Abstract
Three stable microbial consortia, each composed of Bacillus paranthracis and Staphylococcus haemolyticus strains, were isolated from milk of cows diagnosed with mastitis in three geographically remote regions of Russia. The composition of these consortia remained stable following multiple passages on culture media. Apparently, this stability is due to the structure of the microbial biofilms formed by the communities. The virulence of the consortia depended on the B. paranthracis strains. It seems plausible that the ability of the consortia to cause mastitis in cattle was affected by mutations of the cytK gene of B. paranthracis.
Similar content being viewed by others
Introduction
Mastitis, inflammation of the mammary gland, is a common disease in cattle. It is related to the presence of pathogenic microorganisms, the most common of them being E. coli and various Streptococcus and Staphylococcus species1. Non-aureus staphylococci are pathogenic microorganisms frequently isolated in mammary gland infections2,3,4. Another group of microorganisms, which are less frequently isolated from milk but represent a considerable interest, are pathogenic bacilli5. The fact that pathogenic bacilli are capable of sporulation, which allows them to survive thermal treatment of milk and dairy products, further emphasizes their significance. The development of associations composed of different microbial species and their adaptation to their environments are currently increasingly attracting researchers' interest. In the present work, we have for the first time described consortia of Staphylococcus haemolyticus and Bacillus paranthracis isolated from the milk of cows diagnosed with mastitis in three geographically remote regions of Russia.
Results
General genomic characteristics
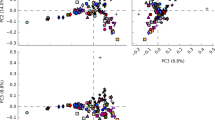
Each of the three studied bacterial consortia (4M, 1702, and 1710) was composed of two pathogenic strains. Genome quality estimation with CheckM showed that all genomes were of high quality (> 98% completeness and < 0.5% contamination). The Genome Taxonomy Database tool kit (GTDB-tk) was used to classify the bacterial genomes. GTDB-tk analysis classified one of strains in each consortium as a member of the Bacillus_A group, namely Bacillus paranthracis. The other strain in each consortium was Staphylococcus haemolyticus based on the taxonomic classification defined by topology and ANI. B. paranthracis strains exhibited a high level of similarity among different consortia, and so did the S. haemolyticus strains (Fig. 1). Each of the B. paranthracis strains possessed ten replicons, including one chromosome and nine plasmids ranging in size from 3124 bp to more than 300 kb. Five larger plasmids apparently had the theta-type replication, whereas the smaller plasmids were of the RCR type. The S. haemolyticus strains contained four replicons: one chromosome and three small plasmids of 6539, 3048, and 2362 bp (Table 1). Genetic features of plasmids found in B. paranthracis and S. haemolyticus strains are listed in Supplementary Data.
To find out in how much the strains composing the consortia differed genetically, pairwise comparison of the B. paranthracis genomes, as well as of the S. haemolyticus genomes, was performed using the ANI calculator. The results of this comparison are shown in Table 2. It can be seen that none of the strains studied was identical to some other strain. Most likely, these consortia originated from a common ancestor consortium and have been evolving independently. The characteristics of the three B. paranthracis genomes were absolutely identical, and so were the characteristics of the three S. haemolyticus genomes; they are listed in Table 3.
Putative virulence genes
The isolated B. paranthracis strains were found to possess several putative virulence genes. The genes nheA, nheB, and nheC encode a pore-forming toxin composed of a cytolytic protein NheA and two associated protein components NheB and NheC, which enhance the biological activity of the cytolytic protein. There were also three homologous reading frames encoding a secreted metalloprotease InhA, which can cleave multiple proteins of the host organism cells. The multiple alignment and the evolutionary tree of the three inaA orthologs are presented in Supplementary Materials, Fig. S1. All B. paranthracis strains possessed two copies of the gene encoding thiol-activated cytolysin ALO, which belongs to the family of cholesterol-dependent cytolysins. ALO is a pore-forming toxin that requires the presence of cholesterol in the membrane for pore formation; to date, the mechanism of this process is not fully understood. In addition, the B. paranthracis genomes included a gene cluster composed of the genes hblA1, hblA2, and hblB, which encodes a component of pore-forming hemolysin BL. However,the most interesting finding in B. paranthracis strains concerned the gene of the pore-forming toxin cytK-2. In all three strains, the sequence of this gene carried a point mutation, an T → A transition at position 357 that produced a TAA stop codon interrupting the reading frame. As a result, instead of the full-size CytK-2 (336 amino acids), these cells probably synthesize its truncated variant of 103 amino acids, which rather resembles leukotoxin LukDv (Fig. 2). The multiple alignment of cytK2 nucleotide sequence of B. paranthracis 4M, B. paranthracis 1710, B. paranthracis 1702, B. cereus E33L, and B. thuringiensis BGSC 4Y1 is presented in Supplementary Materials, Fig. S2. In addition, this truncated protein exhibits 99% homology to subunit C of hemolysin gamma from Streptococcus pneumoniae (GenBank Sequence IDs CKI41873, COH50677, and CJA68379) (Fig. 3).
