Abstract
Many successful vaccines induce persistent antibody responses that can last a lifetime. The mechanisms by which they do so remain unclear, but emerging evidence indicates that they activate dendritic cells via Toll-like receptors (TLRs)1,2. For example, the yellow fever vaccine YF-17D, one of the most successful empiric vaccines ever developed3, activates dendritic cells via multiple TLRs to stimulate proinflammatory cytokines4,5. Triggering specific combinations of TLRs in dendritic cells can induce synergistic production of cytokines6, which results in enhanced T-cell responses, but its impact on antibody responses remain unknown. Learning the critical parameters of innate immunity that program such antibody responses remains a major challenge in vaccinology. Here we demonstrate that immunization of mice with synthetic nanoparticles containing antigens plus ligands that signal through TLR4 and TLR7 induces synergistic increases in antigen-specific, neutralizing antibodies compared to immunization with nanoparticles containing antigens plus a single TLR ligand. Consistent with this there was enhanced persistence of germinal centres and of plasma-cell responses, which persisted in the lymph nodes for >1.5 years. Surprisingly, there was no enhancement of the early short-lived plasma-cell response relative to that observed with single TLR ligands. Molecular profiling of activated B cells, isolated 7 days after immunization, indicated that there was early programming towards B-cell memory. Antibody responses were dependent on direct triggering of both TLRs on B cells and dendritic cells, as well as on T-cell help. Immunization protected completely against lethal avian and swine influenza virus strains in mice, and induced robust immunity against pandemic H1N1 influenza in rhesus macaques.
Similar content being viewed by others
Main
We designed a nanoparticle-based vaccine, similar to a virus in size and composition. A biodegradable synthetic polymer, poly(d,l-lactic-co-glycolic acid) (PLGA)7, was used to synthesize ∼300-nM-sized nanoparticles containing the TLR ligands MPL (TLR4 ligand), R837 (TLR7 ligand), or both ligands, together with an antigen (Supplementary Fig. 1). Immunization of mice with nanoparticles containing MPL and R837 (PLGA(MPL+R837)) plus antigen induced enhanced antibody and T-cell responses, compared to immunization with soluble antigen plus MPL and R837 (data not shown). Consistent with recent observations8,9, delivery of antigen and TLR ligands in separate nanoparticles induced a stronger antibody response than delivery of both in the same nanoparticle (Supplementary Fig. 2). Initially, cohorts of C57BL/6 mice were immunized with nanoparticles containing chicken ovalbumin (OVA) alone (PLGA(OVA)), or (PLGA(OVA)) together with PLGA(MPL), PLGA(R837), or (PLGA(MPL+R837). OVA emulsified in alum was used as a control. Immunization with PLGA(MPL) or PLGA(R837) plus nanoparticles containing 50 μg or 10 μg of OVA (Supplementary Figs 3a, 4a), induced enhanced OVA-specific antibody titres after immunization. Notably, there was a synergistic enhancement of the antibody titres in mice that received PLGA(MPL+R837) (Supplementary Figs 3a, 4a). Secondary immunization with the same immunogen 5 weeks later markedly increased titres in all groups, with the synergy effect with PLGA(MPL+R837) still evident, especially at the lower 10 μg dose (Supplementary Figs 3b, 4b). Thus, in all following experiments we used 10 μg of antigen. In addition to OVA, we also used other antigens including the protective antigen (PA) from Bacillus anthracis10 (Supplementary Fig. 5), and haemagglutinin (HA) from avian influenza H5N1 virus11 (Fig. 1). As observed with OVA, there was a synergistic enhancement in the antigen-specific antibody responses after primary and secondary immunization of mice with PLGA(MPL+R837) plus PLGA(PA) (Supplementary Fig. 5a, b), or PLGA(HA) (Fig. 1a, b).
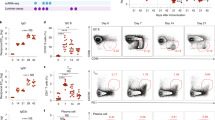
a, b, Antibody titres at 4 weeks after primary and secondary immunization (mean + s.e.m. of four independent experiments, with 4–5 mice per treatment group in each experiment) are shown for IgG2a, IgG2b and IgG1 isotypes. ***P < 0.001, **P < 0.01 (one-way ANOVA with Bonferroni post-hoc test). c, Pooled serum samples from HA-immunized mice at day 28 after secondary immunization boost immunization were tested for their HA-binding avidity using BIAcore SPR-based protein binding assay. Data are representative of plots from one of two independent experiments. RU, maximal response unit. d, Virus neutralization assays were performed with pooled serum samples from treatment groups assayed in duplicates. Results shown are representative titres from one of two independent experiments.
The avidity of antigen–antibody binding is one index of the quality of the antibody response12. We used a surface plasmon resonance (SPR) assay to assess avidity. Sera from mice immunized with PLGA(MPL+R837) plus PLGA(HA) gave the highest binding response (Fig. 1c). The slower dissociation and higher association rates indicate that immunization with PLGA(MPL+R837) plus PLGA(HA) induced a more enhanced high-affinity antibody response than that induced by PLGA(MPL) or PLGA(R837) plus PLGA(HA). A similar trend was also observed with PA as an antigen (Supplementary Fig. 5c). Consistent with the effects on enzyme-linked immunosorbent assay (ELISA) titres and avidity, mice immunized with PLGA(MPL+R837) plus PLGA(HA) had the greatest neutralization antibody titres (Fig. 1d).
Next we assessed the mechanism by which PLGA(MPL+R837) induced synergistic responses. PLGA(MPL+R837) enhanced the secretion of proinflammatory cytokines by dendritic cells in vitro, compared to PLGA(MPL) or PLGA(R837) (Supplementary Fig. 6a). Further, in vivo depletion of dendritic cells in CD11c-DTR transgenic mice13, or Langerhans cells using the Langerin-DTR mice14, resulted in diminished antibody titres (Supplementary Fig. 6b, c). These data demonstrate a critical role for dendritic cells in mediating the antibody response to immunization with PLGA(MPL+R837). Signalling via TLR4 and TLR7 is dependent on the adaptor proteins MyD88 or TRIF (also known as TICAM-1); MPL is reported to signal predominantly via TRIF15. Both MyD88 and TRIF were required for antibody responses stimulated by PLGA(MPL+R837) plus PLGA(OVA) (Supplementary Fig. 7).
