Abstract
Inhibitors of enzymes that inactivate amine neurotransmitters (dopamine, serotonin), such as catechol-O-methyltransferase (COMT) and monoamine oxidase (MAO), are thought to increase neurotransmitter levels and are widely used to treat Parkinson's disease and psychiatric disorders, yet the role of these enzymes in regulating behavior remains unclear. Here, we investigated the genetic loss of a similar enzyme in the model organism Drosophila melanogaster. Because the enzyme Ebony modifies and inactivates amine neurotransmitters, its loss is assumed to increase neurotransmitter levels, increasing behaviors such as aggression and courtship and decreasing sleep. Indeed, ebony mutants have been described since 1960 as "aggressive mutants," though this behavior has not been quantified. Using automated machine learning-based analyses, we quantitatively confirmed that ebony mutants exhibited increased aggressive behaviors such as boxing but also decreased courtship behaviors and increased sleep. Through tissue-specific knockdown, we found that ebony’s role in these behaviors was specific to glia. Unexpectedly, direct measurement of amine neurotransmitters in ebony brains revealed that their levels were not increased but reduced. Thus, increased aggression is the anomalous behavior for this neurotransmitter profile. We further found that ebony mutants exhibited increased aggression only when fighting each other, not when fighting wild-type controls. Moreover, fights between ebony mutants were less likely to end with a clear winner than fights between controls or fights between ebony mutants and controls. In ebony vs. control fights, ebony mutants were more likely to win. Together, these results suggest that ebony mutants exhibit prolonged aggressive behavior only in a specific context, with an equally dominant opponent.
Similar content being viewed by others
The ebony gene encodes an enzyme, a β-alanyl biogenic amine synthase, that catalyzes the addition of β-alanine to biogenic amines (histamine, dopamine, serotonin, and octopamine), which inhibits these neurotransmitters and prepares them for recycling1,2. In a 1960 study of mating behavior, M.E. Jacobs observed that ebony mutants fight more than “light flies”3. Though this manuscript is typically cited as the first genetic study of aggressive behavior in Drosophila4, the aggressive behavior of ebony mutants has not been characterized or quantified. M.E. Jacobs went on to note that ebony mutants also exhibit less courtship behavior3. These two results appear contradictory because courtship and aggression are both positively regulated by biogenic amine neurotransmitters; changes in courtship and aggression are typically correlated, with increases in one behavior normally associated with increases in the other5,6,7,8,9. Since Jacob’s work, no one has characterized ebony aggression; Suh and Jackson10 showed that ebony activity impacts circadian rhythms, possibly via its role in neurotransmitter regulation, but did not examine courtship or aggression10. Here, we quantitatively examined ebony mutants to definitively test the role of this enzyme in regulating three different behaviors affected by biogenic amines: aggression, courtship, and sleep.
Widely used as a phenotypic marker because ebony mutants have very dark pigmentation, Drosophila ebony is expressed in the epidermal epithelium, the oenocytes (abdominal secretory cells), and in several types of glial cells. Importantly, ebony is not expressed in neurons1,11,12,13. Glial cells have long been known to perform essential tasks in the brain, including the provision of metabolic and structural support to neurons14,15,16. Recent research has led to increased appreciation for a more complex role of glial cells in neuronal functions, including the regulation of behavior—in part due to the role of glia in the synthesis, uptake, and regulation of neurotransmitters2,17,18,19. The extent to which glial cells modulate behavior through their regulation of neurotransmission remains understudied; there have only been limited studies directly examining the role of glia-mediated regulation of neurotransmission in complex behaviors (e.g., courtship and aggression)13,18,20. Given the genetic tractability and established behavioral assays in Drosophila, this model is well-equipped to advance our understanding of how glial regulation of neurotransmission affects diverse behaviors.
While there is no mammalian homolog of ebony, several genes might act as functional analogs: mammalian glutamine synthetase, which inactivates the biogenic amine glutamate by converting to glutamine in astrocytes, thereby reducing glutamate levels in the synapse20; catechol-O-methyltransferase (COMT), which inactivates catecholamine neurotransmitters (dopamine, epinephrine, and norepinephrine) by introducing a methyl group21; and monoamine oxidases (MAOs), which inactivate monoamines such as serotonin, dopamine, and tyramine by catalyzing their oxidative deamination22. As stated above, because biogenic amines positively regulate behaviors such as sex drive and aggression23,24, inhibition of enzymes that inactivate these neurotransmitters are predicted to increase these behaviors–for example, increasing aggression, increasing sex drive, and decreasing sleep.
In this work, we used ebony11 mutants to study how the loss of this biogenic amine-inactivating enzyme impacts these behaviors. We used machine learning-based algorithms to quantitatively analyze specific courtship and aggression behaviors (FlyTracker25 and JAABA26; 95.6–100% accuracy with ground truthing). Consistent with previous observations3,27, we found that ebony mutants exhibited decreased courtship behaviors and increased aggression; we also observed increased sleep. Through tissue-specific knockdown of ebony with RNAi, we showed that these behavioral phenotypes can be attributed to its role specifically in glial cells. To understand the basis for these behaviors, we quantified levels of biogenic amines in ebony11 mutant brains and found that they contained significantly decreased levels of histamine, dopamine, and serotonin relative to control brains, with no difference in octopamine levels. This result is consistent with decreased courtship and increased sleep but not with increased aggression. When we examined fights between ebony mutants and controls, we found that ebony mutants were not inherently aggressive; their prolonged fighting appeared to be due to an inability to establish dominance with an equally dominant opponent. Taken together, these findings provide the first quantitative characterization of the role of this glial enzymatic modifier of biogenic amines in regulating behavior, particularly aggression.
Results
ebony mutants lack Ebony protein
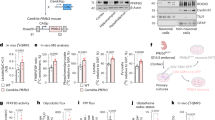
Previous research supports a model in which Ebony protein modifies and inactivates neurotransmitters taken up by glia from the synaptic cleft2. These modified, inactivated neurotransmitters are then transported back to pre-synaptic neurons, where they are reactivated by Tan, a hydrolase which removes the β-alanine group from the neurotransmitter for its re-use in synaptic transmission (Fig. 1A). To explore the role of ebony (e) in behavior, we selected the e11 allele, which we validated for loss of Ebony protein (see our brief discussion of the more commonly used but unvalidated e1 allele in Discussion and Methods). e11 mutants have the dark cuticle color characteristic of all ebony mutants; ebony is expressed in both glia and epidermal cells and loss of Ebony causes accumulation of biogenic amines in epidermal cells, which darkens the cuticle (Figs. 1A, S1A). PCR analysis confirmed that this e11 allele contained the predicted genomic deletion reported by Rossi et al.28. Primers around this expected deletion site amplified a DNA fragment that was ~ 500 bp smaller in e11 mutants than controls (Fig. 1B). While e11 mutants exhibited increased, likely compensatory, ebony mRNA expression relative to controls by qRT-PCR (Fig. S1B), we confirmed by western blot analysis that e11 mutants lacked Ebony protein (Fig. 1C; whole blot in Fig. S1C). Thus, our data confirm the mutation and loss of Ebony protein in e11 mutants (hereafter referred to as ebony mutants).
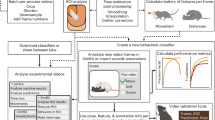
Validation of ebony11 mutant and quantitative analysis pipeline for Drosophila courtship and aggression behaviors. (A) Schematic showing the role of Ebony in recycling biogenic amines via inactivation of neurotransmitters in glial and epidermal cells. (B) PCR of ebony mutants and controls, revealing that e11 mutants have the expected ~500bp deletion. (C) western blot showing that ebony11 mutants lack Ebony protein. (D) schematic of aggression and courtship assays, which were video-recorded and analyzed by FlyTracker and JAABA machine learning classifiers and (E) schematic depicting fly behavioral parameters that were annotated by FlyTracker and used by JAABA machine learning classifiers to quantify specific courtship and aggression behaviors.
Automated quantification of Drosophila courtship and aggression with tracking and machine learning software
Because biogenic amine neurotransmitters are known to influence courtship and aggression7,29,30,31,32,33,34, we tested if our confirmed ebony mutant males exhibited changes in their courtship or aggressive behaviors. To measure courtship and aggression, we used an automated machine-learning based method combining the Caltech FlyTracker25 and the Janelia Automatic Animal Behavior Annotator (JAABA), established by the Branson lab at HHMI Janelia Farm26 (Fig. 1D). We filmed 10-min videos of flies engaged in either courtship or aggression assays and used the Caltech FlyTracker to obtain information about each fly’s position, orientation relative to another fly, poses, and trajectories (Fig. 1E). A machine learning classifier is an algorithm to automatically categorize data within that “class”; for this work, the “class” is a behavior such as chasing or setting wings at a specific angle. We used a subset of annotated videos (training data) to train the JAABA machine learning classifiers to detect the specific, annotated behaviors, validated the trained JAABA classifiers with a second subset of manually-scored control videos, and then used the JAABA classifiers to quantify courtship and aggression behaviors in experiments.