The S. haemolyticus strains lacked genes of pore-forming toxins but possessed several genes responsible for prevention of phagocytosis. These were type 8 capsule genes: capA, capB, capC, capD, and capP. These genes are also involved in formation of biofilms.
Biofilm formation
In addition to the genes capA, capB, capC, capD, and capP mentioned above, the isolated S. haemolyticus strains were found to possess the icaC gene, which is usually a part of the ica operon of intercellular adhesion required for biofilm formation. However, the other genes of this operon, icaADB, were not detected in the S. haemolyticus strains studied. In addition, the S. haemolyticus strains possessed the genes of the adapter proteins MecA and MecB, which play a central role in the process of biofilm formation.
The B. paranthracis genomes included the eps1 cluster comprising 17 open reading frames (Supplementary Materials, Fig. S3); these strains lacked the eps2 cluster. The genomes of B. paranthracis also included the genes of the multifunctional regulators papR, plcR, and sinR, which are of significant importance both for the expression of virulence factors and for biofilm formation. In addition, the genomes of B. paranthracis had other genes involved in biofilm formation: sipW and tasA; furthermore, tasA was represented by three orthologs. The first of them was located remote from sipW, upstream from a gene encoding a serine peptidase of the family S8. Two other tasA orthologs were located immediately downstream from sipW and directly upstream from sinR and inaA_2.
Antibiotic resistance genes
The B. paranthracis strains possessed two chromosomal antibiotic resistance genes. One of them, vmlR, encodes an ATPase that binds to the ribosome and provides resistance to virginiamycin and lincomycin by the target protection mechanism. The other gene, fosB, encodes a magnesium-dependent thiol transferase that degrades fosfomycin.
The chromosome of the S. haemolyticus strains featured two resistance genes: fusC is responsible for inactivation of fusidic acid; mgrA ensures active efflux of a number of antibiotics, such as ciprofloxacin, methicillin, and tetracycline. Furthermore, the 6539-bp plasmid of S. haemolyticus carried two further resistance genes. One of them, aacA–aphD, encodes an aminoglycoside acetyltransferase and provides resistance to aminoglycoside antibiotics (e.g., neomycin, gentamicin, and kanamycin). The other gene was initially described as tetB in Geobacillus stearothermophilus; it is responsible for tetracycline efflux.
Secondary metabolites
The B. paranthracis strains contained two non-ribosomal peptide synthetases responsible for the synthesis of bacillibactin and fengycin, respectively. The S. haemolyticus strains were capable of synthesizing staphyloferrin.
Summary of genes for the virulence, biofilm formation, and antibiotic resistance in B. paranthracis and S. haemolyticus strains is presented in Table 4.
Discussion
Mastitis is a common and economically important disease of cattle. In this work, we used the megasequencing technology to analyze the bacterial communities of the infected milk samples and identify the presence of genes that can participate both in the pathogenic process and in overcoming the host's immune response. It was found that milk samples collected from cows with mastitis consistently contained consortia of two gram-positive bacteria, S. haemolyticus and B. paranthracis. The best-known pathogenic agent associated with cattle mastitis is Staphylococcus aureus, a gram-positive coccus that also causes other clinical infections. It is also recognized as a pathogen provoking food poisoning outbreaks. The pathogenicity of S. aureus is due to a broad range of virulence factors, including hemolysins, leukocidins, enterotoxins, and secretion systems of numerous toxins6,7.
