Abstract
The unfolded protein response (UPR) is activated by endoplasmic reticulum stress resulting from an accumulation of unfolded or mis-folded proteins. The UPR is divided into three arms, involving the activation of ATF-6, PERK and IRE-1, that together act to restrict new protein synthesis and increase the production of chaperones. Recent studies have implicated the PERK and IRE-1 components of the UPR in adipocyte differentiation. In this study, we investigate the importance of ATF6α during adipogenesis using stable knockdown of this protein in the model adipogenic cell line, C3H10T1/2. Reduction of ATF6α expression by >70% resulted in impaired expression of key adipogenic genes and reduced lipid accumulation following the induction of adipogenesis. In contrast, loss of ATF6α did not impair the ability of cells to undergo osteogenic differentiation. Overall, our data indicate that all three arms of the UPR, including ATF6α, must be intact to permit adipogenesis to occur.
Similar content being viewed by others
Introduction
Adipose tissue dysfunction is proposed to contribute significantly to obesity-related metabolic disease.1 Consequently, understanding the molecular basis of adipogenesis may provide therapeutic approaches to improve adipose function and the pathophysiology of obesity. Adipose tissue is not only the predominant site of lipid storage but also functions as an important nutrient sensing and endocrine organ.2 The endoplasmic reticulum (ER) is a major site for protein and lipid synthesis, and in states of overnutrition elevated levels of ER stress have been reported.3 The unfolded protein response (UPR) to cellular stress is activated by the accumulation of unfolded proteins in the ER. The UPR involves three distinct pathways initiated by the activation of the ER membrane kinase PERK, the non-conventional mRNA splicing enzyme IRE-1 and the transcription factor ATF6α. These pathways work together to re-establish cellular homeostasis. Broadly, the PERK and IRE-1 pathways reduce the translation of new proteins, whereas the IRE-1 and ATF6α pathways increase the capacity of the ER to process the accumulated unfolded proteins.4 The importance of the UPR has been demonstrated in highly secretory cells, such as pancreatic β cells5 and plasma B cells, which exhibit elevated levels of the UPR and in B-cells this is essential for terminal differentiation.6 Several studies link ER stress with the pathophysiology of obesity.7, 8 However, given the high secretory capacity and metabolic activity of adipocytes, the involvement of the UPR in normal adipocyte development has also recently been investigated. This has revealed that a physiological level of the UPR may be required for adipogenesis to occur. Specifically, PERK-deficient mouse embryonic fibroblasts and 3T3-L1 preadipocyte cells display reduced adipogenic capacity, reduced expression of lipogenic genes and impaired lipid accumulation.9 A detailed study of the IRE-1 arm of the UPR revealed that this too has an important role in adipogenesis. Expression of mRNA encoding an inactive form of the transcription factor XBP-1 (XBP-1u) is induced by C/EBPβ during early adipogenesis. Upon activation of the UPR, XBP-1u mRNA undergoes non-conventional splicing by IRE-1 to generate an mRNA encoding the active XBP-1 protein (XBP-1s). In turn XBP-1s binds to and activates the promoter of the critical adipogenic transcription factor C/EBPα. Thus, loss of either XBP-1 or IRE-1 in mouse embryonic fibroblasts or 3T3-L1s results in impaired adipogenesis.10
ATF6α is a transmembrane protein that, following activation of the UPR, undergoes proteolytic cleavage by site-specific proteases to produce a free 50 kDa active transcription factor, which upregulates the expression of ER chaperones.4 As both the PERK and IRE-1 arms of the UPR have key roles in adipocyte differentiation, we investigated whether this third arm, involving ATF6α, is also required for adipogenesis.
