Abstract
To identify the methanogenic pathways present in a deep aquifer associated with an accretionary prism in Southwest Japan, a series of geochemical and microbiological studies of natural gas and groundwater derived from a deep aquifer were performed. Stable carbon isotopic analysis of methane in the natural gas and dissolved inorganic carbon (mainly bicarbonate) in groundwater suggested that the methane was derived from both thermogenic and biogenic processes. Archaeal 16S rRNA gene analysis revealed the dominance of H2-using methanogens in the groundwater. Furthermore, the high potential of methane production by H2-using methanogens was shown in enrichments using groundwater amended with H2 and CO2. Bacterial 16S rRNA gene analysis showed that fermentative bacteria inhabited the deep aquifer. Anaerobic incubations using groundwater amended with organic substrates and bromoethanesulfonate (a methanogen inhibitor) suggested a high potential of H2 and CO2 generation by fermentative bacteria. To confirm whether or not methane is produced by a syntrophic consortium of H2-producing fermentative bacteria and H2-using methanogens, anaerobic incubations using the groundwater amended with organic substrates were performed. Consequently, H2 accumulation and rapid methane production were observed in these enrichments incubated at 55 and 65 °C. Thus, our results suggested that past and ongoing syntrophic biodegradation of organic compounds by H2-producing fermentative bacteria and H2-using methanogens, as well as a thermogenic reaction, contributes to the significant methane reserves in the deep aquifer associated with the accretionary prism in Southwest Japan.
Similar content being viewed by others
Introduction
The accretionary prism situated along the Pacific side of Southwest Japan is traceable laterally for some 1800 km and forms a thick sediment that accretes onto the nonsubducting tectonic plate at a convergent plate boundary. The sediment is composed mainly of non- to weakly metamorphosed sequences of sandstone, shale, alternating beds of both, and local associations of chert and greenstone (Taira et al., 1992). The materials are derived from marine sediment scraped from the subducting oceanic crust; therefore, they are rich in complex organic matter (Karasawa and Kano, 1992). The sediment contains layers of water-bearing permeable sandstone and no water-bearing impermeable shale. Groundwater is mainly recharged by rainfall infiltrating into outcrops or faults. The water flows down through the permeable sandstone and is reserved in a deep aquifer. In addition to the groundwater, it has been reported that dissolved natural gases are present in the deep aquifer associated with the accretionary prism in Southwest Japan (Igari and Sakata, 1989).
It is generally believed that natural gas in subsurface environments is formed by microbial production (biogenic origin), nonbiological thermal decomposition of organic matter (thermogenic origin), or geochemical reaction in hydrothermal systems during water–rock interactions (abiogenic origin). These origins have been interpreted based on chemical compositions of gaseous alkanes (for example C1–C3, methane, ethane, propane) and stable carbon and hydrogen isotope ratios of methane (δ13C–CH4 and δD–CH4). The chemical and isotopic data have been interpreted by plotting hydrocarbon gas composition C1/(C2+C3) versus δ13C–CH4 (Bernard et al., 1978) and δD–CH4 versus δ13C–CH4 (Schoell, 1983) on widely used conventional diagrams. Numerous microbiological approaches have also been used to understand the methane production processes in subsurface environments such as petroleum-contaminated aquifers (Gray et al., 2009), geothermal aquifers (Kimura et al., 2005; Mochimaru et al., 2007b), and coalbed aquifers (Shimizu et al., 2007; Strąpoć et al., 2008).
Igari and Sakata (1989) measured stable carbon isotopic ratios of methane in natural gases associated with the accretionary prism in Southwest Japan, and concluded that the methane was of thermogenic origin. Exhaustive geochemical and microbiological studies, however, have not yet been performed to determine the pathways of methane production in situ. The goals of this study were to identify the origin of the methane using the chemical and stable isotopic signatures of natural gases and groundwater and, using culture-dependent and culture-independent methods, to explore the microbial methane production pathway in the deep aquifer associated with the accretionary prism in Southwest Japan.
Materials and methods
Study site and sample collection
Groundwater and natural gas samples were collected from a deep well (Ita-wari well; 34 °52.283′N, 138 °09.150′E) situated in Shimada, Shizuoka Prefecture, Japan (Figure 1). The well is geologically located in the Setogawa group of the Shimanto Belt, which is an ancient accretionary prism situated along the Pacific side of Southwest Japan (Taira et al., 1992). The Setogawa group was deposited 30–40 million years ago with abyssal sediment of the Nankai Trough, and the materials are composed of non- to weakly metamorphosed thick sequences of sandstone, shale, and alternating bed of both (Karasawa and Kano, 1992). The deep well is drilled down to 1500 m and constructed from tight steel casing pipes including strainers at 1188–1489 m. Groundwater flows into the deep well through parts of the strainers and rises up to 250 m underground by natural water pressure. The groundwater is drawn up to ground level by a water pump under anaerobic conditions.
Location of the Shimanto Belt in Southwest Japan. The geological map is according to Kano et al. (1991). ISTL, Itoigawa-Shizuoka Tectonic Line; MTL, Median Tectonic Line.
Groundwater and natural gas were sampled nine times from July 2007 to January 2009. To prevent contamination by air and water from shallow environments, the groundwater was pumped at a flow rate of 116 l min−1 for 24 h before sampling. The groundwater samples were collected under anaerobic conditions into autoclaved glass bottles by using a sterile silicone tube. The concentrations of dissolved natural gases were so high that they were exsolving at ground level. The gases were collected in an inverted funnel underwater and then directed into autoclaved serum bottles. The serum bottles were tightly sealed with sterile butyl rubber stoppers and aluminum crimps underwater to prevent contamination by air.
