Abstract
MicroRNAs (miRNAs) are noncoding, single-stranded RNA molecules that have important roles in a number of physiological and pathological processes. Previous studies have proved that miRNAs targeting ZEB1 and ZEB2 may repress epithelial-to-mesenchymal transition. In this work, we studied the intrarenal expression of miR-200 family, miR-205 and miR-192 in patients with immunoglobulin A (IgA) nephropathy. We studied 43 patients with biopsy-proven IgA nephropathy (IgA group). The intrarenal expression of miRNAs was quantified and compared with that of 15 patients with noninflammatory glomerulosclerosis (GS group) and 20 patients with nephrectomy for kidney cancer as controls (CTL group). The level of intrarenal miR-200c was downregulated, whereas the levels of intrarenal miR-141, miR-205 and miR-192 were upregulated in IgA but not GS group. Proteinuria significantly correlated with the intrarenal expression of miR-200c (r=−0.324, P=0.011) and glomerular filtration rate (GFR) significantly correlated with the intrarenal expression of miR-205 (r=−0.280, P=0.030). The degree of tubulointerstitial scarring correlated with miR-205 expression (r=0.389, P=0.021), whereas glomerulosclerosis correlated with miR-192 expression (r=−0.311, P=0.045). The rate of GFR decline significantly correlated with the intrarenal expression of miR-192 (r=0.373, P=0.015). The intrarenal expression of E-cadherin significantly correlated with the intrarenal expression of miR-200c (r=0.392, P=0.002). The results show that intrarenal expression of miR-200c, miR-141, miR-205 and miR-192 was diversely regulated and correlated with disease severity and progression in patients with IgA nephropathy. These miRNA species may be important in the pathogenesis and progression of IgA nephropathy.
Similar content being viewed by others
Main
Immunoglobulin A (IgA) nephropathy is the most common form of primary glomerulonephritis throughout the world and an important cause of end-stage renal disease (ESRD).1 Indeed, 15–40% of patients with biopsy-proven IgA nephropathy will progress to ESRD in 10–20 years.2, 3
Tubular atrophy and interstitial fibrosis are the most important predictors of poor outcome for IgA nephropathy. Besides preexisting resident fibroblasts, intrinsic renal cells, such as tubular epithelial cells, podocytes, mesangial cells and endothelial cells, probably contribute to renal fibrogenesis by morphological changes to become activated fibroblasts through epithelial-to-mesenchymal transition (EMT), myofibroblasts transdifferentiation or endothelial-to-mesenchymal transition.4, 5, 6, 7 In IgA nephropathy, the expression of fibroblast-specific protein 1, a mesenchymal marker protein, in tubular epithelial cells was shown to correlate well with renal function and disease progression.8 However, the molecular pathogenesis of EMT in IgA nephropathy and other intrinsic kidney diseases remains unclear.
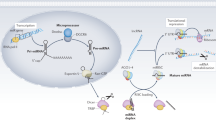
MicroRNAs (miRNAs) are noncoding, single-stranded RNA molecules of about 21–23 nucleotides in length; miRNAs regulate gene expression at post-transcriptional level and have important roles in a number of physiological and pathological processes.9 Although miRNAs were first described 15 years ago, they have not been extensively studied until recent years.10, 11 Previous in vitro studies have shown that members of the miR-200 family (miR-141 and miR-200b) and miR-205 regulate EMT by repressing ZEB1 and ZEB2, major transcription repressors of E-cadherin, a key marker of epithelial cells, and thereby prevent tumor invasion and metastasis.12, 13 Another miRNA, miR-192, was also proved to repress ZEB2 and advance collagen 1-α1 production in mesangial cells.14 However, the role of these miRNAs in intrinsic renal disease has not been explored. In this work, we studied the intrarenal expression of miR-200 family, miR-205 and miR-192, together with gene expression of molecules in the process of EMT in patients with IgA nephropathy.