B cells express and respond to TLRs16,17. Thus, we determined whether direct triggering of TLRs on B cells was essential for antibody responses. In vitro stimulation of naive splenic B cells with PLGA(MPL+R837) synergistically enhanced B-cell proliferation relative to stimulation with PLGA(MPL) or PLGA(R837) (data not shown). To assess this effect in vivo, mice lacking B cells (μMT mice) were reconstituted with B cells from wild-type mice, or MyD88−/− or Trif−/− mice, and then immunized with PLGA(MPL+R837) plus PLGA(OVA) (Fig. 2a). In μMT mice reconstituted with wild-type B cells immunization induced a synergistic enhancement of antibody responses (Fig. 2b). However, μMT mice reconstituted with MyD88−/− or Trif−/− B cells had diminished antibody titres (Fig. 2c), demonstrating that direct TLR triggering on B cells was required for stimulation of antibody responses. We determined then whether the synergy was dependent on co-expression of TLR4 and TLR7 on the same B cell, or whether there could be complementation between B cells lacking different TLRs. We thus transferred B cells from Tlr4−/− mice, or Tlr7−/− mice, or a 1:1 mixture of B cells from Tlr4−/− and Tlr7−/− mice into μMT mice. Immunization with PLGA(MPL+R837) and PLGA(OVA) demonstrated a requirement for co-expression of both TLRs on the same B cell (Fig. 2d).
a, B-cell-deficient mice (μMT mice) were reconstituted with 40 × 106 B cells from C57BL/6 mice or from MyD88−/−, Trif−/−, Tlr4−/− or Tlr7−/− mice, or equal numbers of Tlr4−/− and Tlr7−/− deficient cells to determine whether expression of TLRs and co-expression of TLR4 and TLR7 on the same B cell was necessary for enhancement of antibody responses. Mice were immunized with 10 µg of OVA encapsulated in PLGA nanoparticles and adjuvants. b–d, Mice were bled at day 28 after primary immunization and OVA-specific total IgG antibody responses were determined using ELISA. Antibody titres are shown (mean + s.e.m. of two independent experiments, with 3 mice per treatment group in each experiment). KO, knockout. ***P < 0.001, **P < 0.01 (one-way ANOVA with Bonferroni post-hoc test). e, The magnitude of OVA-specific IFN-γ-producing memory CD4+ T cells in the draining lymph nodes of μMT mice is shown with representative FACS plots. f, The magnitude of OVA-specific IFN-γ-producing memory CD4+ T cells in the draining lymph nodes is dependent on MyD88 and TRIF expression on B cells. Graphs represent mean frequencies ± s.d. of triplicate cultures of pooled lymph node cells from one out of two independent experiments.
Lastly, TLR activation of dendritic cells is known to stimulate antigen-specific CD4+ T-helper cells, which are essential for induction of antibody responses18. Depletion of CD4+ T-helper cells before immunization diminished antibody responses (Supplementary Fig. 8). Consistent with this, immunization with PLGA(MPL+R837) and PLGA(OVA) resulted in an enhanced antigen-specific CD4+ T-cell response (Fig. 2e). Interestingly, we observed (Fig. 2f) that OVA-specific CD4+ T-helper cell responses were substantially reduced in μMT mice transferred with MyD88−/− or Trif−/− B cells (in which the antibody responses were severely compromised; Fig. 2c), indicating a requirement for activated B cells in mediating enhanced activation of CD4+ T-helper cells, as demonstrated previously19.
Antibody responses develop along two anatomically and functionally distinct pathways20. The extrafollicular pathway rapidly generates short-lived, antibody-producing cells (plasma cells), and the germinal-centre pathway generates memory B cells and long-lived plasma cells that secrete high-affinity antibody20. We determined whether immunization with nanoparticles containing different adjuvants differentially regulated the two pathways. Thus, mice immunized with the different adjuvants were killed at day 7, lymph nodes isolated and the presence of antibody-producing plasma cells evaluated by immunohistology. There was no apparent difference in the IgG+ plasma cells, at day 7, between mice immunized with various adjuvants (Supplementary Fig. 9). We investigated also the kinetics of germinal-centre formation following immunization. Notably, mice immunized with both TLR ligands had a greatly enhanced and sustained germinal-centre response compared to those immunized with a single TLR ligand, (Fig. 3a, b and Supplementary Fig. 9). At day 28 there were approximately 10–12 germinal centres per lymph node and at day 42 there were about 6 germinal centres per lymph node, significantly higher than the numbers observed in lymph nodes of mice immunized with single TLR ligands (Fig. 3a, b). By 8 weeks after immunization the numbers of germinal centres were still substantially higher in the PLGA(MPL+R837) group than in the other groups (Fig. 3b). This demonstrates that PLGA(MPL+R837) preferentially enhances the germinal-centre pathway.
a, C57BL/6 mice were immunized with OVA encapsulated in nanoparticles with MPL+R837 plus antigen. Four weeks after primary immunization, draining lymph nodes were excised, tissue sections prepared and stained for germinal centres (GL-7, red; B220, blue; and IgG, green). Images are representative of two independent experiments with draining lymph nodes obtained from 2–3 mice per treatment condition per experiment. Scale bars, 200 μm for first five panels from left and 100 μm for right-hand panel. b, Germinal centres (GCs) were counted in lymph node (LN) sections at the time points indicated and represented as mean ± s.e.m. from 4–6 draining lymph nodes from n = 2–3 mice per treatment group. c, ELISPOT assay. Combination of TLR4 and TLR7 ligands has no effect on the short-lived antibody-secreting cells, relative to single TLR ligands, but stimulates long-lived antibody-secreting cells that persist for ∼1.5 years. Graph represents average spots per 1 × 106 total lymph node cells ± s.e.m. from duplicate cultures per treatment group. Data are representative of at least 2–3 independent experiments per time point indicated.
Further, assessment of antigen-specific IgG-secreting plasma cells by the enzyme-linked immunosorbent spot (ELISPOT) assay, indicated no differences in the numbers of plasma cells at day 7 in the MPL alone versus MPL+R837 groups (Fig. 3c). At day 28 there were enhanced numbers of plasma cells in the PLGA(MPL+R837) group relative to single TLR-ligand groups. Secondary immunization with the same antigen plus adjuvant resulted in a profoundly enhanced and sustained memory response in the PLGA(MPL+R847) group (Fig. 3c). Notably, there was a persistent plasma-cell response in the lymph node for >1.5 years. Interestingly, there was no corresponding enhancement in the numbers of antigen-specific plasma cells in the bone marrow—a known destination for plasma cells21—in the PLGA(MPL+R837) group relative to the single TLR-ligand groups (data not shown). This demonstrates that immunization with PLGA(MPL+R837) preferentially enhances memory B-cell generation. Consistent with this, fluorescence-activated cells sorting (FACS) analysis22 revealed enhanced numbers of isotype-switched, antigen-specific B cells in the PLGA(MPL+R837) group relative to the PLGA(MPL) or PLGA(R837) groups during the memory phase after a secondary boost, but no such enhancement early during the primary response (Supplementary Fig. 10).
The effects of PLGA(MPL+R837) on the germinal-centre pathway might have occurred via early programming of antigen-specific B cells, or as a result of the continued presence of antigen and/or adjuvant. To determine whether there was any early programming of B cells, we isolated isotype-switched, antigen-experienced B cells by FACS at 7 days after immunization with nanoparticles containing various adjuvants plus OVA, and performed microarray analyses to assess their molecular signatures. Notably, there was a great enrichment for genes normally expressed in memory B cells23, in the cells isolated from mice immunized with PLGA(MPL+R837) plus PLGA(OVA); in contrast, in cells isolated from mice immunized with either PLGA(MPL) or PLGA(R837) plus PLGA(OVA), there was no such enrichment (Supplementary Fig. 11a–c). Such genes included Bcl2, Bcl11a, Tank, several type-I interferon (IFN)-related genes, Plcg2 and Cd38, which are known to have key roles in memory B-cell formation, and several genes that regulate the survival, proliferation and differentiation of germinal-centre B cells, such as Il17ra, Il18r1, Pax5, Ifngr2, Bcor and Ikzf1 (Supplementary Fig. 11a, b). These data indicate that immunization with antigen and MPL+R837 stimulates early programming to the germinal-centre/memory pathway.
