Abstract
Mutations in SHANK3 and large duplications of the region spanning SHANK3 both cause a spectrum of neuropsychiatric disorders, indicating that proper SHANK3 dosage is critical for normal brain function. However, SHANK3 overexpression per se has not been established as a cause of human disorders because 22q13 duplications involve several genes. Here we report that Shank3 transgenic mice modelling a human SHANK3 duplication exhibit manic-like behaviour and seizures consistent with synaptic excitatory/inhibitory imbalance. We also identified two patients with hyperkinetic disorders carrying the smallest SHANK3-spanning duplications reported so far. These findings indicate that SHANK3 overexpression causes a hyperkinetic neuropsychiatric disorder. To probe the mechanism underlying the phenotype, we generated a Shank3 in vivo interactome and found that Shank3 directly interacts with the Arp2/3 complex to increase F-actin levels in Shank3 transgenic mice. The mood-stabilizing drug valproate, but not lithium, rescues the manic-like behaviour of Shank3 transgenic mice raising the possibility that this hyperkinetic disorder has a unique pharmacogenetic profile.
Similar content being viewed by others
Main
An increasing number of neuropsychiatric disorders such as autism spectrum disorder (ASD), intellectual disability, schizophrenia, obsessive-compulsive disorder and bipolar disorder are being classified as ‘synaptopathies’1,2,3,4, because the genes mutated in these disorders lead to abnormal synaptic development or function. For example, all members of the SHANK gene family (SHANK1, SHANK2 and SHANK3), which encode for core scaffolding proteins organizing macromolecular complexes at the postsynaptic density (PSD)5, have been linked to human synaptopathies6,7,8. Point mutations in SHANK3 (also called ProSAP2) are seen in non-syndromic autism, intellectual disability and schizophrenia8,9,10,11,12, and deletions of the region containing SHANK3 cause Phelan-McDermid syndrome (22q13 deletion syndrome)13,14. Furthermore, mice modelling SHANK3 deletion mirror the human behavioural phenotypes and display synaptic abnormalities15,16,17.
Interestingly, a few 22q13 duplications spanning SHANK3 have been reported in patients diagnosed with Asperger syndrome, attention deficit hyperactivity disorder (ADHD) or schizophrenia9,10,18, indicating that increased expression of SHANK3 could be also deleterious. Because duplication cases involve large genomic regions (>0.8 Mb) with more than 20 genes, however, it is unclear if it is SHANK3 overexpression that causes a neurological phenotype. Here we describe the generation and characterization of Shank3-overexpressing mice and studies on the role of SHANK3 duplication in hyperkinetic neuropsychiatric disorders in humans. We identify a mechanism that contributes to the neuronal phenotype and report pharmacological therapies that might benefit individuals with SHANK3 overexpression.
Manic-like behaviour in Shank3 transgenic mice
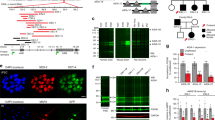
We introduced an amino-terminal enhanced green fluorescent protein (EGFP) tag into mouse Shank3 to generate EGFP–Shank3 transgenic mice (Fig. 1a and Extended Data Fig. 1a). Similar to endogenous Shank3 (ref. 16), EGFP–Shank3 transcript was detected in cortex, hippocampus and striatum (Fig. 1b). The regional and developmental expression of EGFP–Shank3 recapitulated those of endogenous Shank3 (Extended Data Fig. 1b–d). EGFP–Shank3 was targeted to the dendritic spines and co-localized with excitatory postsynaptic marker PSD-95, but not with inhibitory postsynaptic marker gephyrin (Fig. 1c). Compared to wild-type mice, transgenic mice expressed 1.2-fold to twofold higher levels of each of the three major Shank3 isoforms (α, β and γ), with ∼50% more total Shank3 protein (Fig. 1d and Extended Data Fig. 1e, f), which parallels expression in human SHANK3 duplication patients. There was no significant change in levels of other synaptic proteins in the hippocampus and striatum of transgenic mice (Extended Data Fig. 1g, h).
a, Diagrams show the modified Shank3 BAC and EGFP–Shank3. Kozak (ACCATGG) and EGFP-coding sequence were integrated into the start codon of Shank3. b, RNA in situ hybridization with a probe against EGFP detected EGFP–Shank3 in the brain. Cb, cerebellum; Ct, cortex; Hp, hippocampus; St, striatum; TG, transgenic; Th, thalamus; WT, wild type. c, EGFP–Shank3 localizes to excitatory postsynaptic sites in cultured hippocampal neurons. MAP2 is a dendritic marker. Scale bar, 10 μm. d, Quantification of the fold changes of Shank3 (α, β and γ isoforms) in synaptosomal fraction of whole brain (n = 4, biological replicates), hippocampus or striatum (n = 6) from 6-week-old mice. All data are presented as mean ± s.e.m. *P < 0.05; unpaired two-tailed Student’s t-test.
Our Shank3 transgenic mice showed increased locomotor activity and speed compared with wild-type littermates in the open field test (Fig. 2a, b and Extended Data Fig. 2a). Transgenic mice were also hyperactive in their home cages, indicating that their behaviour in the open field was not due to the novel environment (Extended Data Fig. 2b). Given that one patient carrying a large 22q13 duplication spanning SHANK3 has been reported to have ADHD10, we sought to determine whether the hyperactivity in the transgenic mice is related to ADHD by testing whether it could be corrected by amphetamine, a treatment for ADHD. Surprisingly, rather than the expected paradoxical calming effect, an acute injection of amphetamine aggravated the hyperactivity of transgenic mice to a greater degree than wild-type mice (Fig. 2c, d). As increased sensitivity to amphetamine is one characteristic of mania19—bipolar disorder can be misdiagnosed as ADHD, especially in children due to the overlap in diagnostic criteria20—we pursued behavioural tests to determine if there are abnormalities typically associated with a manic state.
a, Transgenic mice show increased locomotor activity in the open field test. b, Transgenic mice did not habituate during the open field assay. c, d, Transgenic mice are hypersensitive to amphetamine. After 30 min of basal activity, amphetamine (2 mg per kg) or saline was administered (arrow) and locomotor activity was monitored for 60 min. e, Transgenic mice spend less time immobile in the tail-suspension test. f, Increased acoustic (120 dB) startle response of transgenic mice. g, Abnormal PPI of transgenic mice. h–j, Abnormal circadian rhythms of transgenic mice. h, Wheel running actograms of wild-type and transgenic female mice. After 8 days of light/dark (L/D) cycle, animals were released into constant darkness (arrow, D/D) for 2 weeks. i, Among 20 transgenic mice tested, 3 displayed complete loss of rhythm during D/D. j, Transgenic mice showed increased period length and activity count, but normal alpha, compared to wild-type mice. All data are presented as mean ± s.e.m. *P < 0.05; **P < 0.01; ***P < 0.001. Statistical analyses for behavioural assays are in Supplementary Table 1.
Immobility during the tail-suspension test has been used as an indicator of despair. Consistent with manic-like behaviour21, the duration of immobility in Shank3 transgenic mice was less than wild-type littermates (Fig. 2e); the transgenic mice also exhibited elevated acoustic startle response with reduced prepulse inhibition (PPI), abnormal circadian rhythms, and hyperphagia-like behaviour (Fig. 2f–j and Extended Data Fig. 2c, d), all of which are behaviours observed in humans during manic episodes22,23.
We normalized Shank3 levels by crossing the transgenic mice with Shank3B+/− mice16 and found that this reversed hyperactivity, reduced immobility in tail-suspension and reduced PPI (Extended Data Fig. 3). Shank3 transgenic mice also displayed decreased social interaction (Extended Data Fig. 2e–g), which has also been seen in patients with large 22q13 duplications9,18. Unlike Shank3 knockout mice16,17, Shank3 transgenic mice did not exhibit repetitive behaviour (Extended Data Fig. 2h–j), but they made fewer calls during the ultrasonic vocalization test at postnatal day 13 (Extended Data Fig. 2k, l). Together, these results suggest that ∼50% increase in Shank3 level causes a hyperkinetic phenotype in mice that resembles mania, and that it is the increased dosage of SHANK3 in 22q13 duplications that probably contributes to hyperkinesia in humans.
SHANK3 duplications in humans
To confirm unequivocally the role of SHANK3 dosage in the neuropsychiatric phenotypes, we queried the array Comparative Genomic Hybridization (aCGH) database at the Molecular Genetics Laboratory of Baylor College of Medicine and identified two cases that carry small duplications (Fig. 3a, b and Supplementary Data).
a, Array plot of an exon-targeted chromosome microarray analysis on DNA from an 11-year-old female with ADHD, seizures and aberrant behaviours. Black dots indicate probes with normal copy number and green dots indicate copy number gain. Solid and dotted lines define the minimum and maximum expected boundaries of the duplication, respectively. b, Array plot of the 35-year-old male with bipolar disorder and epilepsy.
