Abstract
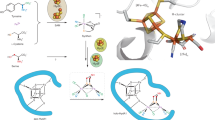
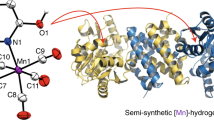
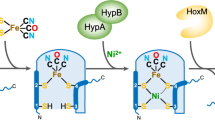
Hydrogenases are abundant enzymes that catalyse the reversible interconversion of H2 into protons and electrons at high rates1. Those hydrogenases maintaining their activity in the presence of O2 are considered to be central to H2-based technologies, such as enzymatic fuel cells and for light-driven H2 production2. Despite comprehensive genetic, biochemical, electrochemical and spectroscopic investigations3,4,5,6,7,8, the molecular background allowing a structural interpretation of how the catalytic centre is protected from irreversible inactivation by O2 has remained unclear. Here we present the crystal structure of an O2-tolerant [NiFe]-hydrogenase from the aerobic H2 oxidizer Ralstonia eutropha H16 at 1.5 Å resolution. The heterodimeric enzyme consists of a large subunit harbouring the catalytic centre in the H2-reduced state and a small subunit containing an electron relay consisting of three different iron-sulphur clusters. The cluster proximal to the active site displays an unprecedented [4Fe-3S] structure and is coordinated by six cysteines. According to the current model, this cofactor operates as an electronic switch depending on the nature of the gas molecule approaching the active site. It serves as an electron acceptor in the course of H2 oxidation and as an electron-delivering device upon O2 attack at the active site. This dual function is supported by the capability of the novel iron-sulphur cluster to adopt three redox states at physiological redox potentials7,8,9. The second structural feature is a network of extended water cavities that may act as a channel facilitating the removal of water produced at the [NiFe] active site. These discoveries will have an impact on the design of biological and chemical H2-converting catalysts that are capable of cycling H2 in air.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Cammack, R., Frey, M. & Robson, R. Hydrogen as a Fuel, Learning from Nature (Taylor & Francis, 2001)
Friedrich, B., Fritsch, J. & Lenz, O. Oxygen-tolerant hydrogenases in hydrogen-based technologies. Curr. Opin. Biotechnol. 22, 358–364 (2011)
Ludwig, M., Cracknell, J. A., Vincent, K. A., Armstrong, F. A. & Lenz, O. Oxygen-tolerant H2 oxidation by membrane-bound [NiFe] hydrogenases of Ralstonia species. Coping with low level H2 in air. J. Biol. Chem. 284, 465–477 (2009)
Saggu, M. et al. Spectroscopic insights into the oxygen-tolerant membrane-associated [NiFe] hydrogenase of Ralstonia eutropha H16. J. Biol. Chem. 284, 16264–16276 (2009)
Cracknell, J. A., Wait, A. F., Lenz, O., Friedrich, B. & Armstrong, F. A. A kinetic and thermodynamic understanding of O2 tolerance in [NiFe]-hydrogenases. Proc. Natl Acad. Sci. USA 106, 20681–20686 (2009)
Lukey, M. J. et al. How Escherichia coli is equipped to oxidize hydrogen under different redox conditions. J. Biol. Chem. 285, 3928–3938 (2010)
Goris, T. et al. A unique iron-sulfur cluster is crucial for oxygen tolerance of a [NiFe]-hydrogenase. Nature Chem. Biol. 7, 310–318 (2011)
Pandelia, M. E. et al. Characterization of a unique [FeS] cluster in the electron transfer chain of the oxygen tolerant [NiFe] hydrogenase from Aquifex aeolicus. Proc. Natl Acad. Sci. USA 108, 6097–6102 (2011)
Schneider, K., Patil, D. S. & Cammack, R. Electron-spin-resonance properties of membrane-bound hydrogenases from aerobic hydrogen bacteria. Biochim. Biophys. Acta 748, 353–361 (1983)
Vignais, P. M. & Billoud, B. Occurrence, classification, and biological function of hydrogenases: an overview. Chem. Rev. 107, 4206–4272 (2007)
Stripp, S. T. et al. How oxygen attacks [FeFe] hydrogenases from photosynthetic organisms. Proc. Natl Acad. Sci. USA 106, 17331–17336 (2009)