To quantify courtship behaviors, we trained JAABA to detect wing extension, chasing, and mounting without copulation, three parameters of courtship (Fig. 1E). We used ground truthing strategy to validate the accuracy of our classifiers for courtship analysis (Fig. S1D). Ground truthing uses all the behavior video frames that have been annotated by the user to make a classifier (training data), and the classifier then quantifies those behaviors for videos it has never seen before (testing data). The user manually annotates the novel “testing” videos, and the computer outputs the accuracy of the classifier’s predictions in comparison. We made a separate classifier to quantify chasing behavior in courtship, as male chasing of females in courtship may have subtle differences in speed and orientation from male chasing of other males in aggression. For courtship, the classifiers agreed with the user for 99.2–100% of the testing data frames. To quantify aggressive male behaviors, we trained classifiers to detect wing threat, chasing, lunging, and boxing (Fig. 1E) and performed the same ground truthing analysis (Fig. S1D). For aggression, these classifiers agreed with the user for 95.6–100% of the testing data frames. Thus, our classifiers for both aggression and courtship were highly accurate and precise, facilitating video analysis at higher throughput than manual analysis.
ebony mutants display decreased courtship behavior towards females
To assay changes in courtship behavior for ebony mutants, we video-recorded the interactions between pairs of 6–8 day-old virgin males and 3–5 day-old wCS (w- Canton S strain) females in a 16 mm bottom-lit chamber for 10 min (Fig. 2A). For each male, we determined the courtship index (CI), a standard metric in the field which reflects the percentage of video frames in which the male exhibited any of several types of courtship behavior. Unless otherwise noted, females were mated to prevent copulation and to ensure a window of courtship behavior extending for the duration of video recording. Using our automated analysis programs, we found that ebony mutant males spent significantly less time exhibiting courtship behavior relative to controls (Fig. 2B). In assessing individual courtship behaviors, we found that ebony mutants spent significantly less time on wing extension (Fig. 2C), chasing (Fig. 2D), and attempted mounting (Fig. 2E) behavior than controls. We also measured the latency to court, or the time elapsed from the start of the recording to the first courtship behavior; ebony males exhibited an average courtship latency of 109 s while CS males had an average courtship latency of 17 s (Fig. 2F). Thus, ebony males exhibited reduced courtship index and were slower to initiate their first courtship attempt compared to CS control males.
ebony mutant males court females, but not males, less than controls. (A) Schematic of our courtship assay with mated wCS females. (B–F) When courting mated wCS females, ebony mutants (green), compared to controls (gray), exhibited: (B) decreased total courtship (p < 0.0001); (C) decreased wingsong behavior (p < 0.0001); (D) decreased chasing behavior (p < 0.0001); (E) decreased attempted mounting (p = 0.0082); and (F) increased latency to first courting event (p < 0.0001). (G) In courtship assays with decapitated wCS virgin females or males, control males courted females (dark gray) more than males (light gray) (p = 0.0211) and more than ebony males courted females (dark green) (p < 0.0001). There was no difference in courtship levels between control males (light gray) and ebony males (light green) with decapitated males (p = 0.2585) or between ebony males with females (dark green; p>0.9999). Each data point represents a single male (n=number of flies); p-values were obtained by Mann-Whitney U test (B–F) and Kruskal-Wallis test with Dunn’s post hoc test (G). Averages are shown with error bars representing SEM. (See Table S1 for n and statistical analysis for each experiment; n.s.= p > 0.05, * = p < 0.05, **= p < 0.01, **** = p < 0.0001).
To confirm that this phenotype is not specific to mated females, we quantified the CI of ebony males with virgin females (Fig. S2A). Virgin females have different cuticular hydrocarbon profiles than mated females, which are known to influence male courtship behavior35,36. Consistent with our results with mated females, ebony mutant males displayed a reduction in total courtship with virgin females (Fig. S2B). ebony males had an average courtship latency of 138 s, while CS males waited, on average, 47 s before initiating courtship (Fig. S2C). We also measured the percentage of pairs with successful copulation within the 10-min observation period. When paired with a virgin female, only 7% of the ebony mutant males achieved copulation within 10 min compared to 67% of the control males (Fig. S2D). Thus, ebony males display defects in courtship toward both mated and virgin females and copulate with virgin females less successfully than control males.
To test if this decrease in courtship behavior was specific to male–female interactions, we tested whether male-male courtship was also affected in ebony mutants. Male-male courtship can be induced by biogenic amine neurotransmitters and substrates of Ebony, such as dopamine30. For this assay, we decapitated both virgin male and female targets to prevent aggression from male targets toward the male being tested. Consistent with our results above, ebony males courted decapitated virgin females significantly less than CS males; in contrast, ebony males courted decapitated virgin males at the same rate as controls (Fig. 2G). Interestingly, while controls showed a courtship preference for females, there was no difference in courtship indices for ebony males with decapitated virgin females or males. Thus, ebony mutants exhibit decreases in male–female courtship relative to controls, with no difference in male-male courtship and no preference for courting either sex.
ebony mutants display increased boxing and wing threat behaviors
While it is widely accepted that ebony mutants are "hyper-aggressive", this characterization appears to be mainly based on two qualitative observations3,27. To quantitatively test this, we video-recorded the interactions between pairs of 6 to 8-day-old virgin males in a 16 mm bottom-lit chamber for 10 min and used the automated analysis described above to measure aggressive behavior. Each chamber contained a small area of food as a territory/resource to induce aggressive behavior (Fig. 3A). To quantify aggressive behaviors, we measured behavioral indices similar to courtship behavioral indices, reflecting the percentage of frames or time spent engaging in this behavior (Fig. 3B-D). We observed that ebony mutants spent dramatically more time engaged in boxing than controls (Fig. 3B); boxing is considered one of the most aggressive behaviors of flies37. We also determined the total number of videos in which we observed boxing; ebony mutants exhibited boxing in 90.3% of their videos, compared to 63.3% for CS controls (Fig. S3A). ebony mutants also displayed increased wing threat behavior relative to controls (Fig. 3C). Unexpectedly, relative to controls, ebony mutants exhibited decreased chasing behavior (Fig. S3B) (not considered a highly aggressive behavior as chasing is also seen in courtship) and decreased lunging (Fig. 3D), though boxing behavior can be considered reciprocal lunging. We also found that ebony mutants had an increased latency for aggressive behaviors relative to controls, similar to the latency in courtship behavior. That is, ebony males were slower to start fighting with each other (latency average of 212 s), than did controls, who fought with each other after an average of 90 s (Fig. 3E). Thus, ebony males exhibited an increase in specific aggressive behaviors such as boxing and wing threat.
ebony mutants spend more time exhibiting “hyper-aggressive” behaviors and show a greater latency to initiate fighting. (A) Schematic of aggression assay. (B–E) In fights between ebony mutants (green), compared to fights between controls (gray), ebony mutants exhibited: (B) increased boxing behavior (p<0.0001); (C) increased wing threat behavior (p<0.0001); (D) decreased lunging behavior (p=0.0497); and (E) greater latency to first fighting event (p=0.0002); each data point represents a video of two flies (n=number of videos). (F–H) In aggression assays using a more ecologically relevant arena (F) with ebony mutants fighting each other (green), compared to controls fighting each other (gray), ebony mutants had: (G) a greater percentage of videos with boxing behavior (p<0.0001, n=number of videos); and (H) increased numbers of lunges per video (p=0.0025; n=number of flies). p-values were obtained by Mann-Whitney U test (B–E, H) and Fisher’s exact test (G); averages are shown with error bars representing SEM. (See Table S1 for n and statistical analysis for each experiment; n.s.= p>0.05, *=p<0.05, ** = p<0.01, *** = p<0.001, **** = p<0.0001.)
Because the small, flat arena required for our automated analysis could itself increase aggressive behavior38, we confirmed these results in an assay with a more ecologically relevant arena with lower throughput analysis. We used a larger chamber with a raised food cup, as described by Fernandez et al.39 (Fig. 3F). Consistent with our results above, in the more ecologically relevant arena, significantly more videos of ebony mutants exhibited boxing relative to videos of controls (Fig. 3G) and ebony mutants exhibited more lunging behavior than controls (Fig. 3H). Lunging is a more commonly observed aggressive behavior than boxing, also exhibited specifically by males37. Latency to lunge, while trending to a shorter time for ebony mutants, was not significantly different from controls (Fig. S3C). Thus, our results suggest that, regardless of arena, ebony males exhibit increased aggression.
ebony mutants exhibit increased sleep duration and sleep consolidation
The four biogenic amines modified by ebony (dopamine, serotonin, histamine, and octopamine) not only promote courtship and aggression, but also suppress sleep40,41,42,43,44,45. To test whether loss of ebony also influenced sleep, we used Drosophila Activity Monitors (DAMs) to measure and compare the sleep of ebony mutants and controls (Fig. 4A). Sleep duration was calculated by adding each subsequent minute after 5 min of inactivity, an analysis based on previous literature46, while a sleep bout is defined as a single, uninterrupted period of sleep (Fig. 4B). Sample activity and sleep plots are shown in Fig. 4C. We found that, while ebony mutants and controls exhibit similar average activity per hour (Fig. 4D), ebony mutants sleep more than controls (Fig. 4E). That is, ebony mutants exhibit increased activity during their wake time relative to controls (Fig. S4). To analyze sleep architecture, we quantified average sleep bout number and length during the day and night. ebony mutants exhibited fewer bouts of sleep during both daytime (Fig. 4F) and nighttime (Fig. 4G) and experienced an increased average sleep bout length (Fig. 4H) during both daytime (Fig. 4I) and nighttime (Fig. 4J). These results show that ebony mutants have fewer interrupted periods of sleep and more consolidated sleep architecture.
ebony mutants have fewer but longer sleep bouts and similar activity levels compared to controls. (A–B) Schematics of: (A) the setup of a Drosophila Activity Monitor (DAM); and (B) typical bouts of activity (wake) and inactivity (sleep) over a 1-hour period. (C) Plots show the average activity (top) and sleep duration per 30 minutes (bottom) over a 24-hour period for control flies (gray) and ebony mutants (green). (D–J) Relative to controls (gray), ebony mutants (green) exhibited: (D) similar average activity per hour (p = 0.4372); (E) increased sleep amount over a 24-hour period (p < 0.0001); (F) decreased average daytime bout number (p = 0.0038); (G) decreased average nighttime bout number (p = 0.0001); (H) increased average bout length (p=0.0002); (I) increased average daytime bout length (p = 0.0006); and (J) increased average nighttime bout length (p=0.0005). Each data point represents a single fly (n = number of flies); p-values were obtained by unpaired Student’s t-test with Welch’s correction (D–G) and Mann-Whitney U test (H–J). Averages are shown with error bars representing SEM. (See Table S1 for n and statistical analysis for each experiment; n.s.= p > 0.05, ** = p < 0.01, *** = p < 0.001, **** = p < 0.0001.)
