Abstract
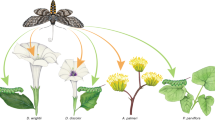
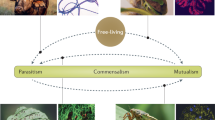
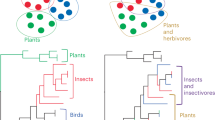
Ecological interactions are crucial to understanding both the ecology and the evolution of organisms1,2. Because the phenotypic traits regulating species interactions are largely a legacy of their ancestors, it is widely assumed that ecological interactions are phylogenetically conserved, with closely related species interacting with similar partners2. However, the existing empirical evidence is inadequate to appropriately evaluate the hypothesis of phylogenetic conservatism in ecological interactions, because it is both ecologically and taxonomically biased. In fact, most studies on the evolution of ecological interactions have focused on specialized organisms, such as some parasites or insect herbivores3,4,5,6,7, belonging to a limited subset of the overall tree of life. Here we study the evolution of host use in a large and diverse group of interactions comprising both specialist and generalist acellular, unicellular and multicellular organisms. We show that, as previously found for specialized interactions, generalized interactions can be evolutionarily conserved. Significant phylogenetic conservatism of interaction patterns was equally likely to occur in symbiotic and non-symbiotic interactions, as well as in mutualistic and antagonistic interactions. Host-use differentiation among species was higher in phylogenetically conserved clades, irrespective of their generalization degree and taxonomic position within the tree of life. Our findings strongly suggest a shared pattern in the organization of biological systems through evolutionary time, mediated by marked conservatism of ecological interactions among taxa.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Price, P. W. Macroevolutionary Theory on Macroecological Patterns (Cambridge Univ. Press, 2003)
Thompson, J. N. The Geographic Mosaic of Coevolution (Univ. Chicago Press, 2005)
Sasal, P., Desdevises, Y. & Morand, S. Host-specialization and species diversity in fish parasites: phylogenetic conservatism? Ecography 21, 639–643 (1998)
Jackson, A. P. & Charleston, M. A. A cophylogenetic perspective of RNA–virus evolution. Mol. Biol. Evol. 21, 45–57 (2004)
Holmes, E. C. The Evolution and Emergence of RNA Viruses (Oxford Univ. Press, 2009)
Gilbert, G. S. & Webb, C. O. Phylogenetic signal in plant pathogen-host range. Proc. Natl Acad. Sci. USA 104, 4979–4983 (2007)
Winkler, I. S. & Mitter, C. in Specialization, Speciation and Radiation: The Evolutionary Biology of Herbivorous Insects (ed. Tilmon, K. J.) 240–263 (Univ. California Press, 2007)
Freckleton, R. P., Harvey, P. H. & Pagel, M. Phylogenetic analysis and comparative data: a test and review of evidence. Am. Nat. 160, 712–726 (2002)
Wiens, J. J. & Graham, C. H. Niche conservatism: integrating evolution, ecology, and conservation biology. Annu. Rev. Ecol. Evol. Syst. 36, 519–539 (2005)
Darwin, C. On the Origin of Species 78–79 (Murray, 1859)
Chase, J. M. & Leibold, M. A. Ecological Niches: Linking Classical and Contemporary Approaches 19–45 (Univ. Chicago Press, 2003)
Revell, L. J., Harmon, L. J. & Collar, D. C. Phylogenetic signal, evolutionary process, and rate. Syst. Biol. 57, 591–601 (2008)
Dunne, J. A., Williams, R. J. & Martinez, N. D. Food-web structure and network theory: the role of connectance and size. Proc. Natl Acad. Sci. USA 99, 12917–12922 (2002)
Bascompte, J. & Jordano, P. Plant-animal mutualistic networks: the architecture of biodiversity. Annu. Rev. Ecol. Evol. Syst. 38, 567–593 (2007)
Guimerà, R. & Amaral, L. A. N. Functional cartography of complex metabolic networks. Nature 433, 895–900 (2005)
Guimerà, R. & Amaral, L. A. N. Cartography of complex networks: modules and universal roles. J. Stat. Mech. P02001 (2005)
Olesen, J. M., Bascompte, J., Dupont, Y. L. & Jordano, P. The modularity of pollination networks. Proc. Natl Acad. Sci. USA 104, 19891–19896 (2007)
