Abstract
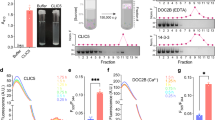
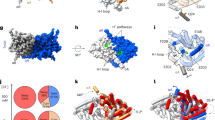
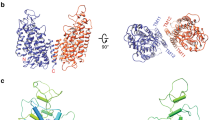
Channels and transporters of the ClC family cause the transmembrane movement of inorganic anions in service of a variety of biological tasks, from the unusual—the generation of the kilowatt pulses with which electric fish stun their prey—to the quotidian—the acidification of endosomes, vacuoles and lysosomes1. The homodimeric architecture of ClC proteins, initially inferred from single-molecule studies of an elasmobranch Cl− channel2 and later confirmed by crystal structures of bacterial Cl−/H+ antiporters3,4, is apparently universal. Moreover, the basic machinery that enables ion movement through these proteins—the aqueous pores for anion diffusion in the channels and the ion-coupling chambers that coordinate Cl− and H+ antiport in the transporters—are contained wholly within each subunit of the homodimer. The near-normal function of a bacterial ClC transporter straitjacketed by covalent crosslinks across the dimer interface and the behaviour of a concatemeric human homologue argue that the transport cycle resides within each subunit and does not require rigid-body rearrangements between subunits5,6. However, this evidence is only inferential, and because examples are known in which quaternary rearrangements of extramembrane ClC domains that contribute to dimerization modulate transport activity7, we cannot declare as definitive a ‘parallel-pathways’ picture in which the homodimer consists of two single-subunit transporters operating independently. A strong prediction of such a view is that it should in principle be possible to obtain a monomeric ClC. Here we exploit the known structure of a ClC Cl−/H+ exchanger, ClC-ec1 from Escherichia coli, to design mutants that destabilize the dimer interface while preserving both the structure and the transport function of individual subunits. The results demonstrate that the ClC subunit alone is the basic functional unit for transport and that cross-subunit interaction is not required for Cl−/H+ exchange in ClC transporters.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Jentsch, T. J. et al. Physiological functions of CLC Cl− channels gleaned from human genetic disease and mouse models. Annu. Rev. Physiol. 67, 779–807 (2005)
Middleton, R. E., Pheasant, D. J. & Miller, C. Reconstitution of detergent-solubilized Cl− channels and analysis by concentrative uptake of 36Cl− and planar lipid bilayers. Methods 6, 28–36 (1994)
Dutzler, R. et al. X-ray structure of a ClC chloride channel at 3.0 Å reveals the molecular basis of anion selectivity. Nature 415, 287–294 (2002)
Dutzler, R., Campbell, E. B. & MacKinnon, R. Gating the selectivity filter in ClC chloride channels. Science 300, 108–112 (2003)
Nguitragool, W. & Miller, C. CLC Cl−/H+ transporters constrained by covalent cross-linking. Proc. Natl Acad. Sci. USA 104, 20659–20665 (2007)
Zdebik, A. A. et al. Determinants of anion-proton coupling in mammalian endosomal CLC proteins. J. Biol. Chem. 283, 4219–4227 (2008)
Bykova, E. A. et al. Large movement in the C terminus of CLC-0 chloride channel during slow gating. Nature Struct. Mol. Biol. 13, 1115–1119 (2006)
Fleming, K. G., Ackerman, A. L. & Engelman, D. A. The effect of point mutations on the free energy of transmembrane α-helix dimerization. J. Mol. Biol. 272, 266–275 (1997)
MacKenzie, K. R. & Fleming, K. G. Association energetics of membrane spanning α-helices. Curr. Opin. Struct. Biol. 18, 412–419 (2008)
Chen, L. et al. Energetics of ErbB1 transmembrane domain dimerization in lipid bilayers. Biophys. J. 96, 4622–4630 (2009)
Yau, W. M. et al. The preference of tryptophan for membrane interfaces. Biochemistry 37, 14713–14718 (1998)
Fang, Y., Kolmakova-Partensky, L. & Miller, C. A bacterial arginine-agmatine exchange transporter involved in extreme acid resistance. J. Biol. Chem. 282, 176–182 (2007)
