Abstract
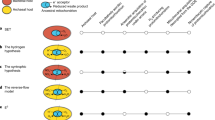
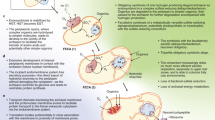
A new hypothesis for the origin of eukaryotic cells is proposed, based on the comparative biochemistry of energy metabolism. Eukaryotes are suggested to have arisen through symbiotic association of an anaerobic, strictly hydrogen-dependent, strictly autotrophic archaebacterium (the host) with a eubacterium (the symbiont) that was able to respire, but generated molecular hydrogen as a waste product of anaerobic heterotrophic metabolism. The host's dependence upon molecular hydrogen produced by the symbiont is put forward as the selective principle that forged the common ancestor of eukaryotic cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Müller, M. The hydrogenosome. J. Gen. Microbiol. 139, 2879–2889 (1993).
Cavalier-Smith, T. Eukaryotes with no mitochondria. Nature 326, 332–333 (1987).
Sogin, M. L., Silberman, J. D., Hinkle, G. & Morrison, H. G. Problems with molecular diversity in the Eukarya. Symp. Soc. Gen. Microbiol. 54, 167–184 (1996).
Whatley, J. M., John, P. & Whatley, F. R. From extracellular to intracellular: the establishment of mitochondria and chloroplasts. Proc. R. Soc. Lond. B 204, 165–187 (1979).
Gray, M. W. & Doolittle, W. F. Has the endosymbiont hypothesis been proven? Microbiol. Rev. 46, 1–42 (1982).
Cavalier-Smith, T. The origin of eukaryote and archaebacterial cells. Ann. NY Acad. Sci. 503, 7–54 (1987).
Cavalier-Smith, T. & Chao, E. E. Molecular phylogeny of the free-living archaezoan Trepomonas agilis and the nature of the first eukaryote. J. Mol. Evol. 43, 551–562 (1996).
Zillig, W. et al. Did eukaryotes originate by a fusion event? Endocytobiosis Cell Res. 6, 1–25 (1989).
Gupta, R. S. & Golding, G. B. The origin of the eukaryotic cell. Trends Biochem. Sci. 21, 166–171 (1996).
Lake, J. A. & Rivera, M. C. Was the nucleus the first endosymbiont? Proc. Natl Acad. Sci. USA 91, 2880–2881 (1994).
Clark, C. G. & Roger, A. J. Direct evidence for secondary loss of mitochondria in Entamoeba histolytica. Proc. Natl Acad. Sci. USA 92, 6518–6521 (1995).
Henze, K. et al. Anuclear gene of eubacterial origin in Euglena reflects cryptic endosymbioses during protist evolution. Proc. Natl Acad. Sci. USA 92, 9122–9126 (1995).
Keeling, P. W. & Doolittle, W. F. Evidence that eukaryotic triosephosphate isomerase is of alpha-proteobacterial origin. Proc. Natl Acad. Sci. USA 94, 1270–1275 (1997).
Doolittle, W. F. Some aspects of the biology of cells and their possible evolutionary significance. Symp. Soc. Gen. Microbiol. 54, 1–21 (1996).
Rosenthal, B. et al. Evidence for the bacterial origin of genes encoding fermentation enzymes of the amitochondriate protozoan parasite Entamoeba histolytica. J. Bacteriol. 179, 3736–3745 (1997).
Searcy, D. G. in The Origin and Evolution of the Cell (eds Hartman, H. & Matsuno, K.) 47–78 (World Scientific, Singapore, (1992)).
de Duve, C. Blueprint for a Cell: the Nature and Origin of Life (Patterson, Burlington, NC, (1991)).
Coombs, G. H. & Müller, M. in Biochemistry and Molecular Biology of Parasites (eds Marr, J. J. & Müller, M.) 33–47 (Academic, London, (1995)).
Müller, M. in Evolutionary Relationships Among Protozoa (eds Coombs, G. H., Vickermann, K., Sleigh, M. A. & Warren, A.) 109–132 (Chapman Hall, London, (1998)).
