide position 517,149 bp in scaffold JH739921) diagnostic for the ALX1 haplotypes associated with blunt and pointed beaks. A standard TaqMan Allele discrimination assay was performed using an Applied Biosystems 7900 HT real-time PCR instrument. The association of individual genotypes with beak shape measurements was evaluated using standard linear regression in R.
Comparison of ALX1 protein sequences among vertebrates
The ALX1 protein sequence for G. fortis was downloaded from NCBI (XP_005421635). This G. fortis protein is a representative for the pointed allele and was edited to create a blunt counterpart by introducing the two amino-acid substitutions (L112P and I208V). ALX1 protein sequences from other species were collected from predicted orthologues of the chicken ALX1 gene in Ensembl51, including representative species from teleosts, reptiles, birds and mammals. The protein sequences were aligned using MUSCLE52 (version 3.8.31) with default settings, and the multiple sequence alignment was viewed and edited using Jalview29,53. The probability of functional consequences of amino-acid substitutions was predicted using SIFT29 with the multiple sequence alignment as input after exclusion of the blunt allele. Both substitutions were predicted to be damaging with probability scores of 0.03, where a score less than 0.05 is considered significant. Both predictions were reported to have a low confidence due to limited divergence in the alignment. However, we argue that because we have sampled orthologues from such a diverse set of species where ALX1 displays considerable conservation, these predictions can be viewed with greater confidence. Protein domains were predicted with Interpro scan54 using the G. fortis ALX1 protein sequence.
Functional annotation of SNPs
NCBI’s genome annotation for the G. fortis assembly (GeoFor1) was downloaded from NCBI’s FTP server (ftp://ftp.ncbi.nlm.nih.gov/genomes/Geospiza_fortis/) in GFF format. The annotation was filtered to include only genes annotated with a coding sequence (13,949 genes with 16,365 transcripts) before using it to build a local SnpEff (version 3.4) database55. The SnpEff database was subsequently used to annotate all detected sequence variants among the Darwin’s finches with putative functional effects according to categories defined in the SnpEff manual. The upstream and downstream categories are regions within 5,000 bp in the respective direction of an annotated gene. SnpEff allows SNPs to be included in multiple categories; for example, a SNP may be intronic in one gene and a synonymous change in another gene residing in the intron of the first gene.
Accession codes
Primary accessions
GenBank/EMBL/DDBJ
Sequence Read Archive
Data deposits
The Illumina reads have been submitted to the short reads archive (http://www.ncbi.nlm.nih.gov/sra) under accession number PRJNA263122 and the consensus sequence for the G. fortis mtDNA has been submitted to GenBank under accession number KM891730.
References
Schluter, D. The Ecology of Adaptive Radiation (Oxford Univ. Press, 2000)
Seehausen, O. African cichlid fish: a model system in adaptive radiation research. Proc. R. Soc. B 273, 1987–1998 (2006)
Lack, D. Darwin’s Finches (Cambridge Univ. Press, 1947)
Grant, P. R. Ecology and Evolution of Darwin’s Finches (Princeton Univ. Press, 1999)
Grant, P. R. & Grant, B. R. How and Why Species Multiply. The Radiation of Darwin’s Finches (Princeton Univ. Press, 2008)
Petren, K., Grant, P. R., Grant, B. R. & Keller, L. F. Comparative landscape genetics and the adaptive radiation of Darwin’s finches: the role of peripheral isolation. Mol. Ecol. 14, 2943–2957 (2005)
Ali, J. R. & Aitchison, J. C. Exploring the combined role of eustasy and oceanic island thermal subsidence in shaping biodiversity on the Galápagos. J. Biogeogr. 41, 1227–1241 (2014)
Geist, D., Snell, H., Snell, H., Goddard, C. & Kurz, M. in The Galápagos: A Natural Laboratory for the Earth Sciences (eds Harpp K. S., Mittelstaedt E., d’Ozouville N., & Graham, D. ) 145–166 (American Geophysical Union, 2014)
Farrington, H. L., Lawson, L. P., Clark, C. M. & Petren, K. The evolutionary history of Darwin’s finches: speciation, gene flow, and introgression in a fragmented landscape. Evolution 68, 2932–2944 (2014)