The first individual is an 11-year-old girl diagnosed with ADHD, combined type. The 22q13 duplication includes the entire SHANK3 and part of the ACR gene (Fig. 3a). ACR encodes acrosin, a protease expressed in sperm facilitating its penetration into oocytes, with no known expression or function in the brain. This individual has seizures and exhibits hyperactivity, poor attention, auditory overstimulation, hyperphagia and kleptomania. Notably, her hyperactivity is resistant to the paradoxical calming effect of stimulant medications seen in most cases of ADHD. The second individual is a 35-year-old man diagnosed with bipolar disorder and epilepsy who has a duplication encompassing SHANK3, ACR and ARSA, the gene encoding arylsulfatase A (Fig. 3b). ARSA deficiency causes metachromatic leukodystrophy, and duplications of ARSA have not been associated with any neurobehavioural phenotype.
These two individuals carry the smallest 22q13 duplications spanning SHANK3 reported so far, and their molecular and clinical findings support the notion that SHANK3 duplications result in hyperkinetic disorders and seizures in humans. More detailed clinical and molecular cytogenetic information about both individuals is provided in the supplementary material.
Synaptic E/I imbalance in Shank3 transgenic mice
Having found that humans with SHANK3 duplications have a hyperkinetic disorder and seizures, we evaluated the Shank3 transgenic mice for seizures and synaptic abnormalities. We observed spontaneous seizures in the Shank3 transgenic mice, but never in wild-type littermates (Supplementary Video 1). We monitored for electrographic seizures using electroencephalography (EEG). The EEG of transgenic mice showed hyperexcitability discharges accompanied by electrographic seizures (Fig. 4a). As alteration in the neuronal excitatory/inhibitory (E/I) balance is considered to cause seizures24, we examined morphological and functional changes of both excitatory and inhibitory synapses. Cultured hippocampal pyramidal neurons from transgenic mice displayed increased VGLUT1-positive PSD-95 (VGLUT1 is also known as SLC17A7), an excitatory synaptic marker, and decreased VGAT-positive (also known as SLC32A1) gephyrin, an inhibitory synaptic marker, puncta density (Fig. 4b).
a, Representative EEG traces from wild-type (n = 5) and transgenic (n = 10) mice. Transgenic mice showed prolonged hyperexcitability discharges and electrographic seizure in all recorded regions. The frequency of epileptiform spikes (dentate gyrus) and electrographic seizure were significantly increased in transgenic mice. b, Increased VGLUT1-positive PSD-95 (n = 28, three independent experiments) and decreased VGAT-positive gephyrin (n = 25) puncta density in cultured hippocampal pyramidal neurons from transgenic mice. Scale bar, 10 μm. c, Frequency, but not amplitude and decay, of mIPSC was decreased in CA1 pyramidal neurons of transgenic mice (wild type n = 14, transgenic n = 18). d, Amplitude, but not frequency and decay, of sEPSC was increased in transgenic neurons (n = 19). e, AMPA/NMDA ratio was normal in transgenic neurons (n = 9). All data are presented as mean ± s.e.m. *P < 0.05; **P < 0.01. Statistical analyses for these data are in Supplementary Table 2 and 3.
We examined functional properties of synapses by whole-cell patch clamping in CA1 pyramidal neurons of acute hippocampal slices. Neuronal density, paired-pulse facilitation ratio, input–output relationship and intrinsic excitability of these neurons were similar in wild-type and transgenic mice, indicating normal basal synaptic transmission at Schaffer collateral-CA1 synapses of transgenic mice (Extended Data Fig. 4a–e). Consistent with decreased density of VGAT-positive gephyrin puncta in cultured hippocampal neurons from transgenic mice, we observed reduction in GABAA receptor-mediated miniature inhibitory postsynaptic current (mIPSC) frequency, but not in amplitude or decay time (Fig. 4c). AMPA receptor-mediated miniature EPSC (mEPSC) was normal, but amplitude of spontaneous EPSC (sEPSC, without tetrodotoxin) was increased in transgenic mice (Extended Data Fig. 4f and Fig. 4d). There was no change in the AMPA/NMDA ratio in CA1 pyramidal neurons of transgenic compared to wild-type mice (Fig. 4e). Consistently, NMDA receptor-dependent synaptic plasticity measured by extracellular field recordings at Schaffer collateral–CA1 pyramidal synapses was normal in trasngenic mice (Extended Data Fig. 4g, h). The spontaneous seizures of transgenic mice, together with morphological and functional alterations in synapses, indicate that the synaptic E/I balance in transgenic neurons is shifted towards excitation.
Arp2/3 complex mediates F-actin increase
To understand the molecular mechanisms underlying the synaptic alterations of Shank3 trasngenic mice, we set out to identify the in vivo interactors of Shank3. We used the EGFP-tag of the Shank3 transgene to isolate Shank3 protein complexes (Extended Data Fig. 5a). Immunoprecipitation followed by mass-spectrometry analysis identified 273 proteins that bind Shank3 in vivo (Supplementary Table 4). There was notable overlap between the proteins on this list, those from our previous yeast two-hybrid screening25, and known mouse or human PSD proteins26,27 (Extended Data Fig. 5b). We combined our two Shank3-focused experimental lists (previous yeast two-hybrid screening25 and current in vivo immunoprecipitation, for a total of 404 proteins called the Shank3 interactome) for further analysis (Extended Data Fig. 5c, d).
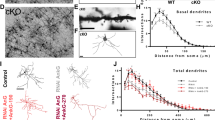
We performed Gene Ontology (GO) and pathway analysis, which suggested that regulation of actin cytoskeleton is a principal role of the Shank3 interacting proteins (Extended Data Fig. 5e and Supplementary Table 5). Based on this analysis, we proposed that abnormal regulation of the actin cytoskeleton could be one major molecular change in the synapses of transgenic mice. Accordingly, we evaluated the effects of a 50% increase in Shank3 on F-actin levels. We found that F-actin levels of excitatory synapses were increased by ∼30% in cultured hippocampal pyramidal neurons from transgenic mice compared to wild-type mice (Fig. 5a), which was restored by expression of Shank3 small interfering RNA (siRNA) (Extended Data Fig. 6a, b). Next, we used our Shank3 interactome to generate a sub-network of actin-related proteins (Fig. 5b) to focus the search on molecular mediators of the elevated F-actin. This network comprises both known and previously unknown Shank3 interactors, which we confirmed by western blotting (Extended Data Fig. 5f). The most interesting new Shank3 direct interactors we identified (based on both yeast two-hybrid screening and in vivo immunoprecipitation) were two subunits of the Arp2/3 complex (ARPC2 and ARPC5L) (Fig. 5b). Arp2/3 complex consists of seven-subunits and binds directly to the F-actin mother filament to initiate nucleation and branching28. Consistent with the interaction analysis, ARPC2 clusters co-localized with EGFP–Shank3 and significantly enlarged in transgenic neurons (Extended Data Fig. 5g and Fig. 5c); the latter observation was reversed by Shank3 siRNA (Extended Data Fig. 6c). To initiate actin polymerization, Arp2/3 complex must be activated by nucleation-promoting factors (NPFs)28. Given that the NPFs WASF1 and cortactin (CTTN) directly interact with Shank3 (refs 29, 30), Shank3 could function as a scaffold that brings WASF1, cortactin and Arp2/3 together, facilitating the formation of F-actin structures (Fig. 5d). Consistent with our hypothesis, co-localization of WASF1 and ARPC2 was enhanced in transgenic neurons (Fig. 5e). Furthermore, the density of dendritic spines was increased in transgenic brains compared to wild-type brains (Fig. 5f).
a, Increased synaptic F-actin in cultured hippocampal pyramidal neurons from transgenic mice (n = 29, three independent experiments). Scale bar, 10 μm. b, Actin cytoskeleton-related sub-network of Shank3 interactome. Y2H, yeast two-hybrid. c, Increased ARPC2 cluster size in transgenic pyramidal neurons (n = 20). d, Diagram shows the proposed role of Shank3 as a platform for F-actin regulating proteins. e, Increased co-localization of ARPC2 and WASF1 in transgenic pyramidal neurons (n = 22). f, Golgi-staining of CA1 pyramidal neurons shows more dendritic spines in transgenic mice (n = 40 neurons from 3 animals per genotype). g, h, Excitatory and inhibitory synaptic distributions of Mena (g) and profilin2 (h) in cultured CA1 pyramidal neurons. Yellow arrowheads indicate protein puncta co-localized with corresponding synaptic markers. i, Quantification of (g) and (h). Density of VGAT-positive Mena puncta is decreased in transgenic neurons (n = 18). VGLUT1-positive profilin2 puncta intensity is increased in transgenic neurons (n = 13), whereas VGAT-positive Mena and profilin2 puncta intensity are decreased in transgenic neurons. All data are presented as mean ± s.e.m. *P < 0.05; **P < 0.01.
Our analysis indicated that hippocampal pyramidal neurons of transgenic mice have a reduced number of inhibitory synapses (Fig. 4b, c), which was unexpected based on the exclusive localization of EGFP–Shank3 in excitatory synapses (Fig. 1c). To understand the underlying mechanism, we first investigated inhibitory neurons, changes of which might secondarily affect inhibitory synapses on pyramidal neurons. However, we found that there was no difference in synapse number on the GAD-6-positive inhibitory neurons of transgenic mice, although these neurons express EGFP–Shank3 (Extended Data Fig. 7).