Müller, M. Energy metabolism of protozoa without mitochondria. Annu. Rev. Microbiol. 42, 465–488 (1988).
Müller, M. in Christian Gottfried Ehrenberg-Festschrift anla¨ßlich der 14. Wissenschaftlichen Jahrestagung der Deutschen Gesellschaft für Protozoologie, 9.-11. Marz 1995 in Delitzsch (Sachsen) (eds Schlegel, M. & Hausmann, K.) 63–76 (Leipziger Universitätsverlag, Leipzig, (1996)).
Iwabe, N., Kuma, K.-I., Hasegawa, M., Osawa, S. & Miyata, T. Evolutionary relationship of archaebacteria, eubacteria and eukaryotes inferred from phylogenetic trees of duplicated genes. Proc. Natl Acad. Sci. USA 86, 9355–9359 (1989).
Woese, C., Kandler, O. & Wheelis, M. L. Towards a natural system of organisms: proposal for the domains Archaea, Bacteria and Eukarya. Proc. Natl Acad. Sci. USA 87, 4576–4579 (1990).
Langer, D., Hain, J., Thuriaux, P. & Zillig, W. Transcription in Archaea: similarity to that in Eukarya. Proc. Natl Acad. Sci. USA 92, 5768–5772 (1995).
Yamamoto, A., Hashimoto, T., Asaga, E., Hasegawa, M. & Goto, N. Phylogenetic position of the mitochondrion-lacking protozoan Trichomonas tenax, based on amino acid sequences of elongation factors 1-α and 2. J. Mol. Evol. 44, 98–105 (1997).
Horner, D. S., Hirt, R. P., Kilvington, S., Lloyd, D. & Embley, T. M. Molecular data suggest an early acquisition of the mitochondrion endosymbiont. Proc. R. Soc. Lond. B 263, 1053–1059 (1996).
Bui, E. T. N., Bradley, P. J. & Johnson, P. J. Acommon evolutionary origin for mitochondria and hydrogenosomes. Proc. Natl Acad. Sci. USA 93, 9651–9656 (1996).
Germot, A., Philippe, H. & Le Guyader, H. Presence of a mitochondrial-type 70-kDa heat shock protein in Trichomonas vaginalis suggests a very early mitochondrial endosymbiosis in eukaryotes. Proc. Natl Acad. Sci. USA 93, 14614–14617 (1996).
Roger, A. J., Clark, C. G. & Doolittle, W. F. Apossible mitochondrial gene in the early-branching amitochondriate protist Trichomonas vaginalis. Proc. Natl Acad. Sci. USA 93, 14618–14622 (1996).
Hrdý, I. & Müller, M. Primary structure and eubacterial relationships of the pyruvate : ferredoxin oxidoreductase of the amitochondriate eukaryote, Trichomonas vaginalis. J. Mol. Evol. 41, 388–396 (1995).
Blattner, F. R. et al. The complete genome sequence of Escherichia coli K-12. Science 277, 1453–1474 (1997).
Kaneko, T. et al. Sequence analysis of the genome of the unicellular cyanobacterium Synechocystis sp. strain PCC6803. II. Sequence determination of the entire genome and assignment of potential protein-coding regions. DNA Res. 3, 109–136 (1996).
Sánchez, L. B. & Müller, M. Purification and characterization of the acetate forming enzyme, acetyl-CoA synthetase (ADP-forming) from the amitochondriate protist, Giardia lamblia. FEBS Lett. 378, 240–244 (1996).
Schönheit, P. & Schäfer, T. Metabolism of hyperthermophiles. World. J. Microbiol. Biotechnol. 11, 26–57 (1995).
Markoŝ, A., Miretsky, A. & Müller, M. Aglyceraldehyde-3-phosphate dehydrogenase with eubacterial features in the amitochondriate eukaryote Trichomonas vaginalis. J. Mol. Evol. 37, 631–643 (1993).