Next we determined whether immunization also enhanced antigen-specific memory T-cell responses. There was a synergistic enhancement of OVA-specific IFN-γ-producing CD4+ T-helper cell responses at 8 weeks after secondary immunization (Supplementary Fig. 12a, b), but not at days 7 or 14 after primary immunization (data not shown), indicating a preferential effect on the generation of memory T cells. Similar results were observed in mice immunized with HA and PA antigen, even 1.5 years after immunization (Supplementary Fig. 12c–f). PLGA(MPL+R837) immunization also enhanced the antigen-specific CD8+ T-cell responses. Although no synergistic enhancement was observed in the frequencies of IFN-γ producing, OVA-specific CD8+ T cells at day 7 after primary immunization, there was an increase at day 7 after secondary immunization (Supplementary Fig. 13). Polyfunctional T cells secreting multiple cytokines such as IFN-γ, TNF-α and IL-2 have been implicated in mediating enhanced protection24. We also observed enhanced numbers of triple (IFN-γ, TNF-α and IL-2) and double cytokine (IFN-γ, IL-2) producing CD8+ T cells in mice immunized with PLGA(MPL+R837) (Supplementary Fig. 14a, b). Thus, PLGA(MPL+R837) enhances the magnitude and quality of the antigen-specific memory CD4+ and CD8+ T cells. To assess the relevance of enhanced B- and T-cell responses for protective immunity, we evaluated efficacy of these vaccines in mediating protection in mice, against the 2009 pandemic H1N1 influenza A virus11 (Supplementary Figs 15, 16) and the H5N1 avian influenza virus11,25 (Supplementary Fig. 17). In each case, there was enhanced antigen-specific humoral immunity and survival against lethal infection of mice (Supplementary Figs 15–17).
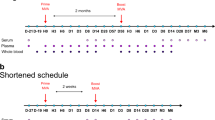
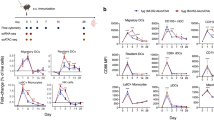
Finally, we assessed the immunogenicity of PLGA(MPL+R837) in non-human primates. In humans and non-human primates, unlike in mice, TLR7 is selectively expressed on plasmacytoid dendritic cells, and not on myeloid dendritic cells2. As multiple dendritic-cell subsets seem to be involved in the stimulation of antibody responses by PLGA(MPL+R837) (Supplementary Fig. 6b, c), we used R848, which signals through both TLR7 and TLR82,26. Further, in humans, although naive B cells do not express TLRs 4, 7 or 8, activated B cells including plasma cells upregulate and respond to these TLRs27. Current human monovalent vaccines against H1N1 influenza contain 15 μg of HA (effectively 45 μg of whole inactivated virus (WIV))28. We therefore used 50 μg of WIV (∼16 μg HA), as well as a fivefold lower dose (∼3 μg HA), with and without adjuvant, to determine whether there was a dose-sparing effect. Four cohorts of four rhesus macaques per cohort were immunized subcutaneously with the indicated doses of WIV, with or without PLGA(MPL+R837) or PLGA(MPL+R848) (Fig. 4a). After a single immunization with 10 μg or 50 μg of WIV without any adjuvants, there was no detectable antibody response (Fig. 4b–d). In contrast, adjuvanting with PLGA(MPL+R848) induced robust antibody responses, as early as 2 weeks (Fig. 4b–d). Immunization with PLGA(MPL+R837) yielded enhanced binding antibodies, but had more modest effects on HA inhibition and neutralization titres (Fig. 4b–d). Notably, the magnitude of HA inhibition and neutralization titres 28 days after a single immunization with PLGA(MPL+R848) was much greater than 1/40, considered the correlate of protection against influenza in humans (Fig. 4c, d)29. There was at least a fivefold dose-sparing effect, as 10 μg of antigen plus PLGA(MPL+R848) yielded a much greater response than that induced by 50 μg of antigen alone (Fig. 4b–d). Secondary immunizations enhanced the antibody responses in all of the groups, and although PLGA(MPL+R848) still induced the strongest response, PLGA(MPL+R837) also induced responses greater than that required for protection (Fig. 4c, d). Further, the dose-sparing effect was still evident after secondary immunization (Fig. 4b–d).
a, Rhesus macaques (n = 4) were immunized with 10 µg of H1N1 WIV with or without nanoparticle-encapsulated MPL+R837 or MPL+R848. Fifty micrograms of MPL and 750 µg of R837 and R848 encapsulated in nanoparticles were used per macaque. One group of 4 animals was also immunized with 50 µg of WIV to determine dose-sparing effects mediated by adjuvants. b, Antibodies against WIV were analysed as described in Methods and results are represented as mean ± s.e.m. c, HA inhibition (HAI) titres were assayed at indicated time points and are represented as mean ± s.e.m. d, Neutralization titres as represented as the reciprocal of the plasma dilution that decreased the number of plaques formed by the live virus by 50%. Statistical significance was analysed by ANOVA (Bonferroni post-hoc test) and indicated on the figures wherever significant. ***P < 0.001, **P < 0.01, *P < 0.05. Data are represented as mean ± s.e.m.
Here we have described a nanoparticle-based vaccine that resembles a virus in size and composition, and that recapitulates the immunogenicity of live viral vaccines5. A notable feature of the immune response stimulated by this vaccine is in the induction of long-lived germinal centres and persistent antigen-specific B- and T-cell responses, similar to that seen with viral infections30. Importantly, although all TLR ligands stimulate primary antibody responses, the combined TLR4 and TLR7 stimulus enhances the germinal-centre pathway of memory B-cell formation and long-lived plasma-cell responses, far more efficiently than stimulation with single TLR ligands (Supplementary Fig. 18). The molecular signatures of antigen-activated B cells isolated early after immunization indicates early programming towards a quasi memory state (Supplementary Fig. 11), but it is also likely that persistent antigen/adjuvants, as evidenced by persistent immune complexes in germinal centres (Fig. 3), have a role. A curious aspect of our data is the requirement for the TLR ligands and antigens to be delivered on two separate particles, consistent with other studies8,9. From a practical perspective, this offers flexibility in coupling a generic adjuvant-containing particle with another particle containing antigen from any pathogen. Further, as each of the components of this vaccine (MPL, R837, PLGA) have been licensed for human use, this vaccine formulation may provide a universal platform for vaccine design against pandemics and emerging infections.
Methods Summary
Synthesis and characterization of nanoparticles
Antigens were encapsulated in PLGA nanoparticles using a water in oil in water (w/o/w) emulsion technique. MPL was encapsulated in PLGA formulations using an oil in water (o/w) single emulsion process as described before with slight modifications8. Sizing of the nanoparticles was conducted using a dynamic light scattering, and protein loading was assessed as described before31.
