Abstract
Topoisomerases regulate DNA topology and are fundamental to many aspects of chromosome metabolism1,2. Their activity involves the transient cleavage of DNA, which, if it occurs near sites of endogenous DNA damage or in the presence of topoisomerase poisons, can result in abortive topoisomerase-induced DNA strand breaks3,4,5. These breaks feature covalent linkage of the enzyme to the DNA termini by a 3′- or 5′-phosphotyrosyl bond and are implicated in hereditary human disease6,7,8, chromosomal instability and cancer4,9, and underlie the clinical efficacy of an important class of anti-tumour poisons3,9,10. The importance of liberating DNA termini from trapped topoisomerase is illustrated by the progressive neurodegenerative disease observed in individuals containing a mutation in tyrosyl-DNA phosphodiesterase 1 (TDP1), an enzyme that cleaves 3′-phosphotyrosyl bonds6,7,8. However, a complementary human enzyme that cleaves 5′-phosphotyrosyl bonds has not been reported, despite the effect of DNA double-strand breaks containing such termini on chromosome instability and cancer6,7,8. Here we identify such an enzyme in human cells and show that this activity efficiently restores 5′-phosphate termini at DNA double-strand breaks in preparation for DNA ligation. This enzyme, TTRAP, is a member of the Mg2+/Mn2+-dependent family of phosphodiesterases. Cellular depletion of TTRAP results in increased susceptibility and sensitivity to topoisomerase-II-induced DNA double-strand breaks. TTRAP is, to our knowledge, the first human 5′-tyrosyl DNA phosphodiesterase to be identified, and we suggest that this enzyme is denoted tyrosyl DNA phosphodiesterase-2 (TDP2).
Similar content being viewed by others
Main
To identify new human tyrosyl DNA phosphodiesterase activities, we exploited the hypersensitivity of Saccharomyces cerevisiae tdp1Δ rad1Δ double-mutant cells to camptothecin (CPT), a topoisomerase I (Top1) poison that induces single-strand breaks (SSBs) with Top1 covalently linked to the 3′-terminus3,10. This strain lacks not only Tdp1 but also Rad1-Rad10 nuclease, which in yeast provides an alternative (endonucleolytic) pathway for removing Top1 from 3′-termini11,12. We transformed this strain with a human complementary DNA library and screened the resulting population of transformants for cellular resistance to CPT. Of six tdp1Δ rad1Δ transformants showing wild-type levels of CPT resistance, three contained cDNA clones encoding TDP1 and three contained cDNA clones encoding TTRAP (TRAF and TNF receptor-associated protein), a protein of unknown function and a putative member of the Mg2+/Mn2+-dependent phosphodiesterase superfamily, with the DNA repair protein apurinic/apyrimidinic (AP) endonuclease-1 (APE-1, also known as APEX1) being its closest relative13,14 (Fig. 1a and Supplementary Fig. 1).
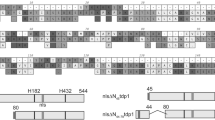
a, Cartoon of TTRAP. b, TTRAP suppresses CPT sensitivity of tdp1Δ rad1Δ yeast. Tenfold serial dilutions of wild-type (WT, left) and tdp1Δ rad1Δ (right) cells containing pACT-TDP1-1, pACT-TTRAP-2 or pACT vector were plated onto media lacking (-) or containing (+) 20 μM CPT. c, Human APE-1 or full length TTRAP suppress the sensitivity of tdp1Δ rad1Δ or apn1Δ apn2Δ tpp1Δ cells to CPT or MMS, respectively. Tenfold serial dilutions of tdp1Δ rad1Δ (left) and apn1 Δapn2Δ tpp1Δ (right) transformed with pGBKT7-TTRAP, pAS-APE-1 or pGBKT7 vector were plated onto media lacking or containing 20 μM CPT or 100 μM MMS. d, Mutant TTRAP fails to suppress CPT sensitivity. Wild-type (left) or tdp1Δ rad1Δ (right) cells transformed with pVTU260 vector or pVTU260 encoding full-length untagged wild-type or mutant (E152A or D262A) TTRAP were analysed for CPT sensitivity as above.
The TDP1 and TTRAP cDNA clones recovered in the genetic screen suppressed the CPT sensitivity of tdp1Δ rad1Δ cells to a similar extent (Fig. 1b and data not shown). Although the pACT-TTRAP clones encoded TTRAP protein that lacked eight (pACT-TTRAP-2, pACT-TTRAP-3) or twenty-two (pACT-TTRAP-1) residues from the amino terminus (data not shown), full-length TTRAP similarly suppressed the CPT sensitivity of tdp1Δ rad1Δ (Fig. 1c, left). In contrast, human APE-1 protein failed to suppress this sensitivity, suggesting that the ability to complement CPT sensitivity in tdp1Δ rad1Δ cells is not a generic feature of metal-dependent phosphodiesterases (Fig. 1c, left). Conversely, whereas human APE-1 suppressed the sensitivity of AP endonuclease-defective apn1Δ apn2Δ tpp1Δ yeast cells to methyl methanesulphonate (MMS)-induced DNA base damage, human TTRAP did not, suggesting that the effect of TTRAP in these experiments was restricted to topoisomerase-mediated DNA damage (Fig. 1c, right). TTRAP contains four highly conserved motifs that putatively assign this protein to the metal-dependent phosphodiesterase superfamily (see Fig. 1a and Supplementary Fig. 1). We thus examined whether mutation of two predicted catalytic residues (Fig. 1a; Glu 152 and Asp 262) within two of these motifs affected the complementation of CPT sensitivity by TTRAP. Indeed, in contrast to wild-type TTRAP protein, neither TTRAP(E152A) nor TTRAP(D262A) restored CPT resistance in tdp1Δ rad1Δ cells (Fig. 1d).