Maduke, M., Pheasant, D. J. & Miller, C. High-level expression, functional reconstitution, and quaternary structure of a prokaryotic ClC-type chloride channel. J. Gen. Physiol. 114, 713–722 (1999)
Walden, M. et al. Uncoupling and turnover in a Cl−/H+ exchange transporter. J. Gen. Physiol. 129, 317–329 (2007)
Accardi, A. & Miller, C. Secondary active transport mediated by a prokaryotic homologue of ClC Cl- channels. Nature 427, 803–807 (2004)
Miller, C. & Nguitragool, W. A provisional transport mechanism for a chloride channel-type Cl−/H+ exchanger. Phil. Trans. R. Soc. B 364, 175–180 (2008)
Elvington, S. M., Liu, C. W. & Maduke, M. C. Substrate-driven conformational changes in ClC-ec1 observed by fluorine NMR. EMBO J. 28, 3090–3102 (2009)
Accardi, A. et al. Synergism between halide binding and proton transport in a CLC-type exchanger. J. Mol. Biol. 362, 691–699 (2006)
Nguitragool, W. & Miller, C. Uncoupling of a CLC Cl−/H+ exchange transporter by polyatomic anions. J. Mol. Biol. 362, 682–690 (2006)
Fu, D. et al. Structure of a glycerol-conducting channel and the basis for its selectivity. Science 290, 481–486 (2000)
Wang, Y. et al. Structure of the formate transporter FocA reveals a pentameric aquaporin-like channel. Nature 462, 467–472 (2009)
Waight, A. B., Love, J. & Wang, D. N. Structure and mechanism of a pentameric formate channel. Nature Struct. Mol. Biol. 17, 31–37 (2010)
Levin, E. J., Quick, M. & Zhou, M. Crystal structure of a bacterial homologue of the kidney urea transporter. Nature 462, 757–761 (2009)
Khademi, S. et al. Mechanism of ammonia transport by Amt/MEP/Rh: structure of AmtB at 1.35 Å. Science 305, 1587–1594 (2004)
Zheng, L. et al. The mechanism of ammonia transport based on the crystal structure of AmtB of Escherichia coli . Proc. Natl Acad. Sci. USA 101, 17090–17095 (2004)
Cowan, S. W. et al. Crystal structures explain functional properties of two E. coli porins. Nature 358, 727–733 (1992)
Shaffer, P. L. et al. Structure and mechanism of a Na+-independent amino acid transporter. Science 325, 1010–1014 (2009)
Theobald, D. L. & Miller, C. Membrane transport proteins: surprises in structural sameness. Nature Struct. Mol. Biol. 17, 2–3 (2010)
Reyes, N., Ginter, C. & Boudker, O. Transport mechanism of a bacterial homologue of glutamate transporters. Nature 462, 880–885 (2009)
Metcalf, D. G. et al. Multiple approaches converge on the structure of the integrin αIIb/β3 transmembrane heterodimer. J. Mol. Biol. 392, 1087–1101 (2009)
Acknowledgements
We are grateful to the scientists at beamline 8.2.1 of the Advanced Light Source, Lawrence Berkeley Laboratory, for much help and advice, to H. Jayaram for help with refinement, and to R. Sah and D. Theobald for comments on the manuscript.
Author information
Authors and Affiliations
Contributions
Experiments were designed by J.L.R. and C.M. and carried out by J.L.R. and L.K.-P., and the manuscript was written by all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-3 with legends and Supplementary Tables 1-2. (PDF 561 kb)
Rights and permissions
About this article
Cite this article
Robertson, J., Kolmakova-Partensky, L. & Miller, C. Design, function and structure of a monomeric ClC transporter. Nature 468, 844–847 (2010). https://doi.org/10.1038/nature09556
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature09556
This article is cited by
-
Structural basis of pH-dependent activation in a CLC transporter
Nature Structural & Molecular Biology (2024)
-
A mechanical-coupling mechanism in OSCA/TMEM63 channel mechanosensitivity
Nature Communications (2023)
-
Comparison of the functional properties of trimeric and monomeric CaiT of Escherichia coli
Scientific Reports (2019)
-
Unfolding of a ClC chloride transporter retains memory of its evolutionary history
Nature Chemical Biology (2018)
-
Dimeric structure of the uracil:proton symporter UraA provides mechanistic insights into the SLC4/23/26 transporters
Cell Research (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.