Abstract
Purpose
Hearing loss is genetically extremely heterogeneous, making it suitable for next-generation sequencing (NGS). We identified a four-generation family with nonsyndromic mild to severe hearing loss of the mid- to high frequencies and onset from early childhood to second decade in seven members.
Methods
NGS of 66 deafness genes, Sanger sequencing, genome-wide linkage analysis, whole-exome sequencing (WES), semiquantitative reverse-transcriptase polymerase chain reaction.
Results
We identified a heterozygous nonsense mutation, c.6881G>A (p.Trp2294*), in the last coding exon of PTPRQ. PTPRQ has been linked with recessive (DFNB84A), but not dominant deafness. NGS and Sanger sequencing of all exons (including alternatively spliced 5′ and N-scan-predicted exons of a putative “extended” transcript) did not identify a second mutation. The highest logarithm of the odds score was in the PTPRQ-containing region on chromosome 12, and p.Trp2294* cosegregated with hearing loss. WES did not identify other cosegregating candidate variants from the mapped region. PTPRQ expression in patient fibroblasts indicated that the mutant allele escapes nonsense-mediated decay (NMD).
Conclusion
Known PTPRQ mutations are recessive and do not affect the C-terminal exon. In contrast to recessive loss-of-function mutations, c.6881G>A transcripts may escape NMD. PTPRQTrp2294* protein would lack only six terminal residues and could exert a dominant-negative effect, a possible explanation for allelic deafness, DFNA73, clinically and genetically distinct from DFNB84A.
Similar content being viewed by others
Introduction
Hearing loss, the most common sensory deficit, affects approximately 1/500 newborns. Most congenital and childhood-onset cases are nonsyndromic (70%) and of monogenic origin, with autosomal-recessive inheritance accounting for about 80%. Approximately 20% of patients have autosomal-dominantly inherited forms (ADNSHL). More than 60 ADNSHL loci have been reported. Targeted next-generation sequencing (NGS) of the known deafness genes’ exons (“gene panels”) is a powerful tool to reveal the disease-causing mutations in patients with hearing loss.
Different mutations in the same Mendelian disease genes may cause distinct—allelic— disorders. They may differ not only clinically but also in their mode of inheritance. Regarding the latter phenomenon, several genes implicated in inherited hearing loss, a condition of extensive genetic heterogeneity, have been associated with both autosomal-recessive and -dominant forms, e.g., GJB2, MYO7A, TMC1, TECTA, MYO6, and TBC1D24.1 Here, we report cosegregation of a C-terminally located heterozygous nonsense mutation in PTPRQ, a gene implicated in autosomal-recessive deafness, with autosomal-dominant progressive hearing loss in a large family. We propose a location-dependent “shift” of the inheritance mode, possibly due to escape of the mutant transcript from nonsense-mediated decay (NMD).
Materials and methods
Patients
Samples were obtained with written informed consent. Clinical investigations were conducted according to the Declaration of Helsinki, and the study was approved by the institutional review boards of the ethics committees of the University Hospital of Dresden and the University Hospital of Cologne.
Clinical examinations
The members of the investigated four-generation Caucasian family from Germany were assessed with regard to their history of hearing (Supplementary Table S1 online). Adult family members III:1, III:2, III:4, and II:2 were investigated by pure-tone audiometry. In children (the index patient, IV:2 and his brother IV:3), pedaudiological investigation included both brain stem electric response audiometry and subjective audiometry. Hearing loss was classified as mild (20–40 dB), moderate (41–60 dB), severe (61–90 dB), or profound (>90 dB) (Supplementary Table S1).