Rezende, E. L., Lavabre, J. E., Guimarães, P. R., Jordano, P. & Bascompte, J. Non-random coextinctions in phylogenetically structured mutualistic networks. Nature 448, 925–928 (2007)
Rezende, E. L., Albert, E. M., Fortuna, M. A. & Bascompte, J. Compartments in a marine food web associated with phylogeny, body mass, and habitat structure. Ecol. Lett. 12, 779–788 (2009)
Oksanen, J. Multivariate Analysis of Ecological Communities in R: Vegan Tutorial (R Project for Statistical Computing, 2008)
Blomberg, S. P., Garland, T. & Ives, A. R. Testing for phylogenetic signal in comparative data: behavioral traits are more labile. Evolution 57, 717–745 (2003)
Pride, D. T., Wassenaar, T. M., Ghose, C. & Blaser, M. J. Evidence of host-virus co-evolution in tetranucleotide usage patterns of bacteriophages and eukaryotic viruses. BMC Genomics 7, 8 (2006)
Price, P. W. in Specialization, Speciation and Radiation: The Evolutionary Biology of Herbivorous Insects (ed. Tilmon, K. J.) 174–187 (Univ. California Press, 2007)
Refrégier, G. et al. Cophylogeny of the anther smut fungi and their caryophyllaceous hosts: prevalence of host shifts and importance of delimiting parasite species for inferring cospeciation. BMC Evol. Biol. 8, 100 (2008)
Maddison, W. P. & Slatkin, M. Null models for the number of evolutionary steps in a character on a phylogenetic tree. Evolution 45, 1184–1197 (1991)
de Nooy, W., Mrvar, A. & Batagelj, V. Exploratory Social Network Analysis with Pajek: Structural Analysis in the Social Sciences (Cambridge Univ. Press, 2005)
Guimerà, R., Sales-Pardo, M. & Amaral, L. A. N. Modularity from fluctuations in random graphs and complex networks. Phys. Rev. E 70, 025101 (2004)
Pagel, M. Inferring the historical patterns of biological evolution. Nature 401, 877–884 (1999)
Maddison, W. P. & Maddison, D. R. Mesquite: a modular system for evolutionary analysis. Mesquite 〈http://mesquiteproject.org〉 (2009)
Acknowledgements
We thank J. Bascompte, J. Bosch, A. González-Megías, P. Jordano, M. Lineham, M. Méndez, I. Reche, E.W. Schupp and S. Strauss for comments on a previous draft, R. Guimerà for kindly providing NETCARTO software, and B. Krasnov, C. Mitter, L. Navarro, J. Ollerton and J. M. Pleguezuelos for providing access to their data set. This work was funded by the Spanish Ministry of Science (J.M.G., M.V. and F.P.) and by the Junta de Andalucía (J.M.G. and F.P.)
Author information
Authors and Affiliations
Contributions
J.M.G., M.V. and F. P. designed the study, J.M.G. compiled the data set and performed the analysis of host use, M.V. performed the phylogenetic analyses, J.M.G. wrote a first version of the manuscript and all authors contributed to the final draft.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Tables 1-6, Supplementary Figure 1 with legend, Supplementary Data 1and Supplementary Notes 1-116 with References. (PDF 897 kb)
Rights and permissions
About this article
Cite this article
Gómez, J., Verdú, M. & Perfectti, F. Ecological interactions are evolutionarily conserved across the entire tree of life. Nature 465, 918–921 (2010). https://doi.org/10.1038/nature09113
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature09113
This article is cited by
-
Changes in herbivory patterns and insect herbivore assemblages associated to canopy of Quercus laurina: importance of oak species diversity and foliar chemical defense
Trees (2023)
-
Effects of tree diversity on insect herbivory
Journal of Forestry Research (2022)
-
Exploring trophic role similarity and phylogenetic relatedness between species in food webs
Community Ecology (2021)
-
Within-individual phenotypic plasticity in flowers fosters pollination niche shift
Nature Communications (2020)
-
Evolutionary conservation of within-family biodiversity patterns
Nature Communications (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.