Immunization of mice and non-human primates
Eight-to-twelve week old BALB/c or C57BL/6 mice (Charles River) were immunized with 10 µg of antigen encapsulated in nanoparticles suspended in 200 µl of PBS subcutaneously at the base of the tail. Ten-to-thirteen year old female Rhesus Macaques (7–10 kg) were used and were immunized subcutaneously in the right leg. All procedures were performed in accordance with the Emory School of Medicine Institutional Animal Care and Use Committee guidelines.
Antigen–antibody affinity binding analysis
SPR binding measurements were carried out on a BIAcore 3000 instrument (BIAcore/GE Healthcare).
H5N1 microneutralization assays
This was performed as described previously25.
B-cell ELISPOT
1 × 106 lymph node cells were serially diluted and cultured overnight in duplicate wells of OVA-coated nitrocellulose-lined 96-well plates (Millipore). Cells were discarded and wells were treated with biotinylated goat anti-mouse total IgG (Southern Biotech) in PBS/0.5% Tween-20 plus 1% FBS for 1.5 h at room temperature (25 °C). Wells were washed and treated with streptavidin alkaline phosphatase (Vector Labs) for another 1.5 h at room temperature. Finally, NBT/BCIP colorimetric substrate for alkaline phosphatase was added to the wells and the reaction was stopped after visualization of purple coloured spots.
Affymetrix GeneChip analysis
Total RNA from sorted B cells was purified using Trizol (Invitrogen). All RNA samples were checked for purity and for integrity and amplified, processed through EXON Module, fragmented and labelled. Labelled targets hybridized to GeneChip Mouse Gene 1.0ST arrays (Affymetrix).
Online Methods
Encapsulation of antigens and TLR ligands in nanoparticles
Antigens were encapsulated in PLGA nanoparticles using a water in oil in water (w/o/w) emulsion technique. Briefly, 200 µl of protein solution (PBS + 0.5% polyvinyl alcohol (PVA) as an excipient (Sigma Aldrich)), OVA grade VI (Sigma Aldrich) at 50 mg ml−1, Bacillus anthracis PA (List Laboratories) at 15 mg ml−1, and A/Vietnam/1203/2004 specific haemagglutinin protein (H5HA; affinity chromatography purified from 293 HEK cells) at 15 mg ml−1 were homogenized with 10% w/v PLGA (RG502H, Bohreinger Ingelheim) in dichloromethane (200 mg in 2 ml) with the Powergen homogenizer (Fisher Scientific) at speed 5 for 1.5 min. The water in oil emulsion (w/o) was then added to 15 ml of 5% w/v solution of PVA for the second emulsion step identical to the first emulsion process described earlier, at speed 5. The water in oil in water (w/o/w) double emulsion was then subjected to solvent evaporation for 4 h at room temperature (25 °C).
MPL was encapsulated in PLGA formulations using an oil in water (o/w) single emulsion process as described before with slight modifications8. MPL (detoxified lipid A, Avanti Lipids) was dissolved in chloroform at 5 mg ml−1 and TLR7 ligand R837 (Invivogen) was dissolved at 10–20 mg ml−1 in DMSO with heating. R848 was dissolved at 12.5 mg ml−1 in dichloromethane. MPL, 0.5 ml at 5 mg ml−1 was added to 200 mg of PLGA polymer dissolved in 2.0 ml of dichloromethane. For particles containing both MPL and R837, 0.5 ml or 5 mg of R837 in DMSO was added to the mixture of PLGA and MPL. For particles containing both MPL and R848, 8 mg of R848 in dichloromethane was added to the mixture of PLGA and MPL. For particles used in non-human primate studies, 0.1 ml of 5 mg ml−1 MPL, 0.5 ml or 10 mg of R837 and 12.5 mg of R848 was added to 200 mg of PLGA polymer in 2.0 ml of dichloromethane. The organic phase containing PLGA with MPL and/or R837 or R848 was homogenized with 15 ml of a 5% w/v PVA solution for 2 min using a speed setting 6 at room temperature. The oil in water emulsion (o/w) was then added to 85 ml of a 5% w/v solution of PVA surfactant to evaporate the organic solvent for 4 h at room temperature. The nanoparticles formed were centrifuged at 3,500g for 20 min and washed with 50 ml of 0.2-µm filter sterilized, deionized water 3 times. Nanoparticles were snap frozen in liquid nitrogen and lyophilized using a Freezone 2.5L benchtop lyophilizer (Labconco).
Nanoparticle characterization
Sizing of the nanoparticles was conducted using a dynamic light scattering based sizer (90PLUS) from Brookhaven Instruments. Sizes are represented as the mean diameter of the volume average size distribution ± standard deviation of different batches. Protein encapsulation levels were estimated as described before using a BCA assay (Pierce Biotechnology)31. UV-Vis scan using a Cary Win50 UV-Vis spectrophotometer yielded a peak absorbance for R837 at 325 nm (327 nm for R848) and encapsulation was estimated using a standard curve of R837 or R848 in DMSO and NaOH/SDS. MPL encapsulated in nanoparticles was used in splenic dendritic-cell stimulation experiments with known concentrations of soluble MPL yielding identical cytokine production and estimated at 100% encapsulation efficiency.
Mouse dendritic cell culture
CD11c+ dendritic cells were isolated from spleens of naive C57BL/6 mice. Spleens were digested with collagenase type IV (Worthington Chemicals), and CD11c+ cells were enriched by positive selection using anti mouse CD11c magnetic beads according to manufacturer’s instructions (Miltenyi Biotech). Dendritic cells (1 × 106 cells ml−1) were cultured in 48-well culture plates with PLGA-encapsulated TLR ligands for 24 h.
Mice
Il6−/− and Ticam11ps2/lps2 strains were obtained from Jackson Labs. Ifnar−/− strain was obtained from S. Speck; MyD88−/− and Tlr7−/− from S. Akira; Tlr4−/− from K. Kobayashi.
Immunization of mice and non-human primates
Eight-to-twelve week old BALB/c or C57BL/6 mice (Charles River Laboratories) were immunized with 10 µg of antigen in nanoparticles (suspended in 200 µl of PBS) subcutaneously at the base of the tail. TLR ligands were co-delivered with protein-encapsulated nanoparticles. Three milligrams of PLGA particles containing MPL, R837 or MPL+R837 containing 37.5 µg of MPL and 60 µg of R837 were used. Imject Alum (Pierce/ThermoScientific) was used to adsorb protein at a 1:1 volume ratio as per the manufacturer’s instructions. Mice were bled via the lateral tail vein at regular intervals after primary and secondary immunizations and serum was isolated for analysis of antibody responses by ELISA assays. CD11c+ DTR mice13 were immunized 24 h after dendritic–cell depletion. Langerin-DTR mice14 were immunized 3 weeks after depletion, when there is replenishment of Langerin+ dendritic cells in the dermis and lymph nodes, but not in the epidermis32. CD4+ T cells were depleted with an anti-mouse CD4+ antibody (clone GK1.5, provided by R. Mittler) injected at 250 µg per mouse as described33. Ten-to-thirteen year old female Rhesus Macaques (7–10 kg) were used. Animals were immunized subcutaneously in the right leg. All animal procedures were performed in accordance with guidelines established by the Emory School of Medicine Institutional Animal Care and Use Committee Guidelines.