We next explored a possible cell-autonomous relationship between excitatory and inhibitory synapses of transgenic pyramidal neurons. We speculated that increased clustering of actin-related proteins in excitatory synapses might reduce some of those proteins in inhibitory synapses, thereby contributing to the decrease of inhibitory synapse number. We focused on actin-related proteins, because the actin cytoskeleton defines the molecular architecture of both E/I synapses31. We picked Mena (also known as Enah) and profilin as candidates for four reasons: they directly interact with gephyrin in inhibitory synapses32; profilin1 and profilin2 are recruited to excitatory synapses by neuronal activity33,34; profilin2 was also identified in our Shank3 in vivo immunoprecipitation (Supplementary Table 4); and Mena and profilin interact with proteins in the Shank3 interactome (Extended Data Fig. 8). Both proteins showed decreased localization to inhibitory synapses, and profilin2 showed enhanced localization to excitatory synapses in transgenic neurons (Fig. 5g–i). Together, these results suggest that increased expression of Shank3 enhances F-actin levels through Arp2/3 complex in excitatory synapses of transgenic neurons. By modulating the synaptic distributions of Mena and profilin2, this contributes, at least in part, to changes in inhibitory synapses.
Shank3 transgenic mice respond to valproate
Having established that Shank3 overexpression causes mania-like behaviour in mice and a hyperkinetic syndrome in humans, we tested whether mood-stabilizing drugs for human mania could rescue the behavioural abnormalities of transgenic mice. First, we examined the effect of lithium, the first Food and Drug Administration (FDA)-approved treatment for bipolar disorder, which inhibits multiple targets including inositol monophosphatase and glycogen synthase kinase 3 (GSK3)35. The basal activity of GSK-3β in transgenic mouse hippocampus and striatum was similar to that of wild-type mice (Fig. 6a). Mice were fed with lithium carbonate-containing chow for three weeks, resulting in serum lithium levels (wild type: 0.82 ± 0.04; transgenic: 0.91 ± 0.08 mmol l−1) comparable to those in the therapeutic range for humans (0.6∼1.2 mmol l−1). Lithium did not mitigate any of the manic-like behaviours of Shank3 transgenic mice (Fig. 6b and Extended Data Fig. 9).
a, Basal activities of GSK-3β and Akt in the hippocampus and striatum of transgenic mice (10-week-old, n = 7) are normal. b, Amphetamine-sensitivity of male transgenic mice was not rescued by lithium. NS, not significant. c, d, Basal locomotor activity and amphetamine-sensitivity of transgenic mice were rescued by valproate. Basal activity of wild-type male mice was decreased by valproate during one 5-min time bin. e–h Acoustic startle response (e, g) and PPI (f, h) of transgenic mice were rescued by valproate. Valproate increased PPI of wild-type male mice. i, Rescue of abnormal EEG in transgenic mice by valproate. During the three consecutive days of tests, EEG was recorded from the three brain regions of transgenic mice (n = 7) for one hour before and after valproate injection. Representative EEG traces measured on day 1 are shown. The number of epileptiform spikes was quantified and normalized to the baseline values before treatment of day 1. All data are presented as mean ± s.e.m. *P < 0.05; **P < 0.01; ***P < 0.001.
Given that our mice display seizures in addition to manic-like behaviour, we decided to test valproate, another FDA-approved mood-stabilizing drug used to treat manic or mixed episodes that also acts as an anticonvulsant, increases GABA neurotransmission and inhibits sodium channels, calcium channels and histone deacetylase 1 (ref. 36). We treated the transgenic mice with acute intraperitoneal injections of either valproate (200 mg per kg) or saline and subjected them to behavioural testing. Valproate reversed the baseline hyperactivity, the amphetamine-hypersensitivity (Fig. 6c, d), and the abnormal acoustic startle response and PPI (Fig. 6e–h) in transgenic mice. The behaviour of wild-type mice, except for a decrease in baseline activity and an increase of PPI in male mice, was not significantly affected by valproate treatment (Fig. 6c–h). Valproate also significantly decreased the frequency of epileptiform spikes in transgenic mice as measured by EEG (Fig. 6i). Valproate treatment changed neither the levels of synaptic proteins in hippocampus or striatum of wild-type and transgenic mice, nor F-actin levels in cultured hippocampal neurons of transgenic mice (Extended Data Fig. 10).
Discussion
There are many genomic loci for which both deletions and duplications cause human neuropsychiatric disorders, indicating that proper dosage of some genes is critical for normal brain function37,38. In most cases, however, the genes causing the respective phenotypes are poorly defined, making it hard to understand pathogenesis and to develop targeted therapies. Chromosome 22q13 spanning SHANK3 is one example. Although SHANK3 loss has been shown to cause ASD in humans, SHANK3 overexpression has never been definitively determined to cause neuropsychiatric disease. In this study, we generated Shank3 transgenic mice that model a human duplication and identified patients with the smallest 22q13 duplications involving SHANK3. The remarkable similarity of the neurobehavioural phenotypes between the mouse model and patients with SHANK3 duplications supports the notion that SHANK3 overexpression causes a hyperkinetic neuropsychiatric disorder that approximates mania. Indeed, either a 50% reduction or 50% increase in Shank3 levels results in marked neuropsychiatric phenotypes and a predisposition to epilepsy.
Notably, the manic-like behaviour of Shank3 transgenic mice was reversed selectively by valproate but not by lithium treatment. It is not uncommon to find human patients with bipolar disorder that are resistant to lithium monotherapy39. Indeed, a subset of individuals with the rapid-cycling form of bipolar disorder is highly resistant to lithium40. As GSK3 is considered the major target for lithium’s efficacy in the treatment of BPD41, the lithium-resistance of Shank3 transgenic mice and their normal GSK-3β activity are congruent with a mechanism independent of GSK-3 and possibly more dependent upon synaptic alterations, including E/I imbalance.
Synaptic E/I imbalance is likely to underlie the pathogenesis for seizures24,42 and a broad spectrum of neuropsychiatric disorders1,43. We found morphological and functional changes in E/I synapses of transgenic neurons favouring excitation, which might account for both seizures and hyperkinetic behaviour. Consistent with this, GABAergic dysfunction in patients with bipolar disorder has been reported44, and valproate (which increases GABA neurotransmission and decreases high-frequency action potential firing) rescued the manic-like behaviours of transgenic mice. In contrast, lithium enhances excitatory postsynaptic potentials in CA1 synapses45, which might explain the resistance of Shank3 transgenic mice to lithium treatment. Mechanistically, enhanced F-actin polymerization through Arp2/3 complex could be responsible for the synaptic changes of transgenic neurons, but other Shank3-dependent molecular pathways cannot be excluded.
In conclusion, our study advances our understanding of the human diseases associated with SHANK3 and the mechanisms that can lead to bipolar disorder. We propose two distinct mechanisms leading to bipolar disorder: one is dysfunction of the well-established GSK-3 pathway that is responsive to lithium treatment; the second is a synaptic dysfunction associated with an E/I balance shifted towards excitation, which could result in hyperkinetic behaviours, and which is resistant to lithium but sensitive to valproate. We believe this sort of pharmacogenetic analysis in model organisms could improve treatment strategies for patients with bipolar disorder as the genetic underpinnings of each patient’s illness are elucidated.
Methods Summary
BAC clone (RP23-278D8) containing entire mouse Shank3 was modified by recombineering techniques. The linearized segment including EGFP–Shank3 plus ∼12 kb in the 5′ direction and ∼2 kb in the 3′ direction was injected into the FVB/N embryos to generate transgenic mice. Behavioural, electrophysiological, biochemical and cell biological characterization of the mice were performed and analysed blind to genotypes. All experimental procedures were reviewed and approved by the Institutional Animal Care and Use Committee for Baylor College of Medicine. The details of experimental procedures, reagents and bioinformatic analyses are described in the Methods.
Online Methods
Generation of Shank3 transgenic mice
To generate Shank3 transgenic mice, we used a BAC clone (RP23-278D8) containing a segment of mouse chromosome 15. This BAC clone was modified by recombineering techniques46 to insert Kozak sequence (ACCATGG) followed by EGFP sequence (cloned from pEGFP-C1) at the first start codon of Shank3 gene (exon 1). The modified BAC clone was double digested with NotI/SwaI, and the ∼75 kb linearized segment with entire Shank3 gene plus ∼12 kb 5′ and ∼2 kb 3′ was injected into the FVB/N embryos (Extended Data Fig. 1a). The primers used for genotyping were designed against EGFP sequence (forward: 5′-ATGGTGAGCAAGGGCGAGGAG-3′, reverse: 5′-GCGGACTTGAAGAAGTCGTGC-3′). All procedures to maintain and use these mice were approved by the Institutional Animal Care and Use Committee for Baylor College of Medicine.
