Abstract
Major rDNA loci, i.e. nucleolus-organizing regions (NORs), were assigned using chromomycin-A3 (CMA3) staining followed by sequential silver (Ag) staining and in situ hybridization (ISH) with a rDNA probe to the chromosomes of the European whitefish (Coregonus lavaretus), the peled (Coregonus peled) and the vendace (Coregonus albula), three closely related coregonine salmonid fishes. One pair of NOR-bearing chromosomes was found in the peled karyotype. Multichromosomal, but stable, locations of rDNA sites on three pairs of chromosomes were observed in the European whitefish karyotype. Multichromosomal polymorphic locations, both in site and number, were observed in the karyotype of the vendace. Several Ag-, CMA3- and ISH-positive regions were found which defined up to seven cytotypes of five NOR-bearing chromosomes. All positive Ag-NORs detected corresponded both to rDNA-ISH- and CMA3-positive signals, which suggests extensive structural polymorphism in the locations of rDNA sites. Stable NOR sites were found at the same location on both homologous elements of the chromosome no. 9 in all individuals, while the remaining NORs were quite variable between individuals, and often present in heterozygous condition. The apparently similar and parallel evolutionary rDNA differentiation patterns in the subfamilies Coregoninae and Salmoninae (family Salmonidae) are observed and discussed.
Similar content being viewed by others
Introduction
The genes coding for the 18S-5.8S-28S rRNA (major rDNA) are organized in long tandem arrays separated by intergenic spacers (IGS), which contain the regulatory elements for rDNA transcription (Moss & Stefanovsky, 1995), and are clustered at specific chromosomal sites called nucleolar organizer regions (NORs). NORs behave as very dynamic chromosome regions in evolutionary terms, with differences in chromosome location being detected between twin species, e.g. family Anostomidae (Galetti et al., 1984), domestic and wild strains of Suidae (Mellink et al., 1992) or populations of the same species, e.g. characid fish Astyanax scarbipinnis (Souza & Moreira-Filho, 1995). Although one NOR-bearing chromosome pair (i.e. simple NORs) is present in most analysed groups (and is considered a plesiomorphic character), multichromosomal locations, i.e. multiple NORs (considered to be derived characters), have been described in a number of different species (e.g. Galetti & Rasch, 1993; Schmid et al., 1995). In some species of amphibians (De Lucchini et al., 1993), fishes (Zhuo et al., 1995) and plants (Schubert & Wobus, 1985), such multichromosomal locations seem to be unstable, with a main constant NOR-bearing pair and some others with variable positions, commonly in heterozygous condition.
The evolutionary tetraploid origin of salmonid fish makes this group an interesting object for investigation of karyotype and chromosome evolution (Hartley, 1987; Phillips & Rab, 2001). Within this group, polymorphisms regarding numbers and locations of rDNA sites have been reported in some species, e.g. brown trout, Salmo trutta (Pendas et al., 1993; Castro et al., 1996; Woznicki et al., 1999) or chars of the genus Salvelinus (Reed & Phillips, 1997; Fujiwara et al., 1998). The patterns observed in these fishes also involve a constant NOR-bearing pair as well as several other (minor) NORs in variable positions.
Fishes of the subfamily Coregoninae (family Salmonidae) present several interesting problems for researchers studying evolution, zoogeography, and taxonomy (e.g. Svardson, 1970; Reist et al., 1998). They show a large morphological variation and plasticity, unusual distribution patterns of populations and species, often related with recent glaciations, and examples of species flocks and sibling species within highly restricted distribution areas (e.g. Bodaly et al., 1991; Bernatchez & Dodson, 1994; Kottelat, 1997).
Coregonid fishes are broadly distributed in Holarctic waters including species of relatively restricted ranges of distribution, such as Coregonus peled, an endemic coregonid species occurring in Siberian waters, and species of circumpolar distribution, e.g. Coregonus lavaretus and Coregonus albula, distributed in the British Isles, the Alpine region, Scandinavia and all the countries of the Baltic Sea watershed.
Chromosome analysis of populations and stocks of coregonids from European lakes have shown no differences in the basic characteristics of their karyotypes within species (chromosome number and fundamental arm number (NF), Jankun et al., 1991, 1995; 1998a,b). We have recently assigned localization of 18S rDNA clusters in two coregonids, the European whitefish and the vendace, by in situ hybridization (Jankun et al. 2000).
