Abstract
Arising from Y. Liu, J. Slotine & A. Barabási Nature 473, 167–173 (2011)10.1038/nature10011; Liu et al. reply
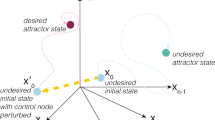
Liu, Slotine and Barabasi1 identify subsets U of nodes in complex networks, which are required to exert full control of these networks. Control in this context means that for each possible state S of the network there exist inputs for all nodes in U, which are sufficient to force the network to state S1. Application of the methodology to gene regulatory networks suggests that roughly 80% of all nodes must be controlled to drive such a network. This seems to contradict recent empirical findings2,3,4,5,6 in the cellular reprogramming field.
Similar content being viewed by others
Main
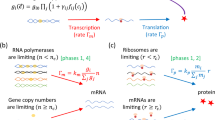
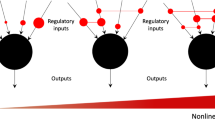
Mammalian cells have been shown to be efficiently reprogrammable towards other cellular phenotypes2,3,4,5,6. Controllability of such complex transitions in transcriptional networks underlying cellular phenotypes appears to be an intrinsic biological property and is being used for the development of novel disease models and cellular therapeutics in regenerative medicine. However, in contrast to technical or social networks, biological networks show a high degree of intrinsic co-regulation. Co-regulation appears to quench the admissible space of states of gene expression networks to a combinatorial expression space with relatively low dimensionality. Recent publications suggest that this subspace is spanned by only a few (five to fifteen) genome-wide differential expression patterns, which are—surprisingly—sufficient to characterize most observable biological phenotypes7,8,9. Each of these patterns is given by weighted sums over the expression of relatively extensive gene sets contributing coherently to the respective pattern7,8,9. Although these empirical studies suggest linear structures underlying the respective subspaces, nonlinear or star-shaped topologies cannot be excluded.
Moreover, genome-wide co-regulation may result in dynamic response features, as observed in chemical networks. Hence, owing to their low dimensionality, full control and reprogramming of biological systems may be achieved by only a few key control factors, seemingly contradicting the more general concept put forward by ref. 1. Recent experimental studies show proof of concept for a wide range of reprogramming of biological systems using overexpression of only one to five transcription factors2,3,4,5,6,10, which effectively regulate only a subset of downstream genes. We note that the number of nodes (five or fewer genes out of about 30,000) needed to fully control a biological system2,3,4,5,6,10 is much less than the estimate of 80% of all nodes1.
Thus, for the special case of biological systems, it may be sufficient to weaken the standard definition of controllability of networks—control of large sets of nodes as suggested by ref. 1—to mean the controllability of restricted, biologically admissible network states.
References
Liu, Y., Slotine, J. & Barabási, A. Controllability of complex networks. Nature 473, 167–173 (2011)
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676 (2006)
Vierbuchen, T. et al. Direct conversion of fibroblasts to functional neurons by defined factors. Nature 463, 1035–1041 (2010)
Ieda, M. et al. Direct reprogramming of fibroblasts into functional cardiomyocytes by defined factors. Cell 142, 375–386 (2010)
Szabo, E. et al. Direct conversion of human fibroblasts to multilineage blood progenitors. Nature 468, 521–526 (2010)
Huang, P. et al. Induction of functional hepatocyte-like cells from mouse fibroblasts by defined factors. Nature 475, 386–389 (2011)
Lukk, M. et al. A global map of human gene expression. Nature Biotechnol. 28, 322–324 (2010)
Müller, F. et al. A bioinformatic assay for pluripotency in human cells. Nature Methods 8, 315–317 (2011)
Dudley, J. T., Tibshirani, R., Deshpande, T. & Butte, A. J. Disease signatures are robust across tissues and experiments. Mol. Syst. Biol. 5 307 10.1038/msb.2009.66 (2009)
Kim, J. B. et al. Oct4-induced pluripotency in adult neural stem cells. Cell 136, 411–419 (2009)
Author information
Authors and Affiliations
Contributions
F.J.M. and A.S. conceptualized, researched and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
Competing financial interests: declared none.
Rights and permissions
About this article
Cite this article
Müller, FJ., Schuppert, A. Few inputs can reprogram biological networks. Nature 478, E4 (2011). https://doi.org/10.1038/nature10543
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10543
This article is cited by
-
The nonlinearity of regulation in biological networks
npj Systems Biology and Applications (2023)
-
NETISCE: a network-based tool for cell fate reprogramming
npj Systems Biology and Applications (2022)
-
On the effects of memory and topology on the controllability of complex dynamical networks
Scientific Reports (2020)
-
Controllability and observability in complex networks – the effect of connection types
Scientific Reports (2017)
-
Energy scaling of targeted optimal control of complex networks
Nature Communications (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.