Abstract
Several members of a three-generation kindred from Sardinia were affected by a maternally inherited syndrome characterized by features of both myoclonus epilepsy with ragged-red fibers (MERRF) and mitochondrial encephalomyopathy with lactic acidosis and stroke-like episodes (MELAS). Clinically, symptoms such as myoclonus epilepsy, neural deafness and ataxia were variably associated with stroke-like episodes and/or migrainous attacks. Morphologically, numerous ME-LAS-associated SDH-stained vessels were observed in muscle biopsies, either alone or in combination with ragged-red fibers, the morphological hallmark of MERRF. Sequence analysis of the mtDNA tRNA genes revealed the presence of a single, heteroplasmic T → C transition at nt 8356, in the region of the tRNALys gene corrsponding to the T-Ψ-C stem. The T → C(8356) transition was exclusively found in the maternal lineage of our family, and the relative amount of the mutant mtDNA species in muscle was correlated with the severity of the clinical presentation. Therefore, we propose that the T → C(8356) transition is responsible for the mitochondrial encephalomyopathy found in our family, and must be added to the expanding list of the pathogenetically relevant mutations of human mtDNA.
Similar content being viewed by others
Introduction
Maternally inherited point mutations of mtDNA are responsible for several neurological syndromes in humans, including Leber’s hereditary optic neuroretinopathy (LHON), neuropathy-ataxia-retinitis pigmentosa (NARP), maternally inherited myopathy and cardiopathy (MIMyCa), myoclonus epilepsy with ragged-red fibers (MERRF), and mitochondrial encephalomyopathy with lactic acidosis and stroke-like episodes (MELAS) [reviewed in ref. 1]. While LHON and NARP can be caused by mutations in mRNA-encoding mtDNA genes, MIMyCa, MERRF and MELAS are well-characterized mitochondrial disorders due to point mutations in tRNA-encoding mtDNA genes. In particular, a heteroplasmic mutation linked to MERRF has been identified as a G → A transition at mtDNA nt 8344, in the region encoding the T-Ψ;-C loop of tRNALys [2]. Several groups, including ours [3], have reported the association of the G → A(8344) mutation in many MERRF families belonging to different ethnic groups; thus, this mutation appears to be the most frequent and widespread, although not the exclusive, cause of MERRF. On the other hand, most of the MELAS cases have been associated with a heteroplasmic A → G transition at mtDNA nt 3243, in the region encoding the DHU loop of tRNALeu(UUR) [4, 5]. A second MELAS mutation has been recently reported at nt 3271, in the anticodon stem of the same tRNA gene [6].
We report here the identification in a large pedigree of a new mtDNA point muation, located in the T-Ψ-C stem of the mtDNA tRNALys gene, etiologically related to a maternally inherited syndrome sharing features of both MERRF and MELAS.
Patients
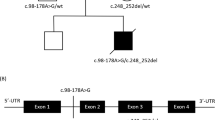
The family originates from a small village in northern Sardinia; the pedigree tree is shown in figure 1. Maternal family members available for this study were divided into three groups: four severely affected patients (I-1, II-2, II-8, III-1), three mildly affected patients (II-4, II-5, II-7), and four clinically asymptomatic relatives (II-1, II-3, II-6, II-9); in addition, a brother of patient I-1, not examined by us, was referred to by his relatives as affected by symptoms compatible with a metabolic myopathy, but no further details could be obtained.
a Pedigree of the propositi. Solid symbols = severely affected individuals; half-solid symbols = moderately affected individuals; Shaded symbols = asymptomatic individuals with mitochondrial abnormalities; open symbols = normal individuals; * = DNA analyzed. b Diagnostic XbaI digestion of PCR-amplified mtDNA fragments. Samples of DNA from each propositus are indicated according to the nomenclature of the pedigree.