Ebony in glia regulates aggression, courtship, and sleep
Because ebony is expressed in both glial cells and in the epidermal epithelium that produces the cuticle, we set out to test if all these observed behavioral changes in ebony mutants (increased aggression, decreased courtship, and increased sleep) are specifically due to Ebony’s function in glia. We first performed whole-body RNAi knock-down of ebony expression by driving UAS-ebony RNAi with a ubiquitous driver, tubulin-Gal4. Consistent with ebony mutants, whole-body RNAi-mediated knockdown of ebony caused the darkened cuticle characteristic of ebony mutants (Fig. 5A). To measure ebony expression in these flies, we performed qPCR analysis using whole flies and found significantly reduced mRNA levels in the whole-body ebony knockdown relative to both controls (Fig. 5B). Thus, UAS-ebony RNAi is an efficient tool to knock down ebony expression and test the role of ebony in glial cells to influence behavior.
Whole body and glial-specific ebony RNAi knockdowns recapitulate ebony mutant behavioral phenotypes. (A) Images of tubulin-Gal4 control (left), UAS-ebony RNAi control (center) and tub-Gal4>UAS-ebony RNAi flies (right) demonstrate the altered pigmentation phenotype resulting from ubiquitous ebony knockdown. (B–H) All Gal4 controls are shown in dark gray, UAS controls in light gray, with tubulin > ebony RNAi in bright green (whole body), and repo > ebony RNAi in dark green (glia-specific). (B) tubulin > ebony RNAi flies exhibited less ebony mRNA expression than controls by qRT-PCR (p < 0.0001, n = number of biological replicates, 15 flies each). (C–E) Ubiquitous ebony RNAi knockdown led to: (C) increased boxing and wing threat aggressive behaviors, compared to tubulin-Gal4 (p = 0.0225) and UAS-ebony RNAi (p < 0.0001) controls; (D) decreased total courtship, compared to tubulin-Gal4 (p < 0.0001) and UAS-ebony RNAi (p < 0.0001) controls; and (E) increased sleep amount over a 24-hour period, compared to tub-Gal4 (p < 0.0001) and UAS-ebony RNAi (p < 0.0001) controls (n = number of videos). (F–H) Glial-specific ebony RNAi knockdown led to: (F) increased boxing and wing threat aggressive behaviors, compared to repo-Gal4 (p < 0.0001) and UAS-ebony RNAi (p = 0.0096) controls; (G) decreased total courtship, compared to repo-Gal4 (p < 0.0001) and UAS-ebony RNAi (p < 0.0001) controls; and (H) increased sleep amount over a 24-hour period, compared to repo-Gal4 (p < 0.0001) and UAS-ebony RNAi controls (p = 0.0099) (n = number of videos). p-values were obtained by ordinary one-way ANOVA (Dunnett’s multiple comparisons; B, E, H) and Kruskal-Wallis test with Dunn’s post hoc test (C–D, F–G). Averages are shown with error bars representing SEM. (See Table S1 for n and statistical analysis for each experiment; n.s.=p > 0.05, * = p < 0.05, ** = p < 0.01, **** = p < 0.0001.)
To confirm that whole-body RNAi-mediated knockdown of ebony expression recapitulates the behavioral phenotypes of ebony mutants, we quantified aggression, courtship, and sleep in flies expressing UAS-ebony RNAi ubiquitously (using tub-Gal4). Consistent with ebony mutants, whole-body RNAi knockdown of ebony (tub-Gal4 > UAS-ebony RNAi) caused significant increases in aggression (as measured by boxing and wing threat) (Fig. 5C), decreases in courtship (Fig. 5D), and increases in sleep (Fig. 5E) relative to controls. These results confirmed that ubiquitous knockdown of ebony expression phenocopied ebony mutants.
To test if Ebony acts specifically in glial cells to affect behavior, we drove UAS-ebony RNAi with the well characterized, glia-specific repo-Gal4 driver (repo-Gal4 > UAS-ebony RNAi)47 and measured aggression, courtship, and sleep. Again, consistent with ebony mutants, glia-specific RNAi-mediated knockdown of ebony led to increased time spent exhibiting aggressive behaviors of boxing and wing threat (Fig. 5F), decreased time spent performing courtship behaviors (Fig. 5G), and increased sleep (Fig. 5H). Thus, taken together, our results show that Ebony protein in glia plays a major role in regulating these complex behaviors.
ebony mutants have decreased neurotransmitter levels
Loss of Ebony is predicted to increase levels of biogenic amine neurotransmitters, which would be consistent with the observed increase in hyper-aggressive behavior in ebony mutants, but not with decreased courtship behavior and increased sleep. To directly test how loss of Ebony affects biogenic amine transmitter levels, we used LC-MS/MS to measure levels of four unmodified (active, Fig. 1A) biogenic amine neurotransmitters (dopamine, histamine, serotonin, and octopamine) in dissected ebony and control brains with intact lamina. The Drosophila optic lobe contains a series of optic ganglia, the most peripheral of which is the lamina. We retained intact lamina because ebony is strongly expressed in glial cells of the lamina11. We found that ebony mutants exhibited decreased levels for three of the four biogenic amines assayed (Fig. 6A–D). Histamine, dopamine, and serotonin were reduced in ebony mutants compared to controls (Fig. 6A–C), with no significant difference in octopamine levels (Fig. 6D). Loss of histamine has been shown to lead to visual defects48. We assayed the specific aggressive behaviors of white-eyed wCS males and white-eyed ebony males (w;;ebony), known to have significant visual defects from the white mutation, to test if visual defects alone cause increased aggression (Fig. S5A–D). White-eyed ebony males exhibited significantly more boxing (Fig. S5A) and wing threat (Fig. S5B) behaviors, a similar level of lunging (Fig. S5C), and less chasing (Fig. S5D) compared to wCS controls. Comparing these aggressive behavioral indices to those of our red-eyed Canton-S control (Figs. 3; S3), wCS males exhibited similar or lower average aggressive behavior indices, particularly a lower average boxing index (Fig. S5A), while white-eyed ebony behavioral indices were comparable to those of red-eyed ebony males (Figs. 3; Fig. S3). Thus, impaired vision does not itself increase aggression. Overall, our finding of reduced biogenic amine levels in ebony mutants is consistent with the observed courtship and sleep phenotypes, but do not explain the elevated boxing and wing threat behaviors observed (Fig. 3B,C).
ebony mutant brains have decreased neurotransmitter levels relative to controls. (A–D) Each graph shows average neurotransmitter levels per brain of ebony mutants (green) and control flies (gray), measured by HPLC/MS (n = number of biological replicates, each consisting of 50 dissected brains with intact lamina). Relative to controls, ebony mutants had: (A) decreased histamine (p = 0.0004); (B) decreased dopamine (p=0.0410); (C) decreased serotonin (p = 0.0457); and (D) similar octopamine levels (p = 0.1084). p-values were obtained by unpaired Student’s t-test with Welch’s correction. Averages are shown with error bars representing SEM. (See Table S1 for n and statistical analysis for each experiment; n.s.= p > 0.05, * = p < 0.05, *** = p < 0.001.)
The genotype of its opponent impacts ebony's aggressive behavioral pattern
To further explore ebony’s aggressive behaviors, we characterized ebony’s specific aggressive behavioral pattern and asked if their aggression patterns in intra-genotype fights (CS vs. CS, ebony vs. ebony) were similar for inter-genotype fights (ebony vs. CS) (Fig. 7A). Similar to the courtship index (CI) above, we determined a comprehensive aggression index (AI) as the percentage of frames per video that a male spent performing any of several aggressive behaviors (chasing, boxing, lunging, or wing threat) to examine overall patterns of behavior. When we viewed the profile of aggressive behavior in this way, we found that ebony mutants exhibited aggressive behaviors for the same amount of time as controls during both intra-genotype and inter-genotype fights, whether or not a hierarchy-based analysis was used (Figs. 7B; S6A; see Methods for description of hierarchy-based analysis). Though total aggression time was identical, ebony mutants had a distinct fighting profile in ebony vs. ebony fights, spending more time boxing and performing wing threat compared to controls (Fig. 7B, intra-genotype fights). This result was recapitulated for intra-genotype fights with RNAi-mediated knockdown of ebony in the whole body (tubulin > ebony RNAi) or glial cells (repo > ebony RNAi), which also exhibited elevated boxing and wing threat relative to respective controls (Fig. S6B). In contrast, in inter-genotype fights, we found a reduction in all four aggressive behaviors.