Primary antibodies
Antibodies used for the western blots and/or immunostainings are Akt (Cell Signaling Technology; 4691), phospho-Akt (Thr 308) (Cell Signaling Technology; 2965), ARPC2 (Millipore; 07-227), β-actin (Abcam; ab20272), CYFIP2 (Abcam; ab95969), GABAAR β3 (NeuroMab; 75-149), GAD-6 (Developmental Studies Hybridoma Bank), Gephyrin (Synaptic Systems; 147 111), GFP (Abcam; ab290, ab13970), GKAP (NeuroMab; 75-156), GluR2 (NeuroMab; 75-002), GSK-3α/β (Millipore; 05-412), phospho-GSK-3β (Ser 9) (Cell Signaling Technology; 9336), Homer (Santa Cruz; sc-15321), MAP2 (Sigma; M9942), Mena/Vasp (Millipore; MAB2635), mGluR5 (Millipore; AB5675), NR1 (Millipore; MAB363), Profilin2 (TU-Braunschweig; 4H5), PSD-95 (NeuroMab; 75-028), pan-Shank (3856; (ref. 47)), Shank1 (NeuroMab; 75-064), Shank2 (NeuroMab; 75-088), Shank3 (Santa Cruz; H-160), VGAT (Synaptic Systems; 131 002), VGlut1 (Synaptic Systems; 135 302) and WASF1 (NeuroMab; 75-048).
Preparation of brain lysates and western blot
S2 (soluble fraction) and P2 (crude synaptosomal fraction) subcellular, and PSD fractions (synaptic fraction after one time Triton X-100 washout) were prepared as described48. For Akt and GSK3 detection, brains were dissected and homogenized in ice-cold lysis buffer (20 mM Tris-HCl, pH 7.4, 150 mM NaCl, 2 mM EDTA, 1% Triton X-100, 10% glycerol with protease and phosphatase inhibitor (Roche)). Western blot images were acquired by LAS 4000 (GE Healthcare) and quantified by an ImageJ software package.
Hippocampal neuron cultures, transfection and immunostaining
Hippocampal neurons were prepared from postnatal day 0-1 FVB/N mice and plated on poly-d-lysine/mouse laminin coated coverslips (BD Biosciences) in Neurobasal medium supplemented with GlutaMAX-I (Invitrogen), B-27 and 1% FBS. At days in vitro (DIV) 7, neurons were transfected with pEGFP-C1 plus either control or Shank3 siRNA (Ambion, s81603 or s81605). At DIV 14, neurons were fixed with 4% formaldehyde/4% sucrose in PBS, and permeabilized with 0.2% Triton X-100 in PBS. PBS with 0.1% BSA and 3% horse serum was used for blocking and antibody incubation. Alexa-conjugated phalloidin (Invitrogen) was used to visualize F-actin. Z-stack images were acquired by LSM710 (Zeiss) confocal microscope under the same parameter settings and the images were quantified (in a blinded manner) using ImageJ.
RNA in situ hybridization
Hybridization probe specific for EGFP sequence was prepared by PCR amplification from the pEGFP-C1 using the following primers (forward: 5′-ATGGTGAGCAAGGGCGAGGAG-3′, reverse: 5′-CTTGTACAGCTCGTCCATGCC-3′). Riboprobe was generated using DIG RNA labelling mix (Roche).
Golgi staining
Standard Golgi-Cox impregnation using the FD Rapid GolgiStain kit (NeuroTechnologies) was performed with brains from 10-week-old FVB/N male mice. Serial sagittal sections (50 μm) were collected and images of dendritic spines on the secondary branches (apical dendrites of CA1 pyramidal neurons) were acquired by LSM710 (Zeiss) confocal microscope under differential interference contrast (DIC) mode. The images were quantified (in blinded manner) using ImageJ.
Cresylviolet staining
Brains from 10-week-old FVB/N male mice were cryosectioned (25 μm) and stained with 0.2% cresyl violet.
Drug treatment
For each experiment, individual was blinded to genotype and littermates were randomly distributed to control and treatment groups as follows; Animal with genotype A was assigned to control, next animal of genotype A to drug, third to control. Same with genotype B. Amphetamine and valproate (Sigma) were dissolved in normal saline to final concentration of 0.2 g l−1 and 20 g l−1, respectively. Mice received intraperitoneal injection of amphetamine (2 mg per kg), valproate (200 mg per kg) or saline in a volume of 10 ml kg−1. Valproate was injected three times before each behavioural assay (10:00 and 17:00 on the day prior to the assay and 9:00 of the day of assay), and assays were performed 30 min after final injection. For lithium treatment, mice were fed either lithium carbonate-containing chow or control chow (Harlan Teklad). The lithium group was initially fed 0.2% lithium carbonate chow for a week followed by 0.4% chow for two more weeks before the behavioural assays. Water with 0.85% sodium chloride was provided to counteract toxicity of lithium. Lithium concentration in serum was measured using lithium assay kit (Crystal Chem) according to the manufacturer’s instruction.
Behavioural assays
All data acquisition and analyses were carried out by an individual blinded to the genotype. For behavioural assays, 2–3-month-old mice (both males and females) of the F1 hybrid (FVB/N × C57BL/6J) were used to mitigate strain specific effects. A sample size of mice was chosen to mitigate genetic background variance. Before each test, mice were habituated in the test room at least for 30 min. Mice with sign of seizures (before, during or after test) were excluded from the analysis. The summary of statistical analysis for behavioural assays is provided in Supplementary Table 1.
Open field
After habituated in the test room (600 lx, 60 dB white noise), mice were placed in the centre of a clear, open Plexiglass chamber (40 × 40 × 30 cm), and the activities were measured by photobeam breaks (Accuscan) for 30 min. To measure amphetamine response, mice received intraperitoneal injection of saline or amphetamine after 30 min test of basal activities, and the activities were measured for additional 60 min.
Home-cage activity
Single-caged mice were habituated in the cage for 12 h, and the activities were measured by photobeam breaks (Accuscan) for 48 h under the light cycle of 07:00–19:00.
Tail suspension
After habituation period in the test room (600 lx, 60 dB white noise), mice were suspended by their tails. Movements were recorded and the immobile time was automatically measured by ANY-maze software (Stoelting). The same parameter setting for the definition of immobility was applied for all the mice tested.
Acoustic startle response and prepulse inhibition
Mice were placed in a test chamber (San Diego Instruments) and habituated for 5 min with 70 dB background white noise. Eight trial types (no stimulus, a 40 ms 120 dB sound as the startle stimulus, and three different prepulse sounds (20 ms 74, 78, 82 dB) either alone or 100 ms before the startle stimulus) were presented in pseudo-random order with six times per each trial type. The interval between each trial type was 10 to 20 s. The maximum startle amplitude during the 65 ms period following the onset of the startle stimulus was used to calculate percentage prepulse inhibition.
Circadian rhythms
Mice were individually housed in cages equipped with a 11.5 cm diameter wheel, and wheel-running activity was recorded by VitalView software (Respironics Mini-Mitter system). Mice were held on a 12:12 light/dark cycle for 8 to 14 days with food, water and temperature at 20 ± 2 °C. Mice were then released into constant dark for 14 days.
Three chamber test
The test was performed as previously described49. The three chamber apparatus is a clear Plexiglass box (24.75 × 16.75 × 8.75 inches) with removable partitions separating the box into left, centre and right chambers. The age- and gender-matched C57BL/6J mice were used as novel partners. Two days before the test, the novel partner mice were habituated to the wire cages (3 inches diameter by 4 inches in height) for 1 h per day. The wire cage with inanimate novel object inside served as a control. The test mouse was habituated in the chamber for 15 min as described49. During the habituation period, neither wild-type nor Shank3 transgenic mice showed a preference for either side of the chamber (data not shown). After the habituation period, the novel partner mouse was placed into one of the wire cages and located randomly in either the right or left side of chamber. The novel object was placed into the wire cage located in the other side of chamber. The test mouse was allowed to explore the three chamber apparatus for 15 min. The movements were recorded and the total amount of time spent in each chamber was automatically measured by ANY-maze software (Stoelting). The close interaction time, defined by rearing, sniffing or pawing at each wire cage, was measured manually.
Grooming
After habituation period in the test room (600 lx, 60 dB white noise), mice were placed into the centre of a Plexiglass cylinder (20 cm in diameter by 30 cm in height) and videotaped for 10 min. The amount of time spent grooming was measured from the videotape.
Ultrasonic vocalization
Separation-induced ultrasonic vocalizations (USV) were measured on pups of postnatal day 6 to 13. A standard housing cage containing one female and her offspring was transferred to a holding room and allowed to acclimate for 30 min. Following this acclimation period, the cage was transferred to a testing room. Pups were placed into a clean and warm (35 °C) cage and allowed to acclimate for 5 min. The mother was removed from the testing room and transferred back to the holding room. After 5 min acclimation period, one pup will be placed individually into a plastic beaker in a testing chamber. A bat detector positioned 10 cm above the beaker transduced ultrasonic signals. USV was quantified for a total testing period of 2 min using sound analysis software (Ultravox, Noldus Information Technology).