Group 1: Severely Affected Patients
Patient I-1, a 54-year-old woman, complained of neural deafness from her childhood, followed by bilateral cataract, and, later, by myoclonus, ataxia and dementia. One of her daughters, patient II-2, died at 32 years of age of a massive cerebral stroke after having suffered during her life from several stroke-like episodes. In addition, she was affected by myoclonus epilepsy and neural deafness from the age of 12. Severe ataxia, frequent migrainous attacks and dementia ensued in the later years. Her 11 -year-old son, patient III-1, was affected by severe myoclonus epilepsy, cerebellar ataxia, and mild neural deafness. Patient II-8, a 16-year-old sister of patient II-2, suffered from severe migrainous attacks from the age of 14, and, more recently, developed myoclonus epilepsy. She was also affected by a multinodular euthyroid goiter. Basal blood lactate was in the upper normal range in patient I-1, and moderately elevated in patients II-8 and III-1 (see table 1). Serum creatine kinase (CK) was also moderately elevated in both patients III-1 and II-8 (301 and 547 U/l, respectively; normal values: 25–190 U/l).
Group 2: Moderately Affected Patients
Patient II-4, a 23-year-old sister of patient II-2, had an isolated, mild neural deafness. Another 17-year-old sister, patient II-7, was affected by both deafness and euthyroid goiter. The latter was the only symptom of patient II-5, a 21-year-old sister of patient II-2. In all three patients the blood lactate and serum CK were normal.
No patient of either group had ptosis, ophthalmoplegia, retinal- or optic-nerve abnormalities. Urinalysis, routine blood tests, lysosomal enzymes in leukocytes, and ECG were all normal.
Group 3: Clinically Asymptomatic Maternal Relatives
Individuals II-1 (30 years old), II-3 (27 years old), II-6 (23 years old), and II-9 (9 years old), brothers of patient II-2, were all clinically normal. Subjects II-6 and II-9, however, both showed EEG abnormalities (sharp theta waves on the parietal regions).
Materials and Methods
Muscle Biopsies, Morphological and Biochemical Studies
A needle muscle biopsy was performed, after informed consent, in the following subjects: I-1, II-3, II-4, II-6, II-7, II-8, II-9 and III-1.
Isolation of blood lymphocytes was carried out using standard procedures.
Muscle biopsies were divided into two portions: one was stored in N2-cooled liquid isopentane for histological and histochemical examination, the other was frozen in liquid nitrogen for mtDNA analysis. Given the small size of the biopsies (approximately 50 mg), biochemical analysis of the mitochondrial respiratory chain could not be performed.
Molecular Genetic Studies
Extraction of total DNA from muscle and lymphocytes, polymerase-chain-reaction (PCR) amplifications and direct sequencing of double-stranded DNA were performed as described previously [7], using suitable pairs of 25-mer primers (sense and antisense), corresponding to portions of human mtDNA flanking the 22 tRNA genes.
The rapid screening of the patients’ mtDNAs was based on mispaired PCR, as described elsewhere [3, 7]. We replaced a G nucleotide with a C nucleotide in an antisense 50-mer primer, corresponding to the mtDNA Cambridge sequence [8] from nt 8357 to nt 8407. The nucleotide substitution corresponded to position 8359 of the antisense mtDNA sequence. The ‘sense’, unmodified primer corresponded to the region from nt 8200 to nt 8225 of the Cambridge sequence. In the presence of the mutant C(8356), but not of the wild-type T(8356), the modification of the primer created a new, diagnostic XbaI restriction site. Since a second XbaI site is normally present at position 8286, the wild-type PCR product was cleaved by XbaI into two fragments of 120 and 86 bp. The latter fragment was also present after XbaI digestion of the mutant PCR product; however, the 120-bp fragment which contained the mutant C(8356), was further cleaved into two fragments of 51 and 69 bp. The cleavage products of the mutant and wild-type PCR products were separated and visualized by electrophoresis on an ethid-ium-bromide-stained 6% acrylamide-TBE gel.
