Abstract
DNA synthesis has been extensively studied, but the chemical reaction itself has not been visualized. Here we follow the course of phosphodiester bond formation using time-resolved X-ray crystallography. Native human DNA polymerase η, DNA and dATP were co-crystallized at pH 6.0 without Mg2+. The polymerization reaction was initiated by exposing crystals to 1 mM Mg2+ at pH 7.0, and stopped by freezing at desired time points for structural analysis. The substrates and two Mg2+ ions are aligned within 40 s, but the bond formation is not evident until 80 s. From 80 to 300 s structures show a mixture of decreasing substrate and increasing product of the nucleotidyl-transfer reaction. Transient electron densities indicate that deprotonation and an accompanying C2′-endo to C3′-endo conversion of the nucleophile 3′-OH are rate limiting. A third Mg2+ ion, which arrives with the new bond and stabilizes the intermediate state, may be an unappreciated feature of the two-metal-ion mechanism.
Similar content being viewed by others
Main
Formation and breakage of chemical bonds underlie all life processes. DNA replication, which is essential for cell proliferation, is but one example1,2. In each reaction cycle, a dNTP complementary to the templating base is incorporated into DNA in a nucleotidyl-transfer reaction catalysed by a polymerase, during which a new bond is formed between the 3′-OH of the primer strand and the α-phosphate of the dNTP, and the phosphodiester bond between the α- and β-phosphates of dNTP is broken (Fig. 1a). As a result the primer strand is extended by one nucleotide and a pyrophosphate is released. This reaction has been shown to require two Mg2+ ions (A and B) and is inferred to be SN2-type, forming a pentacovalent phosphate intermediate3,4. A similar two-metal-ion-dependent mechanism is believed to be shared by all DNA and RNA polymerases and many nucleases5.
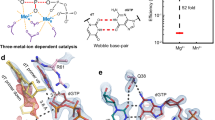
a, Diagrams of DNA synthesis and the procedure of the in crystallo reaction. AH, general acid; B, general base. GS, ground state; PS, product state; RS, reactant state. b, Reaction rate (kobs) is plotted against mixed Mg2+ and Ca2+ at a total concentration of 1 mM. c, The pH- and buffer-dependent reaction rates. d, The ground-state structure. The active site is superimposed with the 2Fo − Fc electron density contoured at 1.2σ (top panel) or with the non-reactive dAMPNPP ternary complex (3MR2, silver, bottom panel). Hydrogen bonds and metal-ion coordination are shown as dashed lines.
DNA synthesis has been analysed by kinetic measurements, dynamic simulations and structural studies2,6,7. Crystal structures of a number of DNA polymerases in substrate-bound forms have been determined using non-reactive substrate analogues or with Ca2+ instead of Mg2+ to prevent the nucleotidyl-transfer reaction5,8,9,10,11,12,13,14,15,16,17,18. These structures show active-site configurations in an array of pre-reaction ground states. Pre-steady-state kinetic studies of several DNA polymerases indicate that, after large conformational changes induced by substrate binding, unspecified subtle changes in the active site are the rate-limiting step19,20,21,22,23,24,25,26,27. The actual process of DNA synthesis has, to our knowledge, never been visualized.
Human DNA polymerase η (Pol η) is specialized in lesion bypass and has a preformed catalytic centre that undergoes limited conformational change during DNA synthesis28. Crystal structures of the catalytic domain of Pol η complexed with DNA and a non-reactive dNTP analogue poised for catalysis have been determined at 1.8 Å resolution29,30. Taking advantage of the slow reaction rate and hindered post-reaction DNA translocation in crystallo, we follow the course of DNA synthesis by X-ray crystallography and report here the structural changes and transient elements that are associated with the reaction.
The ground state of Pol η
The Pol η activity in a reaction buffer containing 0.9 mM Mg2+ and 0.1 mM Ca2+ was only one-third of that with 1 mM Mg2+ (Fig. 1b), indicating that both active-site metal ions have to be Mg2+ for the catalysis. In accordance with the bell-shaped pH-dependence curve characteristic for the acid–base catalysis17,31, the catalytic rate of Pol η is extremely low at pH 6.0 and rises with increasing pH from 6.0 to 8.0 (Fig. 1c). To prevent the nucleotidyl-transfer reaction, crystals of Pol η–substrate complexes were grown at pH 6.0 with only one Ca2+ per protein–DNA–dATP complex (Methods). After soaking in MES (pH 6.0–7.2) or HEPES buffer (pH 7.0–7.5) with Na+ or K+ but no divalent cation for 5 to 30 min, these crystals maintained excellent diffraction up to pH 7.0, but decayed at pH 7.5 with reduced resolution and increased diffuse scattering.