In this study, we have made an extensive cytogenetical analysis of Coregonus lavaretus (European whitefish), C. albula (vendace) and C. peled (peled), species poorly studied from a cytogenetic point of view. We have studied NOR sites by means of Ag and CMA3 staining, and ISH (in situ hybridization). The results presented contribute to understanding evolutionary rDNA differentiation patterns in the subfamilies Coregoninae and Salmoninae.
Materials and methods
Fourteen specimens of the vendace were obtained from Ukiel Lake (Olsztyn District, Poland). Fifteen specimens of the European whitefish were obtained from a population inhabiting the Pomeranian Bay (Baltic Sea), but artificially reared in an experimental fish farm. Peled originated in the Finnish Game and Fisheries Institute in Taivalkoski, Finland (16 specimens).
Cytogenetic techniques
Chromosome preparations were prepared according to Jankun et al. (1998b). Chromomycin-A3 (CMA3) staining and sequential Ag-NOR staining were applied according to Sola et al. (1992) and Howell & Black (1980), respectively. The distributions of Ag-NOR and nucleoli within each individual were scored for all examined individuals of both species. At least 10 metaphase plates and 100 nuclei were used to analyse Ag-NOR, and the nucleoli distribution within each individual, respectively. Some preparations were sequentially stained with Ag-NOR/C-banding in order to visualize C-positive heterochromatin blocks associated with NORs.
ISH
The plasmid pB18′, that includes a long fragment of the human rDNA transcription unit, namely the promoter and most of the 18S gene (Wilson et al., 1978), was used as a probe for ISH. The rDNA probe was labelled with biotin 16-dUTP using a commercially available Biotin Nick Translation Mix (Roche Diagnostics GmbH, Mannheim, Germany). Prior to hybridization, the slides were treated with 100 g/mL RNAse in 2XSSC at 37°C for 1 h, rinsed twice in 2XSSC (5 min), and dehydrated in a graded ethanol series at −20°C. Pretreated slides were denatured for 2 min at 70°C with 70% formamide in 2XSSC, dehydrated through an ethanol series at −20°C and then air dried. After a 10 min denaturation at 100°C, 30μl of the hybridization mixture containing 50% formamide in 2XSSC, 10% dextran sulphate, 100 ng μL−1 of herring sperm DNA and 60 ng of biotinylated probe were applied to each slide under a coverslip. Hybridization was performed overnight at 37°C in a humid chamber. Posthybridization washes were carried out in 2XSSC for 5 min, three times in 50% formamide with 0.05% of Tween 20 for 5 min and twice in 2 X SSC for 5 min (all these washes at 42°C). Hybridization signals were detected using the TSA-Indirect kit (NEN Life Science Products, Boston, MA, USA). Chromosomes were counterstained with Harris haematoxylin.
Results
European whitefish
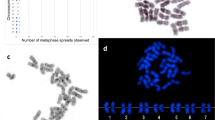
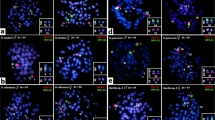
After silver staining one to five different Ag-NORs were found in the karyotype of the European whitefish (Fig. 1b). Chromomycin-A3 staining showed GC rich heterochromatic blocks located in terminal position on the short arm of two subtelocentric pairs (pairs 10 and 11 according to Jankun et al., 1998b; Fig. 1a,b). Two more subtelocentric chromosomes with minor CMA3 signals were also visible in some plates (Fig. 1a). In situ hybridization showed six signals on three small subtelocentric chromosome pairs, apparently the same ones observed after CMA3 staining (pairs 10, 11, and 12; Fig. 1c).
Peled
In all metaphases studied, only two signals of CMA3 and one or two Ag deposits were found (Fig. 2a,b). NORs were located on the short arm of a sub- meta- to subtelocentric chromosome pair. C-banding enabled us to identify this arm as also being C-positive. In three individuals, length polymorphism of this chromosome arm was observed (Fig. 2). One homologue was almost metacentric with Ag, CMA3 or hybridization positive signal along the short arm, whereas the other homologue was subtelocentric. The location of the rRNA genes was confirmed by ISH (Fig. 2c).