Next-generation sequencing of a deafness gene panel
Next-generation sequencing (NGS) of the DNA sample of the index patient, IV:2 (Figure 2a), was conducted for 66 genes (1,259 coding exons) that have been associated with nonsyndromic hearing loss and selected forms of syndromic hearing loss (Supplementary Table S2) on a MiSeq system (Illumina, San Diego, CA), as previously described.2 In brief, sheared DNA was ligated to barcoded adaptors for multiplexing. Exons were targeted by an in-solution customized sequence capture library (NimbleGen, Madison, WI). Amplified enriched DNA was subjected to NGS. Reads were mapped against the hg19 human reference genome using Burrows–Wheeler Aligner3 and processed with SAMtools,4 Picard (http://picard.sourceforge.net) and Genome Analysis Toolkit.5 Variants were filtered against dbNSFP v2.0,6 dbSNP v137, ExAC,7 the Human Gene Mutation Database (HGMD Professional 2013.2),8 and the Cologne Center for Genomics in-house database. The cutoff for the maximum minor allele frequency was set to 1%.9 Nonsense, frameshift and canonical splice site variants were regarded as likely pathogenic. Single-nucleotide variants were assessed using SIFT,10 MutationTaster,11 PolyPhen-2,12 AlignGVGD,13, 14 Pmut,15 NNSPLICE v0.916 and NetGene2.17, 18 SeqPilot SeqNext module (v4.0.1, JSI Medical Systems, Ettenheim, Germany) was used for visualization and final assessment of single-nucleotide variants. Sequence data for PTPRQ (MIM 603317) were compared with the reference sequence NM_001145026.1. Schraders et al.19 have proposed additional PTPRQ exons (Figure 3c) that are not contained in NM_001145026.1. All PTPRQ exons, including those described by Schraders et al. and putative additional exons from N-Scan predictions were covered by panel NGS and subsequent Sanger sequencing. Verification of the PTPRQ mutation c.6881G>A (p.Trp2294*) and segregation analyses were carried out by Sanger sequencing. The novel PTPRQ mutation described herein has been submitted to the Leiden Open Variation Database20 (http://databases.lovd.nl/shared/genes/ptprq).
Genome-wide linkage analysis
DNA was extracted from peripheral blood samples using standard methods. DNA samples of 11 individuals (seven affected individuals from four generations, including the young proband with occasional “borderline” audiometry results, IV:3; Supplementary Table S1) were subjected to genome-wide linkage analysis using the Infinium CoreExome-24 v1.1 BeadChip (Illumina) according to the manufacturer’s protocol. Subsequent data handling was performed using the graphical user interface ALOHOMORA.21 Relationship errors were identified by using the program Graphical Relationship Representation.22 The program PedCheck was applied to find Mendelian errors,23 and data for single-nucleotide polymorphisms (SNPs) with such errors were removed. Non-Mendelian errors were identified by using the program MERLIN24 and unlikely genotypes for related samples were deleted. Linkage analysis was performed assuming autosomal-dominant inheritance, full penetrance, and a disease gene frequency of 0.0001. Multipoint logarithm of the odds (LOD) scores were calculated using the program ALLEGRO.25 Haplotypes were reconstructed with ALLEGRO and presented graphically with HaploPainter.26
Whole-exome sequencing
Whole-exome sequencing (WES) was performed after enrichment with Roche/NimbleGen technology (SeqCap EZ Exome v2.0 Library) in a pool of 10 samples on an Illumina 1500 instrument in rapid mode (TruSeq Rapid SBS chemistry v1). WES data were analyzed with a focus on the mapped candidate region on chromosome 12 including PTPRQ, but also beyond that region (Supplementary Table S2).
Semiquantitative reverse-transcriptase polymerase chain reaction
Fibroblast cell lines derived from skin biopsies of a control subject and patient III:2 were established and maintained in culture at 37 °C, 5% CO2 in Dulbecco's Modified Eagle Medium supplemented with 10% fetal calf serum, 1% penicillin/streptomycin, and 0.75 μg/ml amphotericin B. 2 × 105 fibroblasts were seeded in triplicates in a six-well plate. The following day, RNA was extracted according to the manufacturer’s protocol (RNeasy Mini Kit, Qiagen, Hilden, Germany). 600 ng of fibroblast RNA was retrotranscribed using the QuantiTect Reverse Transcription Kit (Qiagen) and complementary DNA was diluted with TE−4 buffer to 20 ng/μl. Semiquantitative reverse-transcription polymerase chain reaction were carried out using recombinant Taq DNA Polymerase (Invitrogen, Waltham, MA, USA) in a BioRad C1000 Touch Thermal Cycler (Biorad, Hercules, CA, USA). For details regarding polymerase chain reaction conditions and primers, see Supplementary Table S4. Samples were run in a 1.5% agarose gel and visualized with a BioRad Chemidoc XRS system. Analysis was performed using ImageLab 5.2.1 (BioRad) and statistical analysis using MS Excel 2013 (Student’s t-test).