In inter-genotype fights, ebony do not display the aggression and dominance phenotypes seen in intra-genotype fights. (A) Schematic of intra-genotype fights (CS vs. CS or ebony vs. ebony) and inter-genotype fights (CS vs. ebony). (B) Stacked bars show the aggression index for specific aggressive behaviors in intra-genotype fights and inter-genotype fights (wing threat = light pink; chasing = pink; lunging = fuchsia; boxing = purple); note that the percentage of time per behavior per video is greater when quantifying two flies of the same genotype than each fly of different genotypes (n = number of videos). (C) While intra-genotype fights between ebony mutants (dark green) were less likely to result in a winner than intra-genotype fights between controls (gray) (p = 0.0057), inter-genotype fights between ebony mutants and controls (light green) were different from intra-genotype ebony fights (p = 0.0339) and as likely to result in a winner as fights between controls (p > 0.9999); n = number of videos. (D) In inter-genotype fights, ebony mutants won a greater number of fight bouts (p = 0.0175); n = number of videos. p-values were obtained by Kruskal-Wallis test with Dunn’s post hoc test (C) and Mann-Whitney U test (D). Averages are shown with error bars representing SEM. (See Table S1 for n and statistical analysis for each experiment; n.s.= p > 0.05, * = p < 0.05, ** = p < 0.01.)
More strikingly, we observed that, in ebony vs. control fights, the fighting profile of ebony mutants became indistinguishable from controls (Fig. 7B, inter-genotype fights). When fighting controls, ebony mutants had similar total lunge count (Fig. S6C), lunge latency (Fig. S6D), and boxing, wing threat, lunging, and chasing indices (Fig. S6E–H) as controls. Taken together, these results suggest that ebony mutants exhibit a distinct pattern of "hyper-aggression", comprised of increased boxing and wing threat, mainly in intra-genotype fights with each other.
To further understand ebony’s hyper-aggression in ebony vs. ebony fights, we examined their dominance behavior, or their ability to “win” a fight bout. To score each video for a fly’s ability to establish dominance, we counted the percentage of fight bouts that occurred on food and led to a clear winner (i.e., a losing fly was chased/pushed off the food). A fight did not result in dominance if the flies ended a fight while both males remained on the food or left the food simultaneously. If a fight occurred off the food, then the fight was excluded from analysis. Because boxing is a behavior in which flies have reciprocally lunged at one another, essentially exhibiting equal aggression, we hypothesized that ebony mutants may have difficulty establishing a clear dominance relationship with each other in intra-genotype fights. Using this analysis, we found that 40% of ebony fights resulted in dominance compared to 62% of control fights (Fig. 7C). That is, ebony vs. ebony fights were less likely to have a clear winner than control fights, suggesting that ebony mutants have difficulty establishing dominance when fighting each other (Fig. 7C). Similarly, in the more ecologically relevant arena, ebony mutants in intra-genotype fights had reduced probability of establishing dominance compared to control intra-genotype fights; 42% of ebony videos exhibited dominance compared to 63% of control videos, though this was not significant (Fig. S6I).
In contrast, in ebony vs. CS control fights, we found that fights between ebony mutants and controls led to clear winners (58%), at a rate similar (62%) to control fights (Fig. 7C). In fights with ebony vs. controls, the winner was more likely to be ebony mutants, who won 38% of the time compared to 20% of the time for controls (Fig. 7D). Taken together, our results show that ebony mutants do not spend more time in total aggressive behavior than control males nor do they exhibit more aggressive behaviors like boxing when fighting with control males; nonetheless, they are more likely than controls to establish dominance in these intergenotype fights. We hypothesize that this preference for dominance leads to an escalation of aggressive behavior in ebony vs. ebony fights, in which flies had a reduced probability of establishing dominance and seemed to engage in reciprocal, prolonged aggression without retreating.
Discussion
Here we set out to examine the behavior of Drosophila mutants lacking Ebony, a biogenic amine neurotransmitter recycling enzyme expressed in glia and epithelia. Previous literature from the 1960s and ′70s suggested that this mutant exhibited aberrant behavior but these changes in behavior had never been quantified systematically3,27. It was also unclear if ebony specifically in glia controls behavior as ebony is also expressed in other tissues, such as epithelial cells, which synthesize the cuticle, the site of cuticular hydrocarbons, which are important communication signals for Drosophila social behavior. We found, relative to controls, that ebony mutants exhibited increased hyper-aggressive behaviors, such as boxing, decreased courtship, and increased sleep; additionally, we determined that these changes in behavior are due to ebony expression specifically in glia. We quantified four common biogenic amines and substrates of Ebony in the brains of ebony mutants and controls. Contrary to expectation, we found that ebony mutants had lower levels of these neurotransmitters relative to controls. Finally, when we paired ebony and control males together for inter-genotype fights, we found that ebony mutants were no longer hyper-aggressive, though they were more likely to “win” a fight than controls. Thus, we hypothesize that ebony mutants paired together for intra-genotype fights use hyper-aggressive fighting tactics because they are less able to establish dominance.
Previous work characterizing ebony mutants’ behavior generated mixed results. The only published qualitative examination of aggression behavior for ebony mutants in two publications from Jacobs et al.3,27 observed that ebony males “fight more than do light males”3 and were more likely to be the territorial male when paired with control flies. Jacobs et al.27 also noted that ebony males emphasized tussling behavior. Here we show quantitative increases in specific types of aggressive behavior for ebony mutants relative to controls using automated analysis. Regarding ebony courtship, consistent with previous observations that ebony males have defective courtship behaviors3,27,49,50, we found that e11 mutants court significantly less than controls; one group reported increased wing extension duration in ebony mutants, which we did not observe51. Finally, another group previously found that ebony1 mutants exhibited decreased sleep52, in contrast to our finding of increased sleep in ebony11 mutants relative to controls (Fig. 2). We hypothesize that this and other possible differences in behavioral phenotype may be due to differences in ebony mutant alleles; as an example, one published study found that mutants containing different ebony alleles had different circadian locomotor phenotypes53. Because we could not validate two different stocks of the e1 mutants (BDSC#1658 and BDSC#8443) by PCR analysis and we were able to validate the e11 mutation both by PCR and western blot analysis, we used the e11 mutant or ebony RNAi for this work and caution others on using the e1 allele.
It should be noted that others have also previously measured dopamine and histamine levels in ebony mutants, with highly conflicting results. Different reports found higher, similar, or lower levels of different neurotransmitters; these contradictory results may be due to differences in ebony alleles, dissection technique, ages of flies, or quantification technique54,55,56,57,58. In some cases, ebony mutants also had extraneous and confounding mutations that affect neurotransmitter levels, such as white mutations56,59. To reduce these confounding variables, we used outcrossed (6 generations) and validated e11 mutants in an otherwise wild-type background, fully removed the cuticle from their brains, kept the lamina intact, age-matched to controls, used LC-MS/MS analysis (a highly sensitive and accurate technique), and measured all four major biogenic amines. Our definitive results settle this question and show that overall dopamine, histamine, and serotonin levels were significantly decreased in ebony mutants (Fig. 6A–C). Our results are further consistent with the prediction that loss of ebony should affect different biogenic amine neurotransmitters similarly.
Our finding of lower amine neurotransmitter levels in ebony mutants is consistent with two of the three behavioral phenotypes observed: decreased courtship and increased sleep. These two phenotypes are often associated with each other8,9,31,40,41,42,43,44,45,54,60,61,62,63 and also with lower amine neurotransmitter levels, especially dopamine29,31,43,44,45,52,63. These phenotypes are also consistent with low histamine levels, as histamine is required for proper vision in Drosophila64,65. ebony mutants, similar to white-eyed flies and other flies with visual defects66, exhibit less chasing and courtship behavior, likely because these require sighting of the target fly and recognition of other visual cues50.
The finding of decreased amine neurotransmitter levels in ebony mutants is more unexpected with regard to their aggressive behaviors; octopamine, dopamine, and serotonin all positively regulate Drosophila aggression4,31,32,33,34,67. It should be noted that our analysis quantified total neurotransmitter levels, not extracellular neurotransmitter levels. The specific molecular mechanism by which decreased neurotransmitter levels leads to increased aggression remains unclear. One possibility is an effect on pheromone secretion or reception. ebony mutants have been shown to exhibit differences in cuticular hydrocarbons68, pheromones that modulate fly interactions. Cuticular hydrocarbons are made in large secretory cells called oenocytes that are known to express ebony69. Because we showed here that changes in sleep, courtship, and aggression in ebony mutants are due to loss of ebony specifically in glia, our work raises the exciting possibility that ebony mutants may be the first known example of altered glia to oenocyte signaling and regulation. Determining whether and how loss of ebony in glia alters cuticular hydrocarbons to increase aggression will be an interesting question for future investigation.