EEG measurement
All data acquisition and analyses were carried out blinded to genotype. FVB/N 2–3-month-old male mice were anaesthetized with isoflurane. Under aseptic conditions, each mouse was surgically implanted with tungsten electrodes (50 µm diameter) aimed at hippocampal CA1 (P2.0R1.2H1.3) and dentate (P2.0R1.8H1.8) regions. The third EEG electrode made of silver wire (127 µm diameter) was implanted in the subdural space of the right frontal cortex area. A reference/grounding electrode was then positioned in the occipital region of the skull. All electrode wires were attached to a miniature connector (Harwin Connector). After 3 to 5 days of post-surgical recovery, EEG activities (filtered between 0.1 Hz and 5 kHz, sampled at 10 kHz) were recorded for 1 h per day over 3 to 5 days. In the experiments evaluating the effects of valproate on EEG activities, adult male Shank3 transgenic mice with implanted electrode went through recording sessions for 3 consecutive days. On each day, animals received one hour baseline recording before valproate (200 mg per kg, i.p.) followed by another hour of post recording.
EEG data analysis
Electrographic seizure time lasting over 10 s was counted manually and normalized as the percentage of the total recording period. The number of abnormal epileptiform spikes were counted using Clampfit 10 software (Molecular Devices) when the sharp positive deflections exceeding twice the baseline and lasting 25–100 ms50. In the pharmacological tests, spike numbers were normalized to the baseline value before drug administration on day 1.
Whole-cell patch-clamp recordings
All data acquisition and analyses were carried out blinded to genotype. Acute fresh hippocampal slices were prepared from FVB/N male mice at the age of 6–8-weeks old as previously described51. Coronal slices (250-μm thick) containing medial hippocampus were cut with a vibratome (Leica Microsystems) in a chamber filled with chilled (2–5 °C) cutting solution containing (in mM) 110 choline-chloride, 25 NaHCO3, 25 d-glucose, 11.6 sodium ascorbate, 7 MgSO4, 3.1 sodium pyruvate, 2.5 KCl, 1.25 NaH2PO4 and 0.5 CaCl2. The slices were then incubated in artificial cerebrospinal fluid (ACSF, in mM) containing 119 NaCl, 26.2 NaHCO3, 11 d-glucose, 3 KCl, 2 CaCl2, 1 MgSO4, 1.25 NaH2PO4 at the room temperature. The solutions were bubbled with 95% O2 and 5% CO2. Whole-cell recording was made using patchclamp amplifiers (Multiclamp 700B) under infrared–differential interference contrast microscopy (Zeiss). Data acquisition and analysis were performed using digitizers (DigiData 1440A) and analysis software pClamp 10 (Molecular Devices). Signals were filtered at 2 kHz and sampled at 10 kHz. Spontaneous EPSCs (sEPSCs), miniature EPSCs (mEPSCs) and IPSCs (mIPSCs) were recorded from CA1 hippocampal pyramidal neurons at −70 mV in voltage-clamp mode. sEPSCs were recorded in the presence of GABAA receptor blocker, SR95531 (10 μM). mEPSCs were recorded in the presence of tetrodotoxin (TTX, 0.5 μM) and SR95531 (10 μM). Glass pipettes with a resistance of 2.5–5 MΩ were filled with a solution containing (in mM) 140 potassium gluconate, 5 KCl, 10 HEPES, 0.2 EGTA, 2 MgCl2, 4 MgATP, 0.3 Na2GTP and 10 Na2-phosphocreatine, pH 7.2 (with KOH). For recording mIPSCs, potassium gluconate was replaced with KCl (140 μM) in the pipette solution. Besides TTX (0.5 μM), glutamate receptor antagonists 6-cyano-7-nitroquinoxaline-2,3-dione disodium (CNQX) (10 μM) and D-2-amino-5-phosphonovaleric acid (D-AP-5) (25 μM) were also added to the bath. Series resistance (15–30 MΩ) and input resistance (100–200 MΩ) were monitored throughout the whole-cell recording. Evoked EPSCs were recorded in the presence of SR95531 (10 μM). To stimulate presynaptic inputs, stimulation pulses (50 μs duration at 0.1 Hz) were applied to a bipolar tungsten stimulation electrode (WPI) placed along the Schaffer collaterals. Extracellular stimuli were administered using a stimulator system (Master 8). The stimulating and recording pipettes were placed at the same depth in the slice and the distance between them was kept constant (∼300 μm). Data were discarded when the change in the series resistance was >20% during the course of the experiment. The whole-cell recording was performed at 30 ± 1 °C with the help of an automatic temperature controller (Warner Instruments). For measurements of the AMPA-to-NMDA receptor ratio, the CA1 pyramidal neuron was voltage-clamped at 40 mV. First, a stable baseline recording of total EPSCs was obtained. The NMDA receptor antagonist D-AP-5 (50 mM) was then applied to the bath for 5–10 min to isolate fast AMPA-receptor-mediated EPSCs. Digital subtraction of AMPA-receptor-EPSCs from the total EPSCs from the same neuron yielded NMDA-receptor-EPSCs. An average of 12–20 EPSCs were collected for each type of EPSC. The bath solution contained picrotoxin (100 μM). The intracellular solution contained 140mM CsCH3SO3, 10mM HEPES, 1 mM EGTA, 5mM TEA-Cl, 2mM MgCl2, 2.5mM MgATP and 0.3mM GTP, pH 7.2–7.4 (with CsOH).
Extracellular field recordings
All data acquisition and analyses were carried out blinded to genotype. One- to two-month-old FVB/N male mice were anaesthetized with isoflurane and their brains were removed and immersed immediately in ice-cold cutting solution (in mM): 110 sucrose, 3 KCl, 0.5 CaCl2, 60 NaCl2, 7 MgCl2, 1.25 NaH2PO4, 28 NaHCO3, 5 d-glucose saturated with 95% O2 and 5% CO2. 400-µm thick transverse hippocampal slices were prepared with a vibrating microtome Series 1000 (Vibratome) and recovered at 31 ± 0.5 °C for an hour in an interface chamber. Field excitatory postsynaptic potential (fEPSP) were recorded at 31 ± 0.5 °C in an interface chamber perfused at 1 ml min−1 with artificial cerebrospinal fluid (ACSF) containing (in mM): 125 NaCl, 2.5 KCl, 2 CaCl2, 1 MgCl2, 1.25 NaH2PO4, 25 NaHCO3, and 15 d-glucose. Extracellular stimuli were administered along the Schaffer collaterals using Formvar-insulated, bipolar nichrome electrodes controlled by a stimulus isolator (A-M Systems). An ACSF-filled glass-recording electrode was placed in stratum radiatum to record the field potential changes. Electrophysiological traces were amplified with AC-coupled amplifier (model 1800; A-M Systems), digitized using a Digidata 1320A (Molecular Devices), and acquired with pClamp 10 software (Molecular Devices). To assess baseline synaptic transmission, input–output relationships were examined by measuring the rising slope of the fEPSP evoked by 100-μs pulses over various stimulus intensities (1 V to 10 V). The stimulation intensity that evoked a fEPSP whose slope was 30–40% of the maximum fEPSP slope, determined by the input–output recording experiment, was used for the following recording paradigms. Long-term potentiation (LTP) was induced by two trains of 100 stimuli pulses at 100 Hz that are separated 20 s apart. To monitor LTP development, the fEPSPs were recorded every 20 s for 20 min before and 60 min after induction. The magnitude of potentiation was determined by measuring the changes in the slope of the fEPSP. NMDAR-dependent long-term depression (LTD) formation was induced with 900 paired stimuli with 50 ms inter-stimulus interval at 1 Hz frequency. In all experiments, genotypes were confirmed by PCR with tails cut. All data are shown as mean ± s.e.m.
In vivo immunoprecipitation and mass spectrometry
Striatum and hippocampus were dissected from 8 animals (5-weeks-old) per each genotype (wild-type and Shank3 transgenic FVB/N male mice), and crude synaptosomal fraction solubilized with deoxycholic acid (DOC) buffer was prepared as described previously47. Then ∼10 mg of lysates were incubated with GFP-Trap beads (ChromoTek) for 2 h at 4 °C. The beads were briefly washed with binding/dialysis buffer (50 mM Tris-HCl, pH 7.4, 0.1% Triton X-100) and boiled in with 1× NuPAGE LDS sample buffer (Invitrogen) to be loaded on SDS–PAGE (NuPAGE 4–12% Bis-Tris Gel, Invitrogen). The eluted proteins were visualized with Coomassie Brilliant blue stain and excised into 10 gel pieces according to molecular size. The individual gel pieces were destained and subject to in-gel digestion using trypsin. Tryptic peptide was dissolved in 10 μl of loading solution (5% methanol containing 0.1% formic acid) and subjected to nanoflow LC–MS/MS analysis with a nano-LC II (Thermo Scientific) coupled to LTQ Orbitrap Velos (Thermo Scientific) mass spectrometer. The peptides were loaded onto an in-house Reprosil-Pur Basic C18 (3 µm, Dr. Maisch) trap column which was 2 cm × 75 μm size. Then the trap column was washed with loading solution and switched in-line with an in-house 100 mm × 75 μm column packed with Reprosil-Pur Basic C18 equilibrated in 0.1% formic acid/water. The peptides were separated with a 75 min discontinuous gradient of 5–28% acetonitrile/0.1% formic acid at a flow rate of 450 nl min−1. Separated peptides were directly electro-sprayed into LTQ Orbitrap Velos mass spectrometer. The LTQ Orbitrap instrument was operated in the data-dependant mode acquiring fragmentation spectra of the top 50 strongest ions and under direct control of Xcalibur software (Thermo Scientific). Obtained MS/MS spectra were searched against target-decoy mouse refseq database in Proteome Discoverer 1.3 interface (Thermo Fisher) with Mascot algorithm (Mascot 2.3, Matrix Science). The precursor mass tolerance was confined within 20 p.p.m. with fragment mass tolerance of 0.5 Da and a maximum of two missed cleavage allowed. Assigned peptides were filtered with 5% false discovery rate (FDR) and subject to manual verifications. Any protein whose peptide was identified from wild-type brain sample was excluded from the further analysis.