Results
Characterization of hearing loss in a four-generation ADNSHL family
The initial pedaudiological investigation of the index patient, IV:2, at the age of 2½ years was carried out because of delayed language development. A mild to severe bilateral sensorineural hearing loss was found and bilateral conventional hearing aids were put in use. Family members carrying the PTPRQ mutation were found to be affected by hearing loss with variable age of onset (early childhood to third decade) and variable course, as is particularly apparent in the patient’s mother, maternal grandmother, maternal great-grandmother, and great-aunt and her son (Supplementary Table S1,Figure 1). Results from pedaudiologic assessment in a young male carrier of the mutation (IV:3, last audiology exam performed at the age of 4 years) fluctuated between borderline mild hearing impairment and normal hearing (Supplementary Table S1). Individuals III:1, III:2, and IV:1–4 were personally examined by clinical geneticists and had normal body measurements, no specific minor anomalies, and no additional major health issues. The other affected individuals (I:2, II:2, II:3, II:5, and III:4) were reportedly otherwise healthy as well.
A novel PTPRQ nonsense mutation, located in a mapped chromosome 12 candidate region, cosegregates with ADNSHL
Targeted NGS of known autosomal-recessive nonsyndromic hearing loss and ADNSHL genes in the index patient (IV:2) revealed a heterozygous nonsense mutation, c.6881G>A (p.Trp2294*) (Figure 2c), in the last coding exon of PTPRQ (NM_001145026.1), encoding a type III receptor-like protein tyrosine phosphatase (Figure 3a). The p.Trp2294* mutation, which was neither present in ExAC nor in the gnomAD, cosegregated with hearing loss in the family. Additionally, linkage analysis applying genome-wide SNP mapping revealed a single genomic region with the maximum parametric LOD score of 2.4 (when including IV:3 as affected; the LOD score is 2.1 when IV:3 is excluded from the calculation) that includes the PTPRQ gene (12q21.2-12q22) (Figure 2b). Following the identification of the causative mutation in PTPRQ, DFNA73 (MIM 617663) was assigned as a novel locus designation for ADNSHL by OMIM.
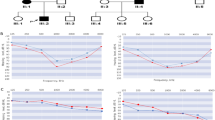
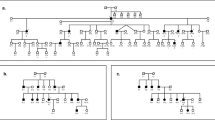
Genetic characterization of a German ADNSHL family segregating a PTPRQ nonsense mutation. (a) Pedigree of the ADNSHL family reported herein. Genotypes (M, p.Trp2294*PTPRQ mutation, WT, wild type) are given below the symbols. Whole-exome sequencing (WES) was conducted with the sample of IV:2. Green asterisks indicate family members whose samples were subjected to genome-wide linkage analysis. (b) Graphical view of the logarithm of the odds (LOD) score calculation from genome-wide single-nucleotide polymorphism mapping for the family: a LOD score of 2.1 was obtained for a region on chromosome 12 (12q21.2-12q22). (c) Electropherogram of the index patient. The nonsense mutation (red “STOP”) is close to the natural termination codon (black “STOP”) of the PTPRQ gene. (d) Graphical presentation of the candidate region from linkage analysis showing the contained genes. WES analysis did not reveal a clearly pathogenic aberration in any of these genes besides PTPRQ.