The same diagnostic procedure was also carried out on 327 DNA samples collected from 167 muscle biopsies and 160 blood lymphocyte pellets, obtained from as many unrelated normal and diseased control individuals. The ‘muscle mtDNA’ control group included 25 normal individuals, 6 patients with Kearns-Sayre syndrome (2 harboring a single mtDNA deletion), 33 patients with progressive external ophthalmoplegia ‘plus’ (2 harboring a mtDNA deletion), 8 patients with MELAS (4 harboring the A → G(3243) mutation), 9 patients with MERRF (7 harboring the A → G(8344) mutation), 3 patients with LHON, 14 patients with Leigh’s syndrome associated with cytochrome C oxidase (COX) deficiency, 28 patients with a nonclassified mitochondrial encephalomyopathy, 13 patients with dyssynergia cerebellaris myoclonica, 29 patients with other neurological syndromes. The ‘lymphocyte mtDNA’ group was composed of DNA samples from 160 individuals originating from Sardinia, including 100 normal individuals, 30 patients affected by multiple sclerosis and 30 patients affected by β-thalassemia.
The proportion of mutant mtDNA was calculated by densitometry of the ethidium-bromide-stained gels as described elsewhere [2], using a Mitsubishi Video Copy Processor (Model P68E), an IV-530 Contour Synthesizer (FOR-A, France) and the Bio-Profil (Vil-ber-Lourmat, France) software package.
Results
Morphological Studies in Muscle
The severely affected patients that could be examined were subjects I-1, II-8 and III-1. Both patients I-1 and III-1 had ragged-red fibers (RRF). Figure 2a shows an example from the muscle biopsy of patient III-1. Moreover, numerous fibers, including the ragged-red ones, were negative for the histochemical reaction of COX (fig. 2b). Atrophic fibers were present in the muscle biopsy of patient I-1. In addition, in the muscle biopsy of patient III-1, the walls of numerous vessels appeared heavily stained at the histochemical reaction for succinic dehydrogenase (SDH) (fig. 2c). These SDH-stained Vessels (SSVs) are considered to be a morphological hallmark of MELAS [9]. Vessels from non-MELAS specimens are virtually unstainable by the histochemical reaction for SDH. Abnormal mitochondrial proliferation and intramitochondrial paracristalline inclusions and concentric cristae were found at the ultra-structural level (not shown). Partial COX depletion, neurogenic changes and numerous SSVs were also present in the muscle biopsy of patient II-8, but RRF were not observed. Among the moderately affected patients, the muscle biopsy of subject II-4 showed SSVs, rare RRF and COX-depeleted fibers and mild neurogenic changes, while the presence of SSVs was the only pathological finding in subject II-7.
Of the three clinically normal subjects (II-3, II-6, II-9), the muscle biopsy of individual II-6 showed rare ragged-red, COX-depleted fibers, as well as SSVs. SSVs and mild signs of neurogenic atrophy were also present in the biopsy of subject II-9, while the biopsy of individual II-3 was normal.
Molecular-Genetic Studies
Identification of a mtDNA tRNALys Mutation. We initially analyzed the muscle mtDNA of patient III-l (mtDNA(III-1). Major rearrangements or known point mutations of mtDNA(III-1) were ruled out (not shown). Nucleotide sequences of tRNA-containing fragments of mtDNA(III-1) were then compared with the corresponding mtDNA Cambridge sequence.
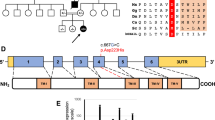
The only difference observed was a cytosine nucleotide replacing the wild-type thymine at position 8356 of the tRNALys (fig. 3). Since the mutant species represented approximately 84% of mtDNA(III-1) (see (table 1), only the C(8356) was detectable in the sequence.