PATIENTS AND METHODS
Subjects
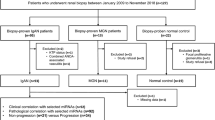
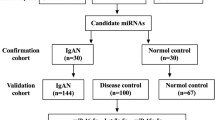
We studied 43 consecutive patients with IgA nephropathy confirmed by kidney biopsy in the Prince of Wales Hospital, Hong Kong, between 2004 and 2007 (IgA group). Patients with other coexisting renal pathology, and recurrent IgA nephropathy after kidney transplantation, were excluded. The study was approved by the clinical research ethical committee of the Chinese University of Hong Kong, all patients provided informed consent. Clinical data including serum creatinine and 24 h urine protein were recorded at the time of kidney biopsy. Glomerular filtration rate (GFR) was estimated by a standard equation.15 We studied the renal biopsy specimen of 15 patients with noninflammatory glomerulosclerosis (eight hypertensive nephrosclerosis, seven focal segmental glomerulosclerosis) (GS group), and normal renal tissue from the nephrectomy specimen of 20 patients with renal cell carcinoma (all clear cell type) as controls (CTL group) for intrarenal miRNA and messenger RNA expression study. All renal tissues were evaluated by standard light microscopy to make sure that specimens were mainly renal cortex and had at least five glomeruli.
After renal biopsy, all patients were followed up every 2 months for at least 12 months. Renal function and the degree of proteinuria were assessed at least every 4 months. Disease progression was measured by the rate of GFR decline, which was calculated by the least-square regression method.16 Treatment for individual patient was determined by responsible physician and not affected by this study. All physicians were masked to the results of miRNA and messenger RNA expression.
Renal Sample Preparation
Immediately after kidney biopsy, the renal tissue was placed in 10% neutral-buffered formaldehyde overnight and then dehydrated in alcohol and embedded in paraffin for intrarenal miRNA and messenger RNA expression. Ten 10 μm sections were cut from the formalin-fixed and paraffin-embedded (FFPE) tissue blocks using a microtome and pooled in a 1.5 ml microcentrifuge tube. The sections were then treated with xylene for 3 min at 50°C and washed twice with 100% ethanol. Then the pellet was air-dried for 30 min at room temperature.
Measurement of miRNA and Messenger RNA Levels
RecoverALL™ total nucleic acid isolation kit (Ambion Inc., Austin, TX, USA) was used for the extraction of total RNA from FFPE tissue, according to the manufacturer's protocol.
TaqMan® MicroRNA Reverse Transcription Kit and High Capacity cDNA Reverse Transcription Kit (both from Applied Biosystems, Foster City, CA, USA) were used for reverse transcription. For miRNA, 5 μl total RNA was mixed with 3 μl specific primers, 0.15 μl 100 mM dNTPs (with dTTP), 1.5 μl 10 × reverse transcription buffer, 1 μl (50 U) MultiScribe™ Reverse Transcriptase, 0.19 μl RNase inhibitor (20 U per μl) and made up to 15 μl with H2O. Reverse transcription was carried out at 16°C for 30 min, 42°C for 30 min and 85°C for 5 min. For messenger RNA, 10 μl total RNA was mixed with 2 μl specific primers, 0.8 μl 100 mM dNTPs (with dTTP), 2 μl 10 × reverse transcription buffer, 1 μl (50 U) MultiScribe™ Reverse Transcriptase, 1 μl RNase inhibitor (20 U/μl) and made up to 20 μl with H2O. Reverse transcription was performed at 25°C for 10 min, 37°C for 120 min and 85°C for 5 min. The resulting cDNA was stored in −80°C until use.