Vendace
Multichromosomal location of NORs was observed in all individuals examined after Ag-staining (Table 1). A constant, non variable NOR site was found at the same location (both homologous elements of the pair no. 9, Jankun et al., 1995) in each individual examined, while other NORs were variable in position, being often present in heterozygous condition (chromosome pairs nos. 1, 5, 8, 10; Table 1). The number of NORs ranged between three and seven per individual. The mean number of nucleoli per nucleus ranged from 1.8 to 4.0, apparently corresponding to the total number of NORs of each individual (Table 1). CMA3 staining and ISH with a rDNA probe detected the same NOR sites in each individual, and correspondingly, rDNA gene clusters were found in all Ag- and CMA3-positive sites (Fig. 3). Simultaneous expression of all NOR sites in the same metaphase was rarely observed. NOR sites appeared stable within each individual because staining with CMA3 and ISH showed the same location of signals in all metaphase plates of any individual.
The largest NOR-bearing pair showed three cytotypes according to NOR location (Fig. 4). Out of 14 individuals analysed, only four lacked NORs in chromosome no. 1, the remaining 10 individuals possessed either one or two NOR-bearing chromosome no. 1, mostly in heterozygous condition (Table 1).
C-banding enabled the identification of one NOR-site located on chromosome no. 5 (Fig. 5). Jankun et al. (1995) observed two forms of this chromosome pair differing by the presence of single or double heterochromatic bands in the distal part of its long arm. In the present study, C-band heteromorphism was not observed in chromosome no. 5. The NOR site on chromosome no. 5 was absent only in one individual. Two individuals had additional weak Ag- and CMA3 signals on the short arm of one homologue of chromosome pair no. 8 (Figs 3,4; Table 1). ISH with pB18′ confirmed the location of rDNA clusters here.
Discussion
The rRNA gene clusters on the fish chromosomes can be identified by Ag staining. This visualizes active NORs, fluorescent staining with antibiotic CMA3 that binds preferentially to GC-rich chromatin segments characteristically present in rDNA sites of teleostean fishes, and/or in situ hybridization with specific rDNA probes (reviewed in Rab et al., 1996). All of these methods were used in this study to characterize NOR locations in the karyotype of the European whitefish, the peled and the vendace. These three species are closely related. Recent genetic evidence (mtDNA) indicates that vendace and peled can be grouped together at a genetic distance of 0.05, and this group showed a genetic distance from whitefishes of 0.20 (Reist et al., 1998). Similar results were obtained by isozyme studies (Bodaly et al., 1991).
The location of NOR sites in the karyotypes of coregonine salmonids remains little known except for preliminary data for some European and North American taxa (Jankun et al., 1995; Phillips et al., 1996; Jankun et al., 1998a). Although the family Salmonidae has evolved from a tetraploid ancestor (Allendorf & Thorgaard, 1984), most species exhibit a single NOR-bearing pair of chromosomes. This situation was found in the peled karyotype (Fig. 2). Our previous studies on chromosomes of whitefish and vendace with Ag- and CMA3-staining suggested a multichromosomal location of NORs in both species (Jankun et al., 1991, 1995; 1998a). In this study, the European whitefish showed a multiple, but not variable, pattern in the number and location of NORs. Our results also confirmed that the four main CMA3- and ISH- positive regions showed Ag-positive, i.e. active NOR sites. Two small additional rDNA clusters were detected on a small subtelocentric pair with both CMA3 and ISH-staining, but rarely showed positive Ag-staining suggesting apparently inactive ribosomal genes as documented in Fig. 1.
Phillips & Rab (2001) included peled in their category I salmonid species with intermediate karyotype; the present results appear to be consistent with such an opinion. NOR location in peled (Coregonus peled) karyotype is described for the first time in this paper. This species shows only one NOR-bearing chromosome pair, that was also confirmed by ISH (Fig. 2). Length polymorphism detected in the short arm of this chromosome pair has been widely described in many organisms, usually being attributed to unequal crossing-over or chromosome rearrangements (Lyckegaard & Clark, 1991). The karyotype of peled from the stock we studied consisted of 76 chromosomes with NF 96 (Kirtiklis, unpublished data). According to other authors who examined Giemsa stained (Kaidanova, 1989) and C-banded karyotypes (Juntunen, 1987), the diploid chromosome number was 74–76 (NF=96–98).
The karyotype of the vendace is very similar to that of the European whitefish (2n=80, NF=98; 2n=80, NF=100, respectively; Jankun et al., 1995, 1998a). These species, however, show very different patterns of NORs (Figs 1,3). Whereas in the European whitefish and peled, the number of NORs appears stable, an extensive polymorphism was observed in the vendace. An apparently homologous NOR-bearing chromosome pair is present in the karyotypes of the three species under study. The main NOR-bearing chromosomes of the European whitefish (no. 10) and peled are similar in size and morphology to the constant NOR-bearing pair in the karyotype of the vendace (no. 9; Figs 1,3).