We next purified recombinant human TTRAP, TDP1 and APE-1 from Escherichia coli (Fig. 2a, lanes 1–3), and incubated the purified proteins with a radiolabelled oligonucleotide duplex containing a nick with a single tyrosine covalently linked to the 3′-terminus by a phosphotyrosyl bond (Fig. 2a, inset). This is an established substrate for TDP1 that mimics the Top1-linked SSBs induced by CPT6,8. As expected, TDP1 cleaved the 3′-phosphotyrosyl bond and thereby converted the 3′-tyrosine terminus to a 3′-phosphate (Fig. 2a, lane 5). In contrast, recombinant human APE-1 failed to do so (Fig. 2a, lane 7). Notably, TTRAP cleaved the 3′-phosphotyrosyl bond, liberating DNA with a 3′-phosphate in a manner similar to TDP1, albeit at much higher enzyme concentrations and less efficiently (Fig. 2a, lane 6). TTRAP was also active on a double-strand break (DSB) substrate containing a 3′-tyrosine (Fig. 2b), albeit ∼50-fold less efficiently and much slower than TDP1 (Fig. 2d, left panel and Supplementary Fig. 2). This activity was greatly reduced or lacking in the absence of Mg2+, consistent with a dependency on metal cofactor for TTRAP activity (Supplementary Fig. 2). Moreover, the TTRAP activity detected here did not reflect a contaminant, because recombinant preparations of TTRAP(E152A) and TTRAP(D262A), purified in parallel with wild-type TTRAP, were largely or entirely inactive (Fig. 2b, lanes 6 and 7). We thus conclude that TTRAP possesses bona fide 3′-tyrosyl DNA phosphodiesterase activity, but that this activity is much weaker than that of TDP1, under the current experimental conditions at least.
a, 3′-TDP activity of TTRAP at SSBs. Lanes 1–3, SDS–PAGE of recombinant proteins. Lanes 4–7, 32P-radiolabelled 43-base-pair (bp) oligonucleotide duplex (50 nM) containing a nick with a tyrosine (Y) linked to the 3′-terminus of the labelled 18-bp oligonucleotide (inset, top) was incubated for 1 h at 37 °C in the absence or presence of the indicated human proteins. Positions of 32P-radiolabelled substrate (18-Y) and product (18-P) are indicated. An 18-bp oligonucleotide with a 3′-OH terminus (18-OH) was included as a marker (3′OH). b, 3′-TDP activity of TTRAP at DSBs. Lanes 1–3, analysis of human TTRAP (wild type), TTRAP(D262A) (D), or TTRAP(E152A) (E) by SDS–PAGE. Lanes 4–8, 32P-radiolabelled 18-bp oligonucleotide duplex (50 nM) containing a DSB with a tyrosine linked covalently to the 3′-terminus of the labelled 18-bp oligonucleotide (inset, top) was incubated as above in the absence or presence of the indicated proteins (1 μM). c, 5′-TDP activity of TTRAP at DSBs. 32P-radiolabelled 19-bp oligonucleotide duplex (50 nM) containing a DSB with a tyrosine linked covalently to the 5′-terminus of the labelled 19-bp oligonucleotide (inset, top) was incubated with of the indicated human proteins (150 nM) as above. Positions of 32P-radiolabelled substrate (Y-19) and repair product (P-19) are indicated. A 19-bp oligonucleotide with a 5′-OH terminus (OH-19) was included as a marker (5′OH). d, Left, the 3′-TDP substrate (50 nM) in b was incubated with human TDP1 (0, 4.5, 9, 36 or 90 nM) or TTRAP (0, 36, 90, 225, 450 or 900 nM) for 1 h at 37 °C. Right, the 5′-TDP substrate (50 nM) in c was incubated with TTRAP (0, 15, 30, 60 or 240 nM) or 400 nM TDP1 for 1 h at 37 °C. Reaction products were quantified using ImageQuant and data are the mean ± s.e.m. of three experiments.
In contrast to Top1, topoisomerase II (Top2) induces DNA DSBs in which the topoisomerase is covalently linked by a phosphotyrosyl bond to the 5′-terminus of the break9. Surprisingly, a human enzyme that can preferentially or efficiently cleave DNA 5′-phosphotyrosyl bonds has not been reported, despite the established affect of this type of break on genetic stability and cancer4,9. Given that the 3′-tyrosyl DNA phosphodiesterase activity of TTRAP was relatively weak, we wondered whether this enzyme might be a 5′-tyrosyl DNA phosphodiesterase. To address this possibility we used a radiolabelled oligonucleotide duplex containing a DSB with a single tyrosine covalently linked to the 5′-terminus by a phosphotyrosyl bond (Fig. 2c, inset). Notably, whereas neither human APE-1 nor TDP1 cleaved the 5′-phosphotyrosyl bond, TTRAP led to the complete conversion of the DSB terminus to a 5′-phosphate (Fig. 2c, lanes 2–4). That the 5′-terminus of the TTRAP reaction product was a phosphate was confirmed using calf intestinal phosphatase, which converted the 5′-terminus of the reaction product to hydroxyl (Supplementary Fig. 3a). Notably, TTRAP activity on the 5′-phosphotyrosyl substrate was largely or entirely absent in reactions that lacked Mg2+ and contained EDTA (Supplementary Fig. 3b), or in reactions containing the phosphodiesterase mutant proteins TTRAP(E152A) or TTRAP(D262A) (Fig. 2c, lanes 5 and 6). Importantly, similar to TDP1 activity on 3′-phosphotyrosyl termini, the activity of TTRAP on 5′-phosphotyrosyl termini was robust and rapid (Fig. 2d and Supplementary Fig. 3c, d). In contrast, TTRAP was not active on a substrate containing an internal abasic site, a preferred substrate of APE-1, or on 5′-AMP DNA termini, a preferred substrate of aprataxin (Supplementary Fig. 4). We conclude from these experiments that TTRAP is a bona fide 5′-tyrosyl DNA phosphodiesterase.