While S. aureus has been described as a major pathogen causing cattle mastitis, coagulase-negative staphylococci (CNS) are also increasingly recognized worldwide as etiologic agents associated with intramammary infections (IMI). However, clinical and pathogenetic significance of their detection in milk sample cultures is still debated. Some researchers believe that CNS are true mastitis-causing agents with major virulence factors, a high level of antimicrobial resistance, and the ability to cause chronic infections. At the same time, others consider them secondary pathogens of cattle.
Bacterial pathogens have to survive, proliferate, and spread within the host; they must be able to adapt to an extremely hostile environment by responding to the existing immune barriers. The interaction between a host and a pathogen should be viewed not as a static phenomenon, but as an arms race in which each competitor tries to act as effectively as possible8. As a consequence, pathogens have developed extremely sophisticated strategies to overcome the host's immune response. In particular, depending on the lifestyle of various microbial pathogens, phagocytosis as such can represent not only an obstacle but also an opportunity for their propagation: some of them employ numerous intricate strategies to enter phagocyte cells and survive in them, whereas other bacteria have developed mechanisms to prevent or evade phagocytosis. For successful infection, pathogens must overcome or avoid the activity of the host's immune system cells. Infection with virulent bacilli is characterized by bacterial proliferation in spite of inflammation at the infection site9. This implies that these bacteria have developed means to counteract inflammatory cells and thus the host immune system.
We performed a comparative analysis of the pathogenicity factors of the other consortium member B. paranthracis, which can attack epithelial cells, as well as overcome and/or suppress the host's immune response. Taxonomically, B. paranthracis is very close to Bacillus cereus, a gram-positive spore-forming bacterium that provokes food poisoning and severe opportunistic infections10,11. B. cereus is present in soil and in food products; it contaminates human skin and nearly all surfaces in hospital environments. It is the second most common cause of mass food-related infection outbreaks after S. aureus12. Therefore, B. cereus sensu lato is of particular interest in terms of food safety and public health because of its ability to spoil food and cause disease due to synthesis of various toxins13. In this study, we determined the presence of pathogenicity factors in the identified consortia. All B. paranthracis strains present in the consortia possessed a cytK-2 gene14 with a mutation disrupting the reading frame. The protein encoded by this new reading frame has a high level of homology to LukDv, a pore-forming toxin protein described in S. pneumoniae. This protein can form pores in the membranes of eukaryotic cells, unbalancing the partial pressure inside the host cells, which results in cell lysis, release of the nutrients into the environment, and ultimately in the death of these cells15. The novel protein is encoded by a gene with a high homology to cytK, which suggests a possibility of horizontal transfer of this gene among various gram-positive microorganisms that are in a physical contact.
The second best-studied genetic determinant of a pore-forming toxin in B. cereus sensu lato is the hbl gene cluster present in all B. paranthracis strains described in this work. Hbl-dependent pore formation requires all Hbl components, which can individually bind to erythrocytes and form a membrane attack complex that ultimately provokes cell lysis16. The presence of the genes encoding PlcR and its inducer PapR in all strains described in our work also serves for positive control of the expression of Hbl-encoding genes. Furthermore, the expression of the gene that encodes the protein highly homologous to LukDv apparently remained under the characteristic positive transcriptional control of the PlcR system17 as described for the wild-type cytK-2 gene.
B. cereus cells can survive in the host organism and cause infection in spite of attracting phagocytes. The genome of B. cereus includes over 350 genes that encode exoproteins; among them, there are at least 50 genes of proteases with several putative pathogenetic functions18. At all stages of infection with pathogenic microorganisms, proteases are key virulence factors. They contribute to colonization of the host by degrading the extracellular matrix of host tissues, including collage and elastin. Together with the cytolytic activity of pore-forming toxins, this protease-mediated degradation promotes bacterial proliferation by providing nutrients, interfering with the host's immune response, damaging the protective endothelium, and disrupting the epithelial barriers19. Among exoproteases of B. cereus, there are two zinc-dependent proteases, InhA1 and NprA, which were quantitatively determined in a study of several exoproteomes20,21,22. InhA1 plays a central role in the virulence of B. cereus sensu lato due to its interactions with both bacterial and host proteins in the course of infection. InhA1 and NprA both feature zinc-binding motifs and include amino acid residues of the active catalytic center common for metalloproteases (HEXXH). InhA1 is lethal when injected into the hemolymph of insects and is capable of cleaving antibacterial peptides, such as cecropin and attacin23. InhA1 also determines the ability of B. cereus cells to evade host macrophages24. The B. paranthracis strains described herein possess genes of three homologous proteins: InhA1, InhA2, and InhA3. NprA enables bacteria to escape from macrophages and therefore represents a key element required to combat the host's immune system. Mutants with inactivated nprA or inhA1 genes cannot escape undamaged if captured by macrophages. NprA degrades host cell components, which can explain the role this proteins plays in the consortium during the escape from macrophages. InhA1-mediated cleavage of NprA is a key step in enhancing NprA activity. Indeed, an active C-terminal domain of NprA suffices to stimulate the release of bacteria from host macrophages after phagocytosis25.