The expression of NORs, as a limited number of the available rDNA loci in the vendace karyotype (Table 1), is thought to be related to transcriptional regulation, as described by Castro et al. (1996) and Fujiwara et al. (1998) in other salmonid species. Sequential C-banding after Ag-staining showed that NOR sites in vendace (Fig. 5) are not strictly associated to the C-positive heterochromatin blocks. A different situation was observed in the peled and other salmonid fishes, where NOR sites always appeared C-positive (Martinez et al., 1991; Fujiwara et al., 1998).
Among salmonids, multichromosomal location of NORs was observed in the genus Salvelinus and in the brown trout, Salmo trutta. The same kind of polymorphism regarding NOR site pattern, including ‘double NOR chromosome’ (1pq in vendace; Fig. 4), was found by Reed & Phillips (1997) on the putative sex chromosomes of the Arctic char, Salvelinus alpinus, and also in some populations of the brown trout (Castro et al., 2001). These authors observed a similar phenomenon to that reported here, i.e., constant NOR sites always present on both homologues of one pair, and other variable sites often present in heterozygous condition. Some NOR sites in these species were apparently bordered or intermingled with multiple copies of telomeric repeats (TTAGGGn), suggesting a possible mechanism for fusion/translocation rearrangements in the origin of new NOR sites (Reed & Phillips, 1995; Abuin et al., 1996). Reed & Phillips (1995) proposed a model that involves homologous chromosomes in translocation rearrangements to explain the variation in the number of NOR-bearing chromosomes observed among individuals in the lake trout, Salvelinus namaycush. Mechanisms such as unequal crossing-over or transposition have been suggested to be responsible for multichromosomal location of rRNA genes (Zhuo et al., 1995).
The characterization of polymorphism of rRNA genes at the population level is the first step to a better understanding of their evolutionary role and to obtain some clues about the mechanism responsible for their origin. The present findings extend our knowledge of the evolution and characteristics of rDNA sites in coregonid fishes, the earliest lineage evolved from a common tetraploid ancestor of salmonids. It seems that evolutionary mechanisms operating at karyotype level are very similar in both the Salmoninae and Coregoninae subfamilies.
References
Abuin, M., Martinez, P. and Sanchez, L. (1996). Localization of the repetitive telomeric sequences (TTAGGG)n in four salmonid species. Genome, 39: 1035–1038.
Allendorf, F. W. and Thorgaard, G. H. (1984). Tetraploidy and the evolution of salmonid fishes. In: Turner, B. J. (ed.) Evolutionary Genetics of Fishes, pp. 1–54. Plenum Press, New York.
Bernatchez, L. and Dodson, J. J. (1994). Phylogenetic relationships among Palearctic and Nearctic whitefish (Coregonus sp.) populations as revealed by mitochondrial DNA variation. Can J Fish Aquat Sci, 51: 240–251.
Bodaly, R. A., Vuorinen, J., Ward, R. D., Luczynski, M. and Reist, J. D. (1991). Genetic comparisons of New and Old World coregonid fishes. J Fish Biol, 38: 37–51.
Castro, J., Rodríguez, S., Pardo, B. G., Sanchez, L. and Martinez, P. (2001). Population analysis of an unusual NOR-site polymorphism in brown trout (Salmo trutta L.). Heredity, 86: 291–302, 10.1046/j.1365-2540.2001.00834.x.
Castro, J., Vinas, A., Sanchez, L. and Martinez, P. (1996). Characterization of an atypical NOR site polymorphism in brown trout (Salmo trutta) with Ag- and CMA3-staining, and fluorescent in situ hybridization. Cytogenet Cell Genet, 75: 234–239.
de Lucchini, S., Nardi, I., Barsacchi, G., Batistioni, R. and Andronico, F. (1993). Molecular cytogenetics of ribosomal (18S+28S and 5S) DNA loci in primitive and advanced urodele amphibians. Genome, 36: 762–763.
Fujiwara, A., Abe, S., Yamaha, E., Yamazaki, F. and Yoshida, M. C. (1998). Chromosomal localization and heterochromatin association of ribosomal regions in salmonid fishes. Chrom Res, 6: 463–471.
Galetti, P. M., Foresti, F., Bertollo, L. A. C. and Moreira-Filho, O. (1984). Characterization of eight species of Anostomidae (Cypriniformes) fish on the basis of the nucleolar organizing region. Caryologia, 37: 401–406.