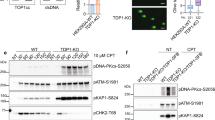
We next examined whether we could detect 5′-tyrosyl DNA phosphodiesterase activity in human whole-cell extracts. Indeed, such activity was readily detected in A549 cell extract (Fig. 3, lanes 1–5). Moreover, this activity was reduced by ∼50% in A549 cells in which TTRAP was depleted by ∼80% by RNA interference (RNAi), and was increased by the addition of recombinant human TTRAP (Fig. 3). However, 5′-tyrosyl DNA phosphodiesterase activity in TTRAP-depleted extracts was not increased by the addition of recombinant TTRAP(D262A) or TTRAP(E152A) (Supplementary Fig. 5). Notably, wild-type and TTRAP-depleted A549 extracts showed similar levels of metal-independent 3′-tyrosyl DNA phosphodiesterase activity, and were equally able to repair a gapped DNA duplex that lacked damaged termini, suggesting that TTRAP-depleted extracts are competent for other types of DNA repair activity (Supplementary Fig. 5). These data indicate that human whole-cell extract possesses robust 5′-tyrosyl DNA phosphodiesterase activity, and suggest that TTRAP is a major source of this activity.
Extract from A549 cells transfected with pSuper (pS) or pSuper-TTRAP (pS-TT) was incubated with a 32P-radiolabelled 20-bp oligonucleotide duplex containing a DSB with a tyrosine linked covalently to the 5′-terminus of the labelled 20-bp oligonucleotide (inset, top). Reaction products were analysed by denaturing PAGE/phosphorimaging. The position of 32P-radiolabelled substrate (Y-20), repair product (P-20), and 20-bp oligonucleotide 5′-OH marker (OH-20) are indicated. The percentage of substrate converted to reaction product was quantified (mean + s.e.m., n = 4) using ImageQuant (bottom right). Top right, western blot of XRCC1 (X) and TTRAP (TT) in the cell extracts used in these experiments. Asterisk denotes a nonspecific band.
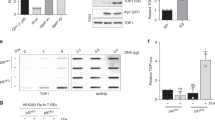
Finally, we examined whether the 5′-tyrosyl DNA phosphodiesterase activity of TTRAP fulfils a role in protecting against topoisomerase-induced DNA damage, in vivo. We first noted that the overexpression of TTRAP, but not of either TDP1 or the mutant proteins TTRAP(D262A) or TTRAP(E152A), measurably increased the resistance of ISE2 cells—a strain of S. cerevisiae with increased drug permeability15—to the Top2 poison etoposide (Fig. 4a and Supplementary Fig. 6). We also noted that enhanced green fluorescent protein (EGFP)-tagged TTRAP accumulated in nuclear promyelocytic leukaemia (PML) bodies, as reported previously16, and more importantly at sites of laser-induced UVA damage (Fig. 4b), consistent with a role for TTRAP in DNA repair. That this accumulation was relatively weak may reflect the fact that topoisomerase-induced DNA breaks are probably only a minor component of UVA laser-induced DNA damage17,18,19. We next examined the effect of TTRAP depletion on the sensitivity of human A549 cells to different types of DNA damage. Whereas TTRAP-depleted cells (see Fig. 3 for a typical level of depletion) had little or no sensitivity to CPT or MMS, these cells were hypersensitive to etoposide (Supplementary Fig. 7). These data suggest that TTRAP promotes the repair of Top2-induced DSBs in human cells. To explore this possibility further, we examined whether TTRAP depletion resulted in the accumulation of DNA strand breaks during exposure to topoisomerase poisons. Notably, TTRAP depletion failed to increase the accumulation of DNA strand breaks during incubation with CPT (a Top1 poison), as measured either by alkaline-comet assays (Supplementary Fig. 7) or by the appearance of γ-H2AX (also known as γ-H2FAX) foci (Fig. 4c, left). This contrasted with the >10-fold increase in DNA strand breaks observed in mouse embryonic fibroblasts (MEFs) lacking Tdp1, which is the primary cellular 3′-tyrosyl DNA phosphodiesterase activity (Supplementary Fig. 7). However, treatment with etoposide resulted in the accumulation of ∼twofold more γ-H2AX foci in TTRAP-depleted cells than in mock-depleted control cells (Fig. 4c, left). Furthermore, the overexpression of wild-type TTRAP, but notably not of TTRAP(D262A) or TTRAP(E152A), reduced levels of etoposide-induced γ-H2AX foci in transiently transfected A549 cells (Fig. 4c, right and Supplementary Fig. 7). Together, these data support the notion that the 5′-tyrosyl DNA phosphodiesterase activity of TTRAP is required for the efficient repair of Top2-induced chromosomal DSBs.