Ogata, H., Lubitz, W. & Higuchi, Y. [NiFe] hydrogenases: structural and spectroscopic studies of the reaction mechanism. Dalton Trans. 7577–7587 (2009)
Vincent, K. A., Parkin, A. & Armstrong, F. A. Investigating and exploiting the electrocatalytic properties of hydrogenases. Chem. Rev. 107, 4366–4413 (2007)
De Lacey, A. L., Fernandez, V. M., Rousset, M. & Cammack, R. Activation and inactivation of hydrogenase function and the catalytic cycle: spectroelectrochemical studies. Chem. Rev. 107, 4304–4330 (2007)
Schwartz, E. & Friedrich, B. in The Prokaryotes (eds Dworkin, M. et al.) 496–563 (Springer, 2006)
Lenz, O. et al. H2 conversion in the presence of O2 as performed by the membrane-bound [NiFe]-hydrogenase of Ralstonia eutropha. ChemPhysChem 11, 1107–1119 (2010)
Volbeda, A. et al. Crystal structure of the nickel-iron hydrogenase from Desulfovibrio gigas. Nature 373, 580–587 (1995)
Volbeda, A. et al. Structural differences between the ready and unready oxidized states of [NiFe] hydrogenases. J. Biol. Inorg. Chem. 10, 239–249 (2005)
Matias, P. M. et al. [NiFe] hydrogenase from Desulfovibrio desulfuricans ATCC 27774: gene sequencing, three-dimensional structure determination and refinement at 1.8 Å and modelling studies of its interaction with the tetrahaem cytochrome c3 . J. Biol. Inorg. Chem. 6, 63–81 (2001)
Ogata, H., Kellers, P. & Lubitz, W. The crystal structure of the [NiFe] hydrogenase from the photosynthetic bacterium Allochromatium vinosum: characterization of the oxidized enzyme (Ni-A state). J. Mol. Biol. 402, 428–444 (2010)
Seefeldt, L. C., Hoffman, B. M. & Dean, D. R. Mechanism of Mo-dependent nitrogenase. Annu. Rev. Biochem. 78, 701–722 (2009)
Peters, J. W. et al. Redox-dependent structural changes in the nitrogenase P-cluster. Biochemistry 36, 1181–1187 (1997)
Knüttel, K. et al. Redox properties of the metal centers in the membrane-bound hydrogenase from Alcaligens eurtophus CH34. Bull. Polish Acad. Sci. 42, 495–511 (1994)
Cammack, R. “Super-reduction” of chromatium high-potential iron-sulphur protein in the presence of dimethyl sulphoxide. Biochem. Biophys. Res. Commun. 54, 548–554 (1973)
Thomson, A. J. et al. Low-temperature magnetic circular-dichroism evidence for the conversion of 4-iron-sulfur clusters in a ferredoxin from Clostridium pasteurianum into 3-iron-sulfur clusters. Biochim. Biophys. Acta 637, 423–432 (1981)
Capozzi, F., Ciurli, S. & Luchinat, C. Coordination sphere versus protein environment as determinants of electronic and functional properties of iron-sulfur proteins. Metal Sites Proteins Models 90, 127–160 (1998)
Carter, C. W., Jr New stereochemical analogies between iron-sulfur electron transport proteins. J. Biol. Chem. 252, 7802–7811 (1977)
Dey, A. et al. Solvent tuning of electrochemical potentials in the active sites of HiPIP versus ferredoxin. Science 318, 1464–1468 (2007)
Heering, H. A., Bulsink, B. M., Hagen, W. R. & Meyer, T. E. Influence of charge and polarity on the redox potentials of high-potential iron-sulfur proteins: evidence for the existence of two groups. Biochemistry 34, 14675–14686 (1995)
Montet, Y. et al. Gas access to the active site of Ni-Fe hydrogenases probed by X-ray crystallography and molecular dynamics. Nature Struct. Biol. 4, 523–526 (1997)
Schubert, T., Lenz, O., Krause, E., Volkmer, R. & Friedrich, B. Chaperones specific for the membrane-bound [NiFe]-hydrogenase interact with the Tat signal peptide of the small subunit precursor in Ralstonia eutropha H16. Mol. Microbiol. 66, 453–467 (2007)
Schink, B. & Schlegel, H. G. The membrane-bound hydrogenase of Alcaligenes eutrophus. I. Solubilization, purification, and biochemical properties. Biochim. Biophys. Acta 567, 315–324 (1979)
Jancarik, J. & Kim, S.-H. Sparse matrix sampling: a screening method for crystallization of proteins. J. Appl. Cryst. 24, 409–411 (1991)