Abzhanov, A., Protas, M., Grant, B. R., Grant, P. R. & Tabin, C. J. Bmp4 and morphological variation of beaks in Darwin’s finches. Science 305, 1462–1465 (2004)
Abzhanov, A. et al. The calmodulin pathway and evolution of elongated beak morphology in Darwin’s finches. Nature 442, 563–567 (2006)
Mallarino, R. et al. Two developmental modules establish 3D beak-shape variation in Darwin’s finches. Proc. Natl Acad. Sci. USA 198, 4057–4062 (2011)
Burns, K. J. et al. Phylogenetics and diversification of tanagers (Passeriformes: Thraupidae), the largest radiation of Neotropical songbirds. Mol. Phylogenet. Evol. 75, 41–77 (2014)
Zhang, G., Parker, P., Li, B., Li, H. & Wang, J. The genome of Darwin’s finch (Geospiza fortis). GigaScience, http://dx.doi.org/10.5524/100040 (3 August 2012)
Ellegren, H. The evolutionary genomics of birds. Annu. Rev. Ecol. Evol. Syst. 44, 239–259 (2013)
Balakrishnan, C. N. & Edwards, S. V. Nucleotide variation, linkage disequilibrium and founder-facilitated speciation in wild populations of the zebra finch (Taeniopygia guttata). Genetics 181, 645–660 (2009)
Swarth, H. S. The avifauna of the Galapagos Islands. Occ. Pap. Calif. Acad. Sci. 18, 1–299 (1931)
Lack, D. The Galapagos finches (Geospizinae): a study in variation. Occ. Pap. Calif. Acad. Sci. 21, 1–159 (1945)
Durand, E. Y., Patterson, N., Reich, D. & Slatkin, M. Testing for ancient admixture between closely related populations. Mol. Biol. Evol. 28, 2239–2252 (2011)
Qvarnstrom, A. & Bailey, R. I. Speciation through evolution of sex-linked genes. Heredity 102, 4–15 (2009)
Li, H. & Durbin, R. Inference of human population history from individual whole-genome sequences. Nature 475, 493–496 (2011)
Rivera-Perez, J. A., Wakamiya, M. & Behringer, R. R. Goosecoid acts cell autonomously in mesenchyme-derived tissues during craniofacial development. Development 126, 3811–3821 (1999)
Rowe, A., Richman, J. M. & Brickell, P. M. Retinoic acid treatment alters the distribution of retinoic acid receptor-β transcripts in the embryonic chick face. Development 111, 1007–1016 (1991)
Uz, E. et al. Disruption of ALX1 causes extreme microphthalmia and severe facial clefting: expanding the spectrum of autosomal-recessive ALX-related frontonasal dysplasia. Am. J. Hum. Genet. 86, 789–796 (2010)
Dee, C. T., Szymoniuk, C. R., Mills, P. E. D. & Takahashi, T. Defective neural crest migration revealed by a zebrafish model of Alx1-related frontonasal dysplasia. Hum. Mol. Genet. 22, 239–251 (2013)
Brugmann, S. A. et al. Comparative gene expression analysis of avian embryonic facial structures reveals new candidates for human craniofacial disorders. Hum. Mol. Genet. 19, 920–930 (2010)
Sommer, P., Napier, H. R., Hogan, B. L. & Kidson, S. H. Identification of Tgfβ1i4 as a downstream target of Foxc1. Dev. Growth Differ. 48, 297–308 (2006)
Wang, J. et al. Factorbook.org: a Wiki-based database for transcription factor-binding data generated by the ENCODE consortium. Nucleic Acids Res. 41, D171–D176 (2013)
Kumar, P., Henikoff, S. & Ng, P. C. Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm. Nature Protocols 4, 1073–1081 (2009)
Grant, P. R. & Grant, B. R. 40 Years of Evolution. Darwin’s Finches on Daphne Major Island (Princeton Univ. Press, 2014)
Boag, P. T. Growth and allometry of external morphology in Darwin’s finches (Geospiza) on Isla Daphne Major, Galápagos. J. Zool. 204, 413–441 (1984)
The Heliconius Genome Consortium Butterfly genome reveals promiscuous exchange of mimicry adaptations among species. Nature 487, 94–98 (2012)
Andersson, L. Molecular consequences of animal breeding. Curr. Opin. Genet. Dev. 23, 295–301 (2013)
Linnen, C. R. et al. Adaptive evolution of multiple traits through multiple mutations at a single gene. Science 339, 1312–1316 (2013)
Siepel, A. et al. Evolutionarily conserved elements in vertebrate, insect, worm, and yeast genomes. Genome Res. 15, 1034–1050 (2005)
Luo, R. et al. SOAPdenovo2: an empirically improved memory-efficient short-read de novo assembler. GigaScience 1, 18 (2012)
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009)