Construction of Shank3 interactome network
To build an interaction network with the union set of Shank3 interactors (in vivo immunoprecipitation plus yeast two-hybrid screening), protein interactions were adopted from iRefIndex database (http://irefindex.org) which provides consolidated protein interactions from the 10 primary interaction databases (BIND, BioGRID, CORUM, DIP, HPRD, IntAct, MINT, MPact, MPPI and OPHID). Only the interactions among human and mouse genes were included. The direct interactions were distinguished from the indirect interactions based on experimental assays. Network graphics were generated with Cytoscape52. Nodes and edges were coloured based on the source and interaction type of the proteins, respectively. Thickness of edges for direct interactions correlates with the number of supportive evidences in the interaction database (Fig. 5b). To simplify the network, orphan nodes, defined by the nodes which have interaction only with Shank3, were excluded from the network (Extended Data Fig. 5c).
Network topology analysis of Shank3 interactome
Shortest path length (the number of links of the shortest path travelling from one protein to another protein) was calculated to measure the network topology of Shank3 interactome. To test the significance, we randomly picked the same number of proteins from the mouse PSP26 and calculated shortest path length of the interactome network generated with the proteins. This re-sampling was repeated 10,000 times and the empirical P value (the number of sample networks whose mean path length is smaller than that of Shank3 interactome/10,000) was calculated.
Gene ontology (GO) analysis
The GO analysis was carried out using DAVID software (version 6.7)53,54. The union set of Shank3 interactors from the in vivo immunoprecipitation and yeast two-hybrid screening was tested against a customized background gene list in which genes expressed at low levels in mouse brain were excluded.
Human subjects
Both individuals were referred to the Medical Genetics Laboratories, Baylor College of Medicine, Houston, USA, for clinical array comparative genomic hybridization (aCGH) analysis. Following informed consent, approved by the Institutional Review Board for Human Subject Research at Baylor College of Medicine, the medical records of the respective individuals were reviewed. Additional clinical information was obtained by phone interview with patient 1’s legal guardian and patient 2, respectively.
Array comparative genomic hybridization
Patient DNA, isolated from peripheral whole blood using the Puregene DNA extraction kit (Gentra) following the manufacturer’s instructions, was analysed using the Baylor College of Medicine V8 OLIGO clinical genomic microarray, described55. Briefly, this is a custom-designed genomic microarray with both genome-wide coverage and supplementary exonic coverage of ∼1,700 known or suspected disease genes, including SHANK3.
Quantification and statistical analysis
For quantification, values from three independent experiments with at least three biological replicates were used. For behavioural assays, all population values appear normally distributed and the variance is similar between the groups. P values were calculated by Student’s t-test or analysis of variance with proper post-hoc tests (GraphPad Prism), as specified in each figure legend or Supplementary Tables 1–3. All data are presented as mean ± s.e.m. *P < 0.05; **P < 0.01; ***P < 0.001.
References
Südhof, T. C. Neuroligins and neurexins link synaptic function to cognitive disease. Nature 455, 903–911 (2008)
Zoghbi, H. Y. Postnatal neurodevelopmental disorders: meeting at the synapse? Science 302, 826–830 (2003)
Bourgeron, T. A synaptic trek to autism. Curr. Opin. Neurobiol. 19, 231–234 (2009)
Ting, J. T., Peca, J. & Feng, G. Functional consequences of mutations in postsynaptic scaffolding proteins and relevance to psychiatric disorders. Annu. Rev. Neurosci. 35, 49–71 (2012)
Sheng, M. & Kim, E. The Shank family of scaffold proteins. J. Cell Sci. 113, 1851–1856 (2000)
Sato, D. et al. SHANK1 Deletions in Males with Autism Spectrum Disorder. Am. J. Hum. Genet. 90, 879–887 (2012)
Berkel, S. et al. Mutations in the SHANK2 synaptic scaffolding gene in autism spectrum disorder and mental retardation. Nature Genet. 42, 489–491 (2010)
Grabrucker, A. M., Schmeisser, M. J., Schoen, M. & Boeckers, T. M. Postsynaptic ProSAP/Shank scaffolds in the cross-hair of synaptopathies. Trends Cell Biol. 21, 594–603 (2011)
Durand, C. M. et al. Mutations in the gene encoding the synaptic scaffolding protein SHANK3 are associated with autism spectrum disorders. Nature Genet. 39, 25–27 (2007)
Moessner, R. et al. Contribution of SHANK3 mutations to autism spectrum disorder. Am. J. Hum. Genet. 81, 1289–1297 (2007)
Gauthier, J. et al. Novel de novo SHANK3 mutation in autistic patients. Am. J. Med. Genet. B. Neuropsychiatr. Genet. 150B, 421–424 (2009)
Gauthier, J. et al. De novo mutations in the gene encoding the synaptic scaffolding protein SHANK3 in patients ascertained for schizophrenia. Proc. Natl Acad. Sci. USA 107, 7863–7868 (2010)
Bonaglia, M. C. et al. Disruption of the ProSAP2 gene in a t(12;22)(q24.1;q13.3) is associated with the 22q13.3 deletion syndrome. Am. J. Hum. Genet. 69, 261–268 (2001)
Bonaglia, M. C. et al. Identification of a recurrent breakpoint within the SHANK3 gene in the 22q13.3 deletion syndrome. J. Med. Genet. 43, 822–828 (2006)
Bozdagi, O. et al. Haploinsufficiency of the autism-associated Shank3 gene leads to deficits in synaptic function, social interaction, and social communication. Mol. Autism 1, 15 (2010)
Peça, J. et al. Shank3 mutant mice display autistic-like behaviours and striatal dysfunction. Nature 472, 437–442 (2011)
Wang, X. et al. Synaptic dysfunction and abnormal behaviors in mice lacking major isoforms of Shank3. Hum. Mol. Genet. 20, 3093–3108 (2011)
Failla, P. et al. Schizophrenia in a patient with subtelomeric duplication of chromosome 22q. Clin. Genet. 71, 599–601 (2007)
Shaltiel, G. et al. Evidence for the involvement of the kainate receptor subunit GluR6 (GRIK2) in mediating behavioral displays related to behavioral symptoms of mania. Mol. Psychiatry 13, 858–872 (2008)
Leibenluft, E. & Rich, B. A. Pediatric bipolar disorder. Annu. Rev. Clin. Psychol. 4, 163–187 (2008)
Martinowich, K., Schloesser, R. J. & Manji, H. K. Bipolar disorder: from genes to behavior pathways. J. Clin. Invest. 119, 726–736 (2009)
Perry, W., Minassian, A., Feifel, D. & Braff, D. L. Sensorimotor gating deficits in bipolar disorder patients with acute psychotic mania. Biol. Psychiatry 50, 418–424 (2001)
Belmaker, R. H. Bipolar disorder. N. Engl. J. Med. 351, 476–486 (2004)
McCormick, D. A. & Contreras, D. On the cellular and network bases of epileptic seizures. Annu. Rev. Physiol. 63, 815–846 (2001)
Sakai, Y. et al. Protein interactome reveals converging molecular pathways among autism disorders. Sci. Transl. Med. 3, 86ra49 (2011)
Collins, M. O. et al. Molecular characterization and comparison of the components and multiprotein complexes in the postsynaptic proteome. J. Neurochem. 97 (Suppl 1). 16–23 (2006)
Bayés, A. et al. Characterization of the proteome, diseases and evolution of the human postsynaptic density. Nature Neurosci. 14, 19–21 (2011)
Campellone, K. G. & Welch, M. D. A nucleator arms race: cellular control of actin assembly. Nature Rev. Mol. Cell Biol. 11, 237–251 (2010)
Proepper, C. et al. Abelson interacting protein 1 (Abi-1) is essential for dendrite morphogenesis and synapse formation. EMBO J. 26, 1397–1409 (2007)
Naisbitt, S. et al. Shank, a novel family of postsynaptic density proteins that binds to the NMDA receptor/PSD-95/GKAP complex and cortactin. Neuron 23, 569–582 (1999)
Sheng, M. & Kim, E. The postsynaptic organization of synapses. Cold Spring Harb. Perspect. Biol. (http://dx.doi.org/10.1101/cshperspect.a005678) (2011)
Giesemann, T. et al. Complex formation between the postsynaptic scaffolding protein gephyrin, profilin, and Mena: a possible link to the microfilament system. J. Neurosci. 23, 8330–8339 (2003)