a, TTRAP increases etoposide resistance in yeast. Tenfold serial dilutions of ISE2 S. cerevisiae transformed with pACT-TDP1-1, pACT-TTRAP-2 or pACT vector were plated onto medium containing 500 μM etoposide. b, GFP–TTRAP accumulates at sites of UVA damage. Human A549 cells transiently transfected with pEGFP–TTRAP were damaged with a UVA laser and EGFP was imaged at the indicated times (min) (arrowheads denote position of laser track). c, TTRAP-depletion increases accumulation of etoposide-induced DSBs. A549 cells transfected with pSuper (pS) or pSuper-TTRAP (pS-TT) were treated with CPT (10 μM) or etoposide (etop, 10 μM) for 16 h and the average number of γ-H2AX foci per cell was quantified. Right, TTRAP overexpression decreases the accumulation of etoposide-induced DSBs. A549 cells transiently transfected with pcDNA3.1His-TTRAP, pcDNA3.1His-TTRAP(D262A) (D), or pcDNA3.1His-TTRAP(E152A) (E) were treated with etoposide as above and the average number of γ-H2AX foci quantified in cells either overexpressing recombinant TTRAP (open bars) or lacking visible recombinant TTRAP (filled bars). Data are the mean ± s.e.m.; n = 3). Asterisks denote statistically significant (paired t-test P ≤ 0.05) differences.
It is noteworthy that the 5′-tyrosyl DNA phosphodiesterase activity of TTRAP can enable the repair of Top2-induced DSBs without the need for nuclease activity, creating a ‘clean’ DSB with 5′-phosphate termini that are ready for ligation. This contrasts with currently established mechanisms for DSB repair, which involve structure-specific nucleases such as CtIP (also known as RBBP8) and the MRE11–RAD50–NBS1 (MRN; NBS1 is also known as NBN) protein complex to ‘trim’ DSB termini20,21. TTRAP may thus provide an ‘error-free’ mechanism for direct end-joining of Top2-induced DSBs, a process that might have particular utility for maintaining genetic stability in long-lived non-cycling cells.
In summary, we have shown that human TTRAP is a 5′-tyrosyl DNA phosphodiesterase that is required for efficient repair of Top2-induced DNA double-strand breaks. Because TDP1 and TTRAP are complementary activities, together providing cells with an ability to remove trapped topoisomerase from both 3′- and 5′-DNA termini, we suggest that TTRAP is also designated tyrosyl DNA phosphodiesterase-2 (TDP2). It will now be of interest to determine how the tyrosyl DNA phosphodiesterase activity of TTRAP relates to the reported involvement of this protein in transcriptional regulation, apoptosis and embryonic development13,22,23,24. Given the central role fulfilled by topoisomerases in chromosome metabolism, our findings suggest that TTRAP may have other important functions in many aspects of cell biology, including the suppression of chromosome instability and cancer.
Methods Summary
Human TTRAP was recovered in a genetic screen for restoration of CPT resistance in tdp1Δ rad1Δ budding yeast cells by transformation with 16 μg of a pACT human cDNA library and plated on appropriate medium supplemented with 20 μM CPT (Sigma). For sensitivity assays, transformants were resuspended in 200 μl sterile H2O and 10 μl drops of tenfold serial dilutions were plated onto medium lacking or containing the appropriate drug. The generation of oligonucleotide duplexes used for in vitro assays, DNA constructs used for expression of TTRAP in yeast, E. coli and mammalian cells, and the pSuper constructs used for short hairpin RNA (shRNA)-mediated depletion of TTRAP, are described in detail in the Methods. The recombinant human proteins APE-1, TDP1 and TTRAP used for in vitro assays were expressed in E. coli and purified by immobilised metal-chelate chromatagoraphy (IMAC) followed by ion-exchange chromatography. The products of in vitro reactions were fractionated by denaturing PAGE and detected by phosphorimaging. The detection of EGFP–TTRAP accumulation at sites of UVA laser damage was achieved by seeding transfected A549 cells onto glass-bottom coverslips, followed by incubation with Hoechst 33258 dye and irradiation with a 351-nm UVA laser (approximately 0.35 J m-2) using a Zeiss Axiovert. For clonogenic survival assays, A549 cells were plated into 10-cm dishes in duplicate and incubated at 37 °C for 4 h. Cells were then mock-treated or treated in medium at 37 °C with genotoxin (Sigma) for 1–3 h, and then incubated at 37 °C in drug-free medium for 14 days to allow the formation of macroscopic colonies. DNA strand breaks were quantified using the alkaline comet assay and by the quantification of γ-H2AX foci.