Antibody ELISA
Ninety-six-well Nunc maxisorp plates were coated with 100 µl of 20 µg ml−1 of OVA, 1 µg ml−1 of PA or 0.5 µg of H5HA protein overnight at 4° C. Plates were washed 3 times with PBS/0.5% Tween-20 using a Biotek auto plate washer and blocked with 200 µl of 4% non-fat dry milk (Biorad) for 2 h at room temperature. Serum samples from immunized mice at the indicated time points were serially diluted in 0.1% non-fat dry milk in PBS/0.5% Tween-20 and incubated on blocked plates for 2 h at room temperature. Detection antibodies were obtained from Southern Biotech. Wells were washed and incubated with anti-mouse IgG2c–HRP (horseradish peroxidase) conjugate (1:2,000), anti-mouse IgG2b–biotin conjugate (1:2,000), anti-mouse IgG1–HRP conjugate (1:5,000) and streptavidin–HRP conjugate (1:5,000) in PBS/0.5% Tween-20 for 2 h at room temperature. Plates were washed and developed using 100 µl per well of tetramethylbenzidine (TMB) substrate (BD Biosciences) and stopped using 2N H2SO4. Plates were analysed using a BioRad plate reading spectrophotometer at 450 nm with correction at 595 nm. Antibody titres were represented as serum reciprocal dilution yielding a ≥0.1 optical density (OD) value at 450 nm. Antibody levels (ng ml−1) in H1N1 WIV-immunized mice were assayed as previously described34. To analyse WIV-specific antibody levels in rhesus macaques, the assay was performed as described for mice. A capture rhesus IgG (clone SB108A) was used to establish a standard curve with the rhesus IgG standards (catalogue no. 0135-01; Southern Biotech). 2 µg ml−1 of WIV was used to coat Nunc maxisorp plates overnight and plates were blocked with 4% non-fat dry milk. Rhesus plasma samples were used at appropriate dilutions for prime and boost and incubated for 2 h at room temperature. Plates were washed 5× with PBS/0.5% Tween-20 using an automated plate washer and a goat anti-rhesus HRP was used at 1:4,000 dilution for 1 h at room temperature, washed and developed with TMB substrate. Antibody concentrations were calculated from the IgG standard curves and represented as μg per ml.
BIAcore assay
SPR binding measurements were carried out on a BIAcore 3000 instrument (BIAcore/GE Healthcare), as described previously35. Serum samples were injected at a 1:50 dilution in PBS for 5 min at a flow rate of 10 μl min−1. Binding to the negative control rPA surface was subtracted from each sample curve and binding response was measured at 15 and 500 s after the end of the injection. As a measure of antigen-specific antibody-binding avidity, maximal response unit (binding RU) and dissociation rates were measured. Maximal binding RU was measured after subtraction of non-specific signal on the control surface at 15 s post-injection and normalized dissociation was calculated as a ratio of late-to-early binding responses over a dissociation phase of 500 s. Following each injection cycle, chip surfaces were regenerated with a short injection of 25 mM NaOH.
H5N1 microneutralization assays
Serum samples from H5HA-immunized mice were tested for their ability to neutralize a recombinant (6:2) A/PR/8/34 influenza A virus expressing the A/Vietnam/1203/2004 H5HA and N1NA in cell culture in vitro as described previously25.
Histology and immunofluorescence
Draining inguinal lymph nodes were isolated and snap frozen in moulds containing OCT medium, dropped into 2-methyl butane cooled with liquid nitrogen. Frozen lymph nodes were sectioned at 5 µm, fixed in ice-cold acetone for 10 min, air dried and stored at −80° C. Sections were fluorescently stained with Dylight488-labelled anti-mouse total IgG (Jackson Immunoresearch), Alexa647-labelled anti-mouse B220 or GL-7 (Ebioscience), followed by Alexa555-conjugated streptavidin (Invitrogen). Fluorescent images were captured using the ×5 and ×20 objectives on a Zeiss Axioscope (Carl Zeiss).
B-cell ELISPOT
1 × 106 lymph node cells were serially diluted and cultured overnight in duplicate wells of OVA-coated nitrocellulose-lined 96-well MAHA ELISPOT plates (Millipore). Cells were discarded and wells were treated with biotinylated goat anti-mouse total IgG (Southern Biotech) in assay buffer (PBS/0.5% Tween-20 +1% FBS) for 1.5 h at room temperature. Wells were washed and treated with streptavidin alkaline phosphatase (Vector Labs) at 1:500 for 1.5 h at room temperature. NBT/BCIP colorimetric substrate for alkaline phosphatase was added to the wells and the reaction was stopped after visualization of purple coloured spots. Number of ELISPOTS per well were counted using an ImmunoSpot ELISPOT reader and represented as number of ELISPOTS (antibody-secreting cells) per total 1 × 106 lymph node cells.
CD4+ T-cell assays
1 × 106 lymph node cells were cultured in a 200 µl volume with 100 µg ml−1 of OVA protein or 5 µg ml−1 of PA or 5 µg ml−1 of H5HA in 96-well round-bottomed plates for 4 days. Cells were transferred to anti-CD3 (10 µg ml−1) and anti-CD28 (2 µg ml−1) coated flat-bottomed 96-well plates for 6 h in the presence of Golgi Plug (1 µg ml−1) (BD Biosciences). Cells were stained for surface CD4 using PerCP anti-mouse CD4 (clone RM4-5) for 30 min at 4 °C. Cells were washed 3 times with FACS buffer, fixed and permeabilized using BD Cytofix/Cytoperm and stained for intracellular IFN-γ using APC-conjugated anti-mouse IFN-γ (clone XMG1.2) for 30 min in 1× BD perm/wash solution at 4 °C. Cells were washed with perm wash followed by FACS buffer and acquired on a FACS Caliber cytometer. FACS data were analysed using the Flow Jo software (Tree Star).
CD8+ T-cell assays
Primary and memory CD8+ T-cell responses were evaluated at day 7 after primary and secondary immunizations. Briefly, peripheral blood mononuclear cells (PBMCs) were enriched using a sucrose density gradient separation (Histopaque; Sigma Aldrich) and cultured with OVA-specific MHC class I restricted peptide at 1 µg ml−1 (SIINFEKL) for restimulation ex vivo in the presence of brefeldin A (5 µg ml−1). Stimulated cells were stained for intracellular cytokines using established protocols from BD Biosciences as explained for CD4+ T-cell staining experiments above. Cells were stained with PerCP-conjugated anti-mouse CD8α (Ly-2) (clone 53-6.7), APC-conjugated anti-mouse IFN-γ (clone XMG1.2), FITC-conjugated anti-mouse TNF-α (clone MP6-XT22) and PE-conjugated anti-mouse IL-2 (clone JES6-5H4). Labelled cells were acquired on a FACS Caliber cytometer and FACS data were analysed using Flow JO software (TreeStar).