One major finding from this work came from our analysis of ebony vs. control fights: we found that ebony males are more likely to establish dominance and win fights against control males, suggesting a basis for their hyper-aggressive behavior in intra-genotype fights. In Drosophila aggression, dominance can be observed in the fly that maintains its position on the food, or the “territory,” over which the males are fighting70. Previous work had suggested that ebony mutants are highly territorial, tending to defend and stay close to their territory (food) more than controls27. Our dominance analysis is consistent with this conclusion and further supports the hypothesis that ebony mutants tend toward dominance and engage in hyper-aggressive behavior when they are also unable to establish dominance. That is, ebony vs. ebony fights tend to escalate as neither fly is willing to concede.
It is worth noting that glial Ebony may play a developmental role in neuronal circuit formation or in building brain structures71 required for normal adult behavior. That is, because ebony mutants lack Ebony for their entire lifespan, including development, loss of Ebony in glia during development, as opposed to or in addition to loss of Ebony during adulthood, could alter neuronal structure, circuit formation, or neuronal activity/function and thereby drive adult behavioral phenotypes. Conditional knockdown of glial ebony only during development or only during adulthood and assaying for subsequent behaviors would clarify the time window during which loss of ebony has greatest impact on adult behavior. Consistent with our work, mouse mutants genetically lacking MAO have also been shown to exhibit increased aggressive behavior72,73,74,75,76. Moreover, specific polymorphisms in both MAO and COMT are associated with aggressive behavior77,78,79,80,81. Thus, our results are consistent with work in mammals and suggest that chronic inhibition of an inactivator of neurotransmitters may lead to decreased neurotransmitter levels and unanticipated effects in genetically predisposed individuals, such as increased aggression in the context of a fight between two dominance-driven opponents. Given the widespread use of pharmacological inhibitors of MAO22 and COMT82, understanding the long-term effects of inhibiting these enzymes with chronic treatment or during development (in utero, childhood, or adolescence) may have a broad and significant impact.
Materials and methods
Fly strains
Genotype | Source | Identifiers |
|---|---|---|
CS;;ebony (ebony mutant) | Bloomington Drosophila Stock Center, outcrossed by Shirasu-Hiza Lab | BDSC #497 |
CS;CS;CS (background of wCS strain) | Shirasu-Hiza Lab | |
UAS-ebony RNAi | Bloomington Drosophila Stock Center | BDSC #28,612 |
Control for UAS-ebony RNAi | Bloomington Drosophila Stock Center | BDSC #36,303 |
wCS;;tubulin-Gal4 | Shirasu-Hiza Lab | |
wCS;;repo-Gal4 | Bloomington Drosophila Stock Center, outcrossed by Shirasu-Hiza Lab | BDSC #7415 |
ebony Mutant selection
While the e1 mutant is more commonly used in ebony research, we chose to work with e11 because we could not molecularly validate two independent e1 stocks obtained from the Bloomington Drosophila Stock Center (BDSC #1658 and BDSC #8443). According to Miller et al.83, e1 mutants are expected to have a transposable element inserted in exon 2 (FlyBase)84, but amplifying the region around the expected insertion site produced a PCR band that was similar in size to that of the wild-type fly. The e11 mutant, on the other hand, produced the expected PCR band containing a ~ 500 bp deletion. Our results are consistent with a previously published report28, which found the expected deletion in the e11 mutant through PCR and DNA sequencing and a wildtype-sized exon 2 region in the e1 mutant. Those authors, like us, also could not sequence that portion of the e1 allele and suggested that an unknown inversion or insertion may have occurred in exon 3.
Sleep and locomotor activity
6 to 8-day-old male flies entrained on a 12:12 Light–Dark cycle were placed in individual 5 mm plastic tubes containing food. Tubes were placed in TriKinetics Drosophila Activity Monitors (DAMs)85 to record their locomotor activity for 5 days in 12:12 Light:Dark (LD). Beam break data was grouped into 1-min bins using DAM File Scan, and pySolo86 (Python-based software) was used to analyze sleep architecture and waking activity. Sleep was defined as a period of at least 5 min of inactivity46.
Courtship assay
Virgin males were collected from crosses within 6 h of eclosion, stored individually in vials for 6–8 days, and entrained on a 12:12 LD cycle. We used 16 mm diameter testing chambers (Greiner Bio-One 24-well no bottom plates) coated with Insect-a-Slip (BioQuip) (also referred to as Fluon) to prevent wall climbing. Two plates were stacked on top of each other with a plastic divider separating them, and the bottom plate was taped underneath with a plastic divider to prevent flies from escaping. Sexually unreceptive wCS females (mated within the last 24 h) were aspirated without anesthesia into the central eight wells of the bottom plate and males were aspirated without anesthesia into the corresponding chambers of the top plate. The plastic divider was removed and the male was dropped into the chamber with the female and videotaped for 10 min using Supereyes B003 + cameras and iSpy software to record courtship behavior towards the female at a frame rate consistent between control and experimental flies within each experiment, of either 10 or 20 frames per second (depending on the computer used for acquisition). Because courtship index represents frames of activity divided by total number of frames, differences in frame rate are normalized. We excluded videos with copulation from our analysis unless otherwise specified (i.e., when examining rates of copulation).
Aggression assay
The same virgin collection conditions and testing chambers were used as for the courtship assay (see above). Using a 1 mL syringe 2–3 h before testing, a single drop of food (Glucose – Archon Scientific) was placed in the center of each well in the bottom plate. One male from each pair was aspirated without anesthesia into one of the central eight wells of the bottom plate. The remaining male from each pair was aspirated without anesthesia into the corresponding well of the top plate. The plastic divider was removed so that the male from the top plate dropped into the bottom well. The males were videotaped for 10 min using Supereyes B003 + cameras and iSpy software to record aggressive behavior at a frame rate consistent between control and experimental flies within each experiment, of either 10 or 20 frames per second (depending on the computer used for acquisition). Because aggression index represents frames of activity divided by total number of frames, differences in frame rate are normalized. Aggression assays performed with the more ecologically relevant setup were done as described in Fernandez et al.39.
Quantification of courtship and aggression behaviors
The CalTech FlyTracker and JAABA softwares were used to automatically quantify the courtship index (CI)/aggression index (AI), or percent of frames spent courting/fighting in a 10-min period, for each male. The Caltech FlyTracker automatically generates data for each video frame such as the location of the male and female, the angle of the wings, the velocity of the flies, etc. We visually inspected each video to ensure tracking accuracy; if a video had tracking errors > 5% of frames we discarded the video. We trained the machine learning algorithm (JAABA) to quantify wing extension, chasing, and attempted mounting behaviors for courtship frame by frame based on the data from the Caltech FlyTracker. We trained the classifiers to detect boxing, wing threat, lunging, and chasing behaviors for aggression in the same way. Because two or more behaviors can co-occur simultaneously (such as wing extension and chasing in courtship, and boxing and wing threat in aggression), it is typical in the field to use a hierarchical code that would not double count frames as “courtship” or “aggression” when computing total courtship and aggression levels (i.e., Fig. 3B and S5A). Our courtship code gives greatest priority to wing extension, followed by chasing and attempted mounting; that is, if wing extension and chasing co-occur in a video frame, JAABA annotates the frame for wing extension only. Our aggression hierarchy gives precedence to boxing, followed by wing threat, chasing, and lunging. For non-hierarchical analysis of total time spent exhibiting aggressive behaviors, which we quantified in Figs. 7B and S5B, we computed the behavioral index for each behavior separately in JAABA and summed.
For aggression videos involving intra-genotype pairing, the aggression index was analyzed for the dominant fly of each fight, determined by which fly exhibited more offensive behavior. In videos with inter-genotype pairing, we manually checked each annotated videos to ensure Caltech FlyTracker accurately tracks the ebony mutant and control fly, which have distinct cuticle colors that can be discerned in our testing chambers.
To assess dominance, we counted the number of fight bouts occurring on food, which can involve a broad range of aggressive behaviors described by Chen et al.37, that led to a clear winner. The fly that successfully chases or nudges the opponent fly off the food is considered the winner of that fight encounter. A fight did not result in dominance if the flies ended a fight and both males remained on the food or left the food simultaneously. We excluded from this analysis any fights that did not occur on the food. Dominance in the more ecologically relevant arena was scored as described in Penn et al.87.
Statistical analysis
We assessed the normality of our data using the D’Agostino-Pearson normality test and the Shapiro–Wilk normality test, which have great power properties over a variety of different statistical distributions88. For data sets that passed both normality tests, we used the unpaired Student’s t-test with Welch’s correction when comparing two groups and the one-way ANOVA (Dunnett’s multiple comparisons) when comparing three or more groups. For data sets that failed either one of the normality tests, we used the Mann–Whitney U test when comparing two groups and the Kruskal–Wallis test with Dunn’s post hoc test when comparing three or more groups. For binary comparisons, we used Fisher’s exact test. See Table S1 for the n, statistical analyses, and justification for statistics for all experiments. All plotted values represent means, with error bars representing SEM.
qPCR
7-day-old male flies previously entrained to 12:12 LD were snap-frozen in liquid nitrogen and stored at −80 °C. RNA was extracted from 15 whole flies for each of 4 biological replicates per genotype with TRIzol (Invitrogen) following the manufacturer’s protocol. Samples were treated with DNaseI (Invitrogen), then heat inactivated. cDNA was synthesized by Revertaid First Strand cDNA Synthesis Kit (Thermo Scientific). PowerUp SYBR Mastermix (Applied Biosystems) was used to perform qRT-PCR using a CFXConnect thermal cycler (BioRad). Primer efficiency and relative quantification of transcripts were determined using a standard curve of serial diluted cDNA. Transcripts were normalized using Actin5C as a reference gene.