The structure of Pol η crystal equilibrated at pH 6.8 was refined at 1.50 Å. A Ca2+ ion clearly occupies the B metal-ion-binding site and is coordinated in the octahedral geometry by oxygen atoms of the dATP and the active-site residues (Fig. 1d). The A metal-ion-binding site has low occupancy of a monovalent cation. The occupancy of Na+ or K+ in the A site increases with the pH (Supplementary Fig. 1a, b), which is probably correlated with deprotonation of the active-site carboxylates. Pol η binds Na+ more readily than K+, particularly below pH 7.0, probably because Na+ is similar to Mg2+ in size and is smaller than K+. As expected, there is no nucleotidyl-transfer reaction, and the structure is termed the ground-state ternary complex.
The ground-state structure is similar to the Pol η ternary complex with a non-reactive dATP analogue dAMPNPP (2′-deoxyadenosine-5′-[(α, β)-imido]triphosophate) and two Mg2+ ions (Protein Data Bank ID 3MR2)29. However, without an A-site divalent cation, the 3′-OH of the primer strand shifts away from the dATP and forms hydrogen bonds with the side chains of the S113 and D115 residues, and the active-site carboxylates D13 and E116 adopt different rotamer conformations (Fig. 1d). Replacement of dAMPNPP with dATP leads to stabilization of R61, which forms bidentate hydrogen bonds with the α-phosphate.
Catalysis in crystallo
The nucleotidyl transfer was initiated by transferring Pol η crystals to a pH 6.8 or 7.0 reaction buffer containing 1 mM Mg2+ but no dATP (Methods). After incubation at 293 K for 40 to 300 s, the reaction was terminated in ∼40-s intervals by freezing crystals in liquid nitrogen at 77 K. Diffraction data were collected to 1.50–1.95 Å Bragg spacings (Supplementary Table 1). The reaction process was monitored by the electron density corresponding to the new chemical bond in the Fo − Fc map compared with the refined ground-state structure (Fig. 2a and Supplementary Fig. 2). The reaction time courses at pH 6.8 and 7.0 in crystallo (Fig. 2b) are approximately 20–100-fold slower than in solution (Fig. 1c), probably owing to the reduced thermal motion. By 40 s, the A site was fully occupied with Mg2+ (Fig. 2a and Supplementary Fig. 1c, d). About 50% of Ca2+ in the B site was replaced by Mg2+, and exchange of the remaining Ca2+ took place slowly (Supplementary Fig. 3). Binding of two Mg2+ ions leads to the alignment of the 3′-OH and dATP18,32. The refined structure is nearly identical to that with dAMPNPP, except for tighter dATP coordination by R61 and closer proximity of the Mg2+ ions (3.4 Å apart versus 3.6 Å). As there is no sign of bond formation, the structure is termed the reactant state.
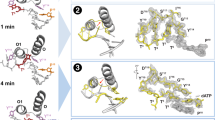
a, Two views (top and bottom diagrams) of Fo(40–230 s) − Fc(40 s) omit maps (4.0σ) superimposed onto the 40-s structure (pH 7.0). The emerging densities are shown by arrows or circled. b, A plot of the absolute peak height of the new bond density versus reaction time at pH 6.8 and 7.0 in AT and TG crystals (Supplementary Table 1a, b). The noise level (1σ) is indicated. c, The 230-second structure refined as the pentacovalent transition state, or d, the reactant state (yellow) and product state (blue) mixture. The 2Fo − Fc (1σ, grey) and Fo − Fc map (± 3.0σ, green and red) are superimposed. Hydrogen bonds and metal-ion coordination are shown as dashed lines.
Electron density corresponding to a new bond between the 3′-OH and α-phosphorus of dATP begun to emerge at 80 s, increased quickly over the next 60 s and reached a maximum after 200 s, when the reaction was 60–70% complete (Fig. 2b). The seemingly reduced rate after 140 s is probably due to the reverse reaction in crystallo. The slight decline of product after 250 s (Fig. 2b) is due to a sideway product translocation, which is clear with longer incubation (Supplementary Fig. 4a). To alleviate an impediment to proper DNA translocation by the crystal lattice, we replaced an AT base pair with a mismatch at the DNA end that formed lattice contacts (Methods and Supplementary Table 2). The DNA with a TG mismatch led to isomorphous crystals and proper translocation of the DNA product (Supplementary Fig. 4b). Interestingly, the time courses of nucleotidyl transfer in the AT and TG crystals are nearly identical and are unaffected by the lattice contacts (Fig. 2b).