Figure 3 shows the location of the T->C(8356) transition in the tRNALys proposed secondary structure. The wild-type T(8356) is normally bound to the A located 18 positions upstream, at nt 8338, as a part of a 5-bp palindrome which forms the T-Ψ-C stem of the tRNALys cloverleaf. Although this T-A pair is only moderately conserved through animal species [2], in mtDNA(III-1) the C(8356) mutation creates a mismatch with the corresponding A(8338), thus disrupting the pair, and presumably affecting the stability of the tRNA T-Ψ-C stem.
Segregation of the T->C(8356) Mutation
Using a diagnostic digestion with XbaI we analyzed the muscle (fig. 1) and lymphocyte (not shown) mtDNA for the presence of the T->C(8356) transition in the available family members.
The second lane of the gel shown in figure 1 shows the wild-type pattern obtained from the muscle mtDNA of a normal family member, II-3. The 120-and 86-bp bands correspond to the cleavage products of the wild-type XbaI site at nt 8286. By contrast, the muscle mtDNAs of the 5 clinically affected patients who were analyzed (patients I-1, II-4, II-7, II-8, III-1), and of 2 out of 3 clinically normal maternal relatives (II-6, II-9) presented a four-band pattern, as would be expected in cases of heteroplasmy for the T->C(8356) transition.
A homoplasmic wild-type two-band pattern was also obtained in all 327 human DNA samples used as controls.
Quantitative Analysis of the T->C(8356) Heteroplasmy. Table 1 shows that the most severely affected patients who were available for this study (subjects I-1 and III-1) had a relative amount of mutated mtDNA well over 80%. Patient II-8, who had less severe myoclonic epilepsy, migrainous attacks and deafness, had 57% of mutated mtDNA in her muscle biopsy, while other subjects, either moderately affected patients or normal individuals, had widely distributed values, ranging from 57 (individual II-6) to 0% (individual II-3). Thus, the degree of mtDNA heteroplasmy seems to correlate with the severity of the clinical presentation and/or with the laboratory indexes of a mitochondrial disorder.
The mutant C(8356) was barely detectable in the lymphocyte mtDNA of the severely affected patients (I-1, II-8, III-1), and absent in the lymphocyte mtDNA of the other family members (not shown).
Discussion
A mtDNA etiology of the syndrome was suggested by the following facts. First, patients had a mitochondrial disorder, demonstrated by the presence of RRF and/or SSVs, abnormal mitochondria at the electron microscope examination, variably elevated levels of blood lactate, and a defective histochemical reaction of COX. Second, with the exception of subject II-3, the maternal relatives had signs of a subclinical mitochondrial disorder, such as proliferation of, and/or histochemical COX deficiency in muscle mitochondria. Third, the pattern of transmission of the trait was compatible with maternal inheritance, because the affected individuals all belonged to the maternal lineage of a single three-generation kindred.
A distinct clinical feature was the presence of thyroid hyperplasia in three members of the family. Although thyroid hyperplasia is endemic in Sardinia, we tentatively attributed this sign to the underlying mitochondrial disorder because we observed the presence of euthyroid goiter in families with mitochondrial encephalomyopathies originating from other Italian regions [in preparation].
A T->C transition, creating a mismatch in the T-Ψ-C stem of the tRNALys mtDNA gene, was identified in the maternal lineage of the kindred. The mutation specifically segregated with the disease: none of 327 normal and diseased controls had the mutation. Moreover, the mutation was heteroplasmic. Heteroplasmy is an indicator of pathogenic mutation: it has never been reported in normal individuals, and it can also explain the variable expressivity of the phenotype. Finally, the proportion of mutant mtDNA in muscle was positively correlated with the severity of the clinical presentation (see table 1 and fig. 1). Incidentally, we noticed that subject II-3, who was completely free of any clinical, morphological or laboratory abnormality, was the only family member whose muscle mtDNA was homoplasmic for the wild-type T(8356). The pathogenicity of the mutation is further supported by the substantial decrease of mutant mtDNA in lymphocytes, suggesting the presence of negative selection against the survival of mitotically active cells harboring the mutation.