Primer Sequences:
ebony-fwd-GTCCGAAGTGGAGAAGAACG
ebony-rev-TCTGTGCTACCATGCTGGTC
Actin5C-fwd-TTGTCTGGGCAAGAGGATCAG.
Actin5C-rev-ACCACTCGCACTTGCACTTTC.
Western blot
Whole-body lysates of 10-day-old male flies (15 flies/sample) were separated by SDS-PAGE on a Tris–Acetate 3–8% gel (NuPAGE, Thermo Scientific), and transferred to a PVDF membrane (Immobilon, Thermo Scientific). The membrane was probed with rabbit polyclonal antibody against Ebony at 1:1000 in 3% BSA (kind gift from Dr. Wittkopp)89. HRP-conjugated anti-rabbit IgG antibody at 1:2000 in 3% BSA was used for signal detection (Cell Signaling, 7074). ECL chemiluminescence reagent (Pierce) was used to visualize horseradish peroxidase activity and detected by CCD camera using a BioRAD Image Station.
Quantification of neurotransmitters
ebony and CS brains, with laminas intact, were dissected in PBS on ice. 50 brains were dissected per sample, with four samples collected for each genotype (200 brains total/genotype). Before freezing, samples were gently spun down and excess PBS was pipetted off. Samples were stored into a −80 °C freezer, then shipped on dry ice to Children’s Hospital of Philadelphia Metabolomic Core for analysis.
Dopamine (DOP), serotonin (SE), octopamine (OCTOP), and histamine (HA) levels were determined using LC–MS/MS the Agilent 1260 Infinity Triple Quad 6410B mass spectrometry (MS) coupled with liquid chromatography (LC) system90,91. Both isotope dilution methodology and standard curve were used to determine neurotransmitter levels. To each sample of brain tissue extract, 2 pM of D4 labeled HA, SR, OCTOP and D3-DOP were added. Then, 160 µL of ethanol and 40 µL of pyridine were added. Derivatization was performed by adding 50 µL of ethyl chloroformate and shaking gently the vial until bubbles were gone. Derivatized neurotransmitters were extracted with ethyl acetate, and the organic phase was dried down and reconstituted in 100 µL of 0.1% of formic acid, spun at 14,000 rpm and transferred into injection vials. 25 µL of the derivatized sample were injected into the LC-MS system. For chromatographic separation, we used Agilent Poroshell 120 column (EC-C18), 3 × 100 µM. Solution A was 0.1% of formic acid and Solution B acetonitrile with 0.1% of formate and 0.005% of tri-fluoro-acetate (TFA). Flow gradient started with 50% of solution B, and then 60% after 3 min, 70%, after 5 min and 100% at 6–15 min. Precursor-product used for each compound was as follows: HA 256-138 and D4-HA 260-142 MRM; OCTOP 280-91 and D4 OCTOP 284-95; SE, 321-203 and D4-SE 325-207 MRM; DOP 370-252 and D3-DOP 373-255 MRM. Concentration of each compound in the sample was calculated based on isotopic enrichment as described92 and normalized to the amount of cellular protein/sample. Simultaneously, the concentration of each compound was verified by using standard curve prepared with a known concentration of each neurotransmitter ranging from 2 to 10 pM/mL and determination of the area under the peak of each compound.
Data availability
The authors declare that all data supporting the findings of this study are available, including replicate experiments, and will be made available upon reasonable request to the corresponding author, Dr. Mimi Shirasu-Hiza.
Change history
11 August 2023
The original online version of this Article was revised: In the original version of this Article, the author name María Paz Fernández was incorrectly indexed. The original Article has been corrected.
References
Perez, M. M., Schachter, J., Berni, J. & Quesada-Allue, L. A. The enzyme NBAD-synthase plays diverse roles during the life cycle of Drosophila melanogaster. J. Insect Physiol. 56, 8–13. https://doi.org/10.1016/j.jinsphys.2009.08.018 (2010).
Richardt, A. et al. Ebony, a novel nonribosomal peptide synthetase for beta-alanine conjugation with biogenic amines in Drosophila. J. Biol. Chem. 278, 41160–41166. https://doi.org/10.1074/jbc.M304303200 (2003).
Jacobs, M. E. Influence of light on mating of Drosophila melanogaster. Ecology 41, 182–188 (1960).
Kravitz, E. A. & Fernandez, M. P. Aggression in Drosophila. Behav. Neurosci. 129, 549–563. https://doi.org/10.1037/bne0000089 (2015).
Zhang, W., Guo, C., Chen, D., Peng, Q. & Pan, Y. Hierarchical control of Drosophila sleep, courtship, and feeding behaviors by male-specific P1 neurons. Neurosci. Bull. 34, 1105–1110. https://doi.org/10.1007/s12264-018-0281-z (2018).
Koganezawa, M., Kimura, K. & Yamamoto, D. The neural circuitry that functions as a switch for courtship versus aggression in Drosophila males. Curr. Biol. 26, 1395–1403. https://doi.org/10.1016/j.cub.2016.04.017 (2016).
Zwarts, L., Versteven, M. & Callaerts, P. Genetics and neurobiology of aggression in Drosophila. Fly 6, 35–48. https://doi.org/10.4161/fly.19249 (2012).
Zhou, C. et al. Molecular genetic analysis of sexual rejection: Roles of octopamine and its receptor OAMB in Drosophila courtship conditioning. J. Neurosci. 32, 14281–14287. https://doi.org/10.1523/JNEUROSCI.0517-12.2012 (2012).
Zhang, S. X., Rogulja, D. & Crickmore, M. A. Dopaminergic circuitry underlying mating drive. Neuron 91, 168–181. https://doi.org/10.1016/j.neuron.2016.05.020 (2016).
Suh, J. & Jackson, F. R. Drosophila ebony activity is required in glia for the circadian regulation of locomotor activity. Neuron 55, 435–447. https://doi.org/10.1016/j.neuron.2007.06.038 (2007).
Richardt, A., Rybak, J., Stortkuhl, K. F., Meinertzhagen, I. A. & Hovemann, B. T. Ebony protein in the Drosophila nervous system: optic neuropile expression in glial cells. J. Comp. Neurol. 452, 93–102. https://doi.org/10.1002/cne.10360 (2002).
Hovemann, B. T. et al. The Drosophila ebony gene is closely related to microbial peptide synthetases and shows specific cuticle and nervous system expression. Gene 221, 1–9. https://doi.org/10.1016/s0378-1119(98)00440-5 (1998).
Ng, F. S., Tangredi, M. M. & Jackson, F. R. Glial cells physiologically modulate clock neurons and circadian behavior in a calcium-dependent manner. Curr. Biol. 21, 625–634. https://doi.org/10.1016/j.cub.2011.03.027 (2011).
Freeman, M. R. & Doherty, J. Glial cell biology in Drosophila and vertebrates. Trends Neurosci. 29, 82–90. https://doi.org/10.1016/j.tins.2005.12.002 (2006).
Volkenhoff, A. et al. Glial glycolysis is essential for neuronal survival in Drosophila. Cell Metab. 22, 437–447. https://doi.org/10.1016/j.cmet.2015.07.006 (2015).
Logan, M. A. Glial contributions to neuronal health and disease: new insights from Drosophila. Curr. Opin. Neurobiol. 47, 162–167. https://doi.org/10.1016/j.conb.2017.10.008 (2017).
Semenoff, D. & Kimelberg, H. K. Autoradiography of high affinity uptake of catecholamines by primary astrocyte cultures. Brain Res. 348, 125–136. https://doi.org/10.1016/0006-8993(85)90368-3 (1985).
Grosjean, Y., Grillet, M., Augustin, H., Ferveur, J. F. & Featherstone, D. E. A glial amino-acid transporter controls synapse strength and courtship in Drosophila. Nat. Neurosci. 11, 54–61. https://doi.org/10.1038/nn2019 (2008).
Rival, T. et al. Physiological requirement for the glutamate transporter dEAAT1 at the adult Drosophila neuromuscular junction. J. Neurobiol. 66, 1061–1074. https://doi.org/10.1002/neu.20270 (2006).
Jackson, F. R. & Haydon, P. G. Glial cell regulation of neurotransmission and behavior in Drosophila. Neuron Glia Biol. 4, 11–17. https://doi.org/10.1017/S1740925X09000027 (2008).
Bilder, R. M., Volavka, J., Lachman, H. M. & Grace, A. A. The catechol-O-methyltransferase polymorphism: Relations to the tonic-phasic dopamine hypothesis and neuropsychiatric phenotypes. Neuropsychopharmacology 29, 1943–1961. https://doi.org/10.1038/sj.npp.1300542 (2004).
Yeung, A. W. K., Georgieva, M. G., Atanasov, A. G. & Tzvetkov, N. T. Monoamine oxidases (MAOs) as privileged molecular targets in neuroscience: Research literature analysis. Front. Mol. Neurosci. https://doi.org/10.3389/fnmol.2019.00143 (2019).
Hull, E. M., Muschamp, J. W. & Sato, S. Dopamine and serotonin: Influences on male sexual behavior. Physiol. Behav. 83, 291–307. https://doi.org/10.1016/j.physbeh.2004.08.018 (2004).