B-cell multicolour flow cytometry
Antibodies used: PE-labelled anti-mouse IgG (Jackson Immunoresearch); PerCP-labelled anti-mouse CD19 (clone 6D5; Biolegend); E-fluor-405-labelled anti-mouse IgD (Ebioscience); Alexa430 succinimidyl ester for live/dead cell discrimination (Invitrogen); biotin-conjugated anti-mouse Ly77/GL7 (clone GL-7; Ebioscience); APC-labelled anti-mouse CD138 (clone 281-2; BD Biosciences); APC-Cy7-labelled anti-mouse TCR-β (clone H57-597; Biolegend); and APC-Cy7-labelled anti-mouse CD11b (clone M1/70, Biolegend). OVA was labelled with Alexa488 2,3,5,6 TFP ester (Invitrogen) as per the manufacturer’s instructions. Unlabelled dye was separated using a 30K cut-off membrane-specified Amicon Ultra4 centrifugal filter (Millipore). Briefly, collagenase-digested lymph nodes from immunized mice were counted for absolute cell numbers. Cells were first stained with Alexa430 succinimidyl ester for dead-cell staining in PBS for 30 min at 4 °C. Cells were washed 2 times with FACS buffer (PBS with 5% FBS) and stained with PE-labelled anti-mouse total IgG for 30 min at 4 °C. Cells were washed 2 times and labelled with the all the above mentioned anti-mouse surface proteins. Qdot655-labelled streptavidin was used to label the biotinylated anti-mouse GL-7 antibodies for another 30 min at 4° C. Stained cell samples were fixed with BD Cytofix (BD Biosciences) and acquired on a LSR-II cytometer (BD Biosciences). FACS data were analysed on Flow JO software. For B-cell sorting assays, cells were fluorescently labelled as described above and sorted on a FACS Aria cell sorter (BD Biosciences).
H1N1 virus stock preparation
Madin-Darby canine kidney (MDCK) cells (ATCC CCL 34, American Type Culture Collection) were maintained in DMEM (Mediatech) containing 10% FBS. Stocks of influenza virus strains were prepared by inoculation with H1N1 swine-origin A/California/04/09 strain in allantoic fluid, in 10- or 11-day-old embryonated hen’s eggs. Virus stocks were harvested from the allantoic fluid. The purity of the virus was determined by SDS–PAGE in combination with Coomassie blue stain and electron microscopy. The HA activity was determined using chicken red blood cells 0.5% w/v in PBS pH 7.2 as previously described36. The purified virus was inactivated with formaldehyde at a final concentration of 0.01% (v/v), incubated for 72 h at 4 °C, and then dialysed against PBS buffer. Inactivation of virus was confirmed by inoculation of the virus into 10-day-old embryonated hen’s eggs and plaque assay in MDCK cells37. We generated mouse adapted A/California/04/09 virus by five serial passages in BALB/c mice. We then determined the LD50 for this virus using the Reed–Muench formula38. For the challenge and infection studies the mice were anesthaetized with isoflurane and then infected with virus by intranasal instillation.
H1N1 HA inhibition assays
Determined as described previously34.
H5N1 and H1N1 lethal challenge infections
Female BALB/c mice immunized with the indicated vaccine schedule were anaesthetized by intraperitoneal injection of ketamine/xylazine and then intranasally infected with 1,000 LD50 of recombinant A/Vietnam/1203/2004 within the enhanced BSL3 Emerging Pathogens Facility at Mount Sinai School of Medicine. To determine survival rates post challenge with mouse adapted A/California/04/09 H1N1 virus, 5 female BALB/c mice per treatment group were challenged 8 weeks after immunization by intranasal instillation of 30 μl of 20× LD50 live virus after anaesthesia with isoflurane and monitored for morbidity and mortality up to 14 days. Animals were humanely killed and reported as dead if body weight loss achieved 25%. All H5N1 animal procedures were performed in accordance with guidelines established by the Mount Sinai School of Medicine Institutional Animal Care and Use Committee and National Institutes of Health for the care and use of laboratory animals.
H1N1 neutralization assays with primate plasma
Sera from non-human primates immunized with swine-origin H1N1 2009 were serially diluted and mixed with 100 p.f.u. of MDCK-grown homologous virus for 1 h at room temperature. The mixture was further added to an MDCK-cell monolayer and incubated for 45 min at room temperature. The inoculum was removed, wells were overlaid with DMEM agar and incubated for 2 days at 37 °C in a 5% CO2 humidified incubator. Then plates were fixed with 0.25% gluteraldeyde and stained with 1% crystal violet in 20% ethanol, and plaques were counted. Neutralizing antibody titres were determined as the reciprocal of the serum dilution that decreased by 50% the number of plaques formed by the live virus.
Affymetrix GeneChip analysis
Total RNA from sorted B cells (1.8 × 104 to 1.2 × 106 cells) was purified using Trizol (Invitrogen). All RNA samples were checked for purity using a ND-1000 spectrophotometer (NanoDrop Technologies) and for integrity by electrophoresis on a 2100 BioAnalyser (Agilent Technologies). The samples were amplified using the Nugen WT Pico Kit (NuGEN Technology) and the target reactions were run with 50 ng of total RNA. The amplification products were processed through the EXON Module (NuGEN Technology), which creates sense-strand cDNA targets. The sense-strand cDNA targets were then fragmented and labelled using NuGEN’s FL-Ovation cDNA Biotin Module V2 (NuGEN Technology). Labelled targets were hybridized to GeneChip Mouse Gene 1.0ST arrays (Affymetrix), following standard Nugen protocols for target hybridization to the Affymetrix Gene Arrays. The hybridizations were run for 16 h at 45 °C, 60 r.p.m. in an Affymetrix Hybridization Oven 640. The Cartridge arrays were washed and stained using the Affyemtrix Fluidics Stations 450, following Affymetrix protocols. Scanning was performed on an Affymetrix GeneChip 3000 7G scanner, and Affymetrix GCOS software was used to perform image analysis and generate raw intensity data. Two independent sets of samples at day 7 post-treatment were used in our analyses. Each set is comprised of B cells from mice immunized with MPL+R837 or from those immunized with antigen plus MPL or R837 alone. Probe sets of all six samples were normalized by RMA, which includes global background adjustment and quantile normalization. Each set of samples was subsequently normalized by z-score (number of standard deviations from mean) and treated as biological replicates. Affymetrix chip annotation of GeneChip Mouse Gene 1.0ST platform was used to annotate and select probe sets that target a known gene (defined as having an Entrez gene ID). Different probe sets that target the same gene were collapsed by taking the probe set with the highest median expression value across all samples. Student’s t-test (P < 0.05) was used to identify genes differentially expressed in mice immunized with antigen plus MPL+R837 compared to those immunized with antigen plus MPL or R837 alone. A meta-analysis was performed using publicly available microarray data of distinct B-cell subsets (plasma, germinal centre and memory B cells)23. Purification strategy, RNA-processing method and hybridization strategy can be found in the original publication23. Raw microarray data (CEL files of samples GSM94747, GSM94762, GSM94763, GSM94764, GSM94765, GSM94766, GSM94767, GSM94768, GSM94769, GSM94771 and GSM94772) were downloaded from the NCBI GEO website (GSE4142) and processed by RMA normalization. Affymetrix chip annotation of the GeneChip Mouse Genome 430 2.0 platform was used to annotate and select probe sets that target a known gene (defined as having an Entrez gene ID). Different probe sets that target the same gene were collapsed by taking the probe set with the highest median expression value across all samples. Student’s t-test (P < 0.05) was used to identify differentially expressed genes (DEGs) between any given two B-cell subsets. Genes were classified as ‘plasma DEGs’ if they were upregulated in plasma compared to germinal-centre B cells and also upregulated in plasma compared to memory B cells. A similar approach was used to identify ‘germinal center DEGs’ and ‘memory DEGs’.