Bacteria of the consortium are constantly exposed to a flow of milk and therefore require the ability to form biofilms. In the biofilm state, they adhere to the udder wall and acquire resistance to stress factors and most antibiotics, while continuing to secrete various pathogenicity factors26. Protection provided by a biofilm enables the bacteria to resist not only the host's defense mechanisms but also the standard antibiotic therapy. In staphylococci, biofilm formation involves the genes of the intercellular adhesion operon (ica), icaADBC, responsible for the synthesis of polysaccharide intercellular adhesin (PIA), the principal component of the exopolysaccharide matrix surrounding bacterial cells within biofilms. Most biofilms found in natural environments are composed of several bacterial species. B. cereus sensu lato is no exception. It is frequently observed in associations with other microorganisms, when biofilms can be described as cooperative consortia where each partner contributes to stability and development of the community27.
Similarly to cytK28, individual genes involved in biofilm formation in B. cereus are regulated by the global transcriptional regulator PlcR together with the oligopeptide encoded by papR, since quorum sensing is critically important not only for toxin expression levels but also for biofilm formation in gram-positive microorganisms. In addition, bacterial adhesion to surfaces, intercellular interactions, cell aggregation, and biofilm formation in B. cereus involve genes of the eps2 cluster, whereas the eps1 gene cluster apparently participates in some sort of social motility of these bacteria. That is, the eps2 cluster is more important for adhesion to epithelial cells29.
In addition, the expression of enterotoxin genes is affected by the phase transition regulator SinR. SinR and PlcR regulate biofilm formation in an interactive manner. SinR was shown to control biofilm formation and swimming motility in B. thuringiensis30. Interestingly, only a small subpopulation of cells in the biofilms expressed hbl, which depended on the expression of sinI30. Two SinR recognition sites were detected upstream from the hbl operon, and one more SinR recognition site was identified upstream from the nhe operon31. It should be mentioned that the spo0A–sinI–sinR regulatory chain in B. cereus AR156 was also shown to be involved in biofilm formation, in cell differentiation, and in host–pathogen interaction32.
Methods
Strains, culturing conditions, and genomic DNA extraction
Microbial consortium 4M was isolated from milk of black-motley cows in the Moscow region. Microbial consortium 1702 was isolated from milk of Simmental cows in the Saratov region. Microbial consortium 1710 was isolated from milk of black-motley Holstein hybrid cows in the Republic of Mordovia. On the map, the sites of sample collection form a triangle with sides of approximately 600, 400, and 300 km (Fig. 4). Based on the results of veterinary and laboratory control, all animals included in the study were diagnosed with the clinical form of mastitis. To obtain pure cultures, milk samples were first inoculated as enrichment cultures in Salt Meat Broth, Azide Dextrose Broth, and Tryptone Soya Broth (HiMedia Laboratories Pvt. Ltd., India) and subsequently transferred onto differential diagnostic media: Baird Parker Agar (HiMedia Laboratories) and Azide Blood Agar Pronadisa (Conda, Spain). For all cultures studied, preliminary microscopy of individual colonies from Baird Parker Agar showed cells of different size and shape (cocci and bacilli) and of the same positive Gram staining. To isolate individual strains and obtain uniform colonies, a series of five passages on the same differential diagnosis medium was performed. After each passage, microscopic analysis revealed the same two types of cells in the field of view.