Connor, D. F., Harrison, R. J. & Melloni, R. H. Biogenic amines and the psychopharmacology of aggression. Expert Opin. Ther. Pat. 8, 349–359. https://doi.org/10.1517/13543776.8.4.349 (1998).
Eyjolfsdottir, E. et al. 772–787 (Springer International Publishing).
Kabra, M., Robie, A. A., Rivera-Alba, M., Branson, S. & Branson, K. JAABA: interactive machine learning for automatic annotation of animal behavior. Nat. Methods 10, 64–67. https://doi.org/10.1038/nmeth.2281 (2013).
Jacobs, M. E. Influence of beta-alanine on mating and territorialism in Drosophila melanogaster. Behav. Genet. 8, 487–502. https://doi.org/10.1007/BF01067478 (1978).
Pérez, M. M., Rossi, F., Quesada-Allué, L. A., Comparison of ebony gene from three ebony mutants. Drosophila Inf. Serv. 97, 30–32 (2014).
Pooryasin, A. & Fiala, A. Identified serotonin-releasing neurons induce behavioral quiescence and suppress mating in Drosophila. J. Neurosci. 35, 12792–12812. https://doi.org/10.1523/JNEUROSCI.1638-15.2015 (2015).
Liu, T. et al. Increased dopamine level enhances male-male courtship in Drosophila. J. Neurosci. 28, 5539–5546. https://doi.org/10.1523/JNEUROSCI.5290-07.2008 (2008).
Andrews, J. C. et al. Octopamine neuromodulation regulates Gr32a-linked aggression and courtship pathways in Drosophila males. PLoS Genet. 10, e1004356. https://doi.org/10.1371/journal.pgen.1004356 (2014).
Alekseyenko, O. V., Chan, Y. B., Li, R. & Kravitz, E. A. Single dopaminergic neurons that modulate aggression in Drosophila. Proc. Natl. Acad. Sci. U S A 110, 6151–6156. https://doi.org/10.1073/pnas.1303446110 (2013).
Dierick, H. A. & Greenspan, R. J. Serotonin and neuropeptide F have opposite modulatory effects on fly aggression. Nat Genet 39, 678–682. https://doi.org/10.1038/ng2029 (2007).
Hoyer, S. C. et al. Octopamine in male aggression of Drosophila. Curr Biol 18, 159–167. https://doi.org/10.1016/j.cub.2007.12.052 (2008).
Everaerts, C., Farine, J. P., Cobb, M. & Ferveur, J. F. Drosophila cuticular hydrocarbons revisited: mating status alters cuticular profiles. PLoS One 5, e9607. https://doi.org/10.1371/journal.pone.0009607 (2010).
Ejima, A. et al. Generalization of courtship learning in Drosophila is mediated by cis-vaccenyl acetate. Curr. Biol. 17, 599–605. https://doi.org/10.1016/j.cub.2007.01.053 (2007).
Chen, S., Lee, A. Y., Bowens, N. M., Huber, R. & Kravitz, E. A. Fighting fruit flies: a model system for the study of aggression. Proc. Natl. Acad. Sci. U S A 99, 5664–5668. https://doi.org/10.1073/pnas.082102599 (2002).
Chowdhury, B., Wang, M., Gnerer, J. P. & Dierick, H. A. The Divider Assay is a high-throughput pipeline for aggression analysis in Drosophila. Commun. Biol. 4, 85. https://doi.org/10.1038/s42003-020-01617-6 (2021).
Fernandez, M. P. et al. Pheromonal and behavioral cues trigger male-to-female aggression in Drosophila. PLoS Biol. 8, e1000541. https://doi.org/10.1371/journal.pbio.1000541 (2010).
Ly, S., Pack, A. I. & Naidoo, N. The neurobiological basis of sleep: Insights from Drosophila. Neurosci. Biobehav. Rev. 87, 67–86. https://doi.org/10.1016/j.neubiorev.2018.01.015 (2018).
Oh, Y., Jang, D., Sonn, J. Y. & Choe, J. Histamine-HisCl1 receptor axis regulates wake-promoting signals in Drosophila melanogaster. PLoS One 8, e68269. https://doi.org/10.1371/journal.pone.0068269 (2013).
Crocker, A., Shahidullah, M., Levitan, I. B. & Sehgal, A. Identification of a neural circuit that underlies the effects of octopamine on sleep:wake behavior. Neuron 65, 670–681. https://doi.org/10.1016/j.neuron.2010.01.032 (2010).
Yuan, Q., Joiner, W. J. & Sehgal, A. A sleep-promoting role for the Drosophila serotonin receptor 1A. Curr. Biol. 16, 1051–1062. https://doi.org/10.1016/j.cub.2006.04.032 (2006).
Crocker, A. & Sehgal, A. Octopamine regulates sleep in Drosophila through protein kinase A-dependent mechanisms. J. Neurosci. 28, 9377–9385. https://doi.org/10.1523/JNEUROSCI.3072-08a.2008 (2008).
Kume, K., Kume, S., Park, S. K., Hirsh, J. & Jackson, F. R. Dopamine is a regulator of arousal in the fruit fly. J. Neurosci. 25, 7377–7384. https://doi.org/10.1523/JNEUROSCI.2048-05.2005 (2005).
Huber, R. et al. Sleep homeostasis in Drosophila melanogaster. Sleep 27, 628–639. https://doi.org/10.1093/sleep/27.4.628 (2004).
Awasaki, T. & Ito, K. Engulfing action of glial cells is required for programmed axon pruning during Drosophila metamorphosis. Curr. Biol. 14, 668–677. https://doi.org/10.1016/j.cub.2004.04.001 (2004).
Borycz, J., Borycz, J. A., Edwards, T. N., Boulianne, G. L. & Meinertzhagen, I. A. The metabolism of histamine in the Drosophila optic lobe involves an ommatidial pathway: Beta-alanine recycles through the retina. J. Exp. Biol. 215, 1399–1411. https://doi.org/10.1242/jeb.060699 (2012).
Rendel, J. M. Mating of ebony vestigial and wild type Drosophila-melanogaster in light and dark. Evolution 5, 226–230. https://doi.org/10.1111/j.1558-5646.1951.tb02781.x (1951).
Crossley, S. & Zuill, E. Courtship behaviour of some Drosophila-melanogaster mutants. Nature 225, 1064–2000. https://doi.org/10.1038/2251064a0 (1970).
Kyriacou, C. P., Burnet, B. & Connolly, K. The behavioural basis of overdominance in competitive mating success at the ebony locus of Drosophila melanogaster. Anim. Behav. 26, 1195–1206 (1978).
Rossi, F. A., Bochicchio, P. A., Quesada-Allué, L. A. & Pérez, M. M. N-β-alanyldopamine metabolism, locomotor activity and sleep in Drosophila melanogaster ebony and tan mutants. Physiol. Entomol. 40(2), 166–174 (2015).
Newby, L. M. & Jackson, F. R. Drosophila-ebony mutants have altered circadian activity rhythms but normal eclosion rhythms. J. Neurogenet. 7, 85–101. https://doi.org/10.3109/01677069109066213 (1991).
Ramadan, H., Alawi, A. A. & Alawi, M. A. Catecholamines in Drosophila melanogaster (wild type and ebony mutant) decuticalarized retinas and brains. Cell Biol. Int. 17, 765–771. https://doi.org/10.1006/cbir.1993.1138 (1993).
Gruntenko, N. et al. The effect of mutations altering biogenic amine metabolism in Drosophila on viability and the response to environmental stresses. Arch Insect. Biochem. Physiol. 55, 55–67. https://doi.org/10.1002/arch.10123 (2004).
Hodgetts, R. B. & Konopka, R. J. Tyrosine and catecholamine metabolism in wild-type Drosophila melanogaster and a mutant, ebony. J. Insect Physiol. 19, 1211–1220. https://doi.org/10.1016/0022-1910(73)90205-9 (1973).
Borycz, J., Borycz, J. A., Loubani, M. & Meinertzhagen, I. A. tan and ebony genes regulate a novel pathway for transmitter metabolism at fly photoreceptor terminals. J. Neurosci. 22, 10549–10557 (2002).
Denno, M. E., Privman, E., Borman, R. P., Wolin, D. C. & Venton, B. J. Quantification of histamine and carcinine in Drosophila melanogaster tissues. ACS Chem. Neurosci. 7, 407–414. https://doi.org/10.1021/acschemneuro.5b00326 (2016).
Borycz, J., Borycz, J. A., Kubow, A., Lloyd, V. & Meinertzhagen, I. A. Drosophila ABC transporter mutants white, brown and scarlet have altered contents and distribution of biogenic amines in the brain. J. Exp. Biol. 211, 3454–3466. https://doi.org/10.1242/jeb.021162 (2008).
Sitaraman, D., Aso, Y., Rubin, G. M. & Nitabach, M. N. Control of sleep by dopaminergic inputs to the Drosophila mushroom body. Front. Neural Circuits 9, 73. https://doi.org/10.3389/fncir.2015.00073 (2015).
Ueno, T. et al. Identification of a dopamine pathway that regulates sleep and arousal in Drosophila. Nat. Neurosci. 15, 1516–1523. https://doi.org/10.1038/nn.3238 (2012).
Lim, J. et al. The mushroom body D1 dopamine receptor controls innate courtship drive. Genes Brain Behav. 17, 158–167. https://doi.org/10.1111/gbb.12425 (2018).
