Abstract
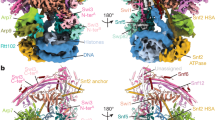
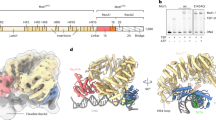
Swi2/Snf2-type ATPases regulate genome-associated processes such as transcription, replication and repair by catalysing the disruption, assembly or remodelling of nucleosomes or other protein–DNA complexes1,2. It has been suggested that ATP-driven motor activity along DNA disrupts target protein–DNA interactions in the remodelling reaction3,4,5. However, the complex and highly specific remodelling reactions are poorly understood, mostly because of a lack of high-resolution structural information about how remodellers bind to their substrate proteins. Mot1 (modifier of transcription 1 in Saccharomyces cerevisiae, denoted BTAF1 in humans) is a Swi2/Snf2 enzyme that specifically displaces the TATA box binding protein (TBP) from the promoter DNA and regulates transcription globally by generating a highly dynamic TBP pool in the cell6,7. As a Swi2/Snf2 enzyme that functions as a single polypeptide and interacts with a relatively simple substrate, Mot1 offers an ideal system from which to gain a better understanding of this important enzyme family. To reveal how Mot1 specifically disrupts TBP–DNA complexes, we combined crystal and electron microscopy structures of Mot1–TBP from Encephalitozoon cuniculi with biochemical studies. Here we show that Mot1 wraps around TBP and seems to act like a bottle opener: a spring-like array of 16 HEAT (huntingtin, elongation factor 3, protein phosphatase 2A and lipid kinase TOR) repeats grips the DNA-distal side of TBP via loop insertions, and the Swi2/Snf2 domain binds to upstream DNA, positioned to weaken the TBP–DNA interaction by DNA translocation. A ‘latch’ subsequently blocks the DNA-binding groove of TBP, acting as a chaperone to prevent DNA re-association and ensure efficient promoter clearance. This work shows how a remodelling enzyme can combine both motor and chaperone activities to achieve functional specificity using a conserved Swi2/Snf2 translocase.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Cairns, B. R. The logic of chromatin architecture and remodelling at promoters. Nature 461, 193–198 (2009)
Li, B., Carey, M. & Workman, J. L. The role of chromatin during transcription. Cell 128, 707–719 (2007)
Dürr, H., Korner, C., Muller, M., Hickmann, V. & Hopfner, K. P. X-ray structures of the Sulfolobus solfataricus SWI2/SNF2 ATPase core and its complex with DNA. Cell 121, 363–373 (2005)
Saha, A., Wittmeyer, J. & Cairns, B. R. Chromatin remodeling by RSC involves ATP-dependent DNA translocation. Genes Dev. 16, 2120–2134 (2002)
Racki, L. R. et al. The chromatin remodeller ACF acts as a dimeric motor to space nucleosomes. Nature 462, 1016–1021 (2009)
Auble, D. T. The dynamic personality of TATA-binding protein. Trends Biochem. Sci. 34, 49–52 (2009)
de Graaf, P. et al. Chromatin interaction of TATA-binding protein is dynamically regulated in human cells. J. Cell Sci. 123, 2663–2671 (2010)
Darst, R. P. et al. Mot1 regulates the DNA binding activity of free TATA-binding protein in an ATP-dependent manner. J. Biol. Chem. 278, 13216–13226 (2003)
Pereira, L. A., van der Knaap, J. A., van den Boom, V., van den Heuvel, F. A. & Timmers, H. T. TAF(II)170 interacts with the concave surface of TATA-binding protein to inhibit its DNA binding activity. Mol. Cell. Biol. 21, 7523–7534 (2001)
Pugh, B. F. Control of gene expression through regulation of the TATA-binding protein. Gene 255, 1–14 (2000)
Auble, D. T. & Hahn, S. An ATP-dependent inhibitor of TBP binding to DNA. Genes Dev. 7, 844–856 (1993)
Mohibullah, N. & Hahn, S. Site-specific cross-linking of TBP in vivo and in vitro reveals a direct functional interaction with the SAGA subunit Spt3. Genes Dev. 22, 2994–3006 (2008)
Pereira, L. A. et al. Molecular architecture of the basal transcription factor B-TFIID. J. Biol. Chem. 279, 21802–21807 (2004)
Darst, R. P., Wang, D. & Auble, D. T. MOT1-catalyzed TBP–DNA disruption: uncoupling DNA conformational change and role of upstream DNA. EMBO J. 20, 2028–2040 (2001)