Online Methods
Yeast strains, screening and sensitivity assays
All yeast strains used in this study have been previously described. Wild type (YW465), tdp1Δ rad1Δ (YW812) and apn1Δ apn2Δ tpp1Δ (YW778) were provided by T. E. Wilson11,25. The ISE2 (JN362a) strain is a gift from M. Neale, originally obtained from J. Nitiss26. Screening of a human cDNA library was performed as follows. Yeast tdp1Δ rad1Δ cells were transformed with 16 μg of pACT human cDNA library27 and plated on Synthetic Complete medium lacking leucine (SC-Leu) supplemented with 20 μM CPT (Sigma). A total of 7.5 × 105 clones were screened, as estimated from the transformation efficiency on SC-Leu lacking CPT. Healthy-looking colonies were isolated after 7 days, rescreened for growth in 20 μM CPT, and their pACT construct was isolated from E. coli. The phenotype was confirmed after retransforming these constructs into tdpΔ rad1Δ cells. Three full-length TDP1 clones were identified (denoted pACT-TDP1-1, -2 and -3), comprising nucleotides 234–2069, 97–2090 and 1–2101, respectively, of the published transcript variant 1 cDNA (NM_018319). Three incomplete clones of TTRAP were identified, one comprising nucleotides 90–1884 (pACT-TTRAP-1), and two comprising nucleotides 49–1889 (pACT-TTRAP-2 and -3) of the published cDNA sequence (NM_016614.2). For sensitivity assays, equal-sized colonies of the corresponding transformants were resuspended in 200 μl sterile H2O, and 10 μl drops of tenfold serial dilutions were plated onto SC-Leu (for pACT transformants), SC-Tryp (for pAS/pGBKT7 transformants), or SC-Ura (for pVTU260 transformants) media lacking or containing the appropriate drug.
DNA constructs
Full-length TTRAP open-reading frame (ORF), flanked by NdeI and BamHI restriction sites, was cloned into pCRII-TOPO (Invitrogen) after PCR amplification from pACT-TTRAP-2 using the following primers (Operon) (the region of TTRAP missing from pACT-TTRAP-2 and included in the primer is underlined): 5′-AGGAAGCATATGGAGTTGGGGAGTTGCCTGGAGGGCGGGAGGGAGGCGGCGG-3′ (forward) and 5′-TGCAACGGATCCAATCAGGGCAAAACCCACAC-3′ (reverse). Mutations were introduced into pCRII-TTRAP with the Quickchange XL site-directed mutagenesis kit (Stratagene) using primers 5′-CCAGATGTGATATTTCTACAGGCAGTTATTCCCCCATATTATAGC-3′ (forward) and 5′-GCTATAATATGGGGGAATAACTGCCTGTAGAAATATCACATCTGG-3′ (reverse) for TTRAP(E152A), and 5′-GCTACAGTTATATTTGCAGGAGCTACAAATCTAAGGGATCGAGAG-3′ (forward) and 5′-CTCTCGATCCCTTAGATTTGTAGCGCCTGCAAATATAACTGTAGC-3′ (reverse) for TTRAP(D262A). pGBKT7 (Clontech) and pET16b (Novagen) constructs expressing TDP1 were previously described6. pGBKT7 (Clontech) and pET16b (Novagen) constructs expressing wild-type TTRAP, TTRAP(E152A) and TTRAP(E262A) were created by sub-cloning the TTRAP ORF from the appropriate pCRII-TOPO construct using NdeI and EcoRI. Untagged full-length human TTRAP yeast expression vectors were created by cloning the appropriate TTRAP ORF from pCRII-TOPO (NdeI-BamHI) into NheI-BamHI of pVTU260 (Euroscarf, provided by E. Hoffmann). pET16b-APE-1 has been described previously28, and the APE-1 ORF was cloned into BamHI-EcoRI sites of pAS2-1 (Clontech). pEGFP-TTRAP was created by cloning a BamHI-EcoRI fragment of pACT-TTRAP-2 into BglII-EcoRI of pEGFP-C3 (Clontech). Mammalian TTRAP expression constructs were created by cloning the appropriate TTRAP ORF from pCRII-TOPO into pcDNA3.1-HisC (Invitrogen) using EcoRI. For pSuper-TTRAP, the following oligonucleotides (Operon) containing an appropriate 19-base-pair (bp) region of homology to TTRAP (underlined) were annealed and sub-cloned into BglII-HindIII sites of pSuper (OligoEngine): 5′-GATCCCCGTACAGCCCAGATGTGATATTCAAGAGATATCACATCTGGGCTGTACTTTTTA-3′ (forward) and 5′-AGCTTAAAAAGTACAGCCCAGATGTGATATCTCTTGAATATCACATCTGGGCTGTACGGG-3′ (reverse).
Purification of recombinant human proteins and human cell extracts
Amino-terminal histidine-tagged recombinant human TTRAP, TTRAP(D262A), TTRAP(E152A), TDP1 and APE-1 were expressed from pET16b in BL21 (DE3) cells and purified by immobilised metal-chelate chromatography (IMAC) and ion-exchange chromatography. Proteins were then dialysed in buffer comprised of 25 mM Tris-HCl, pH 7.5, 1 mM dithiothreitol (DTT), 10% glycerol, and either 100 mM NaCl or 130 mM KCl. Purified human proteins were quantified on Coomassie-blue-stained polyacrylamide gels by comparison to bovine serum albumin (BSA) standards and verified by Bio-Rad Protein Assay Kit. Human whole-cell extracts were prepared from 3 × 106 A549 cells by lysis in 0.1 ml of 20 mM Tris-HCl, pH 7.5, 10 mM EDTA, 1 mM EGTA, 100 mM NaCl, 1% Triton X-100 and protease inhibitor cocktail (Sigma). The extract was clarified by centrifugation and quantified using a Bio-Rad Protein Assay Kit with BSA used as a standard.
