Abstract
Replying to J. M. Dijkstra Nature511,http://dx.doi.org/10.1038/nature13446(2014)
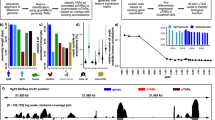
In the accompanying Comment1, Dijkstra has identified gene models potentially encoding cytokines in regions of the Callorhinchus milii genome, whose syntenic counterparts in teleost and tetrapod genomes harbour interleukin genes. These genes were not predicted by our assembly and annotation pipeline2. TBLASTN searches of the genome assembly and RNA-sequencing (RNA-seq) transcriptomes from 10 tissues (including thymus and spleen) of the elephant shark, and RNA-seq transcriptomes from the thymus and spleen of nurse shark using human and teleost fish interleukins as query sequences, were also unable to pick up these genes; although we readily detected other cytokine genes in the same family (for example, interleukin-7 (IL-7) and IL-15). Below are our comments on the candidate genes identified by Dijkstra.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Dijkstra, J. M. TH2 and Treg candidate genes in elephant shark. Nature 511, http://dx.doi.org/10.1038/nature13446 (2014)
Venkatesh, B. et al. Elephant shark genome provides unique insights into gnathostome evolution. Nature 505, 174–179 (2014)
Liu, M. & Grigoriev, A. Protein domains correlate strongly with exons in multiple eukaryotic genomes–evidence of exon shuffling? Trends Genet. 20, 399–403 (2004)
Ohtani, M., Hayashi, N., Hashimoto, K., Nakanishi, T. & Dijkstra, J. M. Comprehensive clarification of two paralogous interleukin 4/13 loci in teleost fish. Immunogenetics 60, 383–397 (2008)
Takizawa, F. et al. Constitutive high expression of interleukin-4/13A and GATA-3 in gill and skin of salmonid fishes suggests that these tissues form Th2-skewed immune environments. Mol. Immunol. 48, 1360–1368 (2011)
Dijkstra, J. M. et al. Identification of a gene for an ancient cytokine, interleukin 15-like, in mammals; interleukins 2 and 15 co-evolved with this third family member, all sharing binding motifs for IL-15Rα. Immunogenetics 66, 93–103 (2014)
Andersen, K. G., Nissen, J. K. & Betz, A. G. Comparative genomics reveals key gain-of-function events in Foxp3 during regulatory T cell evolution. Front. Immunol. 3, 113 (2012)
Van Laethem, F. et al. Lck availability during thymic selection determines the recognition specificity of the T cell repertoire. Cell 154, 1326–1341 (2013)
Dijkstra, J. M., Grimholt, U., Leong, J., Koop, B. F. & Hashimoto, K. Comprehensive analysis of MHC class II genes in teleost fish genomes reveals dispensability of the peptide-loading DM system in a large part of vertebrates. BMC Evol. Biol. 13, 260 (2013)
Author information
Authors and Affiliations
Corresponding authors
PowerPoint slides
Rights and permissions
About this article
Cite this article
Venkatesh, B., Lee, A., Swann, J. et al. Venkatesh et al. reply. Nature 511, E9–E10 (2014). https://doi.org/10.1038/nature13447
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13447
This article is cited by
-
Phylotranscriptomics suggests the jawed vertebrate ancestor could generate diverse helper and regulatory T cell subsets
BMC Evolutionary Biology (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.