Galetti, P. M. and Rasch, E. M. (1993). Chromosome studies in Poecilia latipunctata with NOR polymorphism as shown by silver nitrate and chromomycin A3 (Teleostei: Poeciliidae). Ichthyo Explo Freshwaters, 4: 269–277.
Hartley, S. E. (1987). The chromosomes of salmonid fishes. Biol Rev, 62: 197–214.
Howell, W. M. and Black, D. A. (1980). Controlled silver staining of nucleolus organizer regions with a protective colloidal developer: a 1-step method. Experientia, 36: 1014–1015.
Jankun, M., Boron, A., Kirtiklis, L., Kirchhoffer, A., Woznicki, P. and Luczynski, M. (1998a). Cytogenetic and biochemical studies on european whitefish (Coregonus lavaretus L.) from Switzerland. In: Eckmann, R., Appenzeller, A. and Rosch, R. (eds) Biology and Management of Coregonid Fishes. Arch Hybrobiol Spec Issues Advan Limnol, 50, 363–369.
Jankun, M., Klinger, M. and Woznicki, P. (1995). Chromosome variability in European vendace (Coregonus albula L.) from Poland. Caryologia, 48: 165–172.
Jankun, M., Martinez, P., Padro, B. G., Rab, P., Rabova, M. and Sanchez, L. (2000). rRNA genes map to chromosomes 10, 11 and 12 in European whitefish (Coregonus lavaretus) and to chromosomes 1, 5, 9 and 10 in vendace (Coregonus albula). Chrom Res, 8: 455–455.
Jankun, M., Ocalewicz, K. and Woznicki, P. (1998b). Replication, C- and fluorescent chromosome banding patterns in European whitefish, Coregonus lavaretus L., from Pomeranian Bay, Poland. Hereditas, 128: 195–199.
Jankun, M., Rab, P. and Vuorinen, J. (1991). A karyotype study of Coregonus albula (Pisces, Coregoninae) from Finland. Hereditas, 115: 291–294.
Juntunen, K. (1987). Kromosomimaaritys apuna siikojen taksonomisten ongelmien ratkaisemisessa. Riista-Ja Kalatalouden Tutkimuslaitos Kalantutkimusosasto, 64, 77 p. (in Finnish).
Kaidanova, T. I. (1989). Karyotype. In: Reshetnikov Yu. S and Mukhachev I. S. (eds) Coregonus peled (Gmelin, 1788): Systematics, Morphology, Ecology, pp. 87–94. Nauka, Moskva (in Russian).
Kottelat, M. (1997). European freshwater fishes. Biologia, Bratislava 52/Supplement, 5: 1–271.
Lyckegaard, E. M. S. and Clark, A. G. (1991). Evolution of ribosomal RNA gene copy number on the sex chromosomes of Drosophila melanogaster. Mol Biol Evol, 8: 458–474.
Martinez, P., Vinas, A., Bouza, C., Arias, J., Amaro, R. and Sanchez, L. (1991). Cytogenetical characterization of hatchery stocks and natural populations of sea and brown trout from northwestern Spain. Heredity, 66: 9–17.
Mellink, C. H. M., Bosma, A. A., de Haan, N. A. and Macdonald, A. A. (1992). Numerical variation of nucleolar organizer regions after silver staining in domestic and wild Suidae (Mammalia). Anim Genet, 23: 231–239.
Moss, T. and Stefanovsky, V. Y. (1995). Promotion and regulation of ribosomal transcription by RNA polymerase I. Prog Nucl Acid Res, 50: 207–250.
Pendas, A. M., Moran, P. and Garcia-Vazquez, E. (1993). Multi-chromosomal location of ribosomal RNA genes and heterochromatin associations in brown trout. Chrom Res, 1: 63–67.
Phillips, R. B. and Rab, P. (2001). Chromosome evolution in the Salmonidae (Pisces): an update. Biol Rev, 76: 1–25, 10.1017/s1464793100005613.
Phillips, R. B., Reed, K. M. and Rab, P. (1996). Revised karyotypes and chromosome banding of coregonid fishes from the Laurentian Great Lakes. Can J Zool, 74: 32–329.
Rab, P., Reed, K. M., de Ponce Leon, F. A. and Phillips, R. B. (1996). A new method for detecting nucleolus organizer regions in fish chromosomes using denaturation and propidium iodide staining. Biotech Histochem, 71: 157–162.