Flot, D. et al. The ID23-2 structural biology microfocus beamline at the ESRF. J. Synchrotron Radiat. 17, 107–118 (2010)
Kabsch, W. Xds. Acta Crystallogr. D 66, 125–132 (2010)
Evans, P. Scaling and assessment of data quality. Acta Crystallogr. D 62, 72–82 (2006)
Collaborative Computational Project, Number 4 . The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994)
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Cryst. 40, 658–674 (2007)
Brünger, A. T. et al. Crystallography & NMR system: A new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998)
Vagin, A. A. et al. REFMAC5 dictionary: organization of prior chemical knowledge and guidelines for its use. Acta Crystallogr. D 60, 2184–2195 (2004)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
Laskowski, R. A., MacArthur, M. W., Moss, D. S. & Thornton, J. M. PROCHECK: A program to check the stereochemical quality of protein structures. Appl. Cryst. 26, 283–291 (1993)
Hooft, R. W., Vriend, G., Sander, C. & Abola, E. E. Errors in protein structures. Nature 381, 272 (1996)
McDonald, I. K. & Thornton, J. M. Satisfying hydrogen bonding potential in proteins. J. Mol. Biol. 238, 777–793 (1994)
Wallace, A. C., Laskowski, R. A. & Thornton, J. M. LIGPLOT: a program to generate schematic diagrams of protein-ligand interactions. Protein Eng. 8, 127–134 (1995)
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007)
DeLano, W. L. The PyMOL Molecular Graphics System 〈http://www.pymol.org〉 (2002)
Acknowledgements
We are grateful to U. Müller, M. Weiss and the scientific staff of the BESSY-MX/Helmholtz Zentrum Berlin für Materialien und Energie at beamlines BL 14.1 and BL 14.2, D. von Stetten and A. Royant of the ID29S-Cryobench (ESRF, Grenoble) and the European Synchrotron Radiation Facility (ESRF, Grenoble) at beamlines ID23-1, ID23-2, ID14-1 and ID 14-4, where the data were collected, for support. This work was supported by the EU/FP7 programme Solar-H2 (to J.F.), the DFG Cluster of Excellence ‘Unifying Concepts in Catalysis’ (to S.F., B.F., O.L.), and the Sfb740 (to C.M.T.S.). P.S. acknowledges K. P. Hofmann and his advanced investigator ERC grant (ERC-2009/249910—TUDOR) for support.
Author information
Authors and Affiliations
Contributions
J.F. and P.S. are joint first authors. J.F. optimized cell growth conditions as well as the MBH purification procedure. P.S. conducted the crystallization screening; P.S., J.F. and S.F. optimized MBH crystallization conditions. P.S. and S.K. collected the X-ray diffraction data. P.S. performed data processing, solved and refined the MBH structure. B.F., O.L. and P.S. coordinated the project. J.F., P.S., S.F. and O.L. analysed data. J.F., P.S., S.F., B.F., O.L. and C.M.T.S. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
The file contains Supplementary Figures 1-9 with legends, Supplementary Table 1 and additional references. (PDF 10591 kb)
Rights and permissions
About this article
Cite this article
Fritsch, J., Scheerer, P., Frielingsdorf, S. et al. The crystal structure of an oxygen-tolerant hydrogenase uncovers a novel iron-sulphur centre. Nature 479, 249–252 (2011). https://doi.org/10.1038/nature10505
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10505
This article is cited by
-
Stepwise assembly of the active site of [NiFe]-hydrogenase
Nature Chemical Biology (2023)
-
A personal account on 25 years of scientific literature on [FeFe]-hydrogenase
JBIC Journal of Biological Inorganic Chemistry (2023)
-
O2-tolerant CO dehydrogenase via tunnel redesign for the removal of CO from industrial flue gas
Nature Catalysis (2022)
-
Microbial oxidation of atmospheric trace gases
Nature Reviews Microbiology (2022)
-
Aerobic hydrogen-oxidizing bacteria in soil: from cells to ecosystems
Reviews in Environmental Science and Bio/Technology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.