PTPRQ protein and genomic structure and its analysis by NGS (gene panel and WES). (a) Scheme of the PTPRQ protein with protein domains (FN3, fibronectin type III repeats; TM, transmembrane domain; tyrosine PP, tyrosine protein phosphatase, with active site indicated). All Human Gene Mutation Database–listed mutations are indicated (localizations to scale; truncating mutations indicated above, missense mutations below the scheme). The C-terminal autosomal-dominant nonsyndromic hearing loss (ADNSHL)–associated nonsense mutation described herein is shown in red. (b) Coverage plots from NGS, both gene panel and whole-exome sequencing (WES), related to the PTPRQ RefSeq isoform NM_001145026.1. The 45 exons are completely covered in the panel setup. (c) Coverage plots from NGS, extended gene panel, and WES, related to all N-SCAN–predicted coding regions of a hypothetical “extended“ fusion transcript including both PTPRQ, some interspersed exons, and the adjacent OTOGL gene, NM_173591.3. The red box highlights sequences that are not contained in NM_001145026.1, but which have been proposed as PTPRQ exons assigned by Schraders et al.19
Exclusion of secondary mutations in PTPRQ
NGS and Sanger sequencing of all PTPRQ exons, including previously described alternatively spliced 5′ exons,19 did not identify any other mutation (Table 1, Figure 3b). Furthermore, NGS of all N-SCAN–predicted potential coding regions of a putative “extended” PTPRQ transcript did not detect another pathogenic aberration (Figure 3c).
Assessment of variants in other genes of the mapped 12q locus
WES of the index patient identified another heterozygous nonsense variant, c.4948delA (p.Met1650*), in the LRRIQ1 gene, encoding a leucine-rich repeats and IQ motif containing protein, in the mapped candidate locus on chromosome 12. The c.4948delA (p.Met1650*) variant is listed in dbSNP (rs763597988), but it is rare (one heterozygous individual annotated in the ExAC database, corresponding to a minor allele frequency of 0.0008%). It was present in heterozygous state in two individuals with normal hearing, the youngest sibling (IV:4) of the index patient and his oldest brother (IV:1), and in their father (III:1). Patient III:1 was not aware of his mild hearing loss affecting the high frequencies, which was accidentally diagnosed by us in the course of the audiologic characterization of the family. It was not found in the patient’s mother or her affected relatives. The localization of LRRIQ1 in the mapped candidate locus was thus coincidental and did not reflect linkage with the phenotype in the index patient’s mother’s family. However, it cannot be excluded that it contributes to or even causes the mild hearing loss in the index patient’s father, and that hearing impairment might manifest in the two healthy carriers of the LRRIQ1 variant at an older age.
The WES analysis did not reveal clearly pathogenic variants in known deafness genes (apart from PTPRQ) or in potential candidate genes, particularly not in the chromosome 12 region identified by linkage analysis (Table 1, Supplementary Table S3). Two heterozygous variants were identified in two other genes from the mapped 12q region, SYT1 and TMTC2. Both variants have allele frequencies in the general population above the threshold for autosomal-dominant mutations, and homozygous occurrence in healthy individuals has been documented for both (ExAC database).
PTPRQ transcripts harboring the c.6881G>A variant do not undergo NMD
Due to the low expression of PTPRQ in whole blood samples, we quantified its expression in a patient-derived fibroblast cell line and compared it with a control (Figure 4a). Quantitative analysis for an N-terminal (encompassing exon 2) and a C-terminal PTPRQ (encompassing exon 45) region showed no differential expression between the cell lines. This demonstrates that the mutated allele escapes NMD and supports the hypothesis that a truncated PTPRQ protein is produced.
Semiquantitative reverse-transcription polymerase chain reaction (sqRT-PCR) for PTPRQ exons 2- and 45-spanning regions and quantification. (a) sqRT-PCR of PTPRQ exon 2 and exon 45 region in control and patient-derived fibroblast cell lines. ACTB and HPRT1 are used as reference genes. (b) Quantification of PTPRQ expression, normalized to the average of ACTB and HPRT1. There is no difference between the PTPRQ expression in the control and patient cell lines. Bars show the mean and standard errors. (c) Localization of primers related to the PTPRQ transcript.
Discussion
Biallelic PTPRQ mutations have been reported to cause nonsyndromic autosomal-recessive, but not autosomal-dominant, deafness with vestibular dysfunction (DFNB84A, MIM 613391).19 Here, we report the most C-terminally located PTPRQ mutation to date, c.6881G>A (p.Trp2294*) in the gene’s last exon, exon 45, cosegregating with ADNSHL in a large German family.