Preparation of DNA substrates and in vitro repair reactions
Gel-purified oligonucleotides were 5′-labelled with 32P using [γ-32P]-ATP and T4 PNK or 3′-labelled using [α-32P]-dCTP and Klenow DNA polymerase as described later. For the 19-bp oligonucleotide DSB 5′-phosphotyrosyl substrate, a 5′-Y-18-bp oligonucleotide (5′-Y-TCCGTTGAAGCCTGCTTT-3′) (Midland Certified Reagent Company) was annealed with a 19-bp oligonucleotide (5′-GAAAGCAGGCTTCAACGGA-3′) and the resulting 1-bp 5′ overhang filled-in with [α-32P]-dCTP and Klenow DNA polymerase. For the 20-bp oligonucleotide 5′-phosphotyrosyl DSB substrate (used in reactions with whole-cell extract) the 5′-Y-18-bp oligonucleotide (above) was annealed to the 20-bp oligonucleotide (5′-AGAAAGCAGGCTTCAACGGA-3′), and the resulting 2-bp 5′ overhang filled-in as described earlier but using [α-32P]-dCTP and ddTTP to inhibit degradation of the radiolabelled oligonucleotide by nonspecific nucleases. For the 43-bp oligonucleotide 3′-phosphotyrosyl SSB (nick) substrate, a radiolabelled 3′-Y-18-bp oligonucleotide (32P-5′-TCCGTTGAAGCCTGCTTT-Y-3′) (provided by H. Nash) was annealed with a 25-bp oligonucleotide (5′-GACATACTAACTTGAGCGAAACGGT-3′) and a 43-bp oligonucleotide (5′-CCGTTTCGCTCAAGTTAGTATGTCAAAGCAGGCTTCAACGGAT-3′). For the 18-bp oligonucleotide 3′-phosphotyrosyl DSB substrate, the radiolabelled 3′-Y-18-bp oligonucleotide (above) was annealed with the 19-bp oligonucleotide (5′-AAAGCAGGCTTCAACGGAT-3′). For the AP endonuclease substrate, a 39-bp oligonucleotide (5′-GCGCAGCTAGCGGCGGATGGCXCCGTTGAAGCCTGCTTT-3′) containing an abasic site (X) obtained from Eurofins was 5′-labelled with 32P and then annealed to a 40-bp oligonucleotide (5′-GAAAGCAGGCTTCAACGGAGCCATCCGCCGCTAGCTGCGC-3′). 5′-AMP substrates were prepared as described previously29. Reactions were initiated by mixing 4 μl of an appropriate dilution (in protein dialysis buffer) of the indicated recombinant human protein, or 4 μl of whole-cell extract (16 μg total protein), with 4 μl of 2× Master Mix (50 mM HEPES, pH 8.0, 260 mM KCl, 2 mM DTT and, unless otherwise indicated, 20 mM MgCl2) containing 100 nM (for recombinant protein experiments) or 300 nM (for cell extract experiments) of the appropriate 32P-labelled oligonucleotide duplex. For experiments using cell extract, 150 μM competitor single-stranded oligonucleotide (5′-CTAACTTGAGCGAAACGGT-3′) was present to reduce nonspecific nucleolytic degradation of the duplex substrate. All reactions were terminated by the addition of formamide loading buffer and fractionated by denaturing PAGE; images were analysed by phosphorimaging.
Cells, cell culture and transfection
A549 cells were cultured in DMEM (Gibco, Invitrogen) supplemented with 15% FCS and 1% glutamine. Cells were transfected using Genejuice (Novagene) according to the manufacturer’s instructions. For the generation of wild-type and Tdp1-/- MEFs, Tdp1-/+ animals were mated and 14-day embryos were dissociated, trypsinized and plated in DMEM with 10% FCS. Genotyping for the mutant Tdp1 allele was performed on cell pellets, essentially as described30. For shRNA-mediated depletion of TTRAP, A549 lung carcinoma cells were co-transfected with 2 μg of pCD2E vector encoding G418 resistance and 1 μg of either pSuper or pSuper-TTRAP. After selection in 1.5 mg ml-1 G418 (Gibco, Invitrogen) for 6 days, cells were collected for analysis.
Immunoblotting, immunoflourescence and antibodies
For western blots, cells (10,000 per μl) were lysed in SDS–PAGE loading buffer and incubated at 90 °C for 5 min. Whole-cell extracts were fractionated by SDS–PAGE and transferred to nitrocellulose. Membranes were blocked for 1 h in PBS-Tween 20 (PBST) containing 5% non-fat dried milk (NFDM), and then incubated for 2 h with anti-XRCC1 monoclonal antibody (33-2-5) or affinity-purified anti-TTRAP polyclonal antibody (SY1340) at a 1/100 or 1/200 dilution, respectively, in PBST plus 5% NFDM. Membranes were rinsed in PBST and incubated in PBST plus 5% NFDM containing horseradish-peroxidase-conjugated anti-rabbit IgG or anti-mouse IgG (DAKO), as appropriate, at a 1/3,000 dilution for 1 h at room temperature. Membranes were then rinsed with PBST and antibody complexes detected by enhanced chemiluminescence (Amersham). Note that the anti-TTRAP polyclonal antibody was raised in rabbit by Eurogentec against full-length recombinant human TTRAP expressed in E. coli. For immunofluorescence analysis, cells were grown on coverslips, rinsed in PBS, fixed with 4% paraformaldehyde for 5 min at room temperature, permeabilized with 0.2% Triton X-100, and after rinsing in PBS twice, blocked in PBS plus 5% BSA. Cells were then incubated with appropriate primary antibodies in PBS plus 1% BSA for 1 h at room temperature. Anti-γ-H2AX (Ser 139) monoclonal antibody (Upstate) and anti-TTRAP rabbit polyclonal antibody (Abcam, ab33246) were used at 1/800 and 1/200 dilutions, respectively. After rinsing in PBST three times, cells were incubated for 30 min at room temperature in PBS plus 1% BSA containing appropriate secondary antibody (Alexa Fluor 555 goat anti-mouse IgG and Alexa Fluor 488 goat anti-rabbit IgG, Invitrogen) at a 1/200 dilution, and then rinsed again three times in PBST. Cells were counterstained with 4′,6′-diamidino-2-phenylindole (DAPI) and mounted with Vectashield (Vecta Laboratories).