The intrarenal expression of miR-200 family (miR-200a, miR-200b, miR-200c, miR-141, miR-429), miR-205 and miR-192, together with messenger RNA of ZEB1, SIP1, E-cadherin and vimentin was quantified by RT-QPCR using the ABI Prism 7900 Sequence Detection System (Applied Biosystems). Commercially available TaqMan primers and probes, including two unlabeled PCR primers and one FAM™ dye-labeled TaqMan® MGB probe, were used for all the targets (all from Applied Biosystems). For messenger RNA expression, the primer and probe set were deliberately designed across the intron–exon boundary so as not to detect probable genomic DNA. For RT-QPCR, 10 μl universal master mix, 1 μl primer and probe set, 1.33 μl cDNA and 7.67 μl H2O were mixed to make a 20 μl reaction volume. Each sample was run in triplicate. RT-QPCR were performed at 50°C for 2 min, 95°C for 10 min, followed by 40 cycles at 95°C for 15 s and 60°C for 1 min. β-Glucuronidase (GUSB; Applied Biosystems) and RNU48 (Applied Biosystems) were used as housekeeping genes to normalize the messenger RNA and miRNA expression, respectively.17, 18 Results were analyzed with Sequence Detection Software, version 2.0 (Applied Biosystems). To calculate the differences of expression level for each target among samples, the ΔΔCt method for relative quantitation was used. Average expression level of normal renal tissue from patients with kidney clear cell cancer was used as a calibrator for intrarenal expression and the expression level of targets was a ratio relative to that of the controls.
Assessment of Renal Scarring
Analysis of renal fibrosis was determined on 4 μm paraffin-embedded sections stained by periodic acid Schiff or Jones silver stain. The severity of renal fibrosis was scored by an experienced pathologist who was masked to the results of molecular studies. The severity of glomerulosclerosis was represented by the percentage of sclerotic glomeruli in total glomeruli obtained from biopsy. For tubulointerstitial scarring, 10 microscopic fields were viewed at magnification of × 200 and scored subjectively from 0 to 100% for each patient. The severity of tubulointerstitial scarring was represented by the mean of 10 scores.
Statistical Analysis
Statistical analysis was performed by SPSS for Windows software, version 13.0 (SPSS Inc., Chicago, IL, USA). All the results were presented in mean±s.d. for data normally distributed and median (lower and upper quartiles) for others. As data of gene expression levels were highly skewed, either log-transformation or nonparametric statistical methods were used. We used Mann–Whitney U-test to compare gene expression levels between groups and Spearman's rank-order correlations to test associations between gene expression levels and clinical parameters. When no detectable level of a transcript was found (defined as no detectable level after 40 cycles of RT-QPCR) and there was zero value, a value equal to half of the minimum observed gene expression level was assigned. A P-value below 0.05 was considered statistically significant. All probabilities were two tailed.
RESULTS
The demographic and baseline clinical data of the study subjects are summarized in Table 1. As compared with CTL group, the IgA group had significantly higher level of proteinuria and worse renal function. As compared to the GS group, the IgA group were younger and had less proteinuria, but their serum creatinine and estimated GFR were similar. Histological studies showed that the percentage of glomerulosclerosis and tubulointerstitial scarring of the IgA group were 28.33±26.97 and 25.60±24.06%, respectively, whereas those of the GS group were 25.94±19.84 and 21.00±17.95%, respectively. There was no significant difference in the degree of glomerulosclerosis or tubulointerstitial fibrosis between IgA and GS groups.
Levels of MicroRNAs
The intrarenal expression of various miRNA species is compared and summarized in Figure 1. The intrarenal expression of miR-200c was significantly lower in IgA than in CTL group (0.21 (0.13–0.42) vs 0.53 (0.19–1.93), P=0.007). On the contrary, as compared to CTL group, IgA group had higher intrarenal expression of miR-141 (1.57 (0.95–3.24) vs 0.87 (0.63–1.86), P=0.017), miR-205 (1.99 (0.82–4.05) vs 0.83 (0.35–1.89), P=0.033) and miR-192 (1.77 (1.08–4.20) vs 1.24 (0.54–1.87), P=0.027). The intrarenal expression of miR-200a, miR-200b and miR-429 was similar between IgA and CTL groups (details not shown). In contrast, IgA group had significantly higher intrarenal expression of miR-200a, miR-200b, miR-141, miR-429, miR-205 and miR-192 than the GS group (see Figure 1).