The mammalian NMD surveillance system cannot differentiate between nonsense mutations in the penultimate exon located less than ~55 bp from the final intron or in the last exon and natural stop codons in the last exon.27 In contrast to recessive PTPRQ mutations, which have been proposed to cause DFNB84A via a loss-of-function mechanism,28 messenger RNA carrying c.6881G>A (p.Trp2294*) would probably escape NMD and generate a protein lacking the six C-terminal residues, which may exert a dominant-negative effect. Of note, only one other reported PTPRQ mutation, homozygous c.6775delC in exon 44 in an autosomal-recessive inheritance nonsyndromic hearing loss family,29 affects one of the two last exons. In contrast to the nonsense mutation discussed in our study, a protein generated from this frameshift mutation would be extremely elongated (p.Leu2259Serfs*99), compatible with a recessive loss-of-function allele. We propose a genotype–phenotype correlation reminiscent of (throughout truncating) DIAPH1 mutations causing dominant hearing loss only when located in a short stretch of the penultimate exon or its donor splice site,30 and recessive disease if situated more N-terminally.31
PTPRQ encodes protein tyrosine phosphatase receptor Q and has been found to be identical with the 275-kDa hair-cell antigen, a component of hair-bundle interstereocilial shaft connectors. As such, mouse Ptprq has been shown to be required for the maturation of the hair bundle.28, 32 Ptprq is downregulated in mice with a heterozygous or homozygous point mutation in the seed region of miR-96, a microRNA regulating several genes required for hearing and whose mutations are implicated in inherited hearing loss in mice and humans.33, 34 It is yet unknown if PTPRQ forms dimers, as has been shown for a related protein, PTPRO.35 Interference with the normal protein deduced from the wild-type allele (dominant negative), but also the gain of a novel toxic function, are potential mechanisms of the assumingly autosomal-dominant C-terminal PTPRQ mutation reported herein. The loss of the C-terminal six amino acid residues could also prevent binding of intracellular interaction partners.
When assessing the potential pathogenicity of variants identified via NGS of a disease gene panel, variants in genes that have so far not been linked to the inheritance mode suspected in the given patient should also be considered. We therefore conducted segregation analysis for a heterozygous nonsense mutation in a recessive disease gene, PTPRQ, in a family with apparently dominant transmission of hearing loss. Based on current knowledge, this mutation could have been considered only as a carriership for a recessive mutation, and thus discarded without further verification. Additional WES and linkage analysis support the categorization of the C-terminal PTPRQ mutation as the disease-causing dominant mutation in this family; no other convincing candidate variant was identified, neither in NGS of the deafness gene panel nor in WES (Table 1, Supplementary Table S3), and importantly, the PTPRQ mutation is localized in the mapped chromosomal candidate region. Our findings indicate a “novel” mode of inheritance for PTPRQ mutations. Consequently, PTPRQ should be included in genetic diagnostic testing not only in patients with prelingual recessive deafness, but also in patients with hearing loss of variable age of onset and family history suggesting autosomal-dominant inheritance. Analogous to the apparent genotype–phenotype correlation in DFNA1, truncating mutations in the penultimate and in the last PTPRQ exon should receive special attention in that respect.
Patient IV:3 (4 years of age) carries the mutation and shows fluctuating results (from borderline to normal hearing) in audiologic assessment. However, considering the variable age of onset of the other PTPRQ mutation carriers in the family, he is likely to develop hearing impairment in the future. Our suggestion of PTPRQ as a candidate for autosomal-dominant hearing loss is strongly supported by the results of extensive genetic, but not protein, analyses. Analysis of protein function could further verify PTPRQ as an ADNSHL candidate gene, but would be complicated not only by the large size of the protein, but also because mutant PTPRQTrp2294* would differ from the wild type by just six residues. We therefore quantified PTPRQ expression, and our findings support the hypothesis of a truncated “disease-causing” PTPRQ protein derived from the c.6881G>A (p.Trp2294*) transcript.
References
Hilgert N, Smith RJ, Van Camp G . Forty-six genes causing nonsyndromic hearing impairment: which ones should be analyzed in DNA diagnostics? Mutat Res 2009;681:189–196.
Eisenberger T, Di Donato N, Baig SM et al. Targeted and genomewide NGS data disqualify mutations in MYO1A, the “DFNA48 gene,” as a cause of deafness. Hum Mutat 2014;35:565–570.