Sprouse, R. O., Brenowitz, M. & Auble, D. T. Snf2/Swi2-related ATPase Mot1 drives displacement of TATA-binding protein by gripping DNA. EMBO J. 25, 1492–1504 (2006)
Miller, G. & Hahn, S. A DNA-tethered cleavage probe reveals the path for promoter DNA in the yeast preinitiation complex. Nature Struct. Biol. 13, 603–610 (2006)
Gangaraju, V. K., Prasad, P., Srour, A., Kagalwala, M. N. & Bartholomew, B. Conformational changes associated with template commitment in ATP-dependent chromatin remodeling by ISW2. Mol. Cell 35, 58–69 (2009)
Dasgupta, A., Darst, R. P., Martin, K. J., Afshari, C. A. & Auble, D. T. Mot1 activates and represses transcription by direct, ATPase-dependent mechanisms. Proc. Natl Acad. Sci. USA 99, 2666–2671 (2002)
Klejman, M. P. et al. NC2α interacts with BTAF1 and stimulates its ATP-dependent association with TATA-binding protein. Mol. Cell. Biol. 24, 10072–10082 (2004)
van Werven, F. J. et al. Cooperative action of NC2 and Mot1p to regulate TATA-binding protein function across the genome. Genes Dev. 22, 2359–2369 (2008)
Hsu, J. Y. et al. TBP, Mot1, and NC2 establish a regulatory circuit that controls DPE-dependent versus TATA-dependent transcription. Genes Dev. 22, 2353–2358 (2008)
Geisberg, J. V. & Struhl, K. Cellular stress alters the transcriptional properties of promoter-bound Mot1-TBP complexes. Mol. Cell 14, 479–489 (2004)
Kostrewa, D. et al. RNA polymerase II–TFIIB structure and mechanism of transcription initiation. Nature 462, 323–330 (2009)
Liu, X., Bushnell, D. A., Wang, D., Calero, G. & Kornberg, R. D. Structure of an RNA polymerase II–TFIIB complex and the transcription initiation mechanism. Science 327, 206–209 (2010)
Schluesche, P., Stelzer, G., Piaia, E., Lamb, D. C. & Meisterernst, M. NC2 mobilizes TBP on core promoter TATA boxes. Nature Struct. Biol. 14, 1196–1201 (2007)
Timmers, H. T., Meyers, R. E. & Sharp, P. A. Composition of transcription factor B-TFIID. Proc. Natl Acad. Sci. USA 89, 8140–8144 (1992)
Poon, D., Campbell, A. M., Bai, Y. & Weil, P. A. Yeast Taf170 is encoded by MOT1 and exists in a TATA box-binding protein (TBP)-TBP-associated factor complex distinct from transcription factor IID. J. Biol. Chem. 269, 23135–23140 (1994)
Gkikopoulos, T., Havas, K. M., Dewar, H. & Owen-Hughes, T. SWI/SNF and Asf1p cooperate to displace histones during induction of the Saccharomyces cerevisiae HO promoter. Mol. Cell. Biol. 29, 4057–4066 (2009)
Kim, Y., Geiger, J. H., Hahn, S. & Sigler, P. B. Crystal structure of a yeast TBP/TATA-box complex. Nature 365, 512–520 (1993)
Auble, D. T., Wang, D., Post, K. W. & Hahn, S. Molecular analysis of the SNF2/SWI2 protein family member MOT1, an ATP-driven enzyme that dissociates TATA-binding protein from DNA. Mol. Cell. Biol. 17, 4842–4851 (1997)
Berger, I., Fitzgerald, D. J. & Richmond, T. J. Baculovirus expression system for heterologous multiprotein complexes. Nature Biotechnol. 22, 1583–1587 (2004)
Kagawa, R., Montgomery, M. G., Braig, K., Leslie, A. G. & Walker, J. E. The structure of bovine F1-ATPase inhibited by ADP and beryllium fluoride. EMBO J. 23, 2734–2744 (2004)
Kabsch, W. Automatic processing of rotation diffraction data from crystals of initially unknown symmetry and cell constants. J. Appl. Cryst. 26, 795–800 (1993)
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Cryst. 40, 658–674 (2007)
Bricogne, G., Vonrhein, C., Flensburg, C., Schiltz, M. & Paciorek, W. Generation, representation and flow of phase information in structure determination: recent developments in and around SHARP 2.0. Acta Crystallogr. D 59, 2023–2030 (2003)
Cowtan, K. The Buccaneer software for automated model building. 1. Tracing protein chains. Acta Crystallogr. D 62, 1002–1011 (2006)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
Adams, P. D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. D 58, 1948–1954 (2002)
DeLano, W. L. The PyMOL Molecular Graphics System Version 1.3 r1 (Schrödinger, 2010).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004)