Neuhoff, H. et al. The actin-binding protein profilin I is localized at synaptic sites in an activity-regulated manner. Eur. J. Neurosci. 21, 15–25 (2005)
Ackermann, M. & Matus, A. Activity-induced targeting of profilin and stabilization of dendritic spine morphology. Nature Neurosci. 6, 1194–1200 (2003)
Jope, R. S. Anti-bipolar therapy: mechanism of action of lithium. Mol. Psychiatry 4, 117–128 (1999)
Rosenberg, G. The mechanisms of action of valproate in neuropsychiatric disorders: can we see the forest for the trees? Cell. Mol. Life Sci. 64, 2090–2103 (2007)
Ramocki, M. B. & Zoghbi, H. Y. Failure of neuronal homeostasis results in common neuropsychiatric phenotypes. Nature 455, 912–918 (2008)
Toro, R. et al. Key role for gene dosage and synaptic homeostasis in autism spectrum disorders. Trends Genet. 26, 363–372 (2010)
Gitlin, M. Treatment-resistant bipolar disorder. Mol. Psychiatry 11, 227–240 (2006)
Dunner, D. L. & Fieve, R. R. Clinical factors in lithium carbonate prophylaxis failure. Arch. Gen. Psychiatry 30, 229–233 (1974)
Gould, T. D. & Manji, H. K. Glycogen synthase kinase-3: a putative molecular target for lithium mimetic drugs. Neuropsychopharmacology 30, 1223–1237 (2005)
Maglóczky, Z. & Freund, T. F. Impaired and repaired inhibitory circuits in the epileptic human hippocampus. Trends Neurosci. 28, 334–340 (2005)
Marín, O. Interneuron dysfunction in psychiatric disorders. Nature Rev. Neurosci. 13, 107–120 (2012)
Benes, F. M. et al. Regulation of the GABA cell phenotype in hippocampus of schizophrenics and bipolars. Proc. Natl Acad. Sci. USA 104, 10164–10169 (2007)
Schloesser, R. J., Martinowich, K. & Manji, H. K. Mood-stabilizing drugs: mechanisms of action. Trends Neurosci. 35, 36–46 (2012)
Warming, S., Costantino, N., Court, D. L., Jenkins, N. A. & Copeland, N. G. Simple and highly efficient BAC recombineering using galK selection. Nucleic Acids Res. 33, e36 (2005)
Choi, J. et al. Regulation of dendritic spine morphogenesis by insulin receptor substrate 53, a downstream effector of Rac1 and Cdc42 small GTPases. J. Neurosci. 25, 869–879 (2005)
Han, K. et al. Regulated RalBP1 binding to RalA and PSD-95 controls AMPA receptor endocytosis and LTD. PLoS Biol. 7, e1000187 (2009)
Chao, H. T. et al. Dysfunction in GABA signalling mediates autism-like stereotypies and Rett syndrome phenotypes. Nature 468, 263–269 (2010)
Roberson, E. D. et al. Amyloid-β/Fyn-induced synaptic, network, and cognitive impairments depend on tau levels in multiple mouse models of Alzheimer’s disease. J. Neurosci. 31, 700–711 (2011)
Lu, H., Lim, B. & Poo, M. M. Cocaine exposure in utero alters synaptic plasticity in the medial prefrontal cortex of postnatal rats. J. Neurosci. 29, 12664–12674 (2009)
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003)
Dennis, G. et al. DAVID: database for annotation, visualization, and integrated discovery. Genome Biol. 4, P3 (2003)
Huang D. W, Sherman B. T & Lempicki R. A Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nature Protocols 4, 44–57 (2009)
Boone, P. M. et al. Detection of clinically relevant exonic copy-number changes by array CGH. Hum. Mutat. 31, 1326–1342 (2010)
Acknowledgements
We are indebted to the patients and families who participated in this study; to J. W. Belmont and N. Miller for contributing patients to this study; G. Feng for sharing Shank3B mice; G. Schuster for injection of Shank3 BAC; and C. Spencer for behavioural assays training. This project was supported by The Howard Hughes Medical Institute (H.Y.Z.), National Institutes of Health (NIH) ARRA grant (1R01NS070302) (H.Y.Z.), the Baylor Intellectual and Developmental Disabilities Research Center (P30HD024064) confocal, electrophysiology and mouse neurobehavioral cores, and the Cancer Prevention and Research Institute of Texas (CPRIT) RP110784. J.L.H. was supported by an Early Career Award from the Thrasher Research Fund, NIH 2T32NS043124 and the Ting Tsung and Wei Fong Chao Foundation; C.P.S. was supported by the Joan and Stanford Alexander family, the Ting Tsung and Wei Fong Chao Foundation and the Doris Duke Clinical Scientist Development Award.
Author information
Authors and Affiliations
Contributions
K.H., J.L.H., H.L., H.C., J.T., H.-C.L. and H.Y.Z. designed the experiments. K.H., J.L.H., H.L., H.C., J.T., Z.W., S.H. and H.S. performed the research. K.H., J.L.H., C.P.S., H.L., H.C., H.K., J.T., Z.W., S.H., S.W.C., P.Y., A.M.B., A.P., H.-C.L. and H.Y.Z. collected, analysed and interpreted the data. K.H., J.L.H., C.P.S., H.L., H.K., J.T., H.-C.L. and H.Y.Z. wrote and edited the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Characterization of expression patterns of EGFP–Shank3 and other synaptic proteins in Shank3 transgenic mice.
a, Description of the Shank3 BAC used for transgenic mice generation (image was modified from UCSC genome browser). b, Western blot images show expression of EGFP–Shank3 in the synaptic fraction of Shank3 transgenic brain lysates. EGFP antibody recognized EGFP–Shank3 (∼200 kDa) specifically in synaptic fractions of Shank3 transgenic mice. Asterisk indicates non-specific bands detected by the EGFP antibody. Shank3 antibody detected endogenous Shank3 plus EGFP–Shank3 (arrow) in the transgenic samples. S2, soluble fraction; P2, crude synaptosomal fraction; PSD I, synaptic fraction after one time Triton X-100 washout. c, The brain regional expression pattern of EGFP–Shank3 is similar to that of endogenous Shank3. Cb, cerebellum; Ct, cortex; Hp, hippocampus; St, striatum; Th, thalamus. d, The brain developmental expression pattern of EGFP–Shank3 is similar to that of endogenous Shank3. e, The fold changes of Shank3 proteins in 3-week-old transgenic mice (n = 4) are similar to those of 6-week-old mice (Fig. 1d). f, Female transgenic mice (n = 4) show similar fold changes of Shank3 proteins to male transgenic mice (Fig. 1d). g, h, Expression levels of excitatory (g) and inhibitory (h) synaptic proteins are not significantly altered, except Shank3, in the synaptosomal fraction of 8-week-old transgenic hippocampus and striatum (n = 6). All data are presented as mean ± s.e.m. P values (*P < 0.05) were derived from unpaired, two-tailed Student’s t-test.
Extended Data Figure 2 Behavioural characterization of Shank3 transgenic mice.
a, Transgenic mice showed increased locomotor speed during the 30 min open-field assay. b, Female transgenic mice showed increased home-cage activity. c, Transgenic mice have increased body weight compared to wild-type mice measured at 9-weeks-old (male wild type: 28.1 ± 0.9, male transgenic: 30.6 ± 0.7, female wild type: 20.1 ± 0.3, female transgenic: 21.4 ± 0.4, n = 10–13; *P < 0.05; unpaired two-tailed Student’s t-test) and 15-weeks-old (male wild type: 32.4 ± 1.2, male transgenic: 37.0 ± 1.3, female wild type: 23.6 ± 0.4, female transgenic: 26.6 ± 0.9, n = 10–13; *P < 0.05). d, Increased food intake by transgenic mice. Food intake was measured from single-caged female transgenic or wild-type mice from 6- to 10-weeks-old. At 7-weeks-old (wild type: 3.51 ± 0.05, transgenic: 3.69 ± 0.06, n = 9–11; *P < 0.05; unpaired two-tailed Student’s t-test) and 9-weeks-old (wild type: 3.61 ± 0.07, transgenic: 3.86 ± 0.05, n = 9–11; *P < 0.05), food intake by transgenic mice was significantly higher than wild-type mice. e, f, In the 3-chamber assay, male (e) and female (f) transgenic mice did not show significant preference for novel mice to novel objects. g, Transgenic mice spent significantly less time in close interaction with novel social partners. h, i, Stereotypy time (h) and activity count (i) during the 30 min open-field test were normal in transgenic mice. j, Time spent in grooming during 10 min monitoring was normal in transgenic mice. k, Separation-induced ultrasonic vocalization was measured from wild-type and transgenic mice of postnatal day 6 to 13. Transgenic mice made less calls than wild type at postnatal day 13. l, At postnatal day 13, transgenic mice express more Shank3 proteins than wild-type mice. All data are presented as mean ± s.e.m. *P < 0.05, **P < 0.01, ***P < 0.001. The summary of statistical analyses for behavioural assays is provided in Supplementary Table 1.