Erion, R., DiAngelo, J. R., Crocker, A. & Sehgal, A. Interaction between sleep and metabolism in Drosophila with altered octopamine signaling. J. Biol. Chem. 287, 32406–32414. https://doi.org/10.1074/jbc.M112.360875 (2012).
Hardie, R. C. Is histamine a neurotransmitter in insect photoreceptors?. J. Comp. Physiol. A 161, 201–213. https://doi.org/10.1007/BF00615241 (1987).
Sarthy, P. V. Histamine: A neurotransmitter candidate for Drosophila photoreceptors. J. Neurochem. 57, 1757–1768. https://doi.org/10.1111/j.1471-4159.1991.tb06378.x (1991).
Xiao, C., Qiu, S. & Robertson, R. M. The white gene controls copulation success in Drosophila melanogaster. Sci. Rep. 7, 7712. https://doi.org/10.1038/s41598-017-08155-y (2017).
Asahina, K. Neuromodulation and strategic action choice in Drosophila aggression. Annu. Rev. Neurosci. 40, 51–75. https://doi.org/10.1146/annurev-neuro-072116-031240 (2017).
Massey, J. H. et al. Pleiotropic effects of ebony and tan on pigmentation and cuticular hydrocarbon composition in Drosophila melanogaster. Front. Physiol. 10, 518. https://doi.org/10.3389/fphys.2019.00518 (2019).
Huang, K., Liu, Y. & Perrimon, N. Roles of insect oenocytes in physiology and their relevance to human metabolic diseases. Front. Insect Sci. 2, 7. https://doi.org/10.3389/finsc.2022.859847 (2022).
Kaufmann, J. H. On the definitions and functions of dominance and territoriality. Biol. Rev. 58, 1–20. https://doi.org/10.1111/j.1469-185X.1983.tb00379.x (1983).
Ullian, E. M., Christopherson, K. S. & Barres, B. A. Role for glia in synaptogenesis. Glia 47, 209–216. https://doi.org/10.1002/glia.20082 (2004).
Mejia, J. M., Ervin, F. R., Baker, G. B. & Palmour, R. M. Monoamine oxidase inhibition during brain development induces pathological aggressive behavior in mice. Biol. Psychiatry 52, 811–821. https://doi.org/10.1016/s0006-3223(02)01418-x (2002).
Frau, R., Pardu, A., Godar, S., Bini, V. & Bortolato, M. Combined antagonism of 5-HT(2) and NMDA receptors reduces the aggression of monoamine oxidase a knockout mice. Pharmaceuticals 15, 213. https://doi.org/10.3390/ph15020213 (2022).
Magwai, T. & Xulu, K. R. Physiological genomics plays a crucial role in response to stressful life events, the development of aggressive behaviours, and post-traumatic stress disorder (PTSD). Genes 13, 300. https://doi.org/10.3390/genes13020300 (2022).
Scott, A. L., Bortolato, M., Chen, K. & Shih, J. C. Novel monoamine oxidase A knock out mice with human-like spontaneous mutation. NeuroReport 19, 739–743. https://doi.org/10.1097/WNR.0b013e3282fd6e88 (2008).
Shih, J. C. et al. Ketanserin and tetrabenazine abolish aggression in mice lacking monoamine oxidase A. Brain Res. 835, 104–112. https://doi.org/10.1016/s0006-8993(99)01478-x (1999).
Qayyum, A. et al. The role of the catechol-o-methyltransferase (COMT) GeneVal158Met in aggressive behavior, a review of genetic studies. Curr. Neuropharmacol. 13, 802–814. https://doi.org/10.2174/1570159x13666150612225836 (2015).
Rujescu, D., Giegling, I., Gietl, A., Hartmann, A. M. & Moller, H. J. A functional single nucleotide polymorphism (V158M) in the COMT gene is associated with aggressive personality traits. Biol. Psychiat. 54, 34–39. https://doi.org/10.1016/S0006-3223(02)01831-0 (2003).
Volavka, J., Bilder, R. & Nolan, K. Catecholamines and aggression—The role of COMT and MAO polymorphisms. Youth Violence: Sci. Approaches Prev. 1036, 393–398. https://doi.org/10.1196/annals.1330.023 (2004).
Tosato, S. et al. Effect of COMT genotype on aggressive behaviour in a community cohort of schizophrenic patients. Neurosci. Lett. 495, 17–21. https://doi.org/10.1016/j.neulet.2011.03.018 (2011).
Alia-Klein, N. et al. Brain monoamine oxidase a activity predicts trait aggression. J. Neurosci. 28, 5099–5104. https://doi.org/10.1523/Jneurosci.0925-08.2008 (2008).
Fabbri, M., Ferreira, J. J. & Rascol, O. COMT inhibitors in the management of parkinson’s disease. CNS Drugs 36, 261–282. https://doi.org/10.1007/s40263-021-00888-9 (2022).
Miller, D. E., Cook, K. R., Arvanitakis, A. V. & Hawley, R. S. Third chromosome balancer inversions disrupt protein-coding genes and influence distal recombination events in Drosophila melanogaster. G3-Genes Genomes Genetics 6, 1959–1967. https://doi.org/10.1534/g3.116.029330 (2016).
Larkin, A. et al. FlyBase: Updates to the Drosophila melanogaster knowledge base. Nucleic Acids Res 49, D899–D907. https://doi.org/10.1093/nar/gkaa1026 (2021).
Rosato, E. & Kyriacou, C. P. Analysis of locomotor activity rhythms in Drosophila. Nat. Protoc. 1, 559–568. https://doi.org/10.1038/nprot.2006.79 (2006).
Gilestro, G. F. & Cirelli, C. pySolo: A complete suite for sleep analysis in Drosophila. Bioinformatics 25, 1466–1467. https://doi.org/10.1093/bioinformatics/btp237 (2009).
Penn, J. K., Zito, M. F. & Kravitz, E. A. A single social defeat reduces aggression in a highly aggressive strain of Drosophila. Proc. Natl. Acad. Sci. U S A 107, 12682–12686. https://doi.org/10.1073/pnas.1007016107 (2010).
Yap, B. W. & Sim, C. H. Comparisons of various types of normality tests. J. Stat. Comput. Simul. 81, 2141–2155. https://doi.org/10.1080/00949655.2010.520163 (2011).
Wittkopp, P. J., True, J. R. & Carroll, S. B. Reciprocal functions of the Drosophila yellow and ebony proteins in the development and evolution of pigment patterns. Development 129, 1849–1858 (2002).
Castillero, E. et al. Decreased serotonin transporter activity in the mitral valve contributes to progression of degenerative mitral regurgitation. Sci. Transl. Med. (in press).
van de Merbel, N. C. et al. Quantitative determination of free and total dopamine in human plasma by LC-MS/MS: The importance of sample preparation. Bioanalysis 3, 1949–1961. https://doi.org/10.4155/bio.11.170 (2011).
Weinberg, J. M., Venkatachalam, M. A., Roeser, N. F. & Nissim, I. Mitochondrial dysfunction during hypoxia/reoxygenation and its correction by anaerobic metabolism of citric acid cycle intermediates. Proc. Natl. Acad. Sci. U S A 97, 2826–2831. https://doi.org/10.1073/pnas.97.6.2826 (2000).
Acknowledgements
We thank all members of the Shirasu-Hiza and Canman labs for support, discussions, and feedback. We also acknowledge the Bloomington Drosophila Stock Center for fly stocks (NIH P40OD018537). Anti-Ebony antibody was a kind gift from Dr. Trisha Wittkopp, Department of Ecology and Evolutionary Biology, University of Michigan, Ann Arbor MI. The authors thank Y. Daikhin, O. Horyn, and Ilana Nissim for performing the neurotransmitter measurements in the Metabolomics Core Facility, The Children’s Hospital of Philadelphia. Work was supported by NIH T32GM007088 (M.P.), NIH F31AG074664 (S.J.T.), AFAR Glenn Foundation Postdoctoral Fellowship for Aging Research (M.U.), NIH R01GM117407 (J.C.C.), NIH R01GM130764 (J.C.C.), NSF 1656603 (S.S.), NIH R35GM127049 (M.S.-H.), and NIH R01AG045842 (M.S.-H.).
Author information
Authors and Affiliations
Contributions
M.P., Z.L., S.J.T, and M.S.-H. conceived the experiments. Experiments were performed and analyzed by M.P. (mutant validation, courtship assays, qPCR, brain dissections, sleep, and locomotor activity), Z.L. (courtship and aggression assays, qPCR), S.J.T. (courtship and aggression assays, qPCR), G.T. (aggression assays), M.U. (western blots), R.O. (brain dissections), and I.N. (LC-MS/MS analysis of neurotransmitters). B.Q. wrote MATLAB scripts for automating behavioral analysis. M.P., Z.L., S.J.T., J.S., R.D., S.S., J.C.C., M.P.F., and M.S.-H. made intellectual contributions. M.P., Z.L., S.J.T., J.S., J.C.C., and M.S.-H. designed the figures and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Pantalia, M., Lin, Z., Tener, S.J. et al. Drosophila mutants lacking the glial neurotransmitter-modifying enzyme Ebony exhibit low neurotransmitter levels and altered behavior. Sci Rep 13, 10411 (2023). https://doi.org/10.1038/s41598-023-36558-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-36558-7
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.