Cells from a pure, fresh 50 mL liquid culture from each consortium were harvested by centrifugation at 15,000×g for 3 min and washed three times with sterile Milli Q water under sterile conditions. Total genomic DNA was extracted using the GeneJET Genomic DNA Purification Kit (Thermo Fisher Scientific, Bremen, Germany) following the manufacturer’s instructions. Quality and quantity of the extracted DNA was determined using QubitTM fluorometric quantitation and NanoDrop 2000 (both Thermo Fisher Scientific, Bremen, Germany).
Genome sequencing, assembly and annotation
The genome of both strains in each consortium was sequenced on an Illumina MiSeq platform with a read length of 300 bp (paired end) and Oxford Nanopore MinION. The genome from each member of consortia was assembled using Unicycler from both short and long sequencing reads33. Genome quality estimation was determined with CheckM. Genome circularization was checked manually using BLAST. Genome annotation was performed with Prokka34. Protein coding genes were classified based on the annotation into Cluster of Orthologous Groups (COG) functional categories with the automatic classification COG tool at MicroScope platform35. The data for this study have been deposited in the GenBank under project number PRJNA865942 with accession numbers CP102659-CP102662, and CP113365-CP113402.
Genomes comparison
Classification of the genomes was determined according to the Genome Taxonomy Database (GTDB) using the GTDB-tool kit (GTDB-tk) v.1.1.0 integrated in the MicroScope web-based service35. GTDB-tk provides a taxonomic classification of bacterial and archaeal genomes based on the combination of the GTDB reference tree, the relative evolutionary divergence and the ANI value against reference genomes36. CJ Bioscience's online Average Nucleotide Identity (ANI) calculator was used to compare two prokaryotic genome sequences37. Whole-genome alignments were performed by Mauve v. 2015022638.
Ethics statement
The animal study was reviewed and approved by L.K. Ernst Federal Science Center for Animal Husbandry Bioethics Committee, protocol number 2021-3/4. All methods were carried out in accordance with the approved guidelines and relevant regulations. All methods are reported in accordance with ARRIVE guidelines.
Data availability
The data for this study have been deposited in the GenBank under project number PRJNA865942 with accession numbers CP113365-CP113374 for B. paranthracis 4M; CP102659-CP102662 for S. haemolyticus 4M; CP113375-CP113384 for B. paranthracis 1702; CP113395-CP113398 for S. haemolyticus 1702;CP113385-CP113394 for B. paranthracis 1710, and CP113399-CP113402 for S. haemolyticus 1710.
References
Keane, O. M. Symposium review: Intramammary infections-Major pathogens and strain-associated complexity. J. Dairy Sci. 102, 4713–4726 (2019).
Naushad, S. et al. Comprehensive virulence gene profiling of bovine non-aureus Staphylococci based on whole-genome sequencing data. mSystems https://doi.org/10.1128/mSystems.00098-18 (2019).
De Buck, J. et al. Non-aureus Staphylococci and bovine udder health: Current understanding and knowledge gaps. Front. Vet. Sci. 8, 658031. https://doi.org/10.3389/fvets.2021.658031 (2021).
Condas, L. A. Z. et al. Prevalence of non-aureus staphylococci species causing intramammary infections in Canadian dairy herds. J. Dairy Sci. 100, 5592–5612 (2017).
Farhan, M. G., Abd El-Hamid, M. I. & Hassan, M. N. Propidium monoazide conventional PCR and DNA sequencing: Detection of negative culture bacterial pathogens causing subclinical mastitis. J. Appl. Microbiol. 128, 1595–1605 (2020).
Algammal, A. M., Enany, M. E., El-Tarabili, R. M., Ghobashy, O. I. & Helmy, Y. A. Prevalence, antimicrobial resistance profiles, virulence and enterotoxin-determinant genes of MRSA isolated from subclinical bovine mastitis samples in Egypt. Pathogens https://doi.org/10.3390/pathogens9050362 (2020).
Hashemizadeh, Z., Hadi, N., Mohebi, S., Kalantar-Neyestanaki, D. & Bazargani, A. Characterization of SCCmec, spa types and multi drug resistant of methicillin-resistant Staphylococcus aureus isolates among inpatients and outpatients in a referral hospital in Shiraz, Iran. BMC Res. Notes https://doi.org/10.1186/s13104-019-4627-z (2019).