Li H, Durbin R . Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 2009;25:1754–1760.
Li H, Handsaker B, Wysoker A et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 2009;25:2078–2079.
McKenna A, Hanna M, Banks E et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 2010;20:1297–1303.
Liu X, Jian X, Boerwinkle E . dbNSFP: a lightweight database of human nonsynonymous SNPs and their functional predictions. Hum Mutat 2011;32:894–899.
Lek M, Karczewski KJ, Minikel EV et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature 2016;536:285–291.
Stenson PD, Mort M, Ball EV, Shaw K, Phillips AD, Cooper DN . The Human Gene Mutation Database: building a comprehensive mutation repository for clinical and molecular genetics, diagnostic testing and personalized genomic medicine. Hum Genet 2013;133:1–9.
Bamshad MJ, Ng SB, Bigham AW et al. Exome sequencing as a tool for Mendelian disease gene discovery. Nat Rev Genet 2011;12:745–755.
Ng PC, Henikoff S . SIFT: predicting amino acid changes that affect protein function. Nucleic Acids Res 2003;31:3812–3814.
Schwarz JM, Rodelsperger C, Schuelke M, Seelow D . MutationTaster evaluates disease-causing potential of sequence alterations. Nat Methods 2010;7:575–576.
Adzhubei I, Jordan DM, Sunyaev SR . Predicting functional effect of human missense mutations using PolyPhen-2. Curr Protoc Hum Genet 2013;Chapter 7:Unit7.20.
Tavtigian SV, Deffenbaugh AM, Yin L et al. Comprehensive statistical study of 452 BRCA1 missense substitutions with classification of eight recurrent substitutions as neutral. J Med Genet 2006;43:295–305.
Mathe E, Olivier M, Kato S, Ishioka C, Hainaut P, Tavtigian SV . Computational approaches for predicting the biological effect of p53 missense mutations: a comparison of three sequence analysis based methods. Nucleic Acids Res 2006;34:1317–1325.
Ferrer-Costa C, Gelpi JL, Zamakola L, Parraga I, de la Cruz X, Orozco M . PMUT: a web-based tool for the annotation of pathological mutations on proteins. Bioinformatics. 2005;21:3176–3178.
Reese MG, Eeckman FH, Kulp D, Haussler D . Improved splice site detection in Genie. J Comput Biol 1997;4:311–323.
Brunak S, Engelbrecht J, Knudsen S . Prediction of human mRNA donor and acceptor sites from the DNA sequence. J Mol Biol 1991;220:49–65.
Hebsgaard SM, Korning PG, Tolstrup N, Engelbrecht J, Rouze P, Brunak S . Splice site prediction in Arabidopsis thaliana pre-mRNA by combining local and global sequence information. Nucleic Acids Res 1996;24:3439–3452.
Schraders M, Oostrik J, Huygen PL et al. Mutations in PTPRQ are a cause of autosomal-recessive nonsyndromic hearing impairment DFNB84 and associated with vestibular dysfunction. Am J Hum Genet 2010;86:604–610.
Fokkema IF, Taschner PE, Schaafsma GC, Celli J, Laros JF, den Dunnen JT . LOVD v.2.0: the next generation in gene variant databases. Hum Mutat 2011;32:557–563.
Ruschendorf F, Nurnberg P . ALOHOMORA: a tool for linkage analysis using 10K SNP array data. Bioinformatics 2005;21:2123–2125.
Abecasis GR, Cherny SS, Cookson WOC, Cardon LR . GRR: graphical representation of relationship errors. Bioinformatics 2001;17:742–743.
O’Connell JR, Weeks DE . PedCheck: a program for identification of genotype incompatibilities in linkage analysis. Am J Hum Genet 1998;63:259–266.
Abecasis GR, Cherny SS, Cookson WO, Cardon LR . Merlin: rapid analysis of dense genetic maps using sparse gene flow trees. Nat Genet 2002;30:97–101.
Gudbjartsson DF, Jonasson K, Frigge ML, Kong A . Allegro, a new computer program for multipoint linkage analysis. Nat Genet. 2000;25:12–13.