Extended Data Figure 3 Behavioural phenotypes of Shank3 transgenic mice were rescued by crossing with Shank3B+/− mice.
a, Schematic diagram shows possible genotypic combinations and their Shank3 expression levels (arrow) from the crossing between Shank3 transgenic mice and Shank3B+/− mice. b, Quantification of the levels of Shank3 proteins in each genotype (n = 4). The fold changes were compared to the wild-type (Shank3B+/+) controls. Crossing of Shank3 transgenic mice with Shank3B+/− mice significantly decreased the levels of α isoform and total Shank3 protein. c, d, Locomotor activities of male (c) and female (d) Shank3 transgenic mice were rescued by crossing with Shank3B+/− mice. e, f, Immobile time in tail-suspension test of male (e) and female (f) Shank3 transgenic mice were rescued by crossing with Shank3B+/− mice. g, Prepulse inhibition (by 78 dB) of female Shank3 transgenic mice was rescued by crossing with Shank3B+/− mice. All data are presented as mean ± s.e.m. *P < 0.05; ***P < 0.001.
Extended Data Figure 4 Basal synaptic transmission and NMDA receptor-dependent synaptic plasticity are normal at the hippocampal Schaffer collateral-CA1 synapses of Shank3 transgenic mice.
a, Images of cresylviolet staining show normal cytoarchitecture of transgenic brain. b, Normal paired-pulse facilitation ratio (PPR) of evoked EPSCs (eEPSCs) at transgenic Schaffer collateral-CA1 synapses. c, d, Normal synaptic input-output (I-O) relationship at transgenic Schaffer collateral-CA1 synapses. I-O curve (c) and cumulative curve of I-O slope (d) are shown. Red lines in c represent the fitting curves. e, Left, sample traces of action potentials triggered by 200 pA current injection in hippocampal CA1 region of wild-type and transgenic mice. Middle and right, unaltered number of action potentials trigged by the injection of current at different level (middle) and amplitude of the first action potential triggered by 800-ms-long pulse of 200 pA current (right) show normal intrinsic excitability of hippocampal CA1 pyramidal neurons in transgenic mice. f, Amplitude, frequency and decay of mEPSC are not altered in CA1 pyramidal neurons of transgenic mice. g, h, NMDA receptor-dependent long-term potentiation (g) and long-term depression (h) at Schaffer collateral-CA1 pyramidal synapses are normal in transgenic neurons.
Extended Data Figure 5 Generation and characterization of Shank3 in vivo interactome.
a, Isolation of EGFP–Shank3 protein and its interactors from synaptosomal fraction of transgenic mice. b, Venn diagrams show overlaps among the protein lists from Shank3 in vivo immunoprecipitation, Shank3 yeast two-hybrid screening, and either published mouse PSP (postsynaptic proteome) or human PSD. c, Shank3 interactome network. d, Average path length of Shank3 interactome (3.12, red line) is significantly (P < 0.0001) shorter than that of mouse PSP interactome (mean = 3.91, black line), supporting strong connectivity of Shank3 interactome. e, GO and KEGG pathway analysis of Shank3 interactome reveal enrichment of actin cytoskeleton-related function/pathway. The full result of analysis is in Supplementary Table 5. f, Confirmation of the in vivo interactions between Shank3 and actin-related proteins. g, In cultured hippocampal neurons, ARPC2 proteins are co-localized with F-actin (upper panel) and EGFP–Shank3 (lower panel).
Extended Data Figure 6 Shank3 siRNA reversed increased F-actin levels and ARPC2 cluster size in cultured hippocampal pyramidal neurons from Shank3 transgenic mice.
a, Validation of the two siRNAs against Shank3. HEK293T cells were transfected with HA-Shank3 plus control or Shank3 siRNA. EGFP plasmid was co-transfected as an internal control. After 48 h, expression levels of HA–Shank3 were measured by western blot and quantified (n = 3). b, siRNA targeting Shank3 (si-Shank3) reversed the increased F-actin levels of Shank3 transgenic neurons. Image for si-Shank3 #2 is not shown. c, si-Shank3 reversed the increased ARPC2 cluster size of Shank3 transgenic neurons. All data are presented as mean ± s.e.m. from 20–30 neurons per condition. P values (*P < 0.05, **P < 0.01) were derived from one-way ANOVA with post hoc Tukey’s multiple comparison.
Extended Data Figure 7 Normal excitatory and inhibitory synapse numbers of GAD-6-positive inhibitory neurons of Shank3 transgenic mice.
a, Expression of EGFP–Shank3 in GAD-6-positive inhibitory neurons of Shank3 transgenic mice. The yellow circles of dotted line and solid line indicate neuronal cell bodies of a GAD-6-negative excitatory neuron and a GAD-6-positive inhibitory neuron, respectively. The box 1 shows dendritic segment of the excitatory neuron and the box 2 shows that of the inhibitory neuron. b, Quantification of dendritic EGFP–Shank3 intensity in excitatory and inhibitory transgenic neurons. Inhibitory neurons express less EGFP–Shank3 (excitatory neurons: 1.00 ± 0.06, inhibitory neurons: 0.74 ± 0.09, n = 10; *P < 0.05; unpaired two-tailed Student’s t-test). c, Normal excitatory synapse number of GAD-6-positive inhibitory neurons of Shank3 transgenic mice. d, Normal inhibitory synapse number of GAD-6-positive inhibitory neurons of Shank3 transgenic mice. e, Quantification of (c) (wild type: 1.00 ± 0.10, transgenic: 0.99 ± 0.07, n = 18; P > 0.05; unpaired two-tailed Student’s t-test) and (d) (wild type: 1.00 ± 0.12, transgenic: 0.95 ± 0.09, n = 20; P > 0.05; unpaired two-tailed Student’s t-test).
Extended Data Figure 8 Shank3 interactome is connected with gephyrin-interacting actin-related proteins, Mena, profilin1 and profilin2.
Based on a literature search, we selected candidate actin-related proteins (Mena/VASP, profilin1 and profilin2) that directly interact with inhibitory postsynaptic protein gephyrin. To understand potential interactions among these proteins and the Shank3 interactome, we generated interaction network using the information from our yeast two-hybrid screening and IRefIndex PPI database (http://irefindex.org). The interaction network shows that Mena/VASP, profilin1 and profilin2 are connected with Shank3 interactome mainly through actin-related proteins. Note that one of the gephyrin-interacting proteins, profilin2, was also identified by our Shank3 in vivo immunoprecipitation (Supplementary Table 4).
Extended Data Figure 9 Mania-like behaviours of Shank3 transgenic mice are resistant to lithium treatment.
a, Basal and amphetamine-induced locomotor activities of female transgenic mice were not affected by lithium treatment. b–e, Abnormal acoustic startle response (b, d) and PPI (c, e) of male and female transgenic mice were not rescued by lithium treatment. f, g, Immobile time of male (f) and female (g) transgenic mice in tail-suspension test was not rescued by lithium treatment. Wild-type female mice showed decreased immobile time upon lithium treatment, which is an expected response to high dose lithium treatment in wild-type mice. All data are presented as mean ± s.e.m. *P < 0.05; ***P < 0.001.
Extended Data Figure 10 Valproate treatment does not affect synaptic protein and F-actin levels in neurons of Shank3 transgenic mice.
a, One hour after final valproate injection (200 mg per kg), synaptosomal fraction was prepared from brains, and indicated proteins were detected by western blotting. Neither wild-type nor transgenic mice showed significant change in the levels of synaptic proteins by valproate treatment (n = 7). b, Representative confocal images of Shank3 transgenic cultured hippocampal pyramidal neurons (DIV 14) treated with different concentrations of valproate (0, 0.01, 0.1 and 1 mM) for 30 or 60 min. c, Quantification of (b). There was no significant change in F-actin levels by valproate treatment (n = 16 per condition). All data are presented as mean ± s.e.m.
Supplementary information
Supplementary Data
This file contains Supplementary data and Supplementary Tables 4-5. (PDF 406 kb)
Supplementary Data
This file contains Supplementary Tables 1-3. (XLSX 35 kb)
Spontaneous seizure of Shank3 transgenic mice
This video shows spontaneous seizure from an 8-week old female Shank3 transgenic mouse in the home-cage. (MP4 5938 kb)
Rights and permissions
About this article
Cite this article
Han, K., Holder Jr, J., Schaaf, C. et al. SHANK3 overexpression causes manic-like behaviour with unique pharmacogenetic properties. Nature 503, 72–77 (2013). https://doi.org/10.1038/nature12630
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature12630
This article is cited by
-
Lithium rescues dendritic abnormalities in Ank3 deficiency models through the synergic effects of GSK3β and cyclic AMP signaling pathways
Neuropsychopharmacology (2023)
-
HSPA12A controls cerebral lactate homeostasis to maintain hippocampal neurogenesis and mood stabilization
Translational Psychiatry (2023)
-
Disrupted extracellular matrix and cell cycle genes in autism-associated Shank3 deficiency are targeted by lithium
Molecular Psychiatry (2023)
-
Mutations affecting the N-terminal domains of SHANK3 point to different pathomechanisms in neurodevelopmental disorders
Scientific Reports (2022)
-
SHANK family on stem cell fate and development
Cell Death & Disease (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.