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010)
DePristo, M. A. et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nature Genet. 43, 491–498 (2011)
Van der Auwera, G. A. et al. From FastQ data to high-confidence variant calls: the Genome Analysis Toolkit best practices pipeline. Curr. Protoc. Bioinform. 43, 11.10.1–11.10.33 (2002)
Browning, S. R. & Browning, B. L. Rapid and accurate haplotype phasing and missing-data inference for whole-genome association studies by use of localized haplotype clustering. Am. J. Hum. Genet. 81, 1084–1097 (2007)
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007)
Green, R. E. et al. A draft sequence of the Neandertal genome. Science 328, 710–722 (2010)
Holm, S. A simple sequentially rejective multiple test procedure. Scand. J. Stat. 6, 65–70 (1979)
Rands, C. et al. Insights into the evolution of Darwin’s finches from comparative analysis of the Geospiza magnirostris genome sequence. BMC Genomics 14, 95 (2013)
Weir, J. T. & Schluter, D. Calibrating the avian molecular clock. Mol. Ecol. 17, 2321–2328 (2008)
Watterson, G. A. On the number of segregating sites in genetical models without recombination. Theor. Popul. Biol. 7, 256–276 (1975)
Grant, B. R. & Grant, P. R. Demography and the genetically effective sizes of two populations of Darwin’s finches. Ecology 73, 766–784 (1992)
Nei, M. in Molecular Evolutionary Genetics 276–279 (Columbia Univ. Press, 1987)
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011)
Vilella, A. J. et al. EnsemblCompara GeneTrees: complete, duplication-aware phylogenetic trees in vertebrates. Genome Res. 19, 327–335 (2009)
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004)
Waterhouse, A. M., Procter, J. B., Martin, D. M. A., Clamp, M. & Barton, G. J. Jalview version 2—a multiple sequence alignment editor and analysis workbench. Bioinformatics 25, 1189–1191 (2009)
Jones, P. et al. InterProScan 5: genome-scale protein function classification. Bioinformatics 30, 1236–1240 (2014)
Cingolani, P. et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly (Austin) 6, 80–92 (2012)
Grant, B. R., Grant, P. R. & Petren, K. The allopatric phase of speciation: the sharp-beaked ground finch (Geospiza difficilis) on the Galápagos islands. Biol. J. Linn. Soc. 69, 287–317 (2000)
Grant, P. R., Abbott, I., Schluter, D., Curry, R. L. & Abbott, L. K. Variation in the size and shape of Darwin’s finches. Biol. J. Linn. Soc. 25, 1–39 (1985)
Schluter, D. & Grant, P. R. Ecological correlates of morphological evolution in a Darwin’s finch, Geospiza difficilis. Evolution 38, 856–869 (1984)
Rabosky, D. Diversity-dependence, ecological speciation, and the role of competition in macroevolution. Ann. Rev. Evol. Ecol. Syst. 44, 481–502 (2013)
Acknowledgements
The National Science Foundation (USA) funded the collection of material under permits from the Galápagos and Costa Rica National Parks Services, and in accordance with protocols of Princeton University’s Animal Welfare Committee. The map and images of finch heads are reproduced with permission from Princeton University Press. The project was supported by the Knut and Alice Wallenberg Foundation. Sequencing was performed by the SNP&SEQ Technology Platform, supported by Uppsala University and Hospital, SciLifeLab and Swedish Research Council (80576801 and 70374401). Computer resources were supplied by UPPMAX.
Author information
Authors and Affiliations
Contributions
P.R.G. and B.R.G. collected the material. L.A., P.R.G. and B.R.G. conceived the study. L.A. and M.T.W. led the bioinformatic analysis of data. S.L. and J.B. performed the bioinformatic analysis with contributions from M.S.A., K.M., M.G., A.M.-B., C.-J.R. and N.Z. M.P. and C.W. performed experimental work. L.A., S.L., J.B., B.R.G., P.R.G. and M.T.W. wrote the paper with input from the other authors. All authors approved the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Read depth.
Average read depth in all 120 samples of Darwin’s finches and outgroup species.
Extended Data Figure 2 Genetic diversity among Darwin’s finches.
Heat map illustrating the proportion of shared and fixed polymorphisms among Darwin’s finches and outgroup species.