MicroRNA Levels and Clinical Data
As intrarenal expression of miR-200c, miR-141, miR-205 and miR-192 was significantly different between IgA and CTL groups, we further explore the relation between their level of intrarenal expression and clinical parameters. The result is summarized in Table 2. In short, proteinuria significantly correlated with intrarenal expression of miR-200c (r=−0.324, P=0.011). GFR significantly correlated with intrarenal expression of miR-205 (r=−0.280, P=0.030). There was significant correlation between tubulointerstitial scarring and intrarenal expression of miR-205 (r=0.389, P=0.021). In contrast, glomerular scarring significantly correlated with the intrarenal expression of miR-192 (r=−0.311, P=0.045).
MicroRNA Levels and Renal Function Decline
The average duration of the follow-up was 33.4±12.6 months; the average rate of GFR decline was −0.24±0.62 ml/min per month. The rate of GFR decline of the IgAN group significantly correlated with the intrarenal expression of miR-192 (r=0.373, P=0.015) (Figure 2), but not other targets (Table 2). In contrast, the rate of GFR decline in the GS group did not correlated with the intrarenal miRNA expression (details not shown).
MicroRNA Levels and EMT
As the miRNA targets we identified may affect the process of EMT, we explore the relation between intrarenal miRNA level and the corresponding gene expression of targets related to EMT. The result is summarized in Table 3. In short, the intrarenal expression of vimentin inversely correlated with the intrarenal expression of miR-141 (r=−0.275, P=0.032) whereas the intrarenal expression of E-cadherin significantly correlated with that of miR-200c (r=0.392, P=0.002). The intrarenal expression of ZEB1 and SIP1, however, did not correlate with the intrarenal expression of any miRNA tested in this study (Table 3).
DISCUSSION
Accumulating evidence suggests the involvement of miRNAs in the pathogenesis of kidney diseases. In mice with diabetic nephropathy, intrarenal miR-192 and miR-377 levels were upregulated and facilitated tissue collagen 1-α1 and fibronectin production.14, 19 In rats with polycystic kidney disease (PKD), multiple renal miRNAs were discriminately regulated and were suggested to have an important role in specific pathways underlying PKD.20 Further, microarray studies revealed different miRNA expression profiles in a number of human kidney diseases, such as acute kidney rejection, lupus nephritis and IgA nephropathy.21, 22, 23 In line with the previous pilot study of Dai et al,22 we found that the level of intrarenal miR-200c was downregulated and miR-141, miR-205 and miR-192 was upregulated in the IgA group. In this study, intrarenal expression of all miRNA targets, except miR-200c, was lower in patients with other forms of glomerulosclerosis as compared to IgA nephropathy. Taken together, these studies suggested that various miRNA species might have specific roles in different kidney diseases.
In this study, we found that intrarenal miR-200c is downregulated in the IgA group (and probably GS group, see Figure 1), and its level inversely correlated with proteinuria, suggesting that this miRNA species is lost nonspecifically in the presence of kidney injury. On the contrary, intrarenal miR-205 may be involved in the pathogenesis of IgA nephropathy because it is upregulated in IgA but downregulated in the GS group, and its level inversely correlated with GFR and positively correlated with tubulointerstitial scarring. Further studies would be needed to clarify the role of miRNAs in kidney diseases.
Previous studies proved that the expression of miR-200 family and miR-205 was downregulated in cells undergone EMT and overexpression of these miRNAs prevented development of EMT. Direct inhibition of E-cadherin repressors ZEB1 and SIP1 might be the possible link between these miRNAs and EMT.24, 25 In fact, mRNAs of ZEB1 and SIP1 might be targets of miR-200 family and miR-205. In fact, there were multiple binding sites of ZEB1 and SIP1 for these miRNAs as revealed by computational searches. In this study, we found that intrarenal miR-200c positively correlated with E-cadherin, further suggesting the pathophysiological effects of these miRNAs in IgA nephropathy attribute, at least in part, to the prevention of EMT.