Reed, K. M. and Phillips, R. B. (1995). Molecular cytogenetic analysis of the double-CMA3 chromosome of lake trout, Salvelinus namaycush. Cytogenet Cell Genet, 70: 104–107.
Reed, K. M. and Phillips, R. B. (1997). Polymorphism of the nucleolus organizer region (NOR) on the putative sex chromosomes of Arctic char (Salvelinus alpinus) is not sex related. Chrom Res, 5: 221–227.
Reist, D. J., Maiers, L. D., Bodaly, R. A., Vuorinen, J. A. and Carmichael, T. (1998). The phylogeny of new- and old-world coregonine fishes as revealed by sequence variation in a portion of the d-loop of mitochondrial DNA. In: Eckmann, R., Appenzeller, A. and Rosch, R. (eds) Biology and Management of Coregonid Fishes. Arch Hybrobiol Spec Issues Advan Limnol, 50, 323–339.
Schmid, M., Feichtinger, W., Weimer, R., Mais, C., Bolaños, F. and León, P. (1995). Chromosome banding in Amphibia. XXI. Inversion polymorphism and multiple nucleolus organizer regions in Agalychnis callidryas (Anura, Hylidae). Cytogenet Cell Genet, 69: 18–26.
Schubert, I. and Wobus, U. (1985). In situ hybridization confirms jumping nucleolus organizing regions in Allium. Chromosoma, 92: 143–148.
Sola, L., Rossi, A. R., Iaselli, V., Rasch, E. M. and Monaco, P. J. (1992). Cytogenetics of bisexual/unisexual species of Poecilia. II. Analysis of heterochromatin and nucleolar organizer regions in Poecilia mexicana maxicana by C-banding and DAPI, quinacrine, chromomycin A3, and silver staining. Cytogenet Cell Genet, 60: 229–235.
Souza, I. L. and Moreira-Filho, O. (1995). Cytogenetic diversity in the Astyanax scabripinnis species complex (Pisces, Characidae). I. Allopatric distribution in a small stream. Cytologia, 60: 1–11.
Svardson, G. (1970). Significance of introgression in coregonid evolution. In: Lindsey, C. C. and Woods, C. S. (eds) Biology of Coregonid Fishes, pp. 33–59. University of Manitoba Press, Winnipeg, Canada.
Wilson, G. N., Hollar, B. A., Waterson, R. J. and Schmickel, R. D. (1978). Molecular analysis of cloned human 18S ribosomal DNA segments. Proc Natl Acad Sci USA, 15: 5367–5371.
Woznicki, P., Jankun, M., Kucharczyk, D., Boron, A. and Luczynski, M. (1999). Cytogenetic characterization of sea trout Salmo trutta from Poland. Copeia, 2: 501–505.
Zhuo, L., Reed, K. M. and Phillips, R. B. (1995). Hypervariability of ribosomal DNA at multiple chromosomal sites in lake trout (Salvelinus namaycush). Genome, 38: 487–496.
Acknowledgements
This study was supported by Project No. 5 P06D 024 17, financed by KBN for M.J., Project S 5045011 by CAS for P.R. and M.R. This paper is contribution No. 40 in the Program of Joint Investigation of Holarctic Fishes among Russia, Canada, Finland and Poland.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Jankun, M., Martinez, P., Pardo, B. et al. Ribosomal genes in Coregonid fishes (Coregonus lavaretus, C. albula and C. peled) (Salmonidae): single and multiple nucleolus organizer regions. Heredity 87, 672–679 (2001). https://doi.org/10.1046/j.1365-2540.2001.00964.x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1046/j.1365-2540.2001.00964.x
Keywords
This article is cited by
-
Vendace (Coregonus albula) and least cisco (Coregonus sardinella) are a single species: evidence from revised data on mitochondrial and nuclear DNA polymorphism
Hydrobiologia (2021)
-
Cytogenetic investigation of Cyprinus carpio (Linnaeus, 1758) using giemsa, silver nitrate, CMA3 staining and fluorescence in situ hybridization
The Nucleus (2017)
-
Chromosomal characteristics and distribution of rDNA sequences in the brook trout Salvelinus fontinalis (Mitchill, 1814)
Genetica (2015)
-
Genome differentiation in a species pair of coregonine fishes: an extremely rapid speciation driven by stress-activated retrotransposons mediating extensive ribosomal DNA multiplications
BMC Evolutionary Biology (2013)
-
Cytogenetic characterization of an endemic mahseer, Tor mosal mahanadicus (David, 1953; Teleostei : Cyprinidae)
The Nucleus (2010)