Thiele H, Nurnberg P . HaploPainter: a tool for drawing pedigrees with complex haplotypes. Bioinformatics. 2005;21:1730–1732.
Khajavi M, Inoue K, Lupski JR . Nonsense-mediated mRNA decay modulates clinical outcome of genetic disease. Eur J Hum Genet 2006;14:1074–1081.
Goodyear RJ, Jones SM, Sharifi L, Forge A, Richardson GP . Hair bundle defects and loss of function in the vestibular end organs of mice lacking the receptor-like inositol lipid phosphatase PTPRQ. J Neurosci 2012;32:2762–2772.
Sloan-Heggen CM, Babanejad M, Beheshtian M et al. Characterising the spectrum of autosomal recessive hereditary hearing loss in Iran. J Med Genet 2015;52:823–829.
Neuhaus C, Lang-Roth R, Zimmermann U et al. Extension of the clinical and molecular phenotype of DIAPH1-associated autosomal dominant hearing loss (DFNA1). Clin Genet 2016;91:892–901.
Ercan-Sencicek AG, Jambi S, Franjic D et al. Homozygous loss of DIAPH1 is a novel cause of microcephaly in humans. Eur J Hum Genet 2015;23:165–172.
Goodyear RJ, Legan PK, Wright MB et al. A receptor-like inositol lipid phosphatase is required for the maturation of developing cochlear hair bundles. J Neurosci. 2003;23:9208–9219.
Lewis MA, Quint E, Glazier AM et al. An ENU-induced mutation of miR-96 associated with progressive hearing loss in mice. Nat Genet 2009;41:614–618.
Mencia A, Modamio-Hoybjor S, Redshaw N et al. Mutations in the seed region of human miR-96 are responsible for nonsyndromic progressive hearing loss. Nat Genet 2009;41:609–613.
Hower AE, Beltran PJ, Bixby JL . Dimerization of tyrosine phosphatase PTPRO decreases its activity and ability to inactivate TrkC. J Neurochem 2009;110:1635–1647.
Abe S, Usami S, Shinkawa H, Kelley PM, Kimberling WJ . Prevalent connexin 26 gene (GJB2) mutations in Japanese. J Med Genet 2000;37:41–43.
Carvill GL, Heavin SB, Yendle SC et al. Targeted resequencing in epileptic encephalopathies identifies de novo mutations in CHD2 and SYNGAP1. Nat Genet 2013;45:825–830.
Dreyer B, Tranebjaerg L, Rosenberg T, Weston MD, Kimberling WJ, Nilssen O . Identification of novel USH2A mutations: implications for the structure of USH2A protein. Eur J Hum Genet 2000;8:500–506.
Kelsell DP, Wilgoss AL, Richard G, Stevens HP, Munro CS, Leigh IM . Connexin mutations associated with palmoplantar keratoderma and profound deafness in a single family. Eur J Hum Genet 2000;8:469–472.
Acknowledgements
H.J.B. was supported by the GEERS-Stiftung. We thank the participating members of the family described herein for their support and cooperation, and Annika Greiche for excellent technical assistance.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
T.E., C.D., and C.N. are employees of Bioscientia, which is part of a publicly traded diagnostic company. H. J. B. was an employee of Bioscientia. The other authors declare no conflict of interest.
Additional information
The first two authors contributed equally to this work.
Supplementary material is linked to the online version of the paper at
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Eisenberger, T., Di Donato, N., Decker, C. et al. A C-terminal nonsense mutation links PTPRQ with autosomal-dominant hearing loss, DFNA73. Genet Med 20, 614–621 (2018). https://doi.org/10.1038/gim.2017.155
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gim.2017.155
Keywords
This article is cited by
-
Biallelic mutations in pakistani families with autosomal recessive prelingual nonsyndromic hearing loss
Genes & Genomics (2023)
-
Whole-exome sequencing identified a novel heterozygous mutation of SALL1 and a new homozygous mutation of PTPRQ in a Chinese family with Townes-Brocks syndrome and hearing loss
BMC Medical Genomics (2021)
-
First confirmatory study on PTPRQ as an autosomal dominant non-syndromic hearing loss gene
Journal of Translational Medicine (2019)