There are a few shortcomings of our study. First, we detected the expression levels of the studied miRNAs using whole renal tissue without determining the cellular sources for each of them. The miR-200 family and miR-205 were suggested to be expressed in an epithelial-specific manner and miR-192 expression was previously found in mesangial and tubular epithelial cells.12, 14 Future studies would be necessary to investigate miRNA expression level in specific renal cell type.
Secondly, FFPE tissue was used for the intrarenal expression of miRNAs and mRNAs in this study.26 Good amplification plots could be obtained for both miRNA and mRNA expression from FFPE tissue only if the amplicon of mRNA was shorter than 100 nucleotides. It has been proved that there is good correlation between miRNA expression of RNA extracted from fresh frozen and FFPE samples.27 However, it should be born in mind that nucleic acid modifications and varied degree of nucleic acid fragmentation caused by tissue fixation and embedding might induce errors for relative quantification of the targets.28 Lastly, the sample size was small in the GS group, and our result does not exclude the effect of miRNA on the renal function decline through a nonspecific (ie independent of the diagnosis) mechanism.
It could also be argued that the expression of housekeeping genes may not be the same in the control tissue and the IgA group, where there would be mesangial hypercellularity, glomerular injury and so forth. For this reason, we chose two housekeeping genes (GUSB and RNU48) according to other published reports.17, 18 As the concentration of the total RNA varies between specimens, it is difficult to use exactly the same quantity of the total RNA for RT-QPCR. We did not compare the expression of the two housekeeping genes between the groups.
In summary, we found in this study that the intrarenal expression of miR-200c, miR-141, miR-205 and miR-192 was diversely regulated and correlated with disease severity and progression in patients with IgA nephropathy. The results suggested that these miRNAs might have important roles in the pathogenesis and progression of IgA nephropathy.
References
Donadio JV, Grande JP . IgA nephropathy. N Engl J Med 2002;347:738–748.
Berthoux FC, Mohey H, Afiani A . Natural history of primary IgA nephropathy. Semin Nephrol 2008;28:4–9.
Tan CH, Loh PT, Yang WS, et al. Mycophenolate mofetil in the treatment of IgA nephropathy: a systematic review. Singapore Med J 2008;49:780–785.
Burns WC, Kantharidis P, Thomas MC . The role of tubular epithelial–mesenchymal transition in progressive kidney disease. Cells Tissues Organs 2007;185:222–231.
Li Y, Kang YS, Dai C, et al. Epithelial-to-mesenchymal transition is a potential pathway leading to podocyte dysfunction and proteinuria. Am J Pathol 2008;172:299–308.
Makino H, Kashihara N, Sugiyama H, et al. Phenotypic modulation of the mesangium reflected by contractile proteins in diabetes. Diabetes 1996;45:488–495.
Zeisberg EM, Potenta SE, Sugimoto H, et al. Fibroblasts in kidney fibrosis emerge via endothelial-to-mesenchymal transition. J Am Soc Nephrol 2008;19:2282–2287.
Nishitani Y, Iwano M, Yamaguchi Y, et al. Fibroblast-specific protein 1 is a specific prognostic marker for renal survival in patients with IgAN. Kidney Int 2005;68:1078–1085.
Kloosterman WP, Plasterk RH . The diverse functions of microRNAs in animal development and disease. Dev Cell 2006;11:441–450.
Lee RC, Feinbaum RL, Ambros V . The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 1993;75:843–854.
Ruvkun G . The perfect storm of tiny RNAs. Nat Med 2008;14: 1041–1045.
Gregory PA, Bracken CP, Bert AG, et al. MicroRNAs as regulators of epithelial–mesenchymal transition. Cell Cycle 2008;7:3112–3118.
Paterson EL, Kolesnikoff N, Gregory PA, et al. The microRNA-200 family regulates epithelial to mesenchymal transition. ScientificWorldJournal 2008;8:901–904.
Kato M, Zhang J, Wang M, et al. MicroRNA-192 in diabetic kidney glomeruli and its function in TGF-beta-induced collagen expression via inhibition of E-box repressors. Proc Natl Acad Sci USA 2007;104:3432–3437.