Sprouse, R. O., Brenowitz, M. & Auble, D. T. Snf2/Swi2-related ATPase Mot1 drives displacement of TATA-binding protein by gripping DNA. EMBO J. 25, 1492–1504 (2006)
Darst, R. P., Wang, D. & Auble, D. T. MOT1-catalyzed TBP-DNA disruption: uncoupling DNA conformational change and role of upstream DNA. EMBO J. 20, 2028–2040 (2001)
Chen, H. T. & Hahn, S. Binding of TFIIB to RNA polymerase II: Mapping the binding site for the TFIIB zinc ribbon domain within the preinitiation complex. Mol. Cell 12, 437–447 (2003)
Miller, G. & Hahn, S. A DNA-tethered cleavage probe reveals the path for promoter DNA in the yeast preinitiation complex. Nature Struct. Mol. Biol. 13, 603–610 (2006)
Auble, D. T., Wang, D., Post, K. W. & Hahn, S. Molecular analysis of the SNF2/SWI2 protein family member MOT1, an ATP-driven enzyme that dissociates TATA-binding protein from DNA. Mol. Cell. Biol. 17, 4842–4851 (1997)
Ludtke, S. J., Baldwin, P. R. & Chiu, W. EMAN: semiautomated software for high-resolution single-particle reconstructions. J. Struct. Biol. 128, 82–97 (1999)
van Heel, M., Harauz, G., Orlova, E. V., Schmidt, R. & Schatz, M. A new generation of the IMAGIC image processing system. J. Struct. Biol. 116, 17–24 (1996)
Trabuco, L. G., Villa, E., Mitra, K., Frank, J. & Schulten, K. Flexible fitting of atomic structures into electron microscopy maps using molecular dynamics. Structure 16, 673–683 (2008)
Acknowledgements
We thank the Max-Planck Crystallization Facility Martinsried. We thank M. Lucas, A. Schele, C. Ungewickell, J. Goetzl and Y. Hiruma for help with experimentation. We thank J.-P. Armache and M. Turk for help with electron microscopy data. We are grateful to G. Miller and S. Hahn for advice. We thank the staff at the SLS and ESRF for help with data collection. We thank P. Cramer and members of the Hopfner and Auble laboratories for discussions and comments on the manuscript. This work was supported by the German Research Council (SFB 646 and SFB/TR5) and Excellence Initiative (Center for Integrated Protein Science, Munich) to K.-P.H. and R.B., by DFG grant WE4628/1 to P.Wendler and by NIH grant GM55763 to D.T.A.
Author information
Authors and Affiliations
Contributions
S.C. and M.M. cloned, purified and crystallized EcTBP; S.C. solved its structure. S.C., A.B., M.M. and P.Wollmann cloned, purified and crystallized EcTBP–EcMot1(NTD); P.Wollmann collected data and P.Wollmann, G.W. and K.-P.H. solved the complex structures. R.V. performed FeBABE experiments, M.N.W. conducted yeast molecular biological manipulations and D.T.A. performed gelshifts. P.Wendler, O.B. and R.B. performed and interpreted electron microscopy experiments. P.Wollmann, P.Wendler, R.B., D.T.A. and K.-P.H. planned and interpreted the experiments. D.T.A. and K.-P.H. wrote the manuscript and all authors provided editorial input.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
The file contains Supplementary Figures 1-8 with legends, Supplementary Tables 1-3 and additional references. (PDF 6129 kb)
Rights and permissions
About this article
Cite this article
Wollmann, P., Cui, S., Viswanathan, R. et al. Structure and mechanism of the Swi2/Snf2 remodeller Mot1 in complex with its substrate TBP. Nature 475, 403–407 (2011). https://doi.org/10.1038/nature10215
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10215
This article is cited by
-
Structural convergence endows nuclear transport receptor Kap114p with a transcriptional repressor function toward TATA-binding protein
Nature Communications (2023)
-
mSWI/SNF promotes Polycomb repression both directly and through genome-wide redistribution
Nature Structural & Molecular Biology (2021)
-
Molecular determinants underlying functional innovations of TBP and their impact on transcription initiation
Nature Communications (2020)
-
Multiple direct interactions of TBP with the MYC oncoprotein
Nature Structural & Molecular Biology (2019)
-
Smarca4 ATPase mutations disrupt direct eviction of PRC1 from chromatin
Nature Genetics (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.