Sarantis, H. & Grinstein, S. Subversion of phagocytosis for pathogen survival. Cell Host Microbe. 12, 419–431 (2012).
Hernandez, E., Ramisse, F., Ducoureau, J. P., Cruel, T. & Cavallo, J. D. Bacillus thuringiensis subsp. konkukian (serotype H34) superinfection: Case report and experimental evidence of pathogenicity in immunosuppressed mice. J. Clin. Microbiol. 36, 2138–2139 (1998).
Kavanaugh, D. W., Porrini, C., Dervyn, R. & Ramarao, N. The pathogenic biomarker alcohol dehydrogenase protein is involved in Bacillus cereus virulence and survival against host innate defence. PLoS ONE 17, e0259386. https://doi.org/10.1371/journal.pone.0259386 (2022).
Ramarao, N., Lereclus, D. & Sorokin, A. The Bacillus cereus group. In Molecular Medical Microbiology III, 2nd ed. 1041–1078 (2015).
Stenfors Arnesen, L. P., Fagerlund, A. & Granum, P. E. From soil to gut: Bacillus cereus and its food poisoning toxins. FEMS Microbiol. Rev. 32, 579–606 (2008).
Dietrich, R., Jessberger, N., Ehling-Schulz, M., Märtlbauer, E. & Granum, P. E. The food poisoning toxins of Bacillus cereus. Toxins https://doi.org/10.3390/toxins13020098 (2021).
Porcellato, D. et al. Characterization of Bacillus cereus sensu lato isolates from milk for consumption; phylogenetic identity, potential for spoilage and disease. Food Microbiol. https://doi.org/10.1016/j.fm.2020.103604 (2021).
Koné, K. M., Hinnekens, P., Jovanovic, J., Rajkovic, A. & Mahillon, J. New insights into the potential cytotoxic role of Bacillus cytotoxicus cytotoxin K-1. Toxins https://doi.org/10.3390/toxins13100698 (2021).
Ryan, P. A., Macmillan, J. D. & Zilinskas, B. A. Molecular cloning and characterization of the genes encoding the L1 and L2 components of hemolysin BL from Bacillus cereus. J. Bacteriol. 179, 2551–2556 (1997).
Verdugo-Fuentes, A., Gastélum, G., Rocha, J. & de la Torre, M. Multiple and overlapping functions of quorum sensing proteins for cell specialization in Bacillus species. J. Bacteriol. https://doi.org/10.1128/JB.00721-19 (2020).
Ivanova, N. et al. Genome sequence of Bacillus cereus and comparative analysis with Bacillus anthracis. Nature. 423, 87–91 (2003).
Miyoshi, S. & Shinoda, S. Microbial metalloproteases and pathogenesis. Microbes Infect. 2, 91–98 (2000).
Clair, G., Roussi, S., Armengaud, J. & Duport, C. Expanding the known repertoire of virulence factors produced by Bacillus cereus through early secretome profiling in three redox conditions. Mol. Cell Proteomics. 9, 1486–1498 (2010).
Cadot, C. et al. InhA1, NprA, and HlyII as candidates for markers to differentiate pathogenic from nonpathogenic Bacillus cereus strains. J. Clin. Microbiol. 48, 1358–1365 (2010).
Madeira, J. P., Alpha-Bazin, B., Armengaud, J. & Duport, C. Time dynamics of the Bacillus cereus exoproteome are shaped by cellular oxidation. Front Microbiol. https://doi.org/10.3389/fmicb.2015.00342 (2015).
Dalhammar, G. & Steiner, H. Characterization of inhibitor A, a protease from Bacillus thuringiensis which degrades attacins and cecropins, two classes of antibacterial proteins in insects. Eur. J. Biochem. 139, 247–252 (1984).
Ramarao, N. & Lereclus, D. The InhA1 metalloprotease allows spores of the B. cereus group to escape macrophages. Cell Microbiol. 7, 1357–1364 (2005).
Haydar, A. et al. InhA1-mediated cleavage of the metalloprotease NprA allows Bacillus cereus to escape from macrophages. Front. Microbiol. https://doi.org/10.3389/fmicb.2018.01063 (2018).