Genes that were up- or downregulated in mice immunized with antigen plus MPL+R837 compared to those immunized with antigen plus MPL or R837 alone, were cross-referenced to the genes highly expressed in specific B-cell subsets. Fold enrichment was calculated using the formula (common ZX/deg Z)/(subset X/total), where common ZX = number of genes upregulated in treatment Z (combination of MPL+R837 or individual MPL or R837) and also highly expressed in B-cell subset X; deg Z = number of all genes upregulated in treatment Z; subset X = number of all genes highly expressed in B-cell subset X; and total = number of all genes in the chip common to both platforms. In Supplementary Figure 11a, the labelled genes on the right indicate highly expressed genes in the MPL+R837 group that are known to regulate B-cell survival, the germinal-centre response, and differentiation of memory B cells39,40,41,42,43,44,45,46,47,48,49,50,51,52,53,54,55,56.
References
Pulendran, B. & Ahmed, R. Translating innate immunity into immunological memory: implications for vaccine development. Cell 124, 849–863 (2006)
Kawai, T. & Akira, S. The role of pattern-recognition receptors in innate immunity: update on Toll-like receptors. Nature Immunol. 11, 373–384 (2010)
Pulendran, B. Learning immunology from the yellow fever vaccine: innate immunity to systems vaccinology. Nature Rev. Immunol. 9, 741–747 (2009)
Querec, T. et al. Yellow fever vaccine YF-17D activates multiple dendritic cell subsets via TLR2, 7, 8, and 9 to stimulate polyvalent immunity. J. Exp. Med. 203, 413–424 (2006)
Querec, T. D. et al. Systems biology approach predicts immunogenicity of the yellow fever vaccine in humans. Nature Immunol. 10, 116–125 (2009)
Napolitani, G., Rinaldi, A., Bertoni, F., Sallusto, F. & Lanzavecchia, A. Selected Toll-like receptor agonist combinations synergistically trigger a T helper type 1-polarizing program in dendritic cells. Nature Immunol. 6, 769–776 (2005)
Peek, L. J., Middaugh, C. R. & Berkland, C. Nanotechnology in vaccine delivery. Adv. Drug Deliv. Rev. 60, 915–928 (2008)
Kazzaz, J. et al. Encapsulation of the immune potentiators MPL and RC529 in PLG microparticles enhances their potency. J. Control. Release 110, 566–573 (2006)
Singh, M., Chakrapani, A. & O’Hagan, D. Nanoparticles and microparticles as vaccine-delivery systems. Expert Rev. Vaccines 6, 797–808 (2007)
Young, J. A. & Collier, J. R. Attacking anthrax. Sci. Am. 286, 48–59 (2002)
Chen, G. L. & Subbarao, K. Live attenuated vaccines for pandemic influenza. Curr. Top. Microbiol. Immunol. 333, 109–132 (2009)
Hangartner, L., Zinkernagel, R. M. & Hengartner, H. Antiviral antibody responses: the two extremes of a wide spectrum. Nature Rev. Immunol. 6, 231–243 (2006)
Jung, S. et al. In vivo depletion of CD11c+ dendritic cells abrogates priming of CD8+ T cells by exogenous cell-associated antigens. Immunity 17, 211–220 (2002)
Kissenpfennig, A. et al. Dynamics and function of Langerhans cells in vivo: dermal dendritic cells colonize lymph node areas distinct from slower migrating Langerhans cells. Immunity 22, 643–654 (2005)
Mata-Haro, V. et al. The vaccine adjuvant monophosphoryl lipid A as a TRIF-biased agonist of TLR4. Science 316, 1628–1632 (2007)
Bernasconi, N. L., Traggiai, E. & Lanzavecchia, A. Maintenance of serological memory by polyclonal activation of human memory B cells. Science 298, 2199–2202 (2002)
Pasare, C. & Medzhitov, R. Control of B-cell responses by Toll-like receptors. Nature 438, 364–368 (2005)
Mitchison, N. A. T-cell–B-cell cooperation. Nature Rev. Immunol. 4, 308–312 (2004)
van Essen, D., Dullforce, P., Brocker, T. & Gray, D. Cellular interactions involved in Th cell memory. J. Immunol. 165, 3640–3646 (2000)
McHeyzer-Williams, L. J. & McHeyzer-Williams, M. G. Antigen-specific memory B cell development. Annu. Rev. Immunol. 23, 487–513 (2005)
Slifka, M. K., Antia, R., Whitmire, J. K. & Ahmed, R. Humoral immunity due to long-lived plasma cells. Immunity 8, 363–372 (1998)
McHeyzer-Williams, M. G., McLean, M. J., Lalor, P. A. & Nossal, G. J. Antigen-driven B cell differentiation in vivo. J. Exp. Med. 178, 295–307 (1993)
Luckey, C. J. et al. Memory T and memory B cells share a transcriptional program of self-renewal with long-term hematopoietic stem cells. Proc. Natl Acad. Sci. USA 103, 3304–3309 (2006)
Wille-Reece, U. et al. HIV Gag protein conjugated to a Toll-like receptor 7/8 agonist improves the magnitude and quality of Th1 and CD8+ T cell responses in nonhuman primates. Proc. Natl Acad. Sci. USA 102, 15190–15194 (2005)
Steel, J. et al. Live attenuated influenza viruses containing NS1 truncations as vaccine candidates against H5N1 highly pathogenic avian influenza. J. Virol. 83, 1742–1753 (2009)
Jurk, M. et al. Human TLR7 or TLR8 independently confer responsiveness to the antiviral compound R-848. Nature Immunol. 3, 499 (2002)
Dorner, M. et al. Plasma cell toll-like receptor (TLR) expression differs from that of B cells, and plasma cell TLR triggering enhances immunoglobulin production. Immunology 128, 573–579 (2009)
Clark, T. W. et al. Trial of 2009 influenza A (H1N1) monovalent MF59-adjuvanted vaccine. N. Engl. J. Med. 361, 2424–2435 (2009)
Potter, C. W. & Oxford, J. S. Determinants of immunity to influenza infection in man. Br. Med. Bull. 35, 69–75 (1979)
Bachmann, M. F., Odermatt, B., Hengartner, H. & Zinkernagel, R. M. Induction of long-lived germinal centers associated with persisting antigen after viral infection. J. Exp. Med. 183, 2259–2269 (1996)
Sah, H. A new strategy to determine the actual protein content of poly(lactide-co-glycolide) microspheres. J. Pharm. Sci. 86, 1315–1318 (1997)
Henri, S. et al. CD207+ CD103+ dermal dendritic cells cross-present keratinocyte-derived antigens irrespective of the presence of Langerhans cells. J. Exp. Med. 207, 189–206 (2010)
den Haan, J. M., Kraal, G. & Bevan, M. J. Cutting edge: lipopolysaccharide induces IL-10-producing regulatory CD4+ T cells that suppress the CD8+ T cell response. J. Immunol. 178, 5429–5433 (2007)
Zhu, Q. et al. Immunization by vaccine-coated microneedle arrays protects against lethal influenza virus challenge. Proc. Natl Acad. Sci. USA 106, 7968–7973 (2009)
Staats, H. F. et al. In vitro and in vivo characterization of anthrax anti-protective antigen and anti-lethal factor monoclonal antibodies after passive transfer in a mouse lethal toxin challenge model to define correlates of immunity. Infect. Immun. 75, 5443–5452 (2007)