Results and Discussion
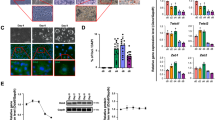
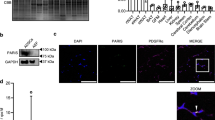
To determine the relative expression of ATF6α in adipose tissue, the levels of ATF6α mRNA were determined in a murine tissue panel (Figure 1a). As might be expected given the ubiquitous nature of the UPR, this revealed detectable levels of ATF6α mRNA in all tissues tested, including abundant expression in mesenteric, subcutaneous and brown adipose tissue. We next examined ATF6α mRNA expression in C3H10T1/2 mesenchymal stem cells following induction of adipogenesis using medium containing fetal bovine serum supplemented with insulin, dexamethasone and IBMX (IDM), as described previously.11 ATF6α mRNA was detected at all time points but did not significantly change as adipogenesis proceeded (Figure 1b). We also determined the expression of ATF6α (p90) protein during adipogenesis in these cells. This revealed a modest increase in ATF6α during the first 5 days of adipogenesis (Figure 1c), which subsequently decreased to undetectable levels at D8 (Figure 1c). The available antibodies were unable to detect the 50 kDa active form of ATF6α (p50). Thus, we are unable to specifically analyse the proteolytic activation of ATF6α during adipogenesis. However, these data demonstrate that ATF6α protein is expressed in these cells and regulated during adipogenesis.
ATF6α tissue distribution and expression in developing adipocytes. (a) ATF6α expression was determined by real-time PCR in a panel of murine tissues. Data are normalised to 18s RNA. Data are expressed as the means±s.e.m. (b) ATF6α expression was determined by real-time PCR in C3H10T1/2 cells grown to confluence then induced to undergo adipogenesis. Total RNA was collected at day 0, 1, 2, 3, 5 and 8 of differentiation and extracted using the RNEasy kit (Qiagen, Crawley, UK). Data are normalised to Cyclophilin A and are expressed as the means±s.e.m. (n=3). * Indicates significant difference from 0 h, P<0.05 by ANOVA with post hoc Tukey's test. (c) Protein lysates were collected from C3H10T1/2 cells induced to differentiate for 0, 1, 2, 3, 5 or 8 days using RIPA buffer containing protease and phosphatase inhibitors. Samples were sonicated and 20 μg subjected to western blotting using antibodies against ATF6α (ab11909, Abcam, Cambridge, UK). Images were quantified using NIH Image J software (http://rsbweb.nih.gov/ij/) and ATF6α intensity normalised to calnexin, n=3, ±s.e.m., * indicates significance of P<0.05 with respect to levels in undifferentiated cells. Representative western blots of ATF6, the adipocyte marker aP2 and the loading control calnexin (ab13504, Abcam) are shown from the same lysate samples.
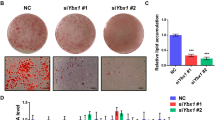
To examine the importance of ATF6α in adipogenesis, we used shRNA to stably inhibit ATF6α expression in C3H10T1/2 cells. Briefly, two shRNA sequences targeting ATF6α (shATF6α-1; exon 1–2; 5′-AAGACTGGGAGTCGACGTTGT-3′ and shATF6α-2; exon 4–5; 5′-AACTCAGCACGTTCCTGAGGA-3′) were designed and cloned into RNAi-ready pSIREN—RetroQ vector (Clontech, Mountain View, CA, USA). Retroviruses were produced by transfecting ATF6α shRNA constructs or a control shRNA vector into BOSC-HEK293 packaging cells. These were used to infect C3H10T1/2 cells, as previously described,11 generating two ATF6α knockdown cell lines. Analysis of the shATF6α expressing cell lines indicated ATF6α mRNA expression was reduced by more than 70% in both cell lines when compared with the control cells across a time course of adipogenesis (Figure 2a). A second isoform of ATF6, ATF6β is highly homologous to ATF6α with a proposed inhibitory role on ATF6α targets.12 We therefore tested whether the ATF6α shRNA affected ATF6β mRNA expression. No significant difference was seen in ATF6β expression levels in cells expressing either ATF6α shRNA when compared with control cells, indicating that both shRNA are specific to ATF6α (Figure 2b). Western blot analysis verified that ATF6α protein levels, were also inhibited by shATF6α-1 and shATF6α-2, respectively, in D0 cells (Figure 2c).