Extended Data Figure 3 Network tree for the Darwin’s finches on the basis of all autosomal sites.
Taxa that showed deviations from classical taxonomy are underscored. Finch heads are reproduced from ref. 5. How and Why Species Multiply: The Radiation of Darwin's Finches by Peter R. Grant & B. Rosemary Grant. Copyright © 2008 Princeton University Press. Reprinted by permission.
Extended Data Figure 4 Taxonomy and rate of speciation.
a, Morphological variation among populations of ground finch (Geospiza) species, scandens, fuliginosa and three others, acutirostris, difficilis and septentrionalis, that were formerly classified as a single species (difficilis). Data are from refs 56, 57, and from ref. 58 for weights and measures of difficilis on Fernandina. b, Morphological variation among populations of G. scandens, conirostris, propinqua and magnirostris assessed by multiple discriminant function analysis in JMP version 9. In a discriminant function analysis of the measured variables, all populations were correctly identified to species (−2 log likelihood P = 0.02). Maximum discrimination was achieved by entering three variables in the sequence beak width, beak length and body size (weight or wing). Substituting beak depth for beak width gave the same result. No other variable entered significantly. Data are from ref. 57, except for scandens and magnirostris data from ref. 30. c, Species accumulation on a log scale as a function of time before the present, dating based on mtDNA. Species are expected to accumulate linearly according to a ‘birth–death’ process, eventually declining under a density- (diversity-) dependent mechanism59.
Extended Data Figure 5 Phylogenies for mtDNA and the sex chromosomes Z and W.
a, Tree based on mtDNA sequences. The dating of the nodes and their variances (in thousands of years) is based on the cytochrome b sequences using the fossil-calibrated divergence rate 2.1% per million years for birds46. This tree based on the full mtDNA sequences shows only minor differences compared with previously published trees based only on the cytochrome b sequence6,9. b, Maximum-likelihood trees based on all Z-linked sites; all nodes having full local support on the basis of the Shimodaira–Hasegawa test are marked by asterisks. c, Tree based on W sequences, only females. Taxa that showed deviations from classical taxonomy are underscored (applies to a–c).
Extended Data Figure 6 ABBA–BABA analysis and demographic history.
a, ABBA–BABA analysis of G. magnirostris, G. conirostris on Española and on Genovesa, and with L. noctis as outgroup. b, Comparison of C. olivacea, C. fusca, a pool of all non-warblers, and with L. noctis as outgroup. The number of informative sites supporting the different trees is indicated both as a percentage and as the actual number. The D statistic and corresponding Holm–Bonferroni-corrected P value are also given for testing the null hypothesis of symmetry in genetic relationships. Finch heads are reproduced from ref. 5. c, PSMC analysis21 of all species except the G. difficilis group. d, PSMC analysis of the G. difficilis group.
Extended Data Figure 7 Sequence conservation of ALX1.
Amino-acid alignment of the complete ALX1 sequence among different vertebrates. Amino-acid substitutions between ALX1 alleles associated with blunt and pointed beaks are highlighted. The homeobox domain is indicated.
Supplementary information
Supplementary Information
This file contains Supplementary Text and References. (PDF 275 kb)
Supplementary Table 1
This file contains read depth in males and females for the identification of scaffolds from chromosome Z and W. Part a shows read depth in males and females for scaffolds assigned to the Z chromosome. Part b shows read depth in males and females for scaffolds assigned to the W chromosome. (XLSX 79 kb)
Supplementary Table 2
This file contains details from ABBA-BABA analyses of Darwin’s finch populations. P-values are two-sided Holm-Bonferroni corrected. (XLSX 59 kb)
Rights and permissions
About this article
Cite this article
Lamichhaney, S., Berglund, J., Almén, M. et al. Evolution of Darwin’s finches and their beaks revealed by genome sequencing. Nature 518, 371–375 (2015). https://doi.org/10.1038/nature14181
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14181
This article is cited by
-
Gene flow and an anomaly zone complicate phylogenomic inference in a rapidly radiated avian family (Prunellidae)
BMC Biology (2024)
-
Recent beak evolution in North American starlings after invasion
Scientific Reports (2024)
-
Odor-regulated oviposition behavior in an ecological specialist
Nature Communications (2023)
-
Molecular mechanisms of adaptive evolution in wild animals and plants
Science China Life Sciences (2023)
-
Survival Versus Attraction Advantages and Talent Selection in Sport
Sports Medicine - Open (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.