From a molecular-genetic point of view, it is interesting to note that the T->C(8356) transition not only affects the same tRNA gene but also the same functional region as the MERRF-associated mutation originally described by Shoffner et al. [2]. However, from a clinical point of view, some members of this family presented features of MERRF, while others shared features of both MERRF and MELAS, the myoclonus epilepsy being variably associated with stroke-like episodes, migrainous attacks and lactic acidosis. Histologically, all examined individuals, except subject II-3, had SSVs, a morphological hallmark of MELAS, either alone or in addition to RRF and COX depletion. As typically found in lesions of mtDNA [1, 10], symptoms ranged from isolated deafness, as in patient II-4, to a moderately severe encephalomyopathy as in patient II-8, to a severe syndrome including muscle wasting, ataxia and myoclonus epilepsy (patient III-1). Dementia was an additional feature in patients I-1 and II-2; the latter died precociously after several stroke-like episodes. Overlap syndromes of MERRF and MELAS have been decribed in the past. MERRF patients may develop overt strokelike episodes or show CT or MRI evidence of silent infarcts [11, 12]. On the other hand, patients with MELAS can have myoclonus [13–16]. However, it is possible that the overlap syndrome of this family may be specifically linked to the new T → C(8356) mutation. In addition, it is known that the concordance between the clinical phenotype and different mtDNA mutations of tRNA genes may vary: for instance, the A->G(3243) MELAS mutation can be often found in patients affected by other mitochondrial encephalomyopathies, including progressive external ophthalmoplegia with RRF, and nonclassified cases [17]. Alternatively, a specific ‘nuclear-genetic background’, or the presence of a second, as yet unidentified, mtDNA mutation which could synergistically act with the T->C(8356) transition, may account for the peculiar combination of symptoms found in our family.
References
Zeviani M, Antozzi C: Defects of mitochondrial DNA. Brain Pathol 1992;2:121–132
Shoffner JM, Lott MT, Lezza AMS, Seibel P, Ballinger SW, Wallace DC: Myoclonic epilepsy and ragged-red fibers disease (MERRF) is associated with a mitochondrial DNA tRNALys mutation. Cell 1990;61:931–937
Zeviani M, Amati P, Bresolin N, Antozzi C, Piccolo G, Toscano A, DiDonato S: Rapid detection of the A → G(8344) mutation of mtDNA in Italian families with myoclonus-epilepsy and ragged-red fibers (MERRF). Am J Hum Genet 1991;48:203–211
Goto Y, Nonaka I, Horai S: A mutation in the tRNAleu(UUR) gene associated with MELAS subgroup of mitochondrial encephalomyopathies. Nature 1990;348:651–653
Ino H, Tanaka M, Ohno K, Hattori K, Ikebe S, Sano T, Ozawa T, Ichici T, Kobayashi M, Wada Y: Mitochondrial leucine tRNA mutation in a mitochondrial encephalomyopathy. Lancet 1991;337:234–235
Goto Y, Nonaka I, Horai S: A new mtDNA mutation associated with mitochondrial myopathy, encephalopathy, lactic acidosis and strokelike episodes (MELAS). Biochim Biophys Acta 1992;1097:238–240
Zeviani M, Gellera C, Antozzi C, Rimoldi M, Morandi L, Villani F, Tiranti V, DiDonato S: Maternally inherited myopathy and cardiomyopathy: Association with mutation in mitochondrial DNA tRNALeu-(UUR). Lancet 1991;338:143–147