Knockdown of ATF6α expression inhibits adipogenesis. C3H10T1/2 cells stably expressing control shRNA (black bars), shATF6α-1 (dark grey bars) or shATF6α-2 (light grey bars) were induced to differentiate for 8 days. Total RNA collected at day 0, 1, 2, 3, 5 and 8 and extracted using the RNEasy kit (Qiagen). mRNA expression of (a) ATF6α, (b) ATF6β, (d) C/EBPβ, (e) PPARγ, (f) SREBP1c, (g) GLUT4 and (h) aP2 were determined using real-time PCR. Data are normalised to Cyclophilin A and are expressed as means±s.e.m. (n=3). * Indicates significant difference in expression from control shRNA at the same timepoint, P<0.05 by ANOVA with post hoc Tukey's test. Inserts show expanded view of early timepoints. (c) Protein lysates were collected from confluent control and ATF6α knockdown cells and western blotted for ATF6α and calnexin as described in Figure 1. (i) Protein lysates were collected from control and ATF6α knockdown cells induced to differentiate for 0, 2, 3, 5 or 8 days, western blotted and aP2 expression quantified as described in Figure 1, n=4, ±s.e.m., * indicates significance of P<0.05 with respect to levels at the same time point in control cells. Representative western blots of aP2 and the loading control calnexin (ab13504, Abcam) are shown from the same lysate samples. (j) Control and ATF6α deficient C3H10T1/2 cells were differentiated in IDM medium for 8 days and lipid accumulation assessed by oil-red O staining, as previously described13 (upper panels). Alternatively, cells were induced to undergo osteogenesis by treatment with Dulbecco's modified Eagle's medium supplemented with 10% FBS, 10 mM Glycerol-2-phosphate, 50 μg ml−1 ascorbic acid, 300 μg ml−1 BMP2 and 1 μM dexamethasone (lower panels). Alkaline phosphatase activity, indicating osteogenic conversion, was visualised using BCIP/NBT (B1911, Sigma-Aldrich, St Louis, MO, USA).
We next examined the effect of ATF6α knockdown of the expression of key adipogenic genes during differentiation. Following induction of adipogenesis with IDM medium, control and ATF6α knockdown cell lines displayed similar increases in the expression of the early adipogenic transcription factor, C/EBPβ (Figure 2d). However, the degree of induction was modestly, but significantly, reduced by ATF6α knockdown. The induction of the adipogenic transcription factor PPARγ was modestly, but significantly, reduced in ATF6α knockdown cells at early time points up to day 3 and more markedly impaired as adipogenesis proceeded (Figure 2e). In addition, expression of the lipogenic transcription factor SREBP1c, the insulin-sensitive glucose transporter, GLUT4 and fatty acid-binding protein, aP2, all exhibited dramatically reduced levels of mRNA expression in the ATF6α knockdown cells (Figures 2f–h). Western blotting of aP2 expression demonstrated that this was not induced following adipogenic treatment of ATF6α knockdown cells, consistent with the mRNA expression data (Figure 2i). Overall, these data demonstrate that, at least in this cellular model of adipogenesis, ATF6α expression is required for the normal induction of adipogenic gene expression. Consistent with the impaired expression of these genes lipid accumulation, as measured by Oil red O staining,13 was markedly reduced in the ATF6α-deficient cells (Figure 2j, upper panels). To ensure that this did not reflect a non-specific defect in cells lacking ATF6α, these cells were also treated with osteogenic induction medium. After 5 days of differentiation alkaline phosphatase activity, a marker of osteogenesis, was visualised with BCIP/NBT. This revealed that the osteogenic capacity of these cells was not impaired by ATF6α knockdown (Figure 2j, lower panels). Thus, ATF6α appears selectively important for adipocyte differentiation.
In conclusion, we have demonstrated that knockdown of ATF6α significantly, and selectively, impairs the ability of C3H10T1/2 cells to differentiate into mature adipocytes. Further study is required to determine the precise mechanisms involved, and to determine whether this is also the case in other cellular and in vivo models of adipocyte development. However, coupled to the previous studies of PERK and IRE-1, these data suggest that all three arms of the UPR must be intact to permit adipogenesis to occur.
References
de Ferranti S, Mozaffarian D . The perfect storm: obesity, adipocyte dysfunction, and metabolic consequences. Clin Chem 2008; 54: 945–955.
Galic S, Oakhill JS, Steinberg GR . Adipose tissue as an endocrine organ. Mol Cell Endocrinol 2010; 316: 129–139.