Laser microirradiation
A549 cells were seeded onto glass-bottom coverslips (MatTek), transfected with 1 μg pEGFP-TTRAP, and 24 h later pre-incubated for 30 min with 10 μg ml-1 Hoechst dye 33258 at 37 °C. Selected cells were then irradiated with a 351-nm UVA laser (approximately 0.35 J m-2) focused through a ×40/1.2-W objective using a Zeiss Axiovert equipped with LSM 520 Meta, and images were taken every minute after irradiation.
Clonogenic survival assays
A549 cells transfected with empty pSuper or pSuper-TTRAP as described earlier were plated into 10-cm dishes (1,000 per plate) in duplicate and incubated at 37 °C for 4 h. Cells were mock-treated or treated in medium at 37 °C with the indicated concentration of CPT or etoposide for 3 h, or with MMS for 1 h. All drugs were from Sigma. Cells were then rinsed in PBS and incubated at 37 °C in drug-free medium for 14 days. Colonies were fixed with 90% ethanol and stained with 1% methylene blue. Survival was calculated as a percentage, using the equation Nt/Nu × 100, in which Nt and Nu are the number of colonies on treated and untreated plates, respectively. Data are shown as the mean and s.e.m. of at least three independent experiments.
Alkaline single-cell agarose gel electrophoresis (comet) assays
DNA strand breaks were quantified using the alkaline comet assay as described previously6. Average tail moments from 100 cells per sample were obtained using Comet Assay III software (Perceptive Instruments), and data are shown as the mean and s.e.m. of at least three independent experiments.
γ-H2AX assays
Cells were mock-treated or treated with the indicated drug at the indicated concentration. After γ-H2AX immunofluorescence staining, the average number of γ-H2AX foci per cell was determined from 50 cells per sample using a Nikon Eclipse 50i microscope at ×100 magnification. Overexpression experiments were carried out as described above except that cells were transfected with 250 ng of the appropriate pcDNA3.1-His construct 1 day before incubation with the drug. Fifteen-to-forty cells per sample were scored, depending on the number of cells highly overexpressing TTRAP (see Supplementary Fig. 7 for representative examples). Data are the mean and s.e.m. of three independent experiments.
References
Champoux, J. J. DNA topoisomerases: structure, function, and mechanism. Annu. Rev. Biochem. 70, 369–413 (2001)
Wang, J. C. Cellular roles of DNA topoisomerases: a molecular perspective. Nature Rev. Mol. Cell Biol. 3, 430–440 (2002)
Li, T. K. & Liu, L. F. Tumor cell death induced by topoisomerase-targeting drugs. Annu. Rev. Pharmacol. Toxicol. 41, 53–77 (2001)
Deweese, J. E. & Osheroff, N. The DNA cleavage reaction of topoisomerase II: wolf in sheep’s clothing. Nucleic Acids Res. 37, 738–748 (2009)
Pourquier, P. & Pommier, Y. Topoisomerase I-mediated DNA damage. Adv. Cancer Res. 80, 189–216 (2001)
El-Khamisy, S. F. et al. Defective DNA single-strand break repair in spinocerebellar ataxia with axonal neuropathy-1. Nature 434, 108–113 (2005)
Takashima, H. et al. Mutation of TDP1, encoding a topoisomerase I-dependent DNA damage repair enzyme, in spinocerebellar ataxia with axonal neuropathy. Nature Genet. 32, 267–272 (2002)
Yang, S. W. et al. A eukaryotic enzyme that can disjoin dead-end covalent complexes between DNA and type I topoisomerases. Proc. Natl Acad. Sci. USA 93, 11534–11539 (1996)
Nitiss, J. L. Targeting DNA topoisomerase II in cancer chemotherapy. Nature Rev. Cancer 9, 338–350 (2009)
Pommier, Y. Topoisomerase I inhibitors: camptothecins and beyond. Nature Rev. Cancer 6, 789–802 (2006)
Vance, J. R. & Wilson, T. E. Yeast Tdp1 and Rad1-Rad10 function as redundant pathways for repairing Top1 replicative damage. Proc. Natl Acad. Sci. USA 99, 13669–13674 (2002)
Liu, C., Pouliot, J. J. & Nash, H. A. Repair of topoisomerase I covalent complexes in the absence of the tyrosyl-DNA phosphodiesterase Tdp1. Proc. Natl Acad. Sci. USA 99, 14970–14975 (2002)
Pype, S. et al. TTRAP, a novel protein that associates with CD40, tumor necrosis factor (TNF) receptor-75 and TNF receptor-associated factors (TRAFs), and that inhibits nuclear factor-κB activation. J. Biol. Chem. 275, 18586–18593 (2000)
Rodrigues-Lima, F., Josephs, M., Katan, M. & Cassinat, B. Sequence analysis identifies TTRAP, a protein that associates with CD40 and TNF receptor-associated factors, as a member of a superfamily of divalent cation-dependent phosphodiesterases. Biochem. Biophys. Res. Commun. 285, 1274–1279 (2001)