Measurement of physical and chemical parameters
Physical and chemical parameters of groundwater were measured at the outflow of the deep well. Temperature was measured with a platinum resistance thermometer (Custom, Tokyo, Japan). Oxidation–reduction potential and pH were measured with RM-20P and HM-20P portable meters (DKK-TOA, Tokyo, Japan), respectively. The oxidation–reduction potential was converted to Eh using a reference electrode. Electric conductivity was measured with a CM-21P portable meter (DKK-TOA). The anion (HCO3−, CI−, Br−, I−, SO42−, acetate, formate) concentrations in the groundwater were analyzed with an IC-2001 ion chromatograph equipped with a TSKgel SuperIC-AZ column (Tosoh, Tokyo, Japan). The cation (Na+, K+, Mg2+, Ca2+) concentrations were analyzed with an IC-2001 ion chromatograph equipped with a TSKgel SuperIC-CR column (Tosoh). Sulfide was analyzed using the methylene blue method. Spectrophotometric analysis of sulfide was performed at the sampling site with a DR/2400 portable spectrophotometer (Hach, Loveland, CO, USA). Dissolved organic carbon in the groundwater filtered through pre-combusted GF/F glass microfiber filters (Whatman, Maidstone, UK) was measured with a TOC-V total organic carbon analyzer (Shimadzu, Kyoto, Japan). Concentrations of H2, N2, O2, CO2, CH4, C2H6, and C3H8 in the natural gas samples were determined on a GC-2014 gas chromatograph (GC) equipped with a thermal conductivity detector and a flame ionization detector (Shimadzu).
Analysis of stable isotopic ratios
The stable hydrogen isotope ratio (D/H) of methane in the natural gas was measured by an online CH4 extraction system, a HP6890 GC (Hewlett-Packard, Palo Alto, CA, USA), a ThermoQuest pyrolysis interface (ThermoFinnigan, Bremen, Germany), and a ThermoQuest DeltaplusXL isotope ratio mass spectrometer (IRMS) (ThermoFinnigan) (Yamada et al., 2003). The same system was used to measure the stable carbon isotope ratio (13C/12C) of methane when the pyrolysis furnace and the ThermoQuest DeltaplusXP were replaced with a combustion furnace (ceramic tube packed with CuO, NiO, and Pt wires) and a Finnigan MAT 252 IRMS (Finnigan MAT, Bremen, Germany). The 13C/12C of total dissolved inorganic carbon (ΣCO2) in groundwater, mainly bicarbonate, was analyzed according to an earlier described method (Miyajima et al., 1995). Groundwater samples for analysis of 13C/12C of ΣCO2 were collected into an autoclaved serum bottles (inner volume, 68.6 ml), fixed with 0.5 ml of saturated HgCl2 solution, and sealed with a sterile butyl rubber stoppers and aluminum crimps with no air bubbles. A 5 ml headspaces were created inside the serum bottles with pure helium gas and acidified by adding 0.5 ml of a CO2-free HCl solution (6.0N). The sample bottles were left in the dark for at least 24 h, during which time the original dissolved CO2 became equilibrated with the headspace gas. The CO2 in this headspace was subsampled, and 13C/12C of CO2 was measured by an online extraction system, HP6890 GC (Hewlett-Packard), the combustion furnace, and a ThermoQuest DeltaplusXP IRMS. Groundwater samples for the analysis of stable hydrogen and oxygen isotope ratios (D/H and 18O/16O) were collected into an autoclaved polycarbonate bottle. The D/H and 18O/16O of water were measured using a conventional IRMS according to the earlier described method (Yoshida and Mizutani, 1986).
These stable isotope ratios are expressed in the conventional δ notation calculated from the equation:

where R is the isotope ratio (D/H, 13C/12C, or 18O/16O). All isotope ratios in this paper are reported relative to the international standards: Vienna Standard Mean Ocean Water for δD and δ18O, and Vienna Pee Dee Belemnite for δ13C. The standard deviations of δD of CH4, δ13C of CH4 and CO2, δD of water, and δ18O of water were found to be ±4‰, ±0.3‰, ±1‰, and ±0.1‰, respectively.
Microbial cell staining and counting
The groundwater samples for total cell counts were fixed with formaldehyde (final concentration 2%). Exactly 10 ml of groundwater sample was filtered using pre-blackened polycarbonate filters (pore size, 0.22 μm; diameter, 25 mm; Millipore, Bedford, MA, USA). Microbial cells collected on the filter were stained with SYBR Green I (1:100 dilution, Molecular Probes, Eugene, OR, USA). The microbial cells were observed under a model BX51 epifluorescence microscope (Olympus, Tokyo, Japan), and over 50 microscopic fields were counted for each sample.
Archaeal and bacterial 16S rRNA gene analyses
Groundwater was collected from the deep well on 25 July 2008. Exactly 6 l of groundwater sample was aseptically filtered with Sterivex-GV filter units (pore size, 0.22 μm; Millipore) using sterilized silicone tubing and a pump. Bulk DNA of microorganisms trapped by the filter units was extracted according to the method described by Kimura et al. (2007). Archaeal 16S rRNA gene fragments were amplified by PCR from the bulk DNA using Archaea-specific primers 109aF (Großkopf et al., 1998) and 915aR (Stahl and Amann, 1991). Bacterial 16S rRNA gene fragments were amplified by PCR from the bulk DNA using Bacteria-specific primers 8bF and prokaryotic universal primer 1512uR (Eder et al., 1999). These reactions were performed using KOD DNA polymerase (Toyobo, Osaka, Japan) in a TaKaRa PCR Thermal Cycler PERSONAL (TaKaRa, Kyoto, Japan) with the following program: 94 °C for 2 min; 25 cycles at 94 °C for 20 s, 56 °C for 30 s, 68 °C for 1 min, and 68 °C for 10 min. PCR products of the archaeal and bacterial 16S rRNA genes were cloned with a Zero Blunt TOPO PCR Cloning Kit (Invitrogen, Carlsbad, CA, USA). The sequences of inserted PCR products selected from recombinant colonies were determined with a model RISA-384 capillary DNA sequencer (Shimadzu). Chimeric sequences were identified using the Bellerophon chimera detection program (Huber et al., 2004). Sequences that were of potential chimeric origin were excluded from further examination.