Levey AS, Bosch JP, Lewis JB, et al. A more accurate method to estimate glomerular filtration rate from serum creatinine: a new prediction equation. Modification of Diet in Renal Disease Study Group. Ann Intern Med 1999;130:461–470.
Mitch W . Measuring the rate of progression of renal insufficiency. In: Mitch WE, Stein J, Brenner BM (eds). The Progressive Nature of Renal Disease. Churchill Livingstone: New York, 1986, pp 203–220.
Bhandari S, Watson N, Long E, et al. Expression of somatostatin and somatostatin receptor subtypes 1–5 in human normal and diseased kidney. J Histochem Cytochem 2008;56:733–743.
Applied Biosystems. Endogenous controls for real-time quantitation of miRNA using TaqMan® microRNA assays. Application Note. TaqMan® MicroRNA Assays: Foster City, CA, USA, 2007.
Wang Q, Wang Y, Minto AW, et al. MicroRNA-377 is up-regulated and can lead to increased fibronectin production in diabetic nephropathy. FASEB J 2008;22:4126–4135.
Pandey P, Brors B, Srivastava PK, et al. Microarray-based approach identifies microRNAs and their target functional patterns in polycystic kidney disease. BMC Genomics 2008;9:624.
Sui W, Dai Y, Huang Y, et al. Microarray analysis of microRNA expression in acute rejection after renal transplantation. Transpl Immunol 2008;19:81–85.
Dai Y, Sui W, Lan H, et al. Comprehensive analysis of microRNA expression patterns in renal biopsies of lupus nephritis patients. Rheumatol Int 2009;29:749–754.
Dai Y, Sui W, Lan H, et al. Microarray analysis of micro-ribonucleic acid expression in primary immunoglobulin A nephropathy. Saudi Med J 2008;29:1388–1393.
Gregory PA, Bert AG, Paterson EL, et al. The miR-200 family and miR-205 regulate epithelial to mesenchymal transition by targeting ZEB1 and SIP1. Nat Cell Biol 2008;10:593–601.
Park SM, Gaur AB, Lengyel E, et al. The miR-200 family determines the epithelial phenotype of cancer cells by targeting the E-cadherin repressors ZEB1 and ZEB2. Genes Dev 2008;22:894–907.
Farragher SM, Tanney A, Kennedy RD, et al. RNA expression analysis from formalin fixed paraffin embedded tissues. Histochem Cell Biol 2008;130:435–445.
Xi Y, Nakajima G, Gavin E, et al. Systematic analysis of microRNA expression of RNA extracted from fresh frozen and formalin-fixed paraffin-embedded samples. RNA 2007;13:1668–1674.
Ambion. RecoverAll Total Nucleic Acid Isolation Kit, Instruction Manual, Applied Biosystems: Foster City, CA, USA, 2008, pp 15–18.
Acknowledgements
This study was supported in part by the CUHK research accounts 6901031 and 8500008.
Author information
Authors and Affiliations
Corresponding author
Additional information
Disclosure/Conflict of interest
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Wang, G., Kwan, BH., Lai, FM. et al. Intrarenal expression of microRNAs in patients with IgA nephropathy. Lab Invest 90, 98–103 (2010). https://doi.org/10.1038/labinvest.2009.118
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.2009.118
Keywords
This article is cited by
-
Blood TGF-β1 and miRNA-21-5p levels predict renal fibrosis and outcome in IgA nephropathy
International Urology and Nephrology (2023)
-
miRNA biomarkers in renal disease
International Urology and Nephrology (2022)
-
Intrarenal microRNA signature related to the fibrosis process in chronic kidney disease: identification and functional validation of key miRNAs
BMC Nephrology (2019)
-
MicroRNA-155-induced T lymphocyte subgroup drifting in IgA nephropathy
International Urology and Nephrology (2017)
-
Renal epithelial miR-205 expression correlates with disease severity in a mouse model of congenital obstructive nephropathy
Pediatric Research (2016)