At the peak of chemical-bond formation between 200 and 250 s, the scissile phosphate can be refined in a pentacovalent transition state without restraints (Fig. 2c); however, the bond distances between the phosphorus and the attacking or leaving oxygen atoms are 2.2–2.5 Å (Fig. 2c), much longer than the expected 2.0 Å observed with transition-state mimics such as AlF4 or MgF3 (refs 33–35). Moreover, in the Fo − Fc map residual electron densities are observed around the new and scissile phosphodiester bonds (Fig. 2c). The same diffraction data, however, can be well fitted as a mixture of the reactant state and the product state immediately before and after the nucleotidyl transfer (Fig. 2d). Because the transition state is transient and unstable, the structures obtained between 80 and 300 s are refined as a mixture of the reactant state and product state at different ratios (Fig. 2b and Supplementary Table 1a). For instance, the 1.52 Å structure at 230 s consists of 40% substrate and 60% product.
Between the reactant and product state, the protein, DNA and dATP are superimposable except for atoms in the reaction centre. Most notably, the α-phosphorus of dATP moves 1.4 Å along a straight line between the attacking and leaving oxygen atoms (separated by 4.6 Å; Fig. 2d). The shift of the α-phosphate is accompanied by alteration of residue R61 of Pol η, which flips away from the scissile phosphate and is replaced by a new metal ion and water molecules (Fig. 2a, d; see details below). On the primer strand, changes are confined to the 3′ nucleotide. The 3′-OH, together with the deoxyribose, moves towards the α-phosphate by 0.5 Å, and the sugar pucker changes from C2′-endo in the reactant state to C3′-endo in the product state (Fig. 2d). With the loss of the nucleophile and α-phosphate as ligands, the A-site Mg2+ dissociates in the product state, as evidenced by the declining occupancy (Supplementary Fig. 1c, d). Concomitantly, D13 assumes a second conformation and forms a hydrogen bond with K224.
The C3′-endo (A form) conformation at the 3′ primer end was observed in ternary complexes with the A-, B- and X-family DNA polymerases and is thought to be important for forming a shallow minor groove for dNTP selection9,11,12,14,15,17,18,32. Among the Y-family polymerases, the primer end has always been observed as C2′-endo28. In the Pol η ground-state and reactant-state complexes, the dATP and the nucleotide 5′ to the primer end have the A-form conformation, but only in the product-state structure does the primer end adopt the C3′-endo conformation to avoid clashes between its C2′ atom and the non-bridging oxygen of dATP during and immediately after the nucleophilic attack (Figs 2d and 3a). Because the electron density is weak for the sugar moiety at the 3′-primer end, we tested the effect of the A-form conformation by using a primer with a ribonucleotide at its 3′ end. The catalytic efficiency (kcat/Km) of Pol η is comparable whether a ribonucleotide or deoxyribonucleotide is at the primer end (Fig. 3b), as observed for DNA pol β (ref. 36), indicating that the A-form conformation is probably necessary for DNA synthesis in general.
a, Superposition of the refined reactant state (80 s, yellow) and product state (300 s, blue). The transient water molecule, probably deprotonating the 3′-OH, is circled. The sugar pucker at the primer 3′ end changes from C2′-endo in the reactant state to C3′-endo in the product state. The clashes between α-phosphate and the C2′ if it is C2′-endo, and between the C2′ and the transient water molecule if the water molecule does not depart, are indicated by red dashed double arrowheads. Hydrogen bonds and metal-ion coordination are shown as dashed lines. b, Wild-type (WT) and S113A-mutant Pol η can both extend primer with a ribonucleotide at the 3′ end (lighter colour) as with pure DNA (darker colour).
A transient water molecule
Two unexpected spheres of electron density appear in the Fo − Fc maps in the course of new bond formation (Fig. 2a). The first is within hydrogen-bonding distance of the 3′-OH. Its electron density peaks at 80 s and declines considerably after 140 s when the product state becomes prominent. Although absent in the ground state and the 40-second reactant-state structure, the electron density is superimposable with a water molecule observed in the Pol η–dAMPNPP ternary complex. This water molecule is hydrogen-bonded with the 3′-OH, the O4′ of dATP and another water molecule in the 80-second reactant-state structure, but it is incompatible with the C3′-endo conformation in the product state (Fig. 3a). The 3′-OH is also hydrogen-bonded to a water molecule in an X-family DNA polymerase crystallized with dUMPNPP (2′-deoxyuridine-5′-[(α, β)-imido]triphosophate)32. A water-mediated and substrate-assisted catalytic mechanism has previously been proposed37,38. In addition to the water molecule, S113 is hydrogen-bonded with the 3′-OH in the ground state and may pick up the proton and pass it to E116 in the reactant state (Fig. 1d). However, the S113A-mutant Pol η retains 95% of kcat with a threefold increase of Km (Fig. 3b and Supplementary Table 3). Interestingly, mutations of the S113 equivalent in the A- and B-family DNA polymerases (highly conserved histidine and threonine, respectively) also have limited impact on catalytic efficiency39,40,41. We therefore propose that 3′-OH is prone to deprotonation when coordinated by Mg2+ and aligned with the incoming nucleotide4. The proton can be passed on to the transient water molecule and then to bulk solvent. When the 3′-end of the primer is a ribonucleotide, the O2′ may relay the proton out instead. An incoming dNTP most probably participates in deprotonation of the nucleophile because, with the nitrogen substitution in dAMPNPP, the water molecule is bound stably to the 3′-OH29,30.