The operational taxonomic units (OTUs) for each clone library were determined using DOTUR software version 1.53 (Schloss and Handelsman, 2005). A 2% distance level between sequences was considered the cutoff to consider distinct OTUs. Each OTU was classified using the Classifier program from the RDP II (Wang et al., 2007), and the nearest relative of each OTU was determined by a FASTA search (Pearson and Lipman, 1988) against the DNA Data Bank of Japan. The coverage of each clone library was calculated by the formula [1−(n1/N)], where n1 is the number of OTUs represented by only one clone and N is the total number of clones examined (Good, 1953). The 16S rRNA gene sequences obtained in this study were deposited in the DNA Data Bank of Japan/EMBL/GenBank database under accession numbers AB477989 to AB478012.
Measurements of potential microbial methane and H2 productions
Exactly 30 ml of groundwater samples were anaerobically injected into autoclaved 70 ml serum bottles that were tightly sealed with sterile butyl rubber stoppers and aluminum crimps used to form a vacuum to prevent contamination by air. To determine the potential for methane production by methanogenic archaea, enrichments using groundwater amended with methanogenic substrates were performed. The groundwater was supplemented with acetate (10 mM), methanol (10 mM), formate (10 mM), or H2+CO2 (80:20, vol/vol; 0.25 MPa). Except for H2+CO2-supplemented bottles, the headspaces of serum bottles were filled with pure N2 gas at 0.25 MPa. The enrichments were anaerobically incubated without shaking at 35, 45, 55, 65, 75, and 85 °C. H2, CH4, and CO2 concentrations in the headspace were measured every 24 h for the initial 6 days, and every 48 h thereafter, with a GC-2014 GC equipped with a thermal conductivity detector (Shimadzu). The enrichments were carried out in duplicate. The 16S rRNA genes of microorganisms in the enrichments in which methane production was observed were analyzed according to the clone library method described above.
Enrichments using groundwater amended with organic substrates were also performed to determine the potential for microbial H2 production through anaerobic biodegradation of organic matter. The groundwater samples were supplemented with 3 ml YPG medium (3.0 g of yeast extract, 3.0 g of peptone, and 0.6 g of glucose per 100 ml of distilled water) and 20 mM of bromoethanesulfonate, an inhibitor of methanogens (Gunsalus et al., 1978). The headspaces of the serum bottles were filled with pure N2 at 0.25 MPa. These enrichments were anaerobically incubated without shaking at 35, 45, 55, 65, 75, and 85 °C. H2, CH4, and CO2 concentrations in the headspace were measured every 12 h for the initial 3 days, and every 24 h thereafter. The enrichments were carried out in duplicate. The 16S rRNA genes of microorganisms in the enrichments in which H2 production was observed were analyzed according to the clone library method described above.
We also measured the potential of methane production by syntrophic biodegradation of fermentative bacteria and methanogenic archaea. The groundwater samples were supplemented with 3 ml of the YPG medium. In these enrichments, the bromoethanesulfonate was not added. The headspaces of the serum bottles were filled with pure N2 at 0.25 MPa. The enrichments were incubated without shaking at 35, 45, 55, 65, 75, and 85 °C, and were performed in duplicate. H2, CH4, and CO2 concentrations in the headspace were measured every 12 h for the initial 4 days, and every 24 h thereafter.
Results
Physical and chemical parameters
Groundwater temperatures ranged from 40.3 to 44.4 °C, and pH was weakly alkaline. Eh ranged from −106 to −54 mV (Supplementary Table S1). Cl−, SO42−, Na+, K+, Mg2+, and Ca2+ in groundwater indicated 10–100 times lower concentrations than those found in surface seawater (Millero et al., 2008). Br− and I− were much lower than those of ancient seawater samples from natural gas fields (Mochimaru et al., 2007a, 2007b). Sulfide was below the detection limit (<0.01 mg l−1). Dissolved organic carbon ranged from 36.5 to 37.7 mg l−1, indicating 15–30 times higher concentrations than those found in coastal seawater (Kimura et al., 2001) (Supplementary Table S2).
Methane was the predominant component of the natural gas (>97%). Other main components were C2H6 and N2 at 1.3–1.4 vol.% and 0.8–1.2 vol.%, respectively. Hydrocarbon gas composition C1/(C2+C3) of the natural gas ranged from 69.8 to 75.0, which indicated that the natural gas included thermogenic C2+ hydrocarbons. H2, O2, and CO2 in natural gas were below the respective detection limits (Supplementary Table S3).
Stable isotopic ratios of natural gas and groundwater
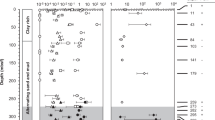
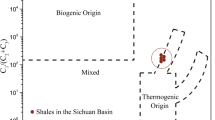
Stable isotope ratios of natural gas and groundwater are summarized in Table 1. Stable carbon isotope ratios of methane (δ13C–CH4) ranged from −39.1‰ to −38.2‰, with an average of −38.8‰. These values are relatively close to those derived from the accretionary prism in Southwest Japan reported earlier: −34.3‰ to −33.8‰ (Igari and Sakata, 1989). When the hydrocarbon gas composition C1/(C2+C3) was plotted versus the δ13C–CH4 diagram according to Bernard et al. (1978), the natural gases examined in this study fell within the range of thermogenic origin (Figure 2a). The stable hydrogen isotope ratio of methane (δD–CH4) ranged from −183‰ to −177‰, with an average of −180‰. When δ13C–CH4 was plotted versus the δD–CH4 diagram according to Schoell (1983), the observed methane fell in the range of thermogenic gases with condensate (Figure 2b).