Di Domenico, E. G. et al. The impact of bacterial biofilms on end-organ disease and mortality in patients with hematologic malignancies developing a bloodstream infection. Microbiol. Spectr. 9, e0055021. https://doi.org/10.1128/Spectrum.00550-21 (2021).
Davey, M. E. & O’toole, G. A. Microbial biofilms: From ecology to molecular genetics. Microbiol. Mol. Biol. Rev. 64, 847–867 (2000).
Gohar, M. et al. The PlcR virulence regulon of Bacillus cereus. PLoS ONE https://doi.org/10.1371/journal.pone.0002793 (2008).
Caro-Astorga, J. et al. Two genomic regions encoding exopolysaccharide production systems have complementary functions in B. cereus multicellularity and host interaction. Sci. Rep. https://doi.org/10.1038/s41598-020-57970-3 (2020).
Fagerlund, A. et al. SinR controls enterotoxin expression in Bacillus thuringiensis biofilms. PLoS ONE. https://doi.org/10.1371/journal.pone.0087532 (2014).
Böhm, M. E., Krey, V. M., Jeßberger, N., Frenzel, E. & Scherer, S. Comparative bioinformatics and experimental analysis of the intergenic regulatory regions of Bacillus cereus hbl and nhe enterotoxin operons and the impact of CodY on virulence heterogeneity. Front. Microbiol. https://doi.org/10.3389/fmicb.2016.00768 (2016).
Xu, S. et al. The spo0A-sinI-sinR regulatory circuit plays an essential role in biofilm formation, nematicidal activities, and plant protection in Bacillus cereus AR156. Mol. Plant Microbe. Interact. 30, 603–619 (2017).
Wick, R. R., Judd, L. M., Gorrie, C. L. & Holt, K. E. Unicycler: Resolving bacterial genome assemblies from short and long sequencing reads. PLoS Comput. Biol. https://doi.org/10.1101/096412 (2017).
Seemann, T. Prokka: Rapid prokaryotic genome annotation. Bioinformatics https://doi.org/10.1093/bioinformatics/btu153 (2014).
Vallenet, D. et al. MicroScope in 2017: An expanding and evolving integrated resource for community expertise of microbial genomes. Nucleic Acids Res. 45, D517-528. https://doi.org/10.1093/nar/gkw1101 (2017).
Chaumeil, P.-A., Mussig, A. J., Hugenholtz, P. & Parks, D. H. GTDB-Tk: A toolkit to classify genomes with the Genome Taxonomy Database. Bioinformatics 36, 1925–1927 (2020).
Yoon, S. H., Ha, S. M., Lim, J. M., Kwon, S. J. & Chun, J. A large-scale evaluation of algorithms to calculate average nucleotide identity. Antonie van Leeuwenhoek. 110, 1281–1286 (2017).
Darling, A. C., Mau, B., Blattner, F. R. & Perna, N. T. Mauve: Multiple alignment of conserved genomic sequence with rearrangements. Genome Res. https://doi.org/10.1101/gr.2289704 (2004).
Acknowledgements
This research was funded by the Russian Science Foundation, grant number 20-16-00106. This research was also supported by the Ministry of Science and Higher Education of the Russian Federation (registration numbers 121052600314-1, and FGGN-2021-0002).
Author information
Authors and Affiliations
Contributions
S.S.—genomes comparison, original draft preparation, editing. F.B.—writing, editing. Al.S.—review preparation, editing. D.N.—microbial consortium isolation, cultures studying. K.F.—genome assembly and annotation. O.A.—laboratory control of animals for diagnosing. E.K.—microbial consortium isolation, pure cultures obtaining. An.S., T.D.—genomes comparison. M.S.—molecular biology methods, cultures studying. A.E.—genome sequencing. K.B.—molecular biology methods, visualization. N.Z.—review, supervision. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sokolov, S., Brovko, F., Solonin, A. et al. Genomic analysis and assessment of pathogenic (toxicogenic) potential of Staphylococcus haemolyticus and Bacillus paranthracis consortia isolated from bovine mastitis in Russia. Sci Rep 13, 18646 (2023). https://doi.org/10.1038/s41598-023-45643-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-45643-w
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.