Compans, R. W. Hemagglutination-inhibition: rapid assay for neuraminic acid-containing viruses. J. Virol. 14, 1307–1309 (1974)
Enioutina, E. Y., Visic, D. & Daynes, R. A. The induction of systemic and mucosal immune responses to antigen-adjuvant compositions administered into the skin: alterations in the migratory properties of dendritic cells appears to be important for stimulating mucosal immunity. Vaccine 18, 2753–2767 (2000)
Reed, L. J. & Muench, H. A simple method of estimating fifty percent endpoints. Am. J. Hyg. 27, 493–497 (1938)
Badr, G. et al. Type I interferon (IFN-α/β) rescues B-lymphocytes from apoptosis via PI3Kδ/Akt, Rho-A, NFκB and Bcl-2/Bcl(XL). Cell. Immunol. 263, 31–40 (2010)
Bekeredjian-Ding, I. B. et al. Plasmacytoid dendritic cells control TLR7 sensitivity of naive B cells via type I IFN. J. Immunol. 174, 4043–4050 (2005)
Thibault, D. L. et al. IRF9 and STAT1 are required for IgG autoantibody production and B cell expression of TLR7 in mice. J. Clin. Invest. 118, 1417–1426 (2008)
Tovey, M. G., Lallemand, C. & Thyphronitis, G. Adjuvant activity of type I interferons. Biol. Chem. 389, 541–545 (2008)
Swanson, C. L. et al. Type I IFN enhances follicular B cell contribution to the T cell-independent antibody response. J. Exp. Med. 207, 1485–1500 (2010)
Liu, H. et al. Functional studies of BCL11A: characterization of the conserved BCL11A-XL splice variant and its interaction with BCL6 in nuclear paraspeckles of germinal center B cells. Mol. Cancer 5, 18 (2006)
Smith, K. G. et al. bcl-2 transgene expression inhibits apoptosis in the germinal center and reveals differences in the selection of memory B cells and bone marrow antibody-forming cells. J. Exp. Med. 191, 475–484 (2000)
Aiba, Y. et al. Preferential localization of IgG memory B cells adjacent to contracted germinal centers. Proc. Natl Acad. Sci. USA 107, 12192–12197 (2010)
Zhou, G. & Ono, S. J. Induction of BCL-6 gene expression by interferon-γ and identification of an IRE in exon I. Exp. Mol. Pathol. 78, 25–35 (2005)
Mitsdoerffer, M. et al. Proinflammatory T helper type 17 cells are effective B-cell helpers. Proc. Natl Acad. Sci. USA 107, 14292–14297 (2010)
Chin, A. I. et al. TANK potentiates tumor necrosis factor receptor-associated factor-mediated c-Jun N-terminal kinase/stress-activated protein kinase activation through the germinal center kinase pathway. Mol. Cell. Biol. 19, 6665–6672 (1999)
Basso, K. & Dalla-Favera, R. BCL6: master regulator of the germinal center reaction and key oncogene in B cell lymphomagenesis. Adv. Immunol. 105, 193–210 (2010)
Kano, G. et al. Ikaros dominant negative isoform (Ik6) induces IL-3-independent survival of murine pro-B lymphocytes by activating JAK-STAT and up-regulating Bcl-xl levels. Leuk. Lymphoma 49, 965–973 (2008)
Ke, N., Godzik, A. & Reed, J. C. Bcl-B, a novel Bcl-2 family member that differentially binds and regulates Bax and Bak. J. Biol. Chem. 276, 12481–12484 (2001)
Airoldi, I. et al. Expression and function of IL-12 and IL-18 receptors on human tonsillar B cells. J. Immunol. 165, 6880–6888 (2000)
Airoldi, I. et al. Heterogeneous expression of interleukin-18 and its receptor in B-cell lymphoproliferative disorders deriving from naive, germinal center, and memory B lymphocytes. Clin. Cancer Res. 10, 144–154 (2004)
Hikida, M. et al. PLC-γ2 is essential for formation and maintenance of memory B cells. J. Exp. Med. 206, 681–689 (2009)
Nera, K. P. & Lassila, O. Pax5—a critical inhibitor of plasma cell fate. Scand. J. Immunol. 64, 190–199 (2006)
Acknowledgements
We thank R. Ahmed and B. Rouse for discussion and comments on the manuscript. We thank B. Norris for assistance with FACS analysis, and H. Oluoch for assistance with cryostat sectioning. The work in the laboratory of B.P. was supported by grants U54AI057157, R37AI48638, R01DK057665, U19AI057266, HHSN266200700006C, NO1 AI50025 and U19AI090023 from the National Institutes of Health and a grant from the Bill & Melinda Gates Foundation. Work in A.G.-S. laboratories was partly funded by grants HHSN266200700010C, U54AI57158) and U01AI070469 from the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
S.P.K. and B.P. designed the study, planned the experiments and analysed the data. B.P. and S.P.K. wrote the manuscript. S.P.K., I.S. and B.P. designed and performed the H1N1 vaccine studies in mice and primates. D.K. assisted with the H1N1 vaccine studies in mice and primates. R.A.A., A.G.-S. and J.S. designed and performed the neutralization assays and challenge experiments with H5N1 vaccine studies in mice. T.H. and R.R. assisted with experiments. H.I.N. performed the microarray analysis. S.S. and M.A. designed and carried out the SPR-based avidity experiments. M.K. assisted with design and execution of mice and non-human primate experiments. N.M. assisted with design of formulations. J.J. assisted with immunohistochemistry and design of experiments. R.J.H. expressed and purified the recombinant H5HA protein. R.C. helped plan and design the H1N1 vaccine study in mice and primates.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Figures
The file contains Supplementary Figures 1-18 with legends. (PDF 3185 kb)
Rights and permissions
About this article
Cite this article
Kasturi, S., Skountzou, I., Albrecht, R. et al. Programming the magnitude and persistence of antibody responses with innate immunity. Nature 470, 543–547 (2011). https://doi.org/10.1038/nature09737
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature09737
This article is cited by
-
Emerging role of microbiota derived outer membrane vesicles to preventive, therapeutic and diagnostic proposes
Infectious Agents and Cancer (2023)
-
SREBP signaling is essential for effective B cell responses
Nature Immunology (2023)
-
A nanoadjuvant that dynamically coordinates innate immune stimuli activation enhances cancer immunotherapy and reduces immune cell exhaustion
Nature Nanotechnology (2023)
-
Immunization against Zika by entrapping live virus in a subcutaneous self-adjuvanting hydrogel
Nature Biomedical Engineering (2023)
-
Toll-like receptor mediated inflammation directs B cells towards protective antiviral extrafollicular responses
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.