Nitiss, J. L. et al. Amsacrine and etoposide hypersensitivity of yeast cells overexpressing DNA topoisomerase II. Cancer Res. 52, 4467–4472 (1992)
Xu, G. L. et al. TTRAP is a novel PML nuclear bodies-associated protein. Biochem. Biophys. Res. Commun. 375, 395–398 (2008)
Mielke, C., Christensen, M. O., Barthelmes, H. U. & Boege, F. Enhanced processing of UVA-irradiated DNA by human topoisomerase II in living cells. J. Biol. Chem. 279, 20559–20562 (2004)
Mielke, C., Kalfalah, F. M., Christensen, M. O. & Boege, F. Rapid and prolonged stalling of human DNA topoisomerase I in UVA-irradiated genomic areas. DNA Repair (Amst.) 6, 1757–1763 (2007)
Kingma, P. S. & Osheroff, N. The response of eukaryotic topoisomerases to DNA damage. Biochim. Biophys. Acta 1400, 223–232 (1998)
Connelly, J. C. & Leach, D. R. Repair of DNA covalently linked to protein. Mol. Cell 13, 307–316 (2004)
Hartsuiker, E., Neale, M. J. & Carr, A. M. Distinct requirements for the Rad32(Mre11) nuclease and Ctp1(CtIP) in the removal of covalently bound topoisomerase I and II from DNA. Mol. Cell 33, 117–123 (2009)
Pei, H. et al. EAPII interacts with ETS1 and modulates its transcriptional function. Oncogene 22, 2699–2709 (2003)
Esguerra, C. V. et al. Ttrap is an essential modulator of Smad3-dependent Nodal signaling during zebrafish gastrulation and left-right axis determination. Development 134, 4381–4393 (2007)
Zucchelli, S. et al. Aggresome-forming TTRAP mediates pro-apoptotic properties of Parkinson’s disease-associated DJ-1 missense mutations. Cell Death Differ. 16, 428–438 (2009)
Vance, J. R. & Wilson, T. E. Repair of DNA strand breaks by the overlapping functions of lesion-specific and non-lesion-specific DNA 3′ phosphatases. Mol. Cell. Biol. 21, 7191–7198 (2001)
Nitiss, K. C., Malik, M., He, X., White, S. W. & Nitiss, J. L. Tyrosyl-DNA phosphodiesterase (Tdp1) participates in the repair of Top2-mediated DNA damage. Proc. Natl Acad. Sci. USA 103, 8953–8958 (2006)
Whitehouse, C. J. et al. XRCC1 stimulates human polynucleotide kinase activity at damaged DNA termini and accelerates DNA single-strand break repair. Cell 104, 107–117 (2001)
Parsons, J. L., Dianova, I. I. & Dianov, G. L. APE1 is the major 3′-phosphoglycolate activity in human cell extracts. Nucleic Acids Res. 32, 3531–3536 (2004)
Reynolds, J. J. et al. Defective DNA ligation during short-patch single-strand break repair in ataxia oculomotor apraxia-1. Mol. Cell. Biol. 29, 1354–1362 (2008)
Katyal, S. et al. TDP1 facilitates chromosomal single-strand break repair in neurons and is neuroprotective in vivo . EMBO J. 26, 4720–4731 (2007)
Acknowledgements
This work was funded by the MRC (G0600776), the BBSRC (BB/C516595/1), and CR-UK (C6563/A10192). S.F.E.K. and F.C.L. were also funded by Fellowships from the Wellcome Trust (S.F.E.K.; 085284), Marie Curie (FCL; 2007-2-1-IEF-221222) and EMBO (FCL; ALTF 956-2006). We thank T. Wilson, H. Nash, M. Neale, J. Nitiss and E. Hoffmann for materials.
Author Contributions F.C.L. developed the genetic screen and conducted the mammalian cell experiments. M.C.Z. and F.C.L. conducted the yeast experiments. K.O. and S.F.E.K. prepared the recombinant proteins, and S.F.E.K. conducted the biochemical experiments. K.W.C., F.C.L. and S.F.E.K. designed and interpreted the experiments. K.W.C. coordinated the project and wrote the manuscript.
Author information
Authors and Affiliations
Corresponding authors
Supplementary information
Supplementary Figures
This file contains Supplementary Figures 1-7 with Legends. (PDF 965 kb)
Rights and permissions
About this article
Cite this article
Ledesma, F., El Khamisy, S., Zuma, M. et al. A human 5′-tyrosyl DNA phosphodiesterase that repairs topoisomerase-mediated DNA damage. Nature 461, 674–678 (2009). https://doi.org/10.1038/nature08444
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature08444
This article is cited by
-
Synthesis of Myrtucommulone D: A Selective Inhibitor of Tyrosyl-DNA Phosphodiesterase 2 Promoting Drug Resistance Reversal in Lung Cancer Cells
Revista Brasileira de Farmacognosia (2024)
-
MUS81 cleaves TOP1-derived lesions and other DNA–protein cross-links
BMC Biology (2023)
-
Replication-associated formation and repair of human topoisomerase IIIα cleavage complexes
Nature Communications (2023)
-
Inactivating TDP2 missense mutation in siblings with congenital abnormalities reminiscent of fanconi anemia
Human Genetics (2023)
-
3-Oxabicyclo[3.3.1]nonenes: synthesis and investigation as tyrosyl-DNA phosphodiesterase 1 inhibitors
Russian Chemical Bulletin (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.