(a) Methane carbon isotope versus hydrocarbon gas composition diagram. The categorization of methane origin is according to Bernard et al. (1978). VPDB, Vienna Pee Dee Belemnite. (b) Methane carbon and hydrogen isotope diagram according to Schoell (1983). TO, thermogenic methane with oil; TC, thermogenic methane with condensate; TD, dry thermogenic methane; VSMOW, Vienna Standard Mean Ocean Water. (c) Carbon isotope compositions of methane and dissolved inorganic carbon (mainly bicarbonate; ΣCO2). The broken lines of equal carbon isotopic fractionation, αC=(δ13C−ΣCO2+103)/(δ13C–CH4+103), are drawn for αC=1.02, 1.04, 1.06, and 1.08. The categorization of methane origin is according to Smith and Pallasser (1996).
Carbon isotopic ratios of total dissolved inorganic carbon (δ13C–ΣCO2) in groundwater were moderately enriched δ13C in the range of +17.7‰ to +18.3‰. We also plotted the observed isotopic values in δ13C–ΣCO2 versus the δ13C–CH4 diagram according to Smith and Pallasser (1996). These values fell on the boundary between biogenic origin by CO2 reduction and mixed biogenic/thermogenic origins (Figure 2c). Stable hydrogen and oxygen isotopic ratios of groundwater ranged from −17.1‰ to −16.4‰ and from +0.8‰ to +0.9‰, respectively, which were in good agreement with those of formation waters obtained from the southeastern Illinois Basin in the mid-continental region of the United States (Taylor, 1974).
Microbial cell density
Microbial cell density in the groundwater sample ranged from 5.8 × 104 to 1.1 × 105 cells ml−1, with an average of 8.0 × 104 cells ml−1 (Supplementary Table S1). The cell density was in good agreement with those in other geothermal aquifers reported by Kimura et al. (2005) and Mochimaru et al. (2007b).
Phylogenetic analysis of archaeal and bacterial 16S rRNA genes
A total of 41 clones in an archaeal 16S rRNA gene clone library were sequenced and divided into four OTUs (SMD-A01 to SMD-A04) (Table 2). The coverage of the clone library was 97.6%. Phylogenetic analysis revealed that 98% of the clones in the library belonged to the order Methanobacteriales (SMD-A01, SMD-A02, and SMD-A03), which is known as a group of methanogenic archaea able to use H2 and CO2 for growth and methanogenesis. SMD-A01 was closely related to the 16S rRNA gene from the cultured organism Methanobacterium aarhusense (94.8% 16S rRNA sequence identity), which is a hydrogenotrophic methanogen (Shlimon et al., 2004). SMD-A02 showed the highest identity to the 16S rRNA gene from the cultured organism M. alcaliphilum (94.8% identity), another hydrogenotrophic methanogen (Worakit et al., 1986). SMD-A03 showed the closest match to the 16S rRNA gene from the cultured organism Methanothermobacter thermautotrophicus (99.4% identity), a thermophilic methanogen that uses H2 and CO2 for methanogenesis (Zeikus and Wolfe, 1972). The remaining OTU, SMD-A04, was closely related to the 16S rRNA gene from the cultured organism Thermococcus gammatolerans (82.3% identity), which is known as a thermophilic heterotrophic archaeon isolated from a deep-sea hydrothermal vent (Jolivet et al., 2003).
A total of 63 clones in a bacterial 16S rRNA gene clone library were sequenced and divided into 16 OTUs (SMD-B01 to SMD-B16) (Table 2). The coverage of the clone library was 90.5%. The most abundant OTU was SMD-B01, which was closely related to the 16S rRNA gene from the cultured organism Thermodesulfovibrio yellowstonii (96.8% 16S rRNA sequence identity), known as sulfate-reducing or fermentative bacterium (Henry et al., 1994). The SMD-B01 clones accounted for 30% of the clones in the library. SMD-B02 showed the highest identity to the 16S rRNA gene from the cultured organism Desulfotomaculum salinum (85.8% identity), which is able to grow by sulfate reduction or fermentation (Nazina et al., 2005), and accounted for 14% of all the clones in the library. SMD-B03 showed the closest match to the 16S rRNA gene from the cultured organism Burkholderia fungorum (99.6% identity), which is known as an aerobic bacterium (Coenye et al., 2001). The SMD-B03 clones accounted for 13% of the clones in the library. SMD-B04 was closely related to the 16S rRNA gene from the cultured organism Thermotoga lettingae (94.6% identity), which is able to degrade methanol to H2 and CO2 (Balk et al., 2002). SMD-B05 was closely related to the 16S rRNA gene from the cultured organism Moorella thermoacetica (86.7% identity), which is known as an anaerobic bacterium-producing H2 and CO2 through fermentation (Jiang et al., 2009). SMD-B06 showed the highest identity to the 16S rRNA gene from the cultured organism Syntrophus gentianae (93.8% identity), which is able to ferment benzoate to H2 and CO2 (Schöcke and Schink, 1998). The remaining low-abundance OTUs showed closest matches to the 16S rRNA genes from anaerobic heterotroph and fermentative bacterium (SMD-B07 to SMD-B16).