The third metal ion
The second emerging sphere of electron density is very close to the α-phosphate on the opposite side of the A- and B-site Mg2+ ions (Fig. 2a). It starts to appear at 140 s and intensifies with the reaction time. By 230 s with the structure refined at 1.52 Å resolution, the octahedral geometry and the short coordination distances indicate that this is a divalent cation and most probably Mg2+ (Fig. 4). This third metal ion is liganded by four water molecules, two of which occupy the space of the departed R61 (Fig. 2d). Two additional ligands are the leaving oxygen (bridging between the α- and β-phosphate) and the non-bridging oxygen of the α-phosphate (Fig. 4). This Mg2+ thus bridges the two reaction products destined to separate and prevents DNA translocation. In the TG crystals with 1 mM Mg2+ in the reaction buffer, the mixed reactant-state and product-state intermediate stays at equilibrium for up to 15 min, but with reduced Mg2+ (5 µM) and addition of dATP (5 µM), product release occurs much faster (Supplementary Information). The physiological role of the third metal ion is probably to neutralize the negative charge built up in the transition state and may also facilitate protonation of the pyrophosphate.
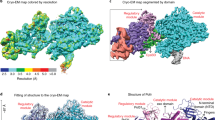
A stereo view of the three metal ions (Mg2+A–C) observed in the intermediate state of 230 s after Mg2+ addition, which consists of a mixture of product state (blue) and reactant state (yellow). The third Mg2+ (Mg2+C) has six ligands, four of which are water molecules. The 2Fo − Fc map contoured at 1.5σ (grey) and Fo − Fc map with the third Mg2+ omitted contoured at 4.0σ (green) are overlaid with the ball-and-stick model. Hydrogen bonds and metal-ion coordination are shown as dashed lines.
Three metal ions have been structurally observed in a number of enzymes catalysing phosphoryl-transfer reactions, including alkaline phosphatase, P1 nuclease and endonuclease IV33,42,43,44. In the case of endonuclease IV, the three Zn2+ ions observed in both the substrate and product state44 are reminiscent of the Mg2+ with Pol η (Supplementary Fig. 5). The two metal ions essential for catalysis flank the scissile phosphate on one side, and the third metal ion bridges the reaction products on the other. A similar arrangement of three metal ions was proposed for group-I introns based on chemical probing45. However, the third metal ion was absent in a crystal structure of a splicing intermediate, probably owing to its transient nature46.
Concluding remarks
Previously, flash–freeze was used to trap covalent intermediates and conformational changes associated with a chemical reaction47,48,49. Here, we extend the technology to record DNA synthesis in real time and at atomic resolution (Fig. 5 and Supplementary Movie 1). The arrival of a water molecule to deprotonate the 3′-OH occurs only after two-Mg2+-ion-induced substrate alignment and in the presence of a correct dNTP but not a non-reactive dAMPNPP. Mg2+-dependent and water-mediated deprotonation may occur in all DNA polymerases. The third metal ion that replaces the side chain of R61 during the nucleotidyl transfer probably stabilizes the transition state and facilitates product release. In the one-metal-ion-dependent TraI-like nucleases and topoisomerases a conserved Lys or Arg also occupies the position equivalent to R61 in Pol η (ref. 50). The involvement of an additional transient metal ion may be a general feature of both the one- and two-metal-ion mechanism.
The B-site metal ion is stably associated with the incoming dNTP and the enzyme. Binding of Mg2+ at the A site aligns the reactants, in particular the 3′-OH, and promotes its deprotonation, which is assisted by dNTP and the transient water molecule. After the reaction is initiated and before the products are released, a third metal ion (C) replaces R61 to stabilize the reaction intermediates. Hydrogen bonds and metal-ion coordination are shown as dashed lines.