Anderson S, Bankier AT, Barrell BG, de Bruijn MHL, Coulson AR, Drouin J, Eperon IC, Nierlich DP, Roe BA, Sanger F, Schreier PH, Smith AJH, Staden R, Young IG: Sequence and organization of the human mitochondrial genome. Nature 1981;290:457–465
Hasegawa H, Matsuoka T, Goto Y, Nonaka I: Strongly succinate dehydrogenase-reactive blood vessels in muscles from patients with mitochondrial myopathy, encephalopathy, lactic acidosis, and stroke-like episodes. Ann Neurol 1991;29:601–605
Wallace DC: Maternal genes: Mitochondrial diseases. Birth Defects 1987;23:137–190
Byrne E, Trounce I, Dennett X, Gilligan B, Morley JB, Marzuki S: Progression from MERRF to MELAS phenotype in a patient with combined respiratory complex I and IV deficiencies. J Neurol Sci 1988;88:327–337
Suzuki T, Koizumi J, Ishikawa N, Ofuku K, Sasaki M, Hori T, Ohkoshi N, Anno I: Mitochondrial encephalomyopathy (MELAS) with mental disorder. CT, MRI and SPECT findings. Neuroradiology 1990;32:74–76
Petty RKH, Harding AE, Morgan-Hughes JA: The clinical features of mitochondrial myopathy. Brain 1986;109:915–938
Peiffer J, Kustermann-Kuhn B, Mortier W, Poremba M, Roggendorf W, Scholte HR, Schröder JM, Wentland B, Wessel K, Zimmermann C: Mitochondrial myopathies with necrotizing encephalopathy of the Leigh type. Pathol Res Pract 1988;183:706–716
Truong DD, Harding AE, Scaravilli F, Smith SJM, Morgan-Hughes JA, Marsden CD: Movement disorders in mitochondrial myopathies. A study of nine cases with two autopsy studies. Mov Disord 1990;5:109–117
Abe K, Inui T, Hirono N, Mezaki T, Kobayashi Y, Kameyama M: Fluctuating MR images with mitochondrial encephalomyopathy, lactic acidosis, stroke-like syndrome (MELAS). Neuroradiology 1990;32:77.
Ciafaloni E, Ricci E, Shanske S, Moraes CT, Silvestri G, Mirano M, Simonetti S, Angelini C, Donati MA, Garcia C, Martinuzzi A, Mosewich R, Servidei S, Zammarchi E, Bonilla E, De Vivo DC, Rowland LP, Schon EA, DiMauro S: MELAS: Clinical features, biochemistry, and molecular genetics. Ann Neurol 1992;31:391–398
Silvestry G, Moraes CT, Shanske S, Oh SJ, DiMauro S: A new mtDNA mutation in the tRNALys gene associated with Myoclonic Epilepsy and Ragged Red Fibers (MERRF). Am J Hum Genet 1992, in press.
Acknowledgements
Supported in part by Telethon-Italy and ARIN (As-sociazione Italiana per la promozione delle Ricerche Neurologiche). NS is a Dottorando di Ricerca in Neurosciences at the University of Naples 1 st School of Medicine.
We are indebted to Dr. Marina Mora for expert technical advice.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Zeviani, M., Muntoni, F., Savarese, N. et al. A MERRF/MELAS Overlap Syndrome Associated with a New Point Mutation in the Mitochondrial DNA tRNALys Gene. Eur J Hum Genet 1, 80–87 (1993). https://doi.org/10.1159/000472390
Received:
Revised:
Accepted:
Issue Date:
DOI: https://doi.org/10.1159/000472390
Key Words
This article is cited by
-
Transfer RNA detection by small RNA deep sequencing and disease association with myelodysplastic syndromes
BMC Genomics (2015)
-
New treatment options for hearing loss
Nature Reviews Drug Discovery (2015)
-
MERRF/MELAS overlap syndrome due to the m.3291T>C mutation
Metabolic Brain Disease (2014)
-
Mitochondrial matters of the brain: mitochondrial dysfunction and oxidative status in epilepsy
Journal of Bioenergetics and Biomembranes (2010)
-
Mitochondrial dysfunction in neurological disorders with epileptic phenotypes
Journal of Bioenergetics and Biomembranes (2010)