Hotamisligil GS . Endoplasmic reticulum stress and the inflammatory basis of metabolic disease. Cell 2010; 140: 900–917.
Ron D, Walter P . Signal integration in the endoplasmic reticulum unfolded protein response. Nat Rev Mol Cell Biol 2007; 8: 519–529.
Scheuner D, Kaufman RJ . The unfolded protein response: a pathway that links insulin demand with {beta}-cell failure and diabetes. Endocr Rev 2008; 29: 317–333.
Gass J . Stressed-out B cells? Plasma-cell differentiation and the unfolded protein response. Trends Immunol 2004; 25: 17–24.
Boden G, Duan X, Homko C, Molina EJ, Song W, Perez O et al. Increase in endoplasmic reticulum stress–related proteins and genes in adipose tissue of obese, insulin-resistant individuals. Diabetes 2008; 57: 2438–2444.
Gregor MF, Yang L, Fabbrini E, Mohammed BS, Eagon JC, Hotamisligil GS et al. Endoplasmic reticulum stress is reduced in tissues of obese subjects after weight loss. Diabetes 2009; 58: 693–700.
Bobrovnikova-Marjon E, Hatzivassiliou G, Grigoriadou C, Romero M, Cavener DR, Thompson CB et al. PERK-dependent regulation of lipogenesis during mouse mammary gland development and adipocyte differentiation. Proc Natl Acad Sci 2008; 105: 16314–16319.
Sha H, He Y, Chen H, Wang C, Zenno A, Shi H et al. The IRE1α-XBP1 pathway of the unfolded protein response is required for adipogenesis. Cell Metab 2009; 9: 556–564.
Payne VA, Grimsey N, Tuthill A, Virtue S, Gray SL, Dalla Nora E et al. The human lipodystrophy gene BSCL2/seipin may be essential for normal adipocyte differentiation. Diabetes 2008; 57: 2055–2060.
Thuerauf DJ, Marcinko M, Belmont PJ, Glembotski CC . Effects of the isoform-specific characteristics of ATF6α and ATF6β on endoplasmic reticulum stress response gene expression and cell viability. J Biol Chem 2007; 282: 22865–22878.
Rochford JJ, Semple RK, Laudes M, Boyle KB, Christodoulides C, Mulligan C et al. ETO/MTG8 is an inhibitor of C/EBPbeta activity and a regulator of early adipogenesis. Mol Cell Biol 2004; 24: 9863–9872.
Acknowledgements
This work was supported by the MRC (New Investigator Research Grant Number GO800203 (to JJR), Program Grant number G09000554 (to SOR)); the Cambridge National Institutes of Health Research Comprehensive Biomedical Research Centre (Grant Number CG50826 METABOLISM (CEL)); the Medical Research Council Centre for Obesity and Related Metabolic Diseases (Grant Number GO600717 (UO)) and the Wellcome Trust (Grant Number 078986/Z/06/Z (to SOR)).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
This work is licensed under the Creative Commons Attribution-NonCommercial-No Derivative Works 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Lowe, C., Dennis, R., Obi, U. et al. Investigating the involvement of the ATF6α pathway of the unfolded protein response in adipogenesis. Int J Obes 36, 1248–1251 (2012). https://doi.org/10.1038/ijo.2011.233
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ijo.2011.233
Keywords
This article is cited by
-
Transcription factor CREB3 is a potent regulator of high-fat diet-induced obesity and energy metabolism
International Journal of Obesity (2022)
-
Endoplasmic reticulum stress is involved in lipid accumulation induced by oleic acid in adipocytes of grass carp (Ctenopharyngodon idella): focusing on the transcriptional level
Fish Physiology and Biochemistry (2022)
-
A lifetime of stress: ATF6 in development and homeostasis
Journal of Biomedical Science (2018)
-
Nonthermal plasma treated solution inhibits adipocyte differentiation and lipogenesis in 3T3-L1 preadipocytes via ER stress signal suppression
Scientific Reports (2018)
-
ER stress signalling through eIF2α and CHOP, but not IRE1α, attenuates adipogenesis in mice
Diabetologia (2013)