Martin, W. & Schnarrenberger, C. The evolution of the Calvin cycle from prokaryotic to eukaryotic chromosomes: a case study of functional redundancy in ancient pathways through endosymbiosis. Curr. Genet. 32, 1–18 (1997).
Fenchel, T. & Finlay, B. J. Ecology and Evolution in Anoxic Worlds (Oxford Univ. Press, Oxford, (1995)).
Gibson, J. L. & Tabita, F. R. The molecular regulation of the reductive pentose phosphate pathway in proteobacteria and cyanobacteria. Arch. Microbiol. 166, 141–150 (1996).
Murrel, J. C. Genetics and molecular biology of methanotrophs. FEMS Microbiol. Lett. 88, 233–248 (1992).
Thauer, R. K., Hedderich, R. & Fischer, R. in Methanogenesis: Ecology, Physiology, Biochemistry and Genetics (ed. Ferry, J. G.) 209–252 (Chapman & Hall, New York, (1993)).
Conrad, R. Soil microorganisms as controllers of atmospheric trace gases (H2, CO, CH4, OCS, N2, and NO). Microbiol. Rev. 60, 609–640 (1996).
Bryant, M. P., Wolin, E. A., Wolin, M. J. & Wolfe, R. S. Methanobacillus omelianskii, a symbiotic association of two species of bacteria. Arch. Microbiol. 59, 20–31 (1967).
Broers, C. A. M., Stumm, C. K., Vogels, G. D. & Brugerolle, G. Psalteriomonas lanterna gen. nov., sp. nov., a free living amoboflagellate isolated from freshwater anaerobic sediments. Eur. J. Protistol. 25, 369–380 (1990).
Embley, T. M. et al. Multiple origins of anaerobic ciliates with hydrogenosomes within the radiation of aerobic ciliates. Proc. R. Soc. Lond. B 262, 87–93 (1995).
Finlay, kB. J., Embley, T. M. & Fenchel, T. Anew polymorphic methanogen, closely related to Methanocorpusculum parvum, living in stable symbiosis within the anaerobic ciliate Trimyema sp. J. Gen. Microbiol. 139, 371–378 (1993).
Stevens, T. O. & McKinley, J. P. Lithoautotrophic microbial ecosystems in deep basalt aquifers. Science 270, 450–454 (1995).
Brinkmann, H. & Martin, W. Higher plant chloroplast and cytosolic 3-phosphoglycerate kinases: a case of endosymbiotic gene replacement. Plant. Mol. Biol. 30, 65–75 (1996).
Kasting, J. F. Earth's early atmosphere. Science 259, 920–926 (1993).
Poole, A. M., Jeffares, D. C. & Penny, D. The path from the RNA world. J. Mol. Evol. 46, 1–17 (1998).
Rospert, S. et al. Methyl-coenzyme M reductase and other enzymes involved in methanogenesis from CO2and H2in the extreme thermophile Methanopyrus kandleri. Arch. Microbiol. 156, 49–55 (1991).
Acknowledgements
We thank H. Brinkmann, M. Embley, K. Henze, R. Herrmann, R. Hensel, D.Oesterheld and L. Sánchez for critical comments on the manuscript and gratefully acknowledge financial support from the Deutsche Forschungsgemeinschaft (W.M.) and the National Institutes of Health (M.M.).
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Martin, W., Müller, M. The hydrogen hypothesis for the first eukaryote. Nature 392, 37–41 (1998). https://doi.org/10.1038/32096
Issue Date:
DOI: https://doi.org/10.1038/32096
This article is cited by
-
Human genetic associations of the airway microbiome in chronic obstructive pulmonary disease
Respiratory Research (2024)
-
Horizontal gene transfer in eukaryotes: aligning theory with data
Nature Reviews Genetics (2024)
-
Actin cytoskeleton and complex cell architecture in an Asgard archaeon
Nature (2023)
-
Eco-evolutionary modelling of microbial syntrophy indicates the robustness of cross-feeding over cross-facilitation
Scientific Reports (2023)
-
Eukaryotes were shaped by Oxygen
Nature Ecology & Evolution (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.