Potential for microbial methane production
Methane production was observed only in the enrichment amended with H2+CO2, similar to earlier observations found in enrichments using formation waters from a gas field in the North Sea (Gray et al., 2009). In particular, high rates of methane production were found in the enrichments incubated at 55 and 65 °C (Figure 3a). Consumption of H2 and CO2 in the enrichments was in good agreement with the values expected from methane production in terms of stoichiometry of H2/CO2-using methanogenesis (4H2+CO2 → CH4+2H2O) (data not shown). The bulk DNA of propagated microorganisms in the enrichment incubated at 55 °C was extracted, and then archaeal 16S rRNA genes were amplified by PCR. The archaeal 16S rRNA gene fragments were then cloned and sequenced. A total of 40 clones were randomly sequenced and were found to belong to only one OTU (H2CO2-A01) (Table 3). H2CO2-A01 was closely related to the 16S rRNA gene from the thermophilic H2-using methanogen M. thermautotrophicus (Zeikus and Wolfe, 1972), and the sequence of the 16S rRNA gene of this OTU was 99.9% identical to the SMD-A03-derived sequence from the clone library for the archaeal community in the groundwater. The SMD-A03 was not the predominant OTU in the clone library for the archaeal community in groundwater. This may be due to the strong selective pressure caused by using very high concentrations of H2 and CO2.
(a) Methane production from the groundwater samples amended with H2+CO2. (b) H2 production from the groundwater samples amended with YPG medium and BES (a methanogen inhibitor). The enrichments were incubated at 35 °C (○), 45 °C (▪), 55 °C (•), 65 °C (□), 75 °C (▵), and 85 °C ( × ). Data points indicate the amount of methane and H2 accumulated in the headspaces of the culture bottles. Errors bars represent the level of variance calculated from measurements from duplicate incubations.
Potential for H2 production by fermentative bacteria
H2 production by fermentative bacteria was observed in the enrichments incubated at 35, 45, 55, 65, and 75 °C. In particular, high H2 production was indicated in enrichments incubated at 55 and 65 °C (Figure 3b). The bulk DNA of propagated microorganisms in the enrichment at 55 °C was extracted, bacterial 16S rRNA genes were amplified by PCR, and the 16S rRNA gene fragments were cloned and sequenced. A total of 37 clones were randomly sequenced and divided into three OTUs (YPGBES-B01, YPGBES-B02, and YPGBES-B03) (Table 3). Nearly 95% of clones in the clone library (YPGBES-B01) were most closely related to the 16S rRNA gene from the cultured organism Clostridium thermocellum, which is known as a thermophilic fermentative bacterium and is able to degrade various organic compounds, including cellulosic materials, to H2 and CO2 (Ng et al., 1977). The remaining 5% of clones (YPGBES-B02 and YPGBES-B03) belonged to the genus Desulfotomaculum. The clones derived from this enrichment were not detected in the clone library for the bacterial community in groundwater. This result may be due to the selective pressure caused by using YPG medium, which contains very high concentrations of organic compounds. Archaeal 16S rRNA genes derived from the enrichment were not amplified by PCR after repeated attempts.
Potential for methane production by syntrophic consortium of H2-producing fermentative bacteria and H2-using methanogens
To confirm whether or not methane is produced from organic matter by cooperative catabolism of H2-producing fermentative bacteria and H2-using methanogens, we performed anaerobic cultivation using groundwater amended with YPG medium. Consequently, methane production was observed in enrichments incubated at 55 and 65 °C (Figures 4a and b). H2 accumulated during the first few days, then gradually decreased to below the detection limit. However, methane generation began while H2 was decreasing, and then methane continued increasing linearly in the stage in which H2 fell below the detection limit. H2 and methane were not detected in enrichments incubated at 35, 45, or 85 °C. In contrast, the 75 °C-incubated enrichment indicated only H2 and CO2 production (Figure 4c).
Discussion
We determined the chemical and isotopic compositions of the natural gas and the groundwater derived from the deep aquifer associated with the accretionary prism in Southwest Japan, and these compositions suggested that the methane was derived from both thermogenic and biogenic processes. The cross-plot of the hydrocarbon gas composition C1/(C2+C3) over δ13C–CH4 typifies methane of thermogenic origin (Figure 2a). Similarly, the δ13C–CH4 versus δD–CH4 plot also indicates methane of thermogenic origin (Figure 2b). However, the carbon isotopic ratio of the total dissolved inorganic carbon (ΣCO2) in the groundwater ranged from +17.7‰ to +18.3‰, whereas the δ13C of CO2 in thermogenic gases ranged from −20‰ to 0‰ (Smith and Pallasser, 1996). Furthermore, this isotopically heavy ΣCO2 is likely to cause high 13C enrichment of methane in H2/CO2-using methanogenesis. We, therefore, took notice of the offsets between δ13C–ΣCO2 and δ13C–CH4 (Table 1). The observed Δ13 CΣCO2−CH4 were in the range of 55.9–57.4‰, which is consistent with the values for H2/CO2-using methanogenesis (55±10‰) rather than thermogenic reactions (Smith and Pallasser, 1996). In addition to the carbon isotopic offsets, hydrogen isotopic offsets were calculated based on δD–H2O and δD–CH4 (Table 1). ranged from 161‰ to 166‰, which agreed with typical fractionation during methanogenesis through CO2 reduction, within the equation produced by Schoell's (1980) equation:

This semi-quantitative empirical relationship has been reported by Aravena et al. (2003) and Strąpoć et al. (2007), and describes the overall hydrogen isotopic fractionation along the transfer of water-derived hydrogen through intermediary reactions into biogenic methane produced by H2-using methanogens. These carbon and hydrogen isotopic fractionations indicate that H2/CO2-using methanogenesis is responsible for the gas composition.
To identify methanogenic archaea in the groundwater samples, we determined the phylogenetic positions of members of the archaeal community using 16S rRNA gene clone libraries. The phylogenetic analysis revealed the dominance of species related to the H2-using methanogens belonging to the order Methanobacteriales. Furthermore, we observed a high rate of methane production in the enrichments using groundwater amended with H2+CO2 (Figure 3a). We confirmed that microorganisms propagated on these enrichments were closely related to M. thermautotrophicus, which is a thermophilic methanogen using H2 and CO2 for growth and methanogenesis (Zeikus and Wolfe, 1972). This combination of phylogenetic and physiological evidence strongly suggests that methane associated with the accretionary prism in Southwest Japan is produced by H2/CO2-using methanogenesis.
Here, one question arises: where does the H2 come from? We determined the phylogenetic positions of bacterial community in the groundwater using 16S rRNA gene clone libraries. We detected several clones that were closely related to fermentative bacteria that can degrade organic matter to H2 and CO2 (SMD-B04, SMD-B05, SMD-B06, and SMD-B11). However, the dominant OTUs in the clone library were sulfate-reducing bacteria related to T. yellowstonii and D. salinum (SMD-B01 and SMD-B02). It is known that these sulfate reducers are able to grow by fermentation in low-sulfate environments (Henry et al., 1994; Nazina et al., 2005). In addition, a few members of the genus Desulfotomaculum degrade organic matter to H2 and CO2 through fermentation process and grow in syntrophic association with hydrogenotrophic methanogens in the absence of sulfate (Imachi et al., 2006). The dominant species related to sulfate-reducing bacteria in the clone library (SMD-B01 and SMD-B02) may grow by fermentation and degrade organic matter to H2 and CO2 in the deep aquifer, because the groundwater has low concentrations of sulfate and sulfide (Supplementary Table S2). We further determined the potential for H2 production by fermentative bacteria based on enrichments using groundwater amended with organic matter (as glucose, peptone, and yeast extract). Consequently, high rates of H2 production were observed in the enrichments incubated at 55 and 65 °C (Figure 3b). Microorganisms propagated in the 55 °C-incubated enrichment were closely related to C. thermocellum (Ng et al., 1977), confirming the growth of fermentative bacteria that degrades organic matter to H2 and CO2. In addition to the microbiological assays, stable carbon isotopic analysis of dissolved inorganic carbon in groundwater (mainly bicarbonate) also suggested bacterial fermentation process in the deep aquifer. The dissolved HCO3− discussed in this study had enriched δ13C values, in the range of +17.7‰ to +18.3‰. Similarly, isotopically heavy dissolved HCO3− has been detected from oil and gas fields, and the primary signature of this HCO3− is thought to be the result of anaerobic biodegradation of organic matter through fermentation processes (Carothers and Kharaka, 1980; Pallasser, 2000). These findings suggest that bacterial fermentation of organic matter in groundwater or the deep aquifer matrix leads to production of H2 and CO2, which can serve as the energy and carbon sources for H2/CO2-using methanogenesis. Although it is unclear what chemical components of the organic matter are actually used in fermentation in the subsurface environments, this can be tested in future studies.
Anaerobic biodegradation of organic matter is achieved by cooperative catabolism of diverse bacteria and archaea; in particular, it is known that syntrophic cooperation (symbioses based on nutritional cooperation) between fermentative bacteria and H2-using methanogens leads to the conversion of diverse organic compounds to methane in anaerobic environments (Schink, 1997; Sakai et al., 2009; Shimoyama et al., 2009). Thus, we measured the potential for methane production by syntrophic biodegradation in enrichments using groundwater amended with organic substrates. Consequently, methane production was high in the enrichments incubated at 55 and 65 °C (Figures 4a and b). The dynamics of H2 and methane consumption/production in these enrichments were obviously similar to earlier observations in syntrophic co-cultures of fermentative bacteria and hydrogenotrophic methanogens (Hattori, 2008). These results strongly suggest that anaerobic biodegradation supports methanogenesis through a coupled mechanism in which H2 released from the anaerobic degradation of organic matter is taken up in the transformation of CO2 to methane, and that these biochemical steps take place simultaneously.
The available microbiological data show that H2-producting fermentation and H2/CO2-using methanogenesis contribute to methane production in the deep aquifer. Overall, our studies conclude that the past and ongoing syntrophic biodegradation of organic matter based on H2-producing fermentative bacteria and H2-using methanogens, as well as thermogenic methane production, contributes to the methane reserves in the deep aquifer associated with the accretionary prism in Southwest Japan.
References
Aravena R, Harrison SM, Barker JF, Abercrombie H, Rudolph D . (2003). Origin of methane in the Elk Valley coalfield, southeastern British Columbia, Canada. Chem Geol 195: 219–227.
Balk M, Weijma J, Stams AJM . (2002). Thermotoga lettingae sp. nov., a novel thermophilic, methanol-degrading bacterium isolated from a thermophilic anaerobic reactor. Int J Syst Evol Microbiol 52: 1361–1368.
Bernard BB, Brooks JM, Sackett WM . (1978). Light hydrocarbons in recent Texas continental shelf and slope sediments. J Geophys Res 83: 4053–4061.
Carothers WW, Kharaka YK . (1980). Stable carbon isotopes of HCO3 in oil-field waters—implications for the origin of CO2 . Geochem Cosmochim Acta 44: 323–332.
Coenye T, Laevens S, Willems A, Ohlen M, Hannant W, Govan JRW et al. (2001). Burkholderia fungorum sp. nov. and Burkholderia caledonica sp. nov., two new species isolated from the environment, animals and human clinical samples. Int J Syst Evol Microbiol 51: 1099–1107.
Eder W, Ludwig W, Huber R . (1999). Novel 16S rRNA gene sequences retrieved from highly saline brine sediments of Kebrit Deep, Red Sea. Arch Microbiol 172: 213–218.
Good IJ . (1953). The population frequencies of species and the estimation of population parameters. Biometrika 40: 237–264.
Gray ND, Sherry A, Larter SR, Erdmann M, Leyris J, Liengen T et al. (2009). Biogenic methane production in formation waters from a large gas field in the North Sea. Extrimophiles 13: 511–519.
Großkopf R, Janssen PH, Liesack W . (1998). Diversity and structure of the methanogenic community in anoxic rice paddy soil microcosms as examined by cultivation and direct 16S rRNA gene sequence retrieval. Appl Environ Microbiol 64: 960–969.
Gunsalus RP, Romesser JA, Wolfe RS . (1978). Preparation of coenzyme M analogs and their activity in the methyl coenzyme M reductase system of Methanobacterium thermoautotrophicum. Biochemistry 17: 2374–2377.
Hattori S . (2008). Syntrophic acetate-oxidizing microbes in methanogenic environments. Microbes Environ 23: 118–127.
Henry EA, Devereux R, Maki JS, Gilmour CC, Woese CR, Mandelco L et al. (1994). Characterization of a new thermophilic sulfate-reducing bacterium Thermodesulfovibrio yellowstonii, gen. nov. and sp. nov.: its phylogenetic relationship to Thermodesulfobacterium commune and their origins deep within the bacterial domain. Arch Microbiol 161: 62–69.
Huber T, Faulkner G, Hugenholtz P . (2004). Bellerophon; a program to detect chimeric sequences in multiple sequence alignments. Bioinformatics 20: 2317–2319.
Igari S, Sakata S . (1989). Origin of natural gas of dissolved-in-water type in Japan inferred from chemical and isotopic compositions: occurrence of dissolved gas of thermogenic origin. Geochem J 23: 139–142.
Imachi H, Sekiguchi Y, Kamagata Y, Loy A, Qiu Y-L, Hugenholtz P et al. (2006). Non-sulfate-reducing, syntrophic bacteria affiliated with Desulfotomaculum cluster I are widely distributed in methanogenic environments. Appl Environ Microbiol 72: 2080–2091.
Jiang B, Henstra AM, Paulo PL, Balk M, van Doesburg W, Stams AJM . (2009). Atypical one-carbon metabolism of an acetogenic and hydrogenogenic Moorella thermoacetica strain. Arch Microbiol 191: 123–131.
Jolivet E, L’Haridon S, Corre E, Forterre P, Prieur D . (2003). Thermococcus gammatolerans sp. nov., a hyperthermophilic archaeon from a deep-sea hydrothermal vent that resists ionizing radiation. Int J Syst Evol Microbiol 53: 847–851.
Kano K, Nakaji M, Takeuchi S . (1991). Asymmetrical melange fabrics as possible indicators of the convergent direction of plates: a case study from the Shimanto Belt of the Akaishi Mountains, central Japan. Tectonophysics 185: 375–388.
Karasawa Y, Kano K . (1992). Formation and deformation process of the slate belt in the Setogawa Group (the Shimanto Belt) of the eastern Akaishi Mountains, easternmost Southwest Japan. J Geol Society Jpn 98: 761–777 (in Japanese with English abstract).
Kimura H, Sato M, Sugiyama C, Naganuma T . (2001). Coupling of thraustochytrids and POM, and of bacterio- and phytoplankton in a semi-enclosed coastal area: implication for different substrate preference by the planktonic decomposers. Aquat Microb Ecol 25: 293–300.
Kimura H, Sugihara M, Yamamoto H, Patel BKC, Kato K, Hanada S . (2005). Microbial community in a geothermal aquifer associated with the subsurface of the Great Artesian Basin, Australia. Extremophiles 9: 407–414.
Kimura H, Ishibashi J, Masuda H, Kato K, Hanada S . (2007). Selective phylogenetic analysis targeting 16S rRNA genes of hyperthermophilic archaea in the deep-subsurface hot biosphere. Appl Environ Microbiol 73: 2110–2117.
Millero FJ, Feistel R, Wright DG, McDougall TJ . (2008). The composition of standard seawater and the definition of the reference-composition salinity scale. Deep-Sea Res I 55: 50–72.
Miyajima T, Yamada Y, Hanba YT, Yoshii K, Koitabashi T, Wada E . (1995). Determining the stable isotope ratio of total dissolved inorganic carbon in lake water by GC/C/IRMS. Limnol Oceanogr 40: 994–1000.
Mochimaru H, Uchiyama H, Yoshioka H, Imachi H, Hoaki T, Tamaki H et al. (2007a). Methanogen diversity in deep subsurface gas-associated water at the Minami-kanto gas field in Japan. Geomicrobiol J 24: 93–100.
Mochimaru H, Yoshioka H, Tamaki H, Nakamura K, Kaneko N, Sakata S et al. (2007b). Microbial diversity and methanogenic potential in a high temperature natural gas field in Japan. Extremophiles 11: 453–461.
Nazina TN, Rozanova EP, Belyakova EV, Lysenko AM, Poltaraus AB, Tourova TP et al. (2005). Description of ‘Desulfotomaculum nigrificans subsp. Salinus’ as a New Species, Desulfotomaculum salinum sp. nov.. Mikrobiologiya 74: 654–662.
Ng TK, Weimer PJ, Zeikus JG . (1977). Cellulolytic and physiological properties of Clostridium thermocellum. Arch Microbiol 114: 1–7.
Pallasser RL . (2000). Recognising biodegradation in gas/oil accumulations through the δ13C compositions of gas components. Org Geochem 31: 1363–1373.
Pearson WR, Lipman DJ . (1988). Improved tools for biological sequence comparison. Proc Natl Acad Sci USA 85: 2444–2448.
Sakai S, Imachi H, Sekiguchi Y, Tseng I-C, Ohashi A, Harada H et al. (2009). Cultivation of methanogens under low-hydrogen conditions by using the coculture method. Appl Environ Microbiol 75: 4892–4896.
Schink B . (1997). Energetics of syntrophic cooperation in methanogenic degradation. Microbiol Mol Biol Rev 61: 262–280.
Schloss PD, Handelsman J . (2005). Introducing DOTUR, a computer program for defining operational taxonomic units and estimating species richness. Appl Environ Microbiol 71: 1501–1506.
Schöcke L, Schink B . (1998). Membrane-bound proton-translocating pyrophosphatase of Syntrophus gentianae, a syntrophically benzoate-degrading fermenting bacterium. Eur J Biochem 256: 589–594.
Schoell M . (1980). The hydrogen and carbon isotopic composition of methane from natural gases of various origins. Geochim Cosmochim Acta 44: 649–661.
Schoell M . (1983). Genetic characterization of natural gases. AAPG Bull 67: 2225–2238.
Shimizu S, Akiyama M, Naganuma T, Fujioka M, Nako M, Ishijima Y . (2007). Molecular characterization of microbial communities in deep coal seam groundwater of northern Japan. Geobiology 5: 423–433.
Shimoyama T, Kato S, Ishii S, Watanabe K . (2009). Flagellum mediates symbiosis. Science 323: 1574.
Shlimon AG, Friedrich MW, Niemann H, Ramsing NB, Finster K . (2004). Methanobacterium aarhusense sp. nov., a novel methanogen isolated from a marine sediment (Aarhus Bay, Denmark). Int J Syst Evol Microbiol 54: 759–763.
Smith JW, Pallasser RJ . (1996). Microbial origin of Australian coalbed methane. AAPG Bull 80: 891–897.
Stahl DA, Amann R . (1991). Development and application of nucleic acid probes in bacterial systematics. In: Stackebrandt E, Goodfellow M. (eds). Nucleic Acid Techniques in Bacterial Systematics. John Wiley and Sons: New York, pp 205–248.
Strąpoć D, Mastalerz M, Eble C, Schimmelmann A . (2007). Characterization of the origin of coalbed gases in southeastern Illinois Basin by compound-specific carbon and hydrogen stable isotope ratios. Org Geochem 38: 267–287.
Strąpoć D, Picardal FW, Turich C, Schaperdoth I, Macalady JL, Lipp JS et al. (2008). Methane-producing microbial community in a coal bed of the Illinois Basin. Appl Environ Microbiol 74: 2424–2432.
Taira A, Byrne T, Ashi J . (1992). Photographic Atlas of an Accretionary Prim. Tokyo University Press: Tokyo, Japan.
Taylor HP . (1974). The application of oxygen and hydrogen isotope studies to problems of hydrothermal alteration and ore deposition. Econ Geol 69: 843–883.
Wang Q, Garrity GM, Tiedje JM, Cole JR . (2007). Naïve Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73: 5261–5267.
Worakit S, Boone DR, Mah RA, Abdel-Samie ME, El-Halwagi MM . (1986). Methanobacterium alcaliphilum sp. nov., an H2-utilizing methanogen that grows at high pH values. Int J Syst Bacteriol 36: 380–382.
Yamada K, Ozaki Y, Nakagawa F, Tanaka M, Yoshida N . (2003). An improved method for measurement of the hydrogen isotope ratio of atmospheric methane and its application to a Japanese urban atmosphere. Atmos Environ 37: 1975–1982.
Yoshida N, Mizutani Y . (1986). Preparation of carbon dioxide for oxygen-18 determination of water by use of a plastic syringe. Anal Chem 58: 1273–1275.
Zeikus JG, Wolfe RS . (1972). Methanobacterium thermoautotrophicus sp. n., an anaerobic, autotrophic, extreme thermophile. J Bacteriol 109: 707–713.
Acknowledgements
We are grateful to Katsuya Yabuzaki and Masataka Saito (Shimada City Hall) for help with the natural gas and groundwater sampling. We thank Julia Maresca for comments on the manuscript. This work was supported by Grant-in-Aid for Young Scientist (B) (No. 18710007) and Scientific Research (A) (No.19201004) from the Ministry of Education, Culture, Sports, Science and Technology (MEXT), the Japan Atomic Energy Agency (JAEA) Cooperative Research Scheme on the Nuclear Fuel Cycle, and the Global Environment Research Fund (B-094) of the Ministry of the Environment, Japan.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on The ISME Journal website (http://www.nature.com/ismej)
Supplementary information
Rights and permissions
About this article
Cite this article
Kimura, H., Nashimoto, H., Shimizu, M. et al. Microbial methane production in deep aquifer associated with the accretionary prism in Southwest Japan. ISME J 4, 531–541 (2010). https://doi.org/10.1038/ismej.2009.132
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2009.132
Keywords
This article is cited by
-
Conceptualization of the recent change in groundwater quality of the shallow aquifer of the hydrocarbon-rich Barmer sedimentary basin, Rajasthan, India
Arabian Journal of Geosciences (2023)
-
Thermophilic endospores associated with migrated thermogenic hydrocarbons in deep Gulf of Mexico marine sediments
The ISME Journal (2018)
-
Potential of biogenic methane for pilot-scale fermentation ex situ with lump anthracite and the changes of methanogenic consortia
Journal of Industrial Microbiology and Biotechnology (2018)
-
Hydrogen and carbon isotope systematics in hydrogenotrophic methanogenesis under H2-limited and H2-enriched conditions: implications for the origin of methane and its isotopic diagnosis
Progress in Earth and Planetary Science (2016)
-
Metabolic stratification driven by surface and subsurface interactions in a terrestrial mud volcano
The ISME Journal (2012)