Methods Summary
Pol η, DNA and dATP complexes mixed with Ca2+ in a 1:1 molar ratio were concentrated to ∼3 mg ml−1 Pol η and crystallized by the hanging-drop vapour-diffusion method29. For in crystallo reactions, crystals were first stabilized in 0.1 M MES (pH 6.8–7.2), 5 µM dATP, 20% (w/v) polyethylene glycol 2000 monomethyl ether (PEG2K-MME) and 1 mM dithiothreitol (DTT) at a specific pH for ∼30 min, transferred in a nylon loop to the reaction buffer containing 0.1 M MES (pH 6.8–7.2), 1 mM MgCl2, 20% (w/v) PEG2K-MME and 1 mM DTT for the desired amount of time, followed by a quick dipping in a cryo-solution supplemented with 20% (w/v) glycerol and flash-cooled in liquid nitrogen. The reaction time courses were determined based on the Fo − Fc maps with all data scaled against and compared to the refined ground-state structures. The intermediate states were refined as a mixture of reactant state and product state, and their ratio at each time point was determined according to the reaction course (Fig. 2b) and difference (Fo − Fc) electron-density maps. Catalytic rates (kobs), kcat and Km were measured using the template/primer listed in Supplementary Table 3.
Online Methods
Protein and crystal preparation
S113A-mutant Pol η was made using QuikChange (Stratagene). Wild-type Pol η (residues 1–432) and S113A-mutant proteins were expressed and purified as described29. The ternary complexes of Pol η, DNA and dATP were mixed with Ca2+ at 1:1 molar ratio and concentrated to a protein concentration of ∼3 mg ml−1. Oligonucleotides used for original (AT pair at the DNA end) and three ‘loosely packed’ crystals (TG or GT mispaired and an unpaired A) are shown in Supplementary Table 2. An unpaired nucleotide at the primer 5′ end was disordered in the original Pol η structures29, and was therefore removed to improve the diffraction quality. All crystals were obtained using the hanging-drop vapour-diffusion method against a reservoir solution containing 0.1 M MES (pH 6.0) and 9–17% (w/v) PEG2K-MME and streak seeding.
Chemical reaction in crystallo
Based on diffraction qualities, the complexes containing the original AT and TG-end DNA were selected for chemical reaction in crystallo. These crystals were first transferred and incubated in a pre-reaction buffer containing 0.1 M MES (pH 6.8–7.2), 5 μM dATP, 20% (w/v) PEG2K-MME and 1 mM dithiothreitol (DTT) for ∼30 min. For the pH 6.8 time course (Fig. 2b, Supplementary Table 1b) and translocation in the TG crystal, the pre-reaction buffer also contained 5 µM MgCl2. The different pre-reaction buffer might cause the larger variations of the pH 6.8 reactions. Chemical reaction was initiated by transferring the crystals with a nylon loop into a reaction buffer containing 0.1 M MES (pH 6.8–7.2), 1 mM MgCl2, 20% (w/v) PEG2K-MME and 1 mM DTT. After incubation for a desired time point (40 s and intervals of ∼40 s), the crystals were quickly dipped in a cryo-solution supplemented with 20% (w/v) glycerol and flash-cooled in liquid nitrogen. For in crystallo reactions, MES buffer was titrated by KOH and not NaOH to avoid binding of Na+ ion at the A site. To observe product release, TG crystals were soaked in 5 µM dATP and 5 µM MgCl2 for 30 min and chased with addition of 1 mM MgCl2 for 1–8 min.
Data collection and structure determination
Diffraction data were collected at 100 K on beam lines 22ID and 22BM of the advanced photon source. Data were processed with HKL200051 or XDS52, and converted to structure factors by TRUNCATE53. All data were isomorphic in the P61 space group and processed with identical unit-cell parameters a = b = 98.8 Å and c = 82.4 Å for structure and electron-density comparison. All structural figures were drawn using PyMOL (http://www.pymol.org).
To monitor the new bond formation (Fig. 2a), Fo − Fc maps are calculated with PHENIX54 using the Fo of 40 s, 80 s, 140 s and 230 s after addition of 1 mM Mg2+ (pH 7.0) and the Fc of the refined reactant-state structure (40 s) with the 3′-OH of the primer end, the A-site Mg2+ and the α-phosphate of dATP omitted. To determine the reaction time courses (Fig. 2b and Supplementary Figs 1–3), all data were scaled against and compared to the refined ground-state structure, and electron densities for the new bond (between the 3′-OH of the primer strand and the α-phosphorus of dATP) and the metal ions were measured in the Fo − Fc map (with the B-site metal ion omitted) at each time point using programs in CCP453. After refinement of the ground-state structure, the intermediate structures were refined as a mixture of reactant state and product state using PHENIX54 and Coot55. The ratio of reactant state and product state at each time point was determined according to the time course of the new bond formation (Fig. 2b) and difference (Fo − Fc) in electron-density maps. For high-resolution refinement (∼1.5 Å resolution), the translation–libration–screw refinement was applied. Data collection and refinement statistics are summarized in Supplementary Table 1.
Kinetic measurements
Kinetic measurements including Km and kcat were done using the template/primer shown in Supplementary Table 3. The basic reaction mixture contained 2.5 nM Pol η, 5 μM 5′-fluorescein-labelled primer and template, and 0–80 µM dNTP in 40 mM Tris–HCl (pH 7.5), 5 mM MgCl2, 10 mM dithiothreitol, 100 mM KCl, 0.1 mg ml−1 bovine serum albumin and 5% glycerol. For the metal-ion-competition assay, Mg2+ was increased from 0 to 1.0 mM in 0.1-mM steps and Ca2+ was decreased to keep the combined concentration of 1.0 mM. The effects of pH were screened from pH 6.0 to pH 8.0. All reactions took place at room temperature for 4 or 8 min and were stopped by addition of 10× formamide loading buffer to the final concentrations of 8% formamide, 1 mM EDTA (pH 8.0) and 0.1 mg ml−1 xylene cyanol. After heating to 90 °C for 3 min and immediately placing on ice, products were resolved on 20% polyacrylamide sequencing gels containing 5.5 M urea. Quantification and curve fitting were carried out as described20. Km, kcat and (kcat/Km) of wild-type and S113A-mutant Pol η catalysing single-nucleotide incorporation using deoxyribonucleotide versus ribonucleotide primer are summarized in Supplementary Table 3 and plotted in Fig. 3b.
References
Echols, H. & Goodman, M. F. Fidelity mechanisms in DNA replication. Annu. Rev. Biochem. 60, 477–511 (1991)
Rothwell, P. J. & Waksman, G. Structure and mechanism of DNA polymerases. Adv. Protein Chem. 71, 401–440 (2005)
Brody, R. S. & Frey, P. A. Unambiguous determination of the stereochemistry of nucleotidyl transfer catalyzed by DNA polymerase I from Escherichia coli . Biochemistry 20, 1245–1252 (1981)
Steitz, T. A. & Steitz, J. A. A general two-metal-ion mechanism for catalytic RNA. Proc. Natl Acad. Sci. USA 90, 6498–6502 (1993)
Yang, W., Lee, J. Y. & Nowotny, M. Making and breaking nucleic acids: two-Mg2+-ion catalysis and substrate specificity. Mol. Cell 22, 5–13 (2006)
Johnson, K. A. Role of induced fit in enzyme specificity: a molecular forward/reverse switch. J. Biol. Chem. 283, 26297–26301 (2008)
Warshel, A. et al. Electrostatic basis for enzyme catalysis. Chem. Rev. 106, 3210–3235 (2006)
Pelletier, H., Sawaya, M. R., Kumar, A., Wilson, S. H. & Kraut, J. Structures of ternary complexes of rat DNA polymerase beta, a DNA template-primer, and ddCTP. Science 264, 1891–1903 (1994)
Doublie, S., Tabor, S., Long, A. M., Richardson, C. C. & Ellenberger, T. Crystal structure of a bacteriophage T7 DNA replication complex at 2.2 Å resolution. Nature 391, 251–258 (1998)
Huang, H., Chopra, R., Verdine, G. L. & Harrison, S. C. Structure of a covalently trapped catalytic complex of HIV-1 reverse transcriptase: implications for drug resistance. Science 282, 1669–1675 (1998)
Li, Y., Korolev, S. & Waksman, G. Crystal structures of open and closed forms of binary and ternary complexes of the large fragment of Thermus aquaticus DNA polymerase I: structural basis for nucleotide incorporation. EMBO J. 17, 7514–7525 (1998)
Franklin, M. C., Wang, J. & Steitz, T. A. Structure of the replicating complex of a pol α family DNA polymerase. Cell 105, 657–667 (2001)
Ling, H., Boudsocq, F., Woodgate, R. & Yang, W. Crystal structure of a Y-family DNA polymerase in action: a mechanism for error-prone and lesion-bypass replication. Cell 107, 91–102 (2001)
Berman, A. J. et al. Structures of phi29 DNA polymerase complexed with substrate: the mechanism of translocation in B-family polymerases. EMBO J. 26, 3494–3505 (2007)
Johnson, S. J., Taylor, J. S. & Beese, L. S. Processive DNA synthesis observed in a polymerase crystal suggests a mechanism for the prevention of frameshift mutations. Proc. Natl Acad. Sci. USA 100, 3895–3900 (2003)
Evans, R. J. et al. Structure of PolC reveals unique DNA binding and fidelity determinants. Proc. Natl Acad. Sci. USA 105, 20695–20700 (2008)
Wang, F. & Yang, W. Structural insight into translesion synthesis by DNA Pol II. Cell 139, 1279–1289 (2009)
Batra, V. K. et al. Magnesium-induced assembly of a complete DNA polymerase catalytic complex. Structure 14, 757–766 (2006)
Patel, S. S., Wong, I. & Johnson, K. A. Pre-steady-state kinetic analysis of processive DNA replication including complete characterization of an exonuclease-deficient mutant. Biochemistry 30, 511–525 (1991)
Joyce, C. M. & Benkovic, S. J. DNA polymerase fidelity: kinetics, structure, and checkpoints. Biochemistry 43, 14317–14324 (2004)
Showalter, A. K. & Tsai, M. D. A reexamination of the nucleotide incorporation fidelity of DNA polymerases. Biochemistry 41, 10571–10576 (2002)
Shah, A. M., Li, S. X., Anderson, K. S. & Sweasy, J. B. Y265H mutator mutant of DNA polymerase β. Proper teometric alignment is critical for fidelity. J. Biol. Chem. 276, 10824–10831 (2001)
Fiala, K. A. & Suo, Z. Mechanism of DNA polymerization catalyzed by Sulfolobus solfataricus P2 DNA polymerase IV. Biochemistry 43, 2116–2125 (2004)
Cramer, J. & Restle, T. Pre-steady-state kinetic characterization of the DinB homologue DNA polymerase of Sulfolobus solfataricus . J. Biol. Chem. 280, 40552–40558 (2005)
Choi, J. Y. & Guengerich, F. P. Adduct size limits efficient and error-free bypass across bulky N2-guanine DNA lesions by human DNA polymerase η. J. Mol. Biol. 352, 72–90 (2005)
Rothwell, P. J., Mitaksov, V. & Waksman, G. Motions of the fingers subdomain of klentaq1 are fast and not rate limiting: implications for the molecular basis of fidelity in DNA polymerases. Mol. Cell 19, 345–355 (2005)
Zhang, H., Cao, W., Zakharova, E., Konigsberg, W. & De La Cruz, E. M. Fluorescence of 2-aminopurine reveals rapid conformational changes in the RB69 DNA polymerase-primer/template complexes upon binding and incorporation of matched deoxynucleoside triphosphates. Nucleic Acids Res. 35, 6052–6062 (2007)
Yang, W. & Woodgate, R. What a difference a decade makes: insights into translesion DNA synthesis. Proc. Natl Acad. Sci. USA 104, 15591–15598 (2007)
Biertümpfel, C. et al. Structure and mechanism of human DNA polymerase η. Nature 465, 1044–1048 (2010)
Zhao, Y., Biertümpfel, C., Gregory, M. T., Hua, Y. J., Hanaoka, F. & Wang, W. Structural basis of human DNA polymerase η-mediated chemoresistance to cisplatin. Proc. Natl Acad. Sci. USA 109, 7269–7274 (2012)
Castro, C. et al. Two proton transfers in the transition state for nucleotidyl transfer catalyzed by RNA- and DNA-dependent RNA and DNA polymerases. Proc. Natl Acad. Sci. USA 104, 4267–4272 (2007)
Garcia-Diaz, M., Bebenek, K., Krahn, J. M., Pedersen, L. C. & Kunkel, T. A. Role of the catalytic metal during polymerization by DNA polymerase lambda. DNA Repair 6, 1333–1340 (2007)
Le Du, M. H. et al. Artificial evolution of an enzyme active site: structural studies of three highly active mutants of Escherichia coli alkaline phosphatase. J. Mol. Biol. 316, 941–953 (2002)
Graham, D. L. et al. MgF3 − as a transition state analog of phosphoryl transfer. Chem. Biol. 9, 375–381 (2002)
Lee, J. Y. & Yang, W. UvrD helicase unwinds DNA one base pair at a time by a two-part power stroke. Cell 127, 1349–1360 (2006)
Cavanaugh, N. A., Beard, W. A. & Wilson, S. H. DNA polymerase β ribonucleotide discrimination: insertion, misinsertion, extension, and coding. J. Biol. Chem. 285, 24457–24465 (2010)
Wang, L., Broyde, S. & Zhang, Y. Polymerase-tailored variations in the water-mediated and substrate-assisted mechanism for nucleotidyl transfer: insights from a study of T7 DNA polymerase. J. Mol. Biol. 389, 787–796 (2009)
Wang, L., Yu, X., Hu, P., Broyde, S. & Zhang, Y. A water-mediated and substrate-assisted catalytic mechanism for Sulfolobus solfataricus DNA polymerase IV. J. Am. Chem. Soc. 129, 4731–4737 (2007)
Polesky, A. H., Dahlberg, M. E., Benkovic, S. J., Grindley, N. D. & Joyce, C. M. Side chains involved in catalysis of the polymerase reaction of DNA polymerase I from Escherichia coli . J. Biol. Chem. 267, 8417–8428 (1992)
Copeland, W. C. & Wang, T. S. Mutational analysis of the human DNA polymerase α. The most conserved region in α-like DNA polymerases is involved in metal-specific catalysis. J. Biol. Chem. 268, 11028–11040 (1993)
Zakharova, E., Wang, J. & Konigsberg, W. The activity of selected RB69 DNA polymerase mutants can be restored by manganese ions: the existence of alternative metal ion ligands used during the polymerization cycle. Biochemistry 43, 6587–6595 (2004)
Kim, E. E. & Wyckoff, H. W. Reaction mechanism of alkaline phosphatase based on crystal structures. Two-metal ion catalysis. J. Mol. Biol. 218, 449–464 (1991)
Romier, C., Dominguez, R., Lahm, A., Dahl, O. & Suck, D. Recognition of single-stranded DNA by nuclease P1: high resolution crystal structures of complexes with substrate analogs. Proteins 32, 414–424 (1998)
Garcin, E. D. et al. DNA apurinic–apyrimidinic site binding and excision by endonuclease IV. Nature Struct. Mol. Biol. 15, 515–522 (2008)
Shan, S., Yoshida, A., Sun, S., Piccirilli, J. A. & Herschlag, D. Three metal ions at the active site of the Tetrahymena group I ribozyme. Proc. Natl Acad. Sci. USA 96, 12299–12304 (1999)
Stahley, M. R. & Strobel, S. A. Structural evidence for a two-metal-ion mechanism of group I intron splicing. Science 309, 1587–1590 (2005)
Ding, X., Rasmussen, B. F., Petsko, G. A. & Ringe, D. Direct structural observation of an acyl-enzyme intermediate in the hydrolysis of an ester substrate by elastase. Biochemistry 33, 9285–9293 (1994)
Scott, W. G., Murray, J. B., Arnold, J. R., Stoddard, B. L. & Klug, A. Capturing the structure of a catalytic RNA intermediate: the hammerhead ribozyme. Science 274, 2065–2069 (1996)
Murray, J. B., Szoke, H., Szoke, A. & Scott, W. G. Capture and visualization of a catalytic RNA enzyme–product complex using crystal lattice trapping and X-ray holographic reconstruction. Mol. Cell 5, 279–287 (2000)
Yang, W. Nucleases: diversity of structure, function and mechanism. Q. Rev. Biophys. 44, 1–93 (2011)
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997)
Kabsch, W. XDS. Acta Crystallogr. D 66, 125–132 (2010)
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010)
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot . Acta Crystallogr. D 66, 486–501 (2010)
Acknowledgements
We thank D. Leahy, M. Gellert and R. Craigie for editing the manuscript. The research was supported by the intramural research program of the National Institute of Diabetes and Digestive and Kidney Diseases, National Institutes of Health (W.Y., T.N. and Y.Z.); the Japan Society for the Promotion of Science Institutional Program for Young Researcher Overseas visits, Kumamoto University, and the Kumayaku Alumni Research Fund (T.N.); Chinese Ministry of Education scholarship (Y.Z.); National Natural Science Foundation of China (Y.-J.H.); and Grant-in-Aid for Scientific Research from the Ministry of Education, Culture, Sports, Science, and Technology of Japan (Y.Y.).
Author information
Authors and Affiliations
Contributions
T.N. and Y.Z. carried out crystallography. Y.Z. did the mutagenesis and kinetic analyses. T.N. handled crystals, refined structures and analysed reaction time courses. W.Y. designed the project. T.N., Y.Z. and W.Y. prepared the manuscript. Y.Y. supervised T.N. and Y.-J.H. advised Y.Z. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Tables 1-3 and Supplementary Figures 1-5. (PDF 2963 kb)
Supplementary Movie 1
This file contains an animation of the chemical bond formation. (MOV 15373 kb)
Rights and permissions
About this article
Cite this article
Nakamura, T., Zhao, Y., Yamagata, Y. et al. Watching DNA polymerase η make a phosphodiester bond. Nature 487, 196–201 (2012). https://doi.org/10.1038/nature11181
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature11181
This article is cited by
-
Monovalent metal ion binding promotes the first transesterification reaction in the spliceosome
Nature Communications (2023)
-
Watching right and wrong nucleotide insertion captures hidden polymerase fidelity checkpoints
Nature Communications (2022)
-
In crystallo observation of three metal ion promoted DNA polymerase misincorporation
Nature Communications (2022)
-
Watching a double strand break repair polymerase insert a pro-mutagenic oxidized nucleotide
Nature Communications (2021)
-
Structural basis of DNA synthesis opposite 8-oxoguanine by human PrimPol